Abstract
Objectives
Classifying lung and colon cancer from histopathological images remains a significant challenge due to the high degree of intra-class feature similarity and complex tissue morphology, particularly in lung cancer cases. While convolutional neural networks (CNNs) have demonstrated strong spatial feature extraction capabilities, they cannot inherently model long-range dependencies and global contextual relationships. Although attention-based methods partially address these limitations, they often suffer from overfitting, limited generalization across heterogeneous datasets, and insufficient interpretability for clinical adoption. To address these challenges, this study presents a Multi-Head Attention-Based Convolutional Neural Network (MHAB-CNN) ensemble framework that captures localized and global feature interactions critical for robust cancer classification.
Methods
A k-fold cross-validation strategy is adopted to train multiple MHAB-CNN models, from which the empirically top-performing ones are selected and aggregated to form a compact ensemble. This approach improves robustness, reduces overfitting, and ensures computational efficiency. Grad-CAM-based visualizations interpret the discriminative regions influencing the model’s predictions.
Results
Experimental evaluation on the LC25000 dataset demonstrates that the proposed framework achieves an average validation accuracy of 99.84% across folds. Furthermore, the E3 ensemble configuration, comprising models M1, M6, and M9, achieves the highest classification score on the held-out test set.
Conclusion
The proposed MHAB-CNN ensemble framework effectively captures localized and global feature interactions critical for robust lung and colon cancer classification, while improving robustness, reducing overfitting, and enhancing interpretability for potential clinical adoption.
1. Introduction
Around the world, lung and colon cancer are the most widespread and fatal types of cancer. According to the World Health Organization (WHO), in 2022, there were 2.3 million new lung cancer cases, resulting in over 1.8 million deaths. Colon cancer, on the other hand, had about 1.9 million new cases and caused more than 935,000 deaths. 1 Lung cancer is expected to rank as the leading cause of cancer deaths around the world in the year 2023, with an accelerated trend of 2.5% each year in regions with high rates of smoking and industrial pollution. 2 These statistics underscore the urgent need for early diagnosis and effective management of lung cancer, particularly in its initial stages, which are increasingly prevalent and crucial for improving patient survival rates.
Analyzing histopathological images plays a vital role in cancer diagnosis by enabling pathologists to identify and classify malignancies based on abnormal cell morphology, structural patterns, and other pathological indicators. 3 However, this manual diagnostic process presents several challenges. It relies heavily on the expertise and judgment of individual pathologists, which can introduce variability and subjectivity into clinical outcomes. Fatigue, interpretative differences, and the subtlety of certain abnormalities may lead to missed or inconsistent diagnoses. 4 Furthermore, the process is time-consuming and labor-intensive, posing significant scalability concerns amid the growing demand for cancer diagnostics. The visual complexity of histopathological patterns often exceeds what can be reliably discerned by the human eye, even experienced professionals. These issues are especially pronounced in resource-constrained settings, where a shortage of trained pathologists leads to further delays in diagnosis and treatment. These challenges collectively underscore the need for advanced technological solutions, particularly artificial intelligence (AI), to enhance cancer diagnosis’s accuracy, efficiency, and scalability.
Machine learning (ML) and deep learning (DL) in medical imaging have significantly advanced diagnostic workflows by offering faster and more accurate analyses. Among various DL techniques, convolutional neural networks (CNNs) have demonstrated exceptional performance, particularly in analyzing histopathological images of lung and colon cancer. 5 Although CNNs effectively identify spatial features within images, they exhibit limitations due to their inherent architectural design. A prominent drawback is their inability to model long-range dependencies, as they predominantly capture local spatial relationships and fail to represent global contextual information. 6 This limitation can result in incomplete or fragmented interpretations, especially in complex medical images where global coherence is essential. Moreover, CNNs are sensitive to scale, shape, and orientation variations due to their fixed kernel sizes and pooling operations, which restrict their adaptability. They also lack mechanisms to selectively emphasize critical features, potentially leading to noisy or inefficient representations. To mitigate these shortcomings, attention mechanisms 7 can be incorporated into CNN architectures,, enabling more nuanced and context-aware analysis of complex and heterogeneous image data.
Attention mechanisms have become a fundamental component of modern DL architectures because they facilitate selective focus on relevant features. Among these, self-attention and its extension, MHA, were first introduced in the transformer architecture. 8 Self-attention enables the model to capture dependencies across all positions in the input, irrespective of distance, making it highly effective for representing global context. MHA enhances this capability by employing multiple parallel attention heads, each learning to focus on different input aspects. This parallel processing results in a richer and more expressive representation of the data. 9 Following the success of MHA in natural language processing, domain-specific attention mechanisms were proposed to address the unique needs of computer vision tasks. Notably, spatial attention 10 and channel attention 11 were developed to highlight important regions and informative feature maps in an image, respectively. While these methods have shown improvements in CNNs, they primarily operate within localized scopes and cannot model long-range dependencies. In contrast, MHA captures global relationships and feature interactions across the entire input, making it particularly advantageous for analyzing complex and diverse data, such as histopathological images.
The unique morphological characteristics of lung and colon cancer histopathological images further emphasize the suitability of Multi-Head Attention (MHA) as a foundational mechanism in diagnostic modeling. These images typically exhibit complex spatial configurations, heterogeneous tissue textures, and subtle variations in nuclear structure and glandular formation.12,13 Such fine-grained patterns, often dispersed across spatially distant regions, pose a significant challenge to traditional CNNs that rely on local feature extraction. 14 In contrast, MHA enables simultaneous attention to multiple image regions, capturing long-range dependencies and contextual cues for distinguishing between benign, malignant, and pre-malignant tissue subtypes. This capability is critical given the high inter-class similarity among cancer subtypes in both lung and colon tissue, where visual differences can be minimal and easily overlooked. MHA has demonstrated effectiveness in such contexts by improving focus on the most discriminative features while suppressing irrelevant regions. For instance, Wen et al. 15 leveraged MHA to enhance facial expression recognition by reducing distraction from non-informative areas. Similarly, Sun et al. 16 showed that MHA facilitates the identification of multiple salient parts within objects, leading to better class separation in visually similar categories. An et al. 17 introduced a repulsive loss function to diversify attention heads, encouraging the model to capture distinct and complementary features. These strategies are particularly valuable when applied to histopathological classification, where nuanced structural cues must be captured to achieve diagnostic reliability.
Although MHA significantly improves model expressiveness and classification accuracy, its flexibility and depth may increase the risk of overfitting, especially when limited training data. This concern is particularly relevant in medical domains where high-quality annotated datasets are scarce.18,19 While both CNN and attention-driven strategies have shown promise, their practical implementations in recent studies expose critical limitations in accuracy, generalizability, and interpretability.
For instance, Hasan et al. 20 proposed a lightweight multi-scale CNN to reduce parameter count while maintaining classification performance, and Al-Jabbar et al. 21 employed hybrid CNN architectures to improve sensitivity and accuracy. Building further, attention-based mechanisms have also been explored to overcome the locality limitations of traditional CNNs. Provath et al. 22 incorporated attention into CNNs, reporting enhanced diagnostic performance. Despite these improvements, several persistent challenges remain, including vulnerability to overfitting,18,19 excessive computational burden, 23 poor generalization to diverse datasets, 24 and lack of interpretability essential for clinical adoption. 25 Moreover, current attention-based models often underutilize ensemble learning strategies, which could otherwise enhance reliability and robustness. 22 To address current shortcomings, we suggest a new approach to ensemble learning that integrates CNNs with Multi-Head Attention MHA to improve the classification processes of lung and colon cancers from histopathological images.
Most current ensemble techniques in analyzing histopathological images are based on output synthesis from different architectures (e.g., VGG, ResNet) using simple averaging or voting. These methods are unfocused on model performance in ensemble formation and lack fold-based training.26,27
Traditional ensemble techniques, including bagging, stacking, and snapshot ensembles, often feature all models, regardless of the model’s performance, which deteriorates overall performance.28,29 Additionally, applying interpretable tools such as Grad-CAM, which are mainly focused on individual models, suffers from ensemble effects. 30
Oppositely, our approach has a uniform MHAB-CNN architecture trained in 10 folds, and only the top-3 models based on validation accuracy are selected to construct the ensemble. This selection technique improves model integration because weaker models are excluded, enhancing generalization and robustness. Model interpretability is also advanced at the ensemble level through Weighted and Intersection Grad-CAM heatmaps, strengthening knowledge in this domain. While attention mechanisms address the spatial locality constraints of CNNs, their complexity often increases the risk of overfitting, particularly in limited-data regimes common in medical imaging. To counter this, our approach incorporates a k-fold cross-validation strategy to train multiple lightweight CNN-MHA models, from which the n top-performing models are selected and aggregated. This ensemble strategy enhances predictive reliability, captures complementary representations, and reduces computational overhead by limiting the ensemble size. We conduct comprehensive ablation studies to optimize the number of attention heads used within each model to ensure a trade-off between performance and efficiency. Moreover, advanced visualization techniques are employed to qualitatively assess the discriminative capacity of individual models in the ensemble. The key contributions of this work are as follows: • We propose a novel and lightweight ensemble learning framework that integrates CNNs with MHA mechanisms to effectively capture spatially diverse and morphologically complex patterns in histopathological images. This combination enhances the model’s capability to discern fine-grained tissue characteristics across long-range dependencies, which is particularly vital in cancer subtype differentiation. • To mitigate overfitting and improve model generalization, especially in data-scarce medical settings, we employ a k-fold cross-validation scheme and construct an ensemble from the empirically top n performing CNN-MHA models. This strategy balances diversity and computational efficiency, ensuring that only the most robust models contribute to final inference. • We conduct extensive ablation studies to identify the optimal number of attention heads used within each MHA-augmented CNN. This analysis ensures the ensemble achieves a desirable trade-off between classification accuracy and computational cost, a critical requirement for practical deployment in clinical environments. • We employ Grad-CAM-based visualization techniques to qualitatively interpret the decision-making behavior of each model in the ensemble. This analysis enables a deeper understanding of the learned feature representations and provides insight into how the ensemble generalizes across diverse histopathological patterns. • Experimental results on lung and colon histopathological image datasets demonstrate the effectiveness of the proposed framework. The model achieved an average validation accuracy of 99.84% across 10-fold cross-validation, indicating strong and consistent performance. Furthermore, the ensemble achieved a 100% classification accuracy on the held-out test set, highlighting its generalization capability.
The rest of the paper is organized as follows. Section 2 presents the Related Works associated with our work. Section 3 represents the Methodology of the proposed work. Result and Discussion associated with the work is presented in Section 4 and Section 5, respectively. Finally, we present the Conclusion in Section 6.
2. Related Works
Lung and colon cancer classification using histopathological images has gained considerable attention in recent years, leveraging advanced deep-learning architectures, ensemble learning techniques, and optimization methods. Below, we provide a detailed review of key studies in this domain.
2.1. CNN based
Several recent works on lung and colon cancer diagnosis use CNN-based architectures with distinct design strategies that balance accuracy, interpretability, and computational efficiency. For example, Hasan et al. 20 proposed a lightweight multi-scale CNN (LW-MS CNN) that contains only 1.1 million parameters for the classification of lung and colon cancers with a multi-class prediction accuracy of 99.20%. The addition of explainable AI tools, Grad-CAM and SHAP, strengthened the interpretability and trustworthiness of the model. Although best suited for real-time applications, the light nature may also hinder grip over complicated patterns in heterogeneous datasets.
Summary of methods and limitations in lung and colon cancer classification studies.

2.2. Attention Based Method
Beyond standard convolutional models, attention mechanisms have also been incorporated to enhance feature representation and contextual learning in cancer diagnosis. Provath et al. 22 proposed a global context attention-based convolutional neural network, achieving an impressive image-level accuracy of 99.76% and patient-level accuracy of 96.5%. The model also reduced computational complexity, making it suitable for mobile deployment. However, the lack of ensemble methods limits their potential for improved robustness and generalization.
Similarly, Indumathi and Siva 33 introduced a hybrid CNN-BiLSTM model with a multi-head self-attention mechanism for predicting lung disorders using medical imaging datasets, achieving high classification accuracies of 94.9%, 97.8%, 97.9%, and 94.6% on the Chest X-ray, PET/CT, CECT, and JSRT datasets, respectively. Despite its superior performance compared to existing models, the computational complexity may hinder its real-time applicability in clinical environments.
2.3. Ensemble based learning
Ensemble-based approaches have also gained traction for cancer diagnosis, aiming to improve robustness, accuracy, and generalization by integrating multiple models or learning strategies. Sünnetci and Alkan 31 proposed an automated lung cancer diagnosis framework using machine learning algorithms, probabilistic majority voting, and optimization techniques. The method utilized Bag of Features for feature extraction and employed Linear Discriminant, Optimizable Support Vector Machine, and Optimizable K-Nearest Neighbor classifiers, achieving an accuracy of 99.28%. A detailed theoretical framework for majority voting was provided, and a user-friendly graphical interface was developed to aid radiologists. However, the study was confined to lung cancer detection and did not explore extending the approach to other cancer types, such as colon cancer.
Expanding the coverage to multi-class classification, Abd El-Aziz et al. 23 proposed a deep learning fusion model for multi-class lung and colon cancer classification. The model combines three pre-trained architectures: ResNet-101V2, NASNetMobile, and EfficientNet-B0, thereby leveraging feature fusion to boost classification accuracy. Therefore, it stands out with exceptional performance metrics, such as 99.94% accuracy on the LC25000 dataset. However, the reliance on multiple Convolutional neural networks (CNNs) increased computational demands, posing challenges for deployment in resource-constrained environments.
Mengash et al. 32 introduced the MPADL-LC3 approach, combining the marine predators algorithm with deep learning to classify lung and colon cancer. The supplementary preprocessing of this approach is based on CLAHE to augment contrast in images; then, feature extraction is performed by MobileNet, and classification by deep belief networks DBN. Hyperparameters were optimized using MPA, which thus increased the performance of the model to an accuracy of 99.27%. However, since it relied on a heuristic optimization algorithm, MPADL-LC3 could have volatile outcomes depending on parameter settings.
Talukder et al. 26 provide a hybrid ensemble model for cancer detection. The authors combine deep feature extraction with ensemble learning, achieving very accurate detection results using the ensemble technique to improve predictive performance. The model’s effectiveness is tested on the LC25000 lung and colon cancer dataset, detecting accuracies of 99.05% lung cancer, 100% colon cancer, and 99.30% combined lung and colon cancer. Nevertheless, computational complexity in merging several ensemble techniques hinders scalability while pointing to optimization that may enhance efficiency and applicability in clinical settings.
Razmjouei et al. 3 proposed a metaheuristic-based two-stage ensemble deep learning architecture to classify lung and colon cancers, thereby obtaining remarkable accuracies of 99.85% on two-class colon cancer and 98.96% on combined lung/colon cancer. However, hyperparameter sensitivity requires extensive tuning, which the model cannot generalize to different datasets. Moreover, multiple CNNs with ML models add non-interpretability; hence, it becomes difficult to justify what led the model to make a particular decision. This opacity is a major hindrance to its use in medical diagnosis.
Syal et al., 25 in a bid to improve the detection of lung and colorectal cancer, came up with an ensemble technique. The model was reliable and resulted in an accuracy of 0.96 percent. However, the paper does not provide specific explainability, which is a critical aspect in medical uses.
Unlike previous works that individually focused on CNNs, attention modules, or ensemble strategies, our proposed framework unifies these components into a single lightweight and interpretable system designed specifically for histopathological cancer diagnosis. While many earlier methods either lacked generalization across cancer types, introduced excessive computational overhead through complex ensemble combinations, or ignored interpretability aspects, our model addresses all three challenges simultaneously. By integrating MHA into compact CNN backbones, selecting top-performing models via k-fold validation, and employing Grad-CAM-based visual explanations, we ensure both high accuracy and transparency. Furthermore, the framework consistently performs well across validation folds and achieves perfect accuracy on the test set, demonstrating strong robustness and practical applicability in clinical scenarios.
3. Methods
The methodology adopted in this research is designed to systematically address the classification of lung and colon cancer using the LC25000 dataset.
34
The overall workflow, illustrated in Figure 1, outlines the key steps involved in the process, from data collection and preprocessing to model training, ensemble creation, and evaluation. Each major step in the workflow is detailed in the subsequent subsections, ensuring a clear understanding of the approach. Workflow of the proposed framework with multi-head attention-based CNN.
The workflow begins with collecting and preprocessing the data, including shuffling and splitting it into training and testing subsets. A fixed portion of the data is kept as an independent held-out test set, while the remaining data are used for training and validation. The training subset is further utilized for 10-fold cross-validation to train multiple instances of the proposed MHAB-CNN model. Models with the best validation performance are selected and combined into ensembles using mean and voting strategies to enhance classification reliability. These ensembles are then evaluated on the independent held-out test set to assess their performance. To further ensure interpretability, Grad-CAM visualizations are applied, highlighting the robustness and clarity of the proposed approach. Algorithm 1, outlines the workflow of a MHA-based CNN model, incorporating dataset splitting, k-fold cross-validation, ensemble creation, testing, and Grad-CAM visualization. The following sections elaborate on each of these steps, providing a comprehensive view of the methodology.


3.1. Dataset description
Class-wise sample distribution in the LC25000 dataset.


Sample images from the dataset, LC2500.
3.2. Data preprocessing
Several preprocessing steps were applied to prepare the dataset for training. First, all images were resized to a consistent dimension of 224 × 224 pixels. This ensures that every image has the same size, making them compatible with the model’s input requirements. Next, all images were converted to RGB format, ensuring they have the same color structure, regardless of their original format. The images were then transformed into tensors, scaling their pixel values to the range [0, 1], which is essential for efficient processing by the deep learning model. Additionally, labels were assigned to each image based on the folder structure of the dataset. Each class was mapped to a unique numerical value, making it easier for the model to understand and differentiate between the categories during training.
3.3. Dataset splitting and cross-validation
The dataset is partitioned into training and testing sets, ensuring class balance in both splits (Figure 3). Specifically, 10% of the data (with each class contributing 2% of the total samples) is set aside as the test set for final evaluation. The remaining 90% of the data (with each class contributing 18% of the total samples) is used for training. To ensure a thorough and reliable model evaluation, the training set undergoes 10-fold cross-validation, where it is evenly divided into 10 folds, with each fold serving as a validation set in turn while the others are used for training. For example: • In the first fold (F1), fold 1 is used for validation, and folds 2 through 10 are used for training. • In the last fold (F10), fold 10 is used for validation, and folds 1 through 9 are used for training. Dataset Splitting approach.

This process produces 10 trained models, each validated on a distinct fold, mitigating overfitting and ensuring the generalizability of the proposed model.
3.4. Proposed model: MHAB-CNN
Model architecture summary.

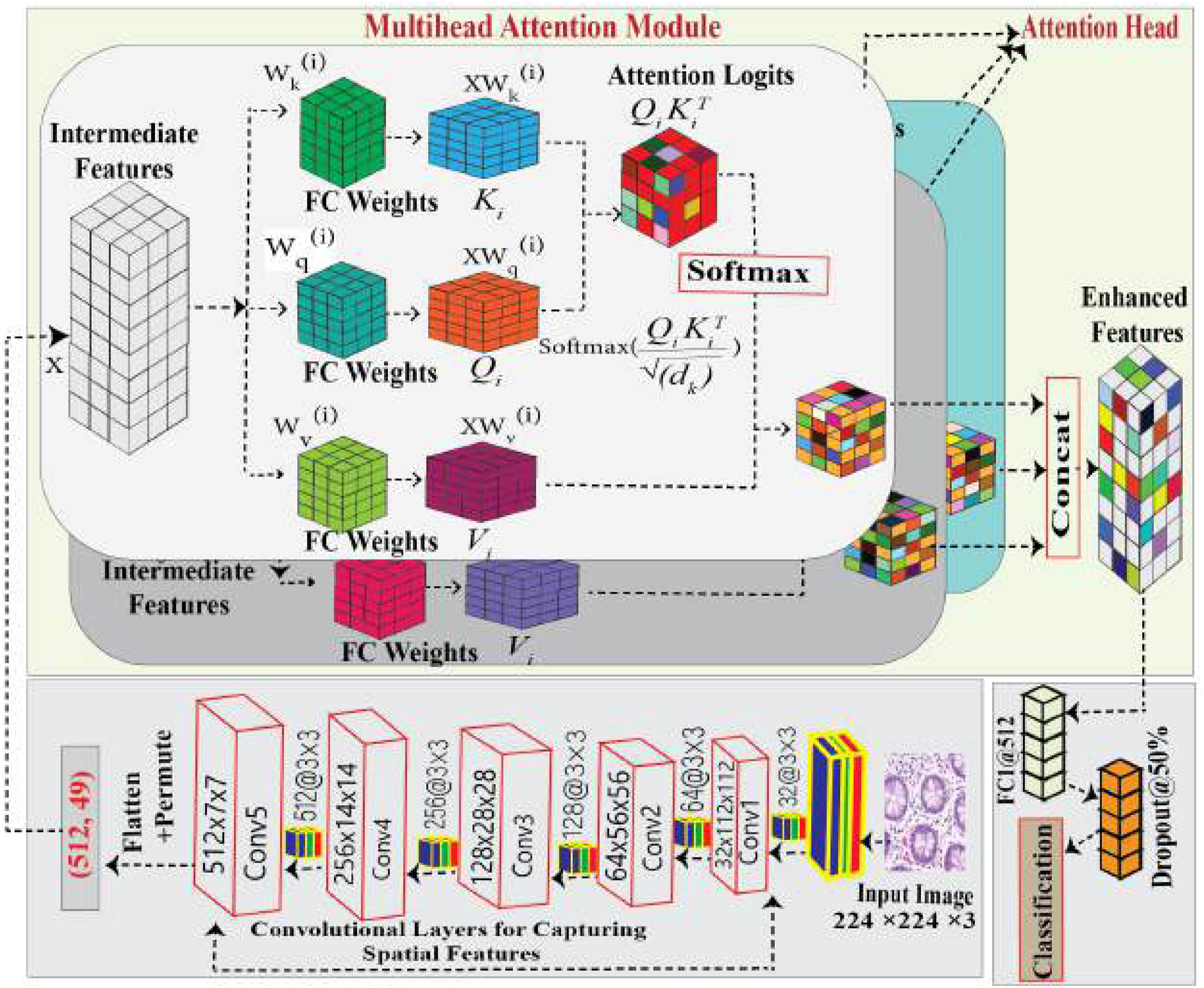
Overview of the Multi-Head Attention-Based CNN architecture for Lung and Colon Cancer Histopathology Classification.
3.4.1. Spatial feature extraction
The input images are processed sequentially through a series of layers designed to extract spatial features. This sequence comprises convolutional layers, batch normalization, ReLU activation, and max pooling, applied in blocks as illustrated in Figure 4. Each block progressively refines the spatial features, capturing essential patterns such as edges, textures, and structures. Below, we provide a detailed explanation of each component involved in the spatial feature extraction process.


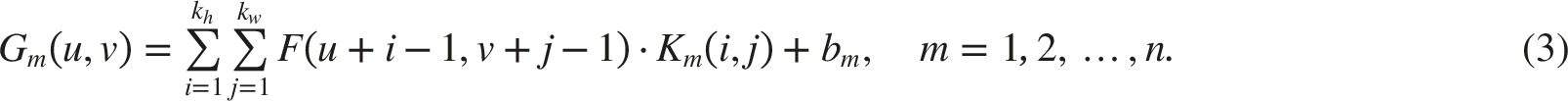
This process enables CNNs to learn diverse features across multiple layers, making them highly effective for tasks such as image recognition and object detection.35–37



ReLU activation keeps all positive values and zeros out all negative ones. It plays a role in alleviating the vanishing gradient problem, which helps improve gradient flow during training. ReLU is often used after each convolution to yield non-linear outputs. 38
The input feature map 

After extracting spatial features through the sequential convolutional blocks, it is essential to enhance the model’s ability to capture complex dependencies and relationships within the feature maps. While convolutional layers excel at local feature extraction, they may struggle to establish long-range dependencies, critical for distinguishing subtle patterns in medical images. To address this limitation, we integrate a multi-head attention, which enables the model to selectively focus on the most relevant regions of the extracted features.
3.4.2. Multi-head attention for enhanced cancer detection
In the context of lung and colon cancer detection, accurately identifying fine-grained details within medical images is crucial for improving diagnostic performance. We incorporate the MHA mechanism after convolutional blocks in the model pipeline to address this challenge. This approach enables the network to focus on critical regions within the feature maps generated by the sequential convolutional layers, ensuring a more comprehensive understanding of the complex patterns in medical imagery. Figure 4 shows the multi-head-based proposed CNN Architecture.
Architecture of the single head.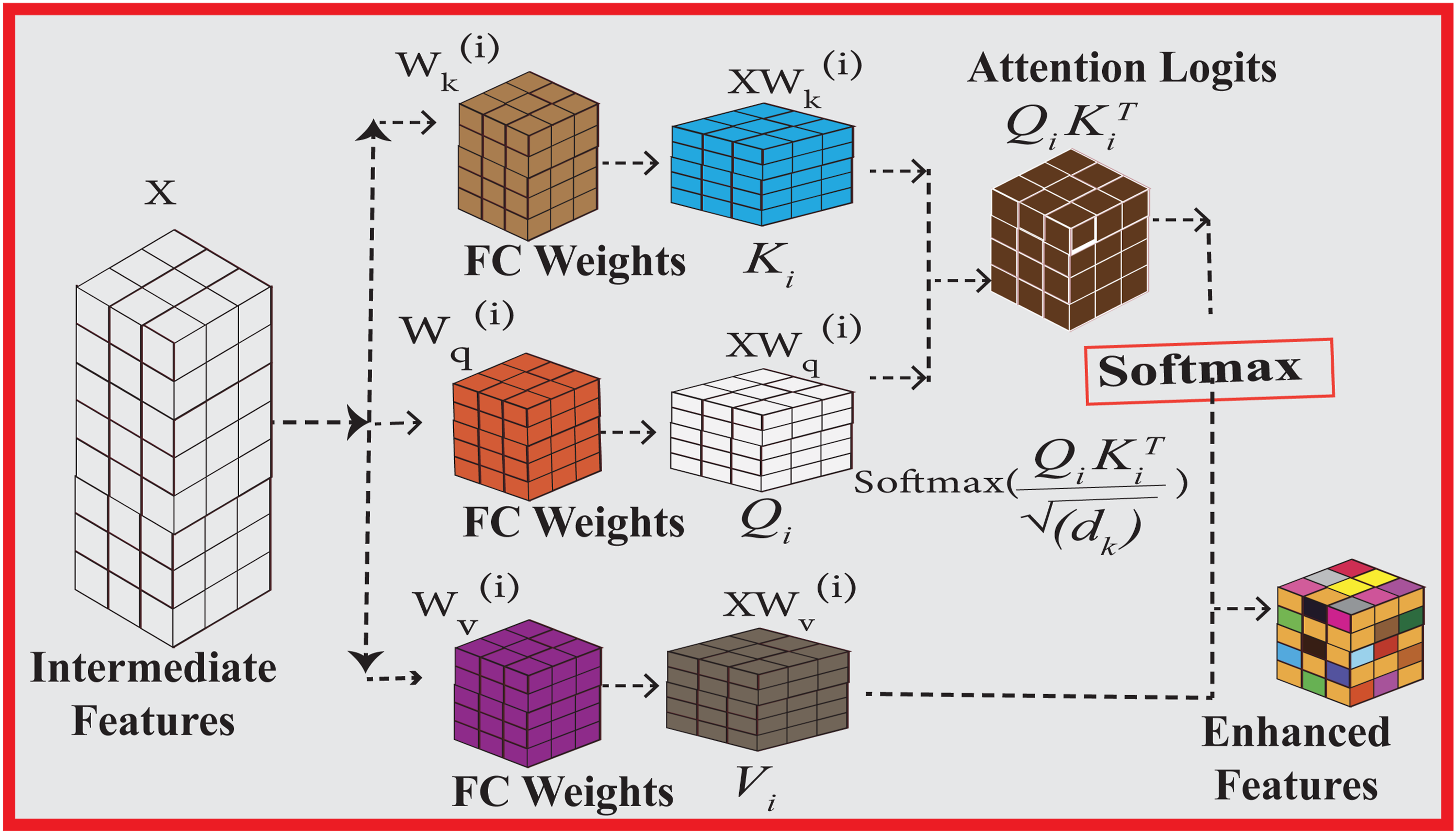
3.4.2.1. Scaled dot-product attention
The multi-head attention mechanism builds upon the scaled dot-product attention to compute relevance scores between different parts of the input feature space. Given the input feature matrix 
Here,
The scaled dot-product attention is computed as:
The term
3.4.2.2. Multi-head attention for feature refinement
To enable the model to attend to information from different representation subspaces jointly, multiple attention heads are used in parallel. For each head j = 1, 2, …, h, separate projections are computed:

The outputs of all heads are concatenated and projected to the final feature space:
3.4.2.3. Feed-forward network for feature processing
Following the multi-head attention, a position-wise feed-forward network (FFN) further processes the output. It consists of two linear transformations with a non-linear activation in between:
Here,
3.4.3. Classification steps
CNN models are composed of two sequential steps: feature extraction and classification. In the feature extraction step, a convolution operation is performed on the input image, resulting in a three-dimensional matrix or a tensor containing multiple image characteristics. In the classification step, fully connected layers, fluent with one-dimensional data, take the lead as ANN.


In our model, which is derived from MHAB-CNN architecture, all the fully connected layers had a dropout rate of 0.5. This approach inflicted 50% random deactivation of the neurons during the training, resulting in the network obtaining persistent and diverse feature representation. This assisted the network in learning the significant attributes within the particular classes and the general attributes that are requisite in distinguishing between the lung and colon cancer subtypes. The dropout experiments were performed deliberately, and in fact, this helped in generalization because it reduced the overfitting, which would make the model unable to perform on unseen data. In this instance, the dropout approach helped enhance the model performance against the LC25000 dataset; in this instance, increased accuracy and overfitting insensitivity were noted.
In this equation,




This method helps the model decide which class the input belongs to.
3.5. Individual model training and validation
The experimental dataset is split into k-Folds, where k ∈ {1, 2, …, 10} and each fold has its train and validation subset. These training and testing sets from each fold have been used to train the proposed MHAB-CNN, and after that, the performance of the trained model is verified using the testing set. In this way, ten distinct models (M k ) were created from every 10-folds where k ∈ {1, 2, …, 10}. Each of these trained models’ validation accuracy is recorded for consideration in creating an ensemble approach.
3.6. Ensemble model construction
An empirical analysis was conducted to select the top-performing models based on validation accuracy from the set of individual models {M1, M2, …, M10}. To improve overall classification performance and robustness, these selected models were combined to form ensemble models by aggregating their predictions. The selection of models for the ensemble was guided strictly by validation performance, ensuring rigorous evaluation and avoiding data leakage from the held-out test set. In this study, two common ensemble strategies—Mean Aggregation and Voting—were applied to evaluate the effectiveness of combining multiple models in enhancing predictive performance.
3.6.1. Mean aggregation
This approach works by taking the mean probability output from the selected models. In the end, the average probability for each class is calculated, and the top class is output as the final prediction. There are two straightforward procedures: 1. 2.


This approach incorporates the subjective probabilities of each model’s output, resulting in improved outcomes. Total certainty is taken into account from all models when probabilities are averaged.
3.6.2. Voting ensemble
The final class label in this approach is obtained using an ensemble technique, known as majority voting. In the selection, each model has the power to issue a vote for the class that the model predicts, and this is done in a series of steps as follows: 1. Count the total votes for each class 2. Determine the final predicted class by selecting the class with the highest vote count:


The chosen ensemble voting method is used per the consensus the models provide. The decision that is favored by the votes is selected as the outcome. Voting ensemble and mean aggregation are balancing techniques. While means aggregation offers probabilistic decisions on which prediction is most probably correct, the voting ensemble looks at the cast votes to make decisions, making it more robust to outliers or noise. Combining these methods to improve the accuracy and reliability of ensemble predictions is possible since they use different models.
3.7. Evaluation of ensembles
A test set held out from the training data is used to evaluate the constructed ensembles. This evaluation is intended to be both an objective and an accurate assessment of the performance of the ensembles constructed. The collection of measures used for assessment is exhaustive with respect to the aspects of classification effectiveness. Accuracy computes the proportion of the instances that have been correctly classified out of all the cases. In contrast, precision indicates the fraction of true positives from the tested optimistic predictions made by the model, which demonstrates the model’s accuracy. Recall or sensitivity quantifies the model correctly identifying positive instances, indicating how complete a model is in its specification. The F1-score which is defined as the average of the precision and recall may be helpful in such situations about class recognition. Cohen’s Kappa, in addition, allows for determining consensus that may occur purely by chance, even if or when the failure to reach an agreement has enabled specific measures’ results to be more complex. Together, these measures provide a good and comprehensive view of the effectiveness of the ensemble models trained in the cancer classification problem.
3.8. Visualization and robustness analysis
In verifying the interpretability and reliability of the developed ensemble models, the Grad-CAM visualizations are then used on the test samples. It uses the gradients of the output with respect to the feature maps and tries to focus on the image regions deemed necessary by the model in a given instance. Figure 12 presents representative results of Grad-CAM visualizations for the test samples of the models applying the proposed methodology, which supports the validity of the developed method.
4. Results
This section commences with a brief overview of the Environment Setup and Experimental Settings, before focusing on the Evaluation Metrics. In what follows, we provide the Numerical Results of our work. Then we turn to Grad-CAM Visualizations and Ensemble Justification. The final subsection of the section is dedicated to the Ablation Study.
4.1. Environment Setup and experimental settings
Training parameters.
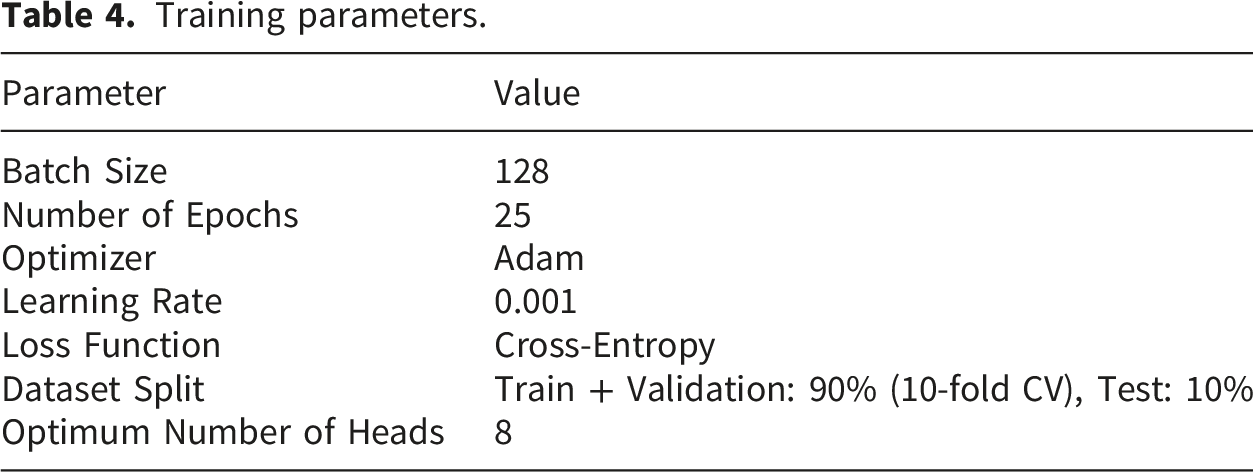
4.2. Evaluation matrices
In this study, we discuss how the performance of our model can be evaluated using multiple important metrics. The suite of metrics that we discuss includes accuracy, precision, recall, F1-score, Cohen’s Kappa, confusion matrix, ROC curves, accuracy curves and loss curves against epochs. Cohen’s Kappa is an index used to adjust the classification accuracy in the context of chance level agreement when comparing how predicted labels relate to actual labels. There are many instances of class imbalance, and accuracy is often misleading, and this metric is rather useful in those cases. If the class labels are not uniformly distributed, Cohen’s Kappa offers an advantage over the other metrics in that it measures how effective the model is while adjusting for the chance agreement
40
:
The Receiver Operating Characteristic (ROC) curve describes the graphical representation of the relationship between the sound range output of the recall and the false error rate at different threshold levels, which have been set. The area under the curve (AUC) is a global metric indicating the overall model’s capability to discriminate, and the model is ranked with respect to AUC – the higher the AUC value, the better the model. 41 With these metrics combined, we aim to deliver a qualitative and quantitative evaluation of our model’s performance from all angles.
4.3. Numerical Results
Ten implementations of the MHAB-CNN model were trained one at a time on a selected fold of a 10-fold dataset. Each model learned the assigned fold during training for 25 epochs. Different patterns have been captured because the folds are not strictly overlapping. Ensembles have been constructed after training all ten models. Based on the validation accuracy ranking obtained during 10-fold cross-validation, models M1, M6, and M9 were selected to form the E3 ensemble. The metrics of interest are precision, loss, and ROC curves.
Models M1, M6, and M9 have been trained and validated for 25 epochs as indicated in Figure 6. The first column displays accuracy as a function of epoch, whereas the second column displays loss as a function. Figure 6(a) shows the evolution of the training and validation accuracy of model M1 versus the number of epochs. In terms of metrics, training, and validation, there is a constant enhancement throughout epoch 2. In the third epoch, there is a short drop in the validation accuracy, followed by a rise. Subsequent validations display an increasing trend but with slight oscillation, notably around the fifth epoch, where the accuracy is roughly highest. However, in all epochs, the training accuracy also advances smoothly, but with minor oscillations. The validity accuracy at the end of the final epochs shows an upward trend despite some oscillations. Figure 6(b) displays the loss curves for the training and validation datasets of model M1, which shows a similar pattern in that the losses were also reducing over time. Figure 6(c) demonstrates the accuracy curve of the model M6 and the trend of training and validation accuracy with respect to the epochs. The loss curve of model M6 is presented in Figure 6(d), which shows the trend of training and validation losses throughout the training. Similarly, M9 is illustrated in Figure 6(e) accuracy curve, whereas its loss curve is presented in Figure 6(f). In composite, these subfigures narrate the learning of the M6 and M9 models as they display the accuracy and loss measures over epochs. So, all models M1, M6, and M9 succeed in learning, as we notice training and validation accuracies getting higher and higher, while training and validation losses getting lower and lower, but these models were at times hard on the validation set. Training and validation curves for models M1, M6, and M9 over 25 epochs. The left column shows accuracy vs. epoch, while the right column shows loss vs. epoch.
Figure 7(a) and 7(b) show the ROC curves for the training and testing sets of model M1 across five classes: Co_aca, Co_n, Lu_aca, Lu_n and Lu_scc. The AUC values take on a value of 1.0 except for Lu_aca and Lu_scc in both validation and test, indicating a slightly poor separation for these classes. Models M6 and M9 show a comparable performance trend, and their training and test ROC curves are located in Figure 7(c)–7(f), respectively. Again, these models have relatively high AUC values, except for Lu_aca and Lu_scc, where slight variations are noted. In Figure 8, the mean and voting ensemble of M1, M6, and M9 models’ performances improved greatly, resulting in AUC of 1.00 across all five classes. This further demonstrates the strength of the ensemble strategy in improving classification performance, especially for difficult classes. ROC Curves for models M1, M6, and M9 on validation and test sets. The left column shows ROC curves for the validation set, while the right column shows ROC curves for the test set. ROC curve on test data of E3 (a) mean ensemble (b) voting.

Performance metrics across different folds.

Performance of nine top models.

Composition of ensemble models and their constituent individual models.

Performance of voting and mean-based ensemble approaches for different numbers of models.

In group E5, all metrics reached perfect scores using the mean method. Additionally, the voting method performed strongly, with most scores centered around 0.9996. However, the mean method was more consistent for this group. The results for E7 and E9 remained stable, as most metrics returned consistently high scores. Accuracy, precision, recall, and F1-score were 0.9996, while the Cohen’s kappa value was 0.9995, and the AUC remained constant at 1.0000. Therefore, this confirms that these ensemble methods are reliable.
In conclusion, all ensemble methods performed strongly. The mean method performed best for E5, while the voting method showed a slight advantage for E3. It is also noted that all ensemble groups achieved an AUC of 1.0000, confirming the strong classification ability of these ensembles.
Performance Metrics of Ensemble Models (4 different combinations of M1, M6, M9, M10).

Performance metrics comparison between Ensemble of MHAB-CNN and S-CNN across different classes on the Test set.

M1, M6, and M9 with their evolution ensembles E3 (means and ensembles voting) techniques have resulted in the confusion matrices shown in Figure 9. The dataset has 25,000 test samples divided into 5 classes. These classes are Co_aca, Co_n, Lu_aca, Lu_n and Lu_scc. Model M1 had only two misclassifications. One Lu_scc sample is misclassified as Lu_aca, and another Lu_aca sample was misclassified as Lu_scc. Model M6 had three misclassifications, one Lu_aca sample was misclassified as Lu_scc and two Lu_scc samples were misclassified as Lu_aca. Model M9 performed poorly and had five misclassifications where three Lu_scc samples were misclassified as Lu_aca, one Co_aca sample was misclassified as Co_n and then a Lu_scc sample was misclassified as Lu_aca. Using ensemble M3 methods using both the mean and voting techniques, all 25000 test samples were correctly classified, meaning minimal errors were present and especially none. This proves the efficiency of ensemble methods, as with the combination of individual model strengths, their accuracy is increased while errors are reduced. The results show how ensembles increase robustness and generalisation, and prove they enhance overall classification performance. Confusion matrices for models M1, M6, M9, and their ensembles.
In Figure 10, we present the number of instances per class where the ensemble mechanism successfully corrected errors compared to the worst-performing individual model among three models (M1, M6, and M9). For each class, the maximum number of incorrect predictions across the individual models was used as the reference, and the number of instances where the ensemble produced the correct prediction was counted. From the results, we observe that no improvements were observed for the Co_aca and Lu_n categories, while one instance each was corrected for Co_n and Lu_scc. The most significant benefit was recorded for the Lu_aca category, where the ensemble corrected four instances compared to the worst individual model. The intrinsic characteristics of the LC25000 dataset can explain this pattern. Lung adenocarcinoma (Lu_aca) samples exhibit substantial morphological heterogeneity, including gland formation, nuclear size, and tissue architecture variations. Furthermore, Lu_aca shares visual similarities with both normal lung tissues (Lu_n) and lung squamous cell carcinoma (Lu_scc), increasing the likelihood of misclassification when relying on a single model. The ensemble mechanism leverages the diverse strengths of different models, mitigating individual weaknesses and achieving more robust predictions. As a result, ensemble learning proves particularly effective for complex and heterogeneous classes such as Lu_aca. Comparison of the number of instances per class where ensemble learning improved prediction accuracy over the worst-performing individual model.
4.4. Evaluation metrics and statistical validation of top-model ensemble
The deep learning models were assessed using 10-fold cross-validation. Based on the validation accuracy of the folds, we chose the top three models (M1, M6, and M9) to create ensemble classifiers utilizing weighted voting and Strict Majority Voting. To ensure consistency, the models were assessed on an identical testing dataset of n=2500 images.
Performance metrics with 95% confidence intervals for top models and ensembles.

The 95% Confidence Intervals (CIs) were calculated using the normal approximation method: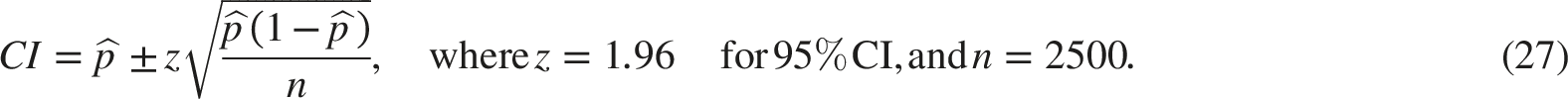
For the ensemble models, all test samples were correctly classified. Therefore, the estimated proportion
Additionally, to further examine the stability of the results, we applied bootstrap resampling with 1000 resamples. This approach estimates the variability of the evaluation metrics across different resampled datasets and provides more robust confidence interval estimates.
To determine whether the performance improvement from the ensemble models is statistically significant, we performed both the Paired T-Test and the Wilcoxon Signed-Rank Test using per-fold accuracy scores of the individual and ensemble models. The resulting p-values were: • Paired T-Test: p = 0.0634 • Wilcoxon Signed-Rank Test: p = 0.2500
While these p-values are above the commonly used significance level of 0.05, this outcome is expected and can be explained by two main reasons. First, the statistical tests were conducted using the accuracy scores of each fold obtained from the 10-fold cross-validation process. This results in a relatively small sample size for the hypothesis test. Second, the models being compared already demonstrate very high performance. As a result, there is very little variance among their results. Therefore, the gap between the standalone models and the ensemble models becomes very small and difficult to detect. In situations like these, statistical tests may not have enough power to detect such small differences. As such, the relatively large p-values should be interpreted with respect to the limited number of folds and the closely matching performance values across the models. Despite this, the ensemble models achieved the best performance across all evaluation metrics, which again supports the effectiveness and reliability of the proposed ensemble method.
5. Discussion
Comparison with state-of-the-art model.

Comparing Our Proposed Method with Previous Studies provides a detailed comparison between the outcomes achieved by our novel approach and those reported in earlier research efforts.
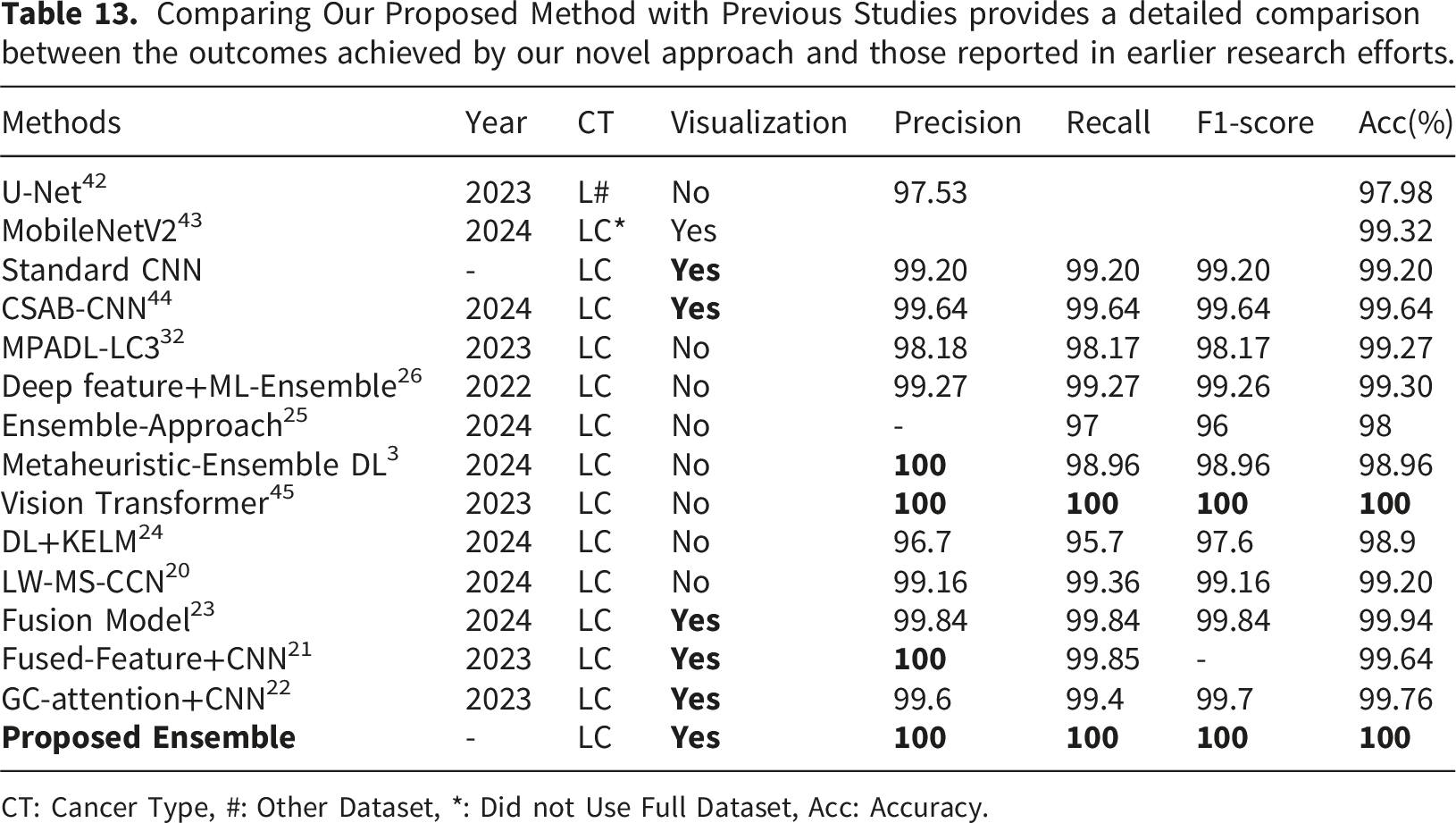
CT: Cancer Type, #: Other Dataset, *: Did not Use Full Dataset, Acc: Accuracy.
The LC25000 dataset is one of the most recognized benchmarks for the classification of histopathology images. However, past analyses of the dataset using t-SNE visualizations show that while some histopathology classes overlap and intermingle, other classes form relatively clear clusters. 44 Such a structured distribution of features may help explain the near-perfect performance results obtained in our experiments. In future work, we plan to further assess the generalizability of the proposed MHAB-CNN ensemble by testing it on a wider range of diverse datasets that more closely resemble real-world imaging conditions.
5.1. Ablation study
As illustrated in Figure 11(a), the mean ensemble approach shows a significant improvement in performance metrics (accuracy, precision, recall, F1-score, and Cohen’s kappa) as the number of attention heads, h, is doubled, following values h = [2,4,8,16,32]. At lower values of h, particularly 2 and 4, the model achieves moderate performance, with an accuracy of 0.6376 and 0.7588, respectively. Precision, recall, F1-score, and Cohen’s kappa also show similar trends of gradual improvement. However, at h = 8, there is a sharp rise, with metrics nearing optimal values (accuracy, precision, recall, F1-score all at approximately 0.9996 and kappa at 0.9995), marking a threshold beyond which further increases in head count have negligible impact, with all metrics remaining nearly constant at their peak values. This indicates that, for the mean ensemble, h = 8 is a critical point where performance stabilizes, suggesting that further increases in computational resources for more heads may not yield significant improvements. Similarly, Figure 11(b) demonstrates the impact of increasing head count on the voting ensemble performance. At lower values of h (2 and 4), metrics are moderate, with accuracy rising from 0.6416 to 0.7508, and corresponding increases in precision, recall, F1-score, and kappa. At h = 8, all metrics reach maximum values, achieving perfect accuracy and precision of 1.0, with recall, F1 − score, and kappa similarly reaching their upper bounds. Like the mean ensemble, further increases to h = 16 and h = 32 do not contribute additional performance gains. This indicates that the voting ensemble also benefits from an increased number of heads only up to h = 8. These results suggest that while both ensemble methods benefit from a higher number of heads, particularly up to h = 8, further doubling yields diminishing returns in performance. Impact of increasing number of heads on both mean and voting E3 model.
5.2. Grad-CAM visualization and ensemble justification
To verify applicability and strengthen interpretability of the ensemble model E3, composed of M1, M6, and M9, a Grad-CAM method was used to display images that contain the regions of importance that contribute the most to the predictions made by the model. This section details how Grad-CAM was computed for each of the individual models, how the ensemble visualizations have been created, and how these have been used to support the performance of the ensemble model. Grad-CAM generates for each model Mi a class-specific localization map 

To facilitate the interpretation of Grad-CAM, the heat map Visualization of Grad-CAM heatmaps for selected models and ensemble techniques.


Throughout the remaining samples, models that agree on the same predicted class (via voting) consistently highlight similar regions in their Grad-CAM heatmaps. The WE heatmap represents the averaged relevant regions emphasized by models that voted for the same class. In contrast, the IE heatmap focuses on the intersection of these regions, showcasing only the most consistent features. These heatmaps provide a valuable tool for quick and reliable pathological analysis by highlighting averaged and common areas critical for classification decisions.
5.3. Deployment feasibility
Inference time.

Overall, both models show efficiency. Nevertheless, the ensemble model exhibited improved accuracy and only suffered a small increase of time taken for inference.
6. Conclusion
In conclusion, our research demonstrates the effectiveness of MHA-based CNNs combined with ensemble learning for classifying lung and colon tissues, including benign and cancerous types. By utilizing advanced attention mechanisms and ensemble techniques, we achieved perfect performance metrics across all folds on the LC25000 dataset, including a train accuracy, validation accuracy, precision, recall, and kappa score of 100%. Grad-CAM visualizations further supported the model’s reliability by highlighting relevant regions for classification; however, formal validation of these visual explanations by expert pathologists remains an important direction for future work. These results signify a meaningful advancement in automated histopathological analysis, offering valuable insights for clinical diagnosis and treatment planning. Despite these achievements, challenges remain. The availability of diverse and comprehensive datasets is crucial for improving the model’s generalizability to a broader range of pathological conditions. Additionally, while Grad-CAM aids interpretability, further integration of explainable AI techniques is necessary to enhance trust in AI-based diagnostic tools. Future efforts could also focus on integrating the model with multi-modal datasets, including genetic or radiological data, to improve diagnostic accuracy. Other promising directions are exploring lightweight versions of the model for deployment in resource-constrained settings and incorporating continual learning mechanisms to adapt to evolving data. These advancements could bridge the gap between research and practical clinical applications, enhancing patient outcomes.
Footnotes
Author contributions
Conceptualization, A.K.Z.R.R., S.M.M.R.S., A.D.R., S.B., S.K. K.G.K.; Methodology, A.K.Z.R.R., S.M.M.R.S., A.D.R.; Software, S.B., S.K., K.G.K.; Validation, K.G.K., A.W.R., A.K.B., S.A.; Formal analysis, S.M.M.R.S., A.W.R., A.K.B., M.A.M.; Investigation, A.K.Z.R.R., A.D.R., S.B.; Resources, S.M.M.R.S., A.W.R., A.K.B.; Data curation, A.K.Z.R.R., A.D.R., S.K., K.G.K.; Writing—original draft, A.K.Z.R.R., S.A., A.D.R., S.B., S.K., K.G.K.; Writing—review & editing, M.A.M. S.B., S.A., K.G.K., A.W.R.; Visualization, A.K.Z.R.R., S.M.M.R.S., A.D.R., S.B.; Supervision, A.W.R., M.A.M; Funding acquisition, S.A., M.A.M. All authors have read and agreed to the published version of the manuscript.
Funding
This work was funded by the Ongoing Research Funding program, (ORF-2026-623), King Saud University, Riyadh, Saudi Arabia.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The dataset used during the current study is publicly available from the LC25000 dataset. 34
