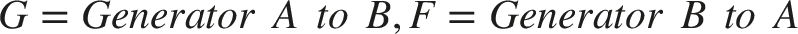Abstract
Objective
Stain color variations caused by differences in staining environments and scanning devices pose a major challenge for deep learning–based analysis of digital histopathological images. This study aims to develop a robust stain normalization framework that preserves structural information while enabling stable color-domain conversion across heterogeneous stain domains.
Methods
We propose a generative adversarial network (GAN)–based training and testing framework, termed I-GAN, which integrates StainGAN and Stain-to-Stain Translation (STST). The method incorporates identity loss within an RGB–grayscale training strategy and applies RGB images during testing to preserve original stain information. Performance was evaluated on the MITOS-ATYPIA 14 dataset using SSIM, PSNR, and DeltaE-ITP, and further assessed on downstream classification tasks using Camelyon17 and the ICIAR2018 BACH Challenge datasets.
Results
On MITOS-ATYPIA 14, I-GAN achieved an SSIM of 0.980, a PSNR of 29.579, and a DeltaE-ITP of 46.284, indicating superior structural preservation and color fidelity. For classification tasks, I-GAN obtained an average precision of 0.964 on Camelyon17 and an accuracy of 0.87, precision of 0.86, and recall of 0.87 on the ICIAR2018 BACH dataset.
Conclusions
The proposed I-GAN framework improves stain normalization for hematoxylin and eosin–stained digital histopathology images by preserving structural integrity and achieving accurate color-domain conversion. These results demonstrate the robustness and practical applicability of the proposed approach for medical image analysis.
1. Introduction
The number of studies applying deep-learning techniques to the field of medical image analysis has increased enormously in recent years. The reason for this is because of the ability of deep learning models to process large and complex medical image data of various types, especially because of the increasing popularity of whole-slide image (WSI) applications. For example, Hegde et al. 1 and Chen et al. 2 applied deep learning to WSI retrieval, whereas Shao et al. 3 and Khened et al. 4 focused on segmentation and classification of medical WSI medical images. These automatic analytical results can assist clinical physicians and laboratory scientists to diagnose diseases effectively and efficiently. However, owing to the inconsistency of pathological tissue staining techniques or images scanned by different scanners, the accuracy of automatic identification of pathological tissue images by deep learning prediction models is corresponding lower. To effectively solve these problems, several color transformation techniques have been developed specifically for stain normalization in digital pathological images.5,6 Hoque et al. 5 summarized that variation in scanners, tissue preparation protocols, staining reagents, dye concentration, pH, and staining time were all contributing factors to variability in H&E appearance, and Xie et al. 7 demonstrated that residual water and high-humidity storage conditions could affect protein integrity and cause detectable changes in eosin staining intensity.
Staining pathological tissue on slides can lead to inconsistent staining results owing to differences in humidity, temperature, and dye concentrations. In addition, the staining difference caused by different scanning devices can also influence the staining results. Although this does not have a major impact on manual interpretation by pathologists, it can easily cause misjudgments in computer-aided automatic judgment. Thus, if deep learning technologies are used to create automatic prediction models, stain normalization preprocessing technologies for pathology images in general can be used to train and create automatic identification models for digital pathology image analysis.
Several methods have been proposed to solve the problem of inconsistencies in the staining results of pathology images. These various approaches can be divided into conventional algorithms and deep-learning-based techniques for stain normalization. Conventional stain normalization methods include the techniques proposed by Vahadane et al., 8 Reinhard et al., 9 and Macenko et al. 10 Vahadane’s method performed stain separation in the optical density (OD) space using sparse non-negative matrix factorization (SNMF), Macenko’s method was also working in OD space and estimated stain vectors via singular value decomposition (SVD), whereas Reinhard’s method performed color transfer by matching the per-channel mean and standard deviation in the Lab color space. As for deep-learning-based approaches, the StainGAN 11 technique was first proposed by Shaban et al. based on the CycleGAN 12 framework. StainGAN successfully applied GAN technology to stain normalization by training the GAN to convert different color domains. Subsequently, StainNet 13 was proposed by Kang et al. to improve the stain normalization performance of StainGAN. Munien et al. 14 applied the conventional stain normalization techniques of Reinhard and Macenko to StainNet and proved that stain normalization can be effectively improved in terms of precision, recall, and accuracy for cancer identification models by combining conventional algorithms and deep learning techniques to construct automatic identification systems for pathology image diagnosis. It is worth noting that while stain normalization should improve the generalization ability of prediction model, stain-normalized images sometimes lead to a performance degradation in real-world applications. Cong et al. proposed the Color Adaptive Generative Adversarial Network (CAGAN), 15 which combined supervised learning on paired target-domain images with unsupervised learning from source-domain images via a dual-decoder architecture to achieve more robust stain normalization across heterogeneous domains. Salvi et al. 16 introduced GCC-GAN and GSN-GAN, generative models that formulated color normalization in digital pathology and dermatology as a pixel-wise image-to-image translation. However, Hoque et al. 5 pointed out that during stain normalization, important information might be lost or artifacts might appear in the newly normalized images, which affected the prediction accuracy of pathological image analysis.
Conventional stain normalization techniques for pathology images and deep-learning approaches typically assumed a single source–target setting, where images from one stain domain were mapped to a predefined target stain domain. This performed reasonably well when data came from only these two domains, but in realistic multi-center and multi-scanner scenarios, many heterogeneous stain styles might exist simultaneously. In such cases, pairwise source–target designs became difficult to scale, because a separate normalization model or a target selection was required for each new domain. In particular, traditional stain normalization techniques must manually select the required target images one by one, which made them especially sensitive to the choice of reference images; an inappropriate target selection could lead to suboptimal or even failed in stain normalization.17,18 As shown in Figure 1, different target images were selected and applied to Vahadane’s color-domain conversion technique, which significantly affect the results. Figure 1 illustrates that even when three very similar target images were selected, the original source images were significantly converted to different color domains. Vahadane’s method has shown that the conversion results would lead significantly different color domains when selecting similar target images (Vahadane1, Vahadane2, Vahadane3).
When deep-learning techniques were used for stain color space conversion, most existing methods still relied on explicitly defining one source stain domain and one target stain domain for training. For example, the training processes of StainGAN required two different stain domains to be specified in advance, and the generator was then trained to map images from domain A to domain B. This is effective when there was a single, well-defined source and target pair. However, in cross-center and cross-scanner settings, data might come from many stain domains. If there were n different stain domains and all of them were normalized to the same target domain, in principle, requiring up to (n − 1) separated source–target generators.
Another popular deep-learning technique for stain normalization is the STST technique proposed by Salehi et al. 19 Recently, several stain normalization approaches have adopted training schemes that use grayscale source images paired with their corresponding RGB target images, thereby avoiding the need to explicitly define two stain domains during training.18,19 The STST requires only a single target color domain for training. The original image and the corresponding grayscale image were fed into the Pix2Pix architecture for training. This method can convert grayscale images into the Hematoxylin and Eosin (H&E) color domain, overcoming the disadvantages of StainGAN, which requires two different color domains to be defined in advance. However, due to the input image is a grayscale image during the testing procedures, the originally stained information is lost, resulting in the quality of the converted images being worse than that of StainGAN in terms of evaluation indicators, such as the structural similarity index measure (SSIM) metric.
To simultaneously solve the problems of selecting two stain domains required by conventional training and testing methods and the issue of maintaining image quality caused by using grayscale images, a new training-testing method based on StainGAN is proposed. We prove that our proposed model can outperform the conventional training and testing methods in terms of the evaluation indicators of the SSIM and peak signal-to-noise ratio (PSNR). Our proposed I-GAN framework only requires the selection of one target stain domain to be specified during training. The generator is trained with RGB–Gray pairs from this target domain together with Identity Loss, so it learns to reproduce the target stain appearance, and which can effectively solve the problem of facing multiple different color domains simultaneously and avoid the subsequent misidentification caused by the loss of part of the image information when training using only the original grayscale image. The model architecture is explained in detail in the following sections.
2. Materials and methods
2.1. Dataset
Three datasets, including MITOS-ATYPIA 14 Challenge, 20 Camelyon17 Challenge, 21 and ICIAR2018 BACH Challenge 22 were used for comparison in this study. The results of the final converted histopathological images from the different approaches were compared and analyzed using SSIM, PSNR, DeltaE-ITP, and classification performance.
The first MITOS-ATYPIA 14 Challenge dataset was generated by scanning each slide using the Hamamatsu and Aperio scanners. The images generated using the Hamamatsu scanner were resampled and aligned according to the resolution of the Aperio scanner. These original images were then segmented into sub-images with a size of 256 × 256 pixels, generating 38,400 training images and 15,874 test images. Simultaneously, each training image was transformed into its corresponding grayscale image for the subsequent model training. The second Camelyon17 Challenge dataset was obtained from five medical centers, each of which provide 20 pathologist-labeled WSIs of breast cancer annotations and normal regions, for a total of 100 cancer WSIs. We also segmented the original images into sub-images of 256 × 256 pixels. Using the provided pixel-level annotation masks, we calculated the proportion of tumor pixels within each patch and removed patches in which the cancerous region occupied less than 25% of the total patch area. After these preprocessing steps, 12, 840 training, 2, 545 validations, and 1, 118 test images were generated. Again, the corresponding grayscale images were generated for the subsequent GAN training. The third ICIAR2018 BACH Challenge is a breast cancer image dataset with four different categories: normal, benign, in situ carcinoma, and invasive carcinoma. It contains 100 images with a size of 2048 × 1536 pixels. They were also divided into training, validation, and test datasets at a ratio of 60:25:15. Finally, each image was resampled to a size of 512 × 512 pixels for subsequent training procedures. For cross-dataset evaluation, the ICIAR2018 BACH images were normalized using the I-GAN model trained on Camelyon17 256 × 256 patches, and the obtained stain-normalized images were fed into the ResNet50 classifier. From the first MITOS-ATYPIA 14 challenge dataset (15,874 images), which were used to measure the SSIM and PSNR metrics between the original and stain-normalized images and to evaluate the SSIM and DeltaE-ITP of the true and stain-normalized images.
In this study, the Camelyon17 and ICIAR2018 BACH Challenge datasets were used to verify image classification applications. Camelyon17 contains a binary classification dataset and the ICIAR2018 BACH Challenge contains a four-class classification dataset. The Camelyon17 dataset and its grayscale images were used to train the Camelyon17 stain normalization model. In the ICIAR2018 BACH Challenge the model trained on Camelyon17 was directly used for stain normalization. This can be used to check the robustness of the stain-conversion model for different datasets.
2.2. Methods
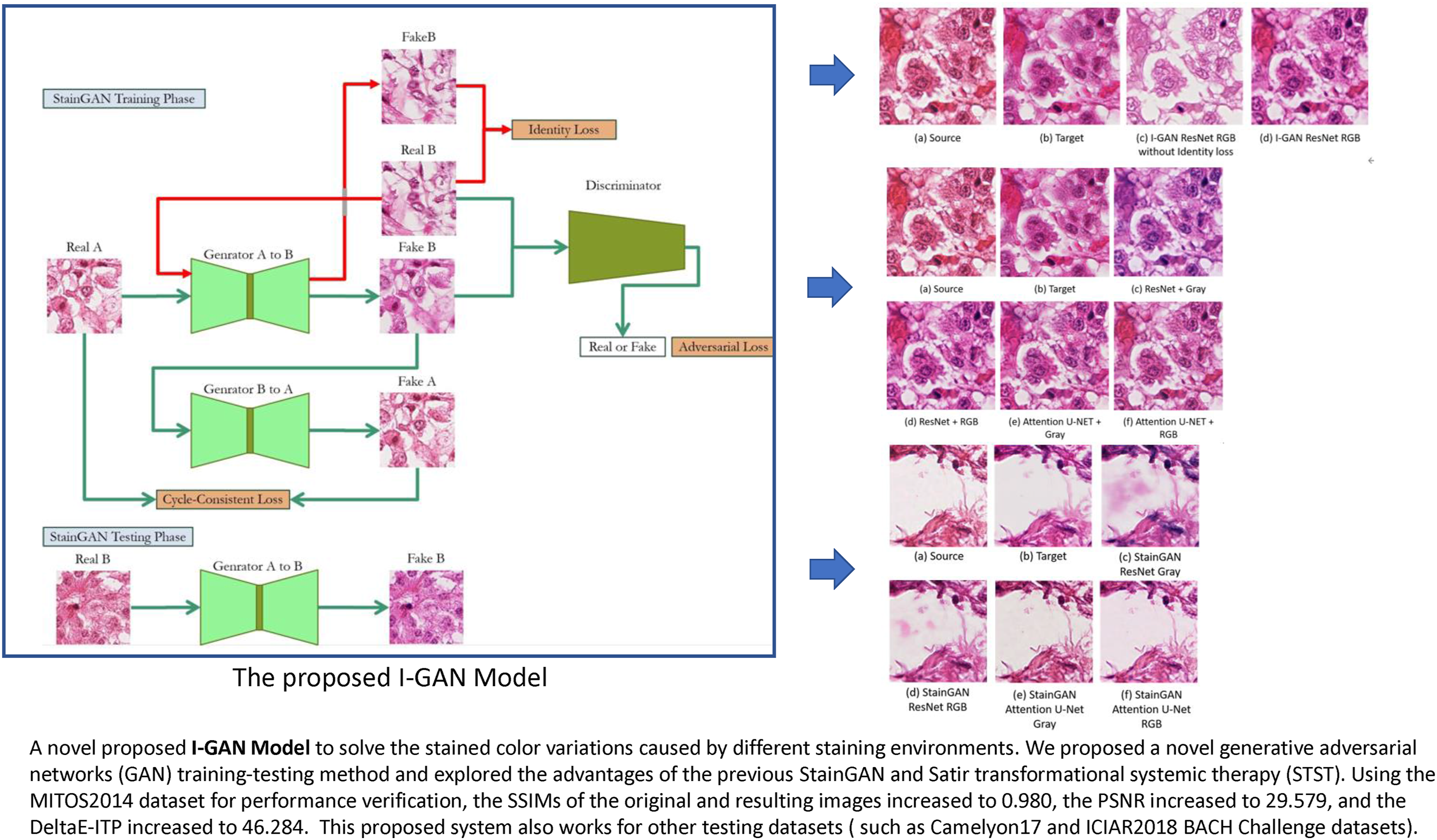
The architecture of the proposed model is shown in Figure 2. The training data of StainGAN was first changed to the STST mechanism. The MITOS-ATYPIA 14 Challenge Hamamatsu images and their grayscale images were sent to the model for training. However, the StainGAN test method was used for the testing procedures. The Aperio RGB images were input to Generator A and B, and automatically converted to Hamamatsu color space images after the model computation. Based on this training and testing mechanisms, this architecture is called the Identity Loss GAN (I-GAN) training-testing framework. I-GAN training-testing framework.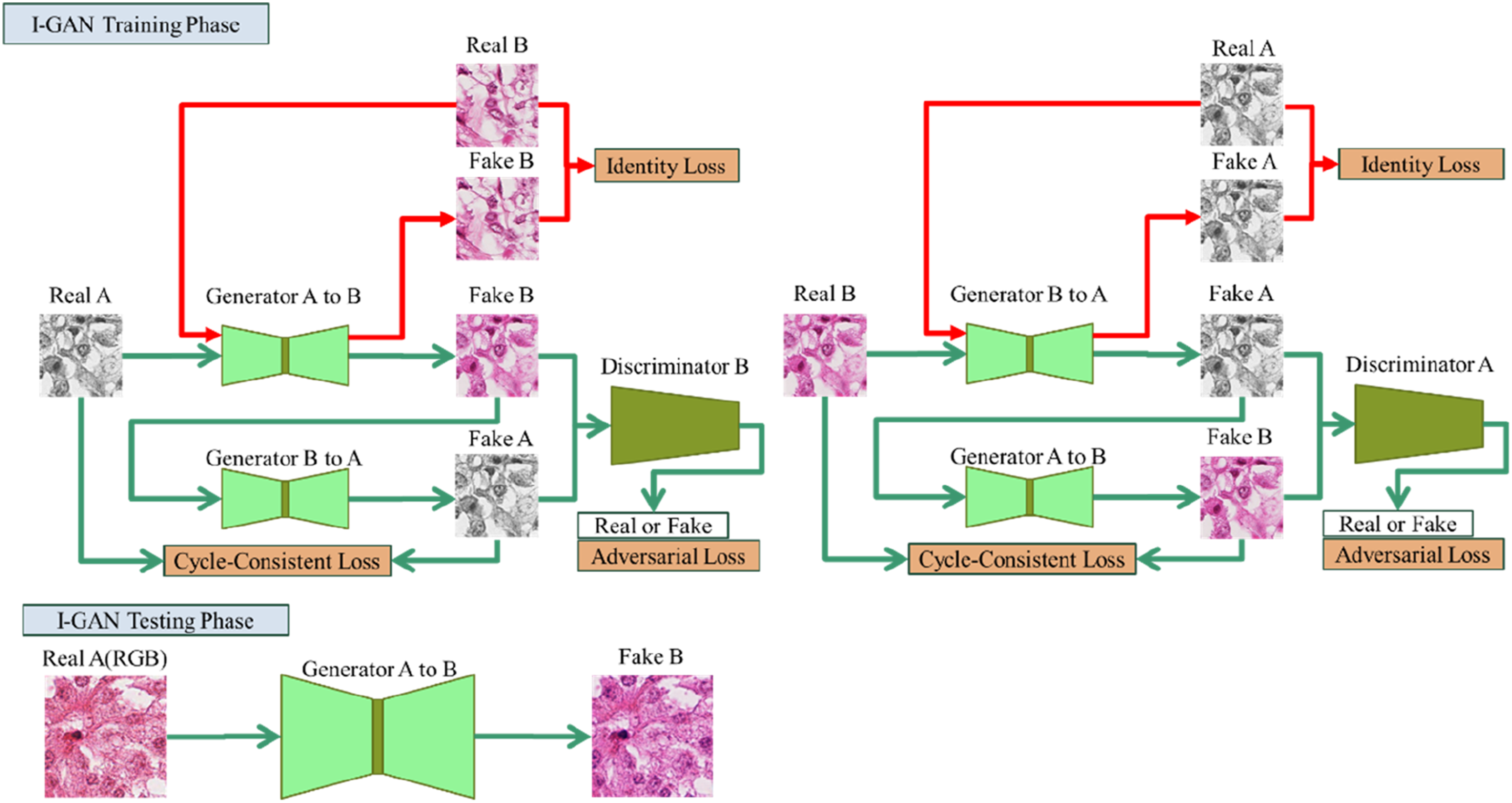
Because we input RGB images during the testing procedures, which is different from the original method of inputting grayscale images, the input images retained the original staining information of Hematoxylin and Eosin (e.g., nuclei vs. cytoplasm contrast). It is expected that the images converted by this system will provide a more suitable structure for distinguishing between the two different dyes, Hematoxylin and Eosin (H&E), so the color space conversion results are expected to be closer to the true H&E staining images.
To improve the performance of the proposed I-GAN model, we replaced the generator Resnet architecture with an attentional U-Net architecture. 23 The Attention U-Net GAN was successfully applied by Bai et al. 24 to convert autofluorescent staining images to immunohistochemistry (IHC). It is expected that this study can also improve the conversion results by effectively converting the staining images into grayscale images and H&E staining images, and also avoid checkerboard effects and GAN-generated artifacts after GAN conversion. The SSIM and PSNR evaluation metrics can be improved simultaneously.
The color space conversion system architecture proposed in this study uses three types of loss functions: Adversarial, Cycle-Consistency, and Identity Loss.
25
Adversarial loss allows the model to learn to convert original images to real images, and the learning processes can learn the distributions of real images. The Cycle-Consistency loss can keep the output results of the two generators consistently. After converting from domain A to domain B, the result remains consistent after converting to domain A again. The last type of Loss Function is Identity Loss, which can make the color style of the converted image consistent with the target image results. Among them, Identity Loss is of particular importance to our model architecture. We will further analyze it in Section 3.2.
For RGB images, all reconstruction-based loss terms were computed channel-wise. The L1 distances in the cycle-consistency and identity losses were calculated independently for each channel (R, G, and B), and then averaged across channels.
During training, grayscale images were converted into three-channel RGB format by replicating the single intensity channel across R, G, and B. Therefore, the same per-channel loss computation was consistently applied to both grayscale and RGB inputs.
In the validation phase using the first MITOS-ATYPIA 14 dataset, we designed six different validation methods for the model architecture and training method, respectively, to verify the actual impact of different model architectures and training-testing methods on the experimental results. For the Camelyon17 dataset, this study randomly selected two patients’ WSIs as the target color space for the training processes. In summary, the key contribution of this work is a novel use of Identity Loss in the RGB–Grayscale training–testing framework, which enables a single StainGAN-based model to learn the target stain style and generalize to unseen RGB stain domains.
2.3. Implementation details
For the stain normalization tasks, we set a learning rate of 0.0002 and a batch size of 4. The model was trained for 20 epochs, and the weights from the 20th epoch were used as the final stain normalization model for the best performance.
The number of training epochs (20 epochs) was determined empirically based on the convergence behavior observed during training processes. In our experiments, both generator and discriminator losses gradually stabilized within approximately 15–20 epochs, indicating that the adversarial training processes had reached a stable equilibrium.
For the classification tasks, we used ResNet50, 26 a widely adopted convolutional neural network backbone in histopathological image analysis, pre-trained on ImageNet, for binary classification on the Camelyon17 dataset and four-class classification on the ICIAR2018 BACH Challenge dataset. Using a fixed and well-established architecture allowed the effect of different stain normalization methods to be evaluated for a fair comparison, without introducing confounding factors from classifier design. We adopted the Adam optimizer with a learning rate of 0.0001 and a batch size of 8, together with a cosine annealing scheduler that decreased the learning rate from 0.0001 to 0 over 50 epochs.
3. Experiments and results
3.1. Evaluation metrics
The success of image color space conversion can be evaluated using three similarity metrics: the Structural Similarity index (SSIM), 27 Peak Signal-to-Noise-Ratio (PSNR), and DeltaE-ITP. 28 The value of Structural Similarity index (SSIM) is between [0, 1], and higher values represent a higher similarity of the images. The PSNR value represents the degree of image distortion, and higher values reflect less distortion after conversion. The calculation process of using DeltaE-ITP is to calculate the color differences between the corresponding pixels of the two images using a pixel-by-pixel evaluation method.
As for the color evaluation standards, this study adopts the specifications of the BT.2124-0 recommendation published by ITU-R in 2019. Including DeltaE-ITP, which converts the RGB color space into the LMS color space. The main purpose was to achieve a better adaptation for fitting the cone cells of human eyes. Therefore, in this study, the images converted to the color space were also compared pixel by pixel with the ground truth images, and the average values of the top 10%, 30%, and 50% of the difference values were calculated.
To statistically analyze the performance of the classification task after color space conversion, the Camelyon17 dataset was applied for illustration. The classification results were compared based on the basis of average precision (AP) and shown in Eq. (1).
29
For the color space conversion performance on the third ICIAR2018 BACH Challenge dataset, the precision, recall, and accuracy measurement were calculated as indicators for the different models and shown in Eq. (2).
3.2. Effects of identity loss
This section discusses the importance of Identity Loss for our model. In our Loss Function, Identity Loss allows the use of domain B images to generate another domain B image during training, so that the model has the ability to generate domain B style images. The training-testing method used in this study is equivalent to calculating Identity Loss once. By using Identity Loss to generate the characteristics of the domain B style, the model can also convert other RGB images into domain B style images. Figure 3(a) represents as the source image, Figure 3(b) shows as the target-domain image, and Figure 3(c) and (d) represent the results produced by the I-GAN model without and with identity loss, respectively. The model with identity loss generates more realistic H&E images and preserves better tissue structure compared with the model trained without identity loss. Effects of applying Identity Loss and their corresponding results: (a) a source image; (b) a target image; (c) the image generated by I-GAN ResNet without Identity Loss; (d) the image generated by I-GAN ResNet with Identity Loss.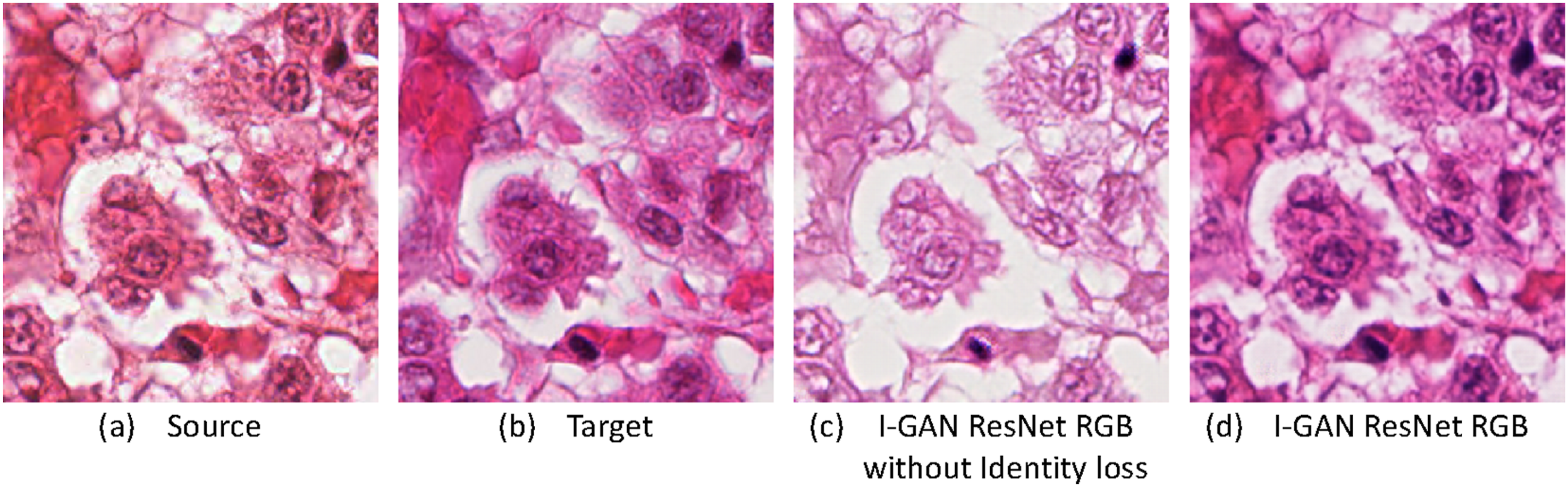
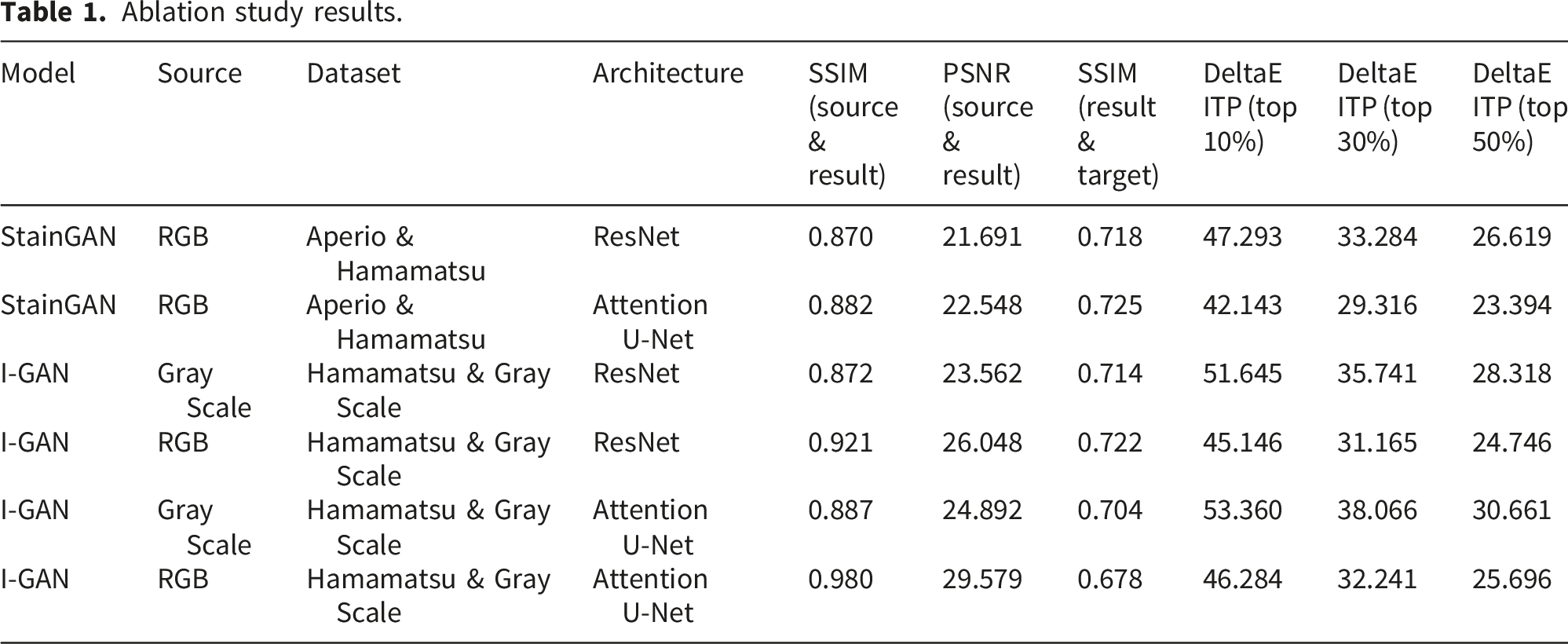
3.3. Ablation study
To discuss the effects of using different color space conversion models (StainGAN/I-GAN), using different input image formats (RGB/GRAY), or using different generators (Resnet/Attention U-Net), because the StainGAN color space conversion models were required two explicitly defined stain domains (A and B) with RGB images in both domains, so this study designed a total of six different test combinations to investigate the performance of different color space conversion technologies. The MITOS-ATYPIA 14 dataset and the corresponding grayscale images were used for the ablation study. By comparing the results of these six different combination experiments, it is possible to compare and verify the effects of different training methods and model architectures on the final performance of color space conversion.
Ablation study results.
In Figure 4, one representative example is used to compare the four I-GAN configurations listed in Table 1 and to illustrate the effects of the input type (grayscale vs. RGB) and generator backbone (ResNet vs. Attention U-Net) on H&E appearance. Figure 4(a) and (b) show the source and target images, respectively. When using a grayscale input with a ResNet generator (Figure 4(c)), the generated image shows weak H&E separation, resulting in low contrast between hematoxylin-stained nuclei and eosin-stained cytoplasm, and the overall appearance is less consistent with the target stain style. In contrast, switching the input to RGB while keeping the ResNet generator (Figure 4(d)) better preserves stain cues and makes eosin-rich cytoplasmic regions more distinguishable. Similarly, under the Attention U-Net generator, RGB input produces clearer nuclear–cytoplasm contrast than grayscale input (Figure 4(e) and (f)). Therefore, we recommend the I-GAN configuration with RGB input and an Attention U-Net generator as the best performance setting for stain normalization in our experiments. Comparison of the differences in the results of the model using RGB and grayscale: (a) source image; (b) target image; (c) converted image using ResNet architecture and grayscale images; (d) ResNet architecture and RGB image; (e) attention U-Net architecture and grayscale images; (f) attention U-Net architecture and RGB image.
3.4. GAN-generated artifacts
Our methods also demonstrate the artifacts generated by four different combination settings of different generators (Resnet/Attention U-Net) and different input images (RGB/GRAY). Figure 5(a) and (b) show the source and target images, respectively. It can be observed from Figure 5(c) and (d) that using RGB images as input can more effectively slow down the generation of artifacts than using grayscale images as input. However, if comparing Figure 5(c) and (e) (using grayscale images) with Figure 5(d) and (f) (using RGB images), it can be found that using the Attention U-Net generator can significantly reduce the generation of artifacts compared to using Resnet. Again, the proposed combination of I-GAN RGB images with the attention U-Net generator provided the best performance in terms of artifact generation during color space conversion. Comparison of artifacts generated by different models: (a) a source image; (b) a target image; (c) a converted image with ResNet + Gray; (d) a converted image with ResNet + RGB; (e) a converted image with attention U-Net + gray; (f) a converted image with attention U-Net + RGB.
3.5. MITOS-ATYPIA 14
Comparison of five stain normalization methods on MITOS-ATYPIA 14 challenge SSIM, PSNR, and DeltaE-ITP values.


An example of different models for MITOS-ATYPIA 14 dataset: (a) source image; (b) target image; (c) Vahadane; (d) STST; (e) StainGAN; (f) I-GAN.
3.6. Camelyon17
This experiment compared the average performance of five different techniques in binary classification using the Resnet model classifier after stain normalization using the Camelyon17 dataset. The evaluation and comparison were performed using the average precision (AP) metric. The analysis results and detailed performance are shown in supplemental Table A and B. All results shown in the tables were set to Center4 as the same target for stain normalization. As shown in supplemental Table A, the proposed I-GAN model achieved the highest average AP value of 0.962 among the five medical centers after stain normalization procedures, outperforming all other stain normalization techniques. An example of using the Camelyon17 dataset for comparison was shown in Figure 7 for comparison. An example of different models for Camelyon17 dataset: (a) source image; (b) target image; (c) Vahadane; (d) STST; (e) StainGAN; (f) I-GAN.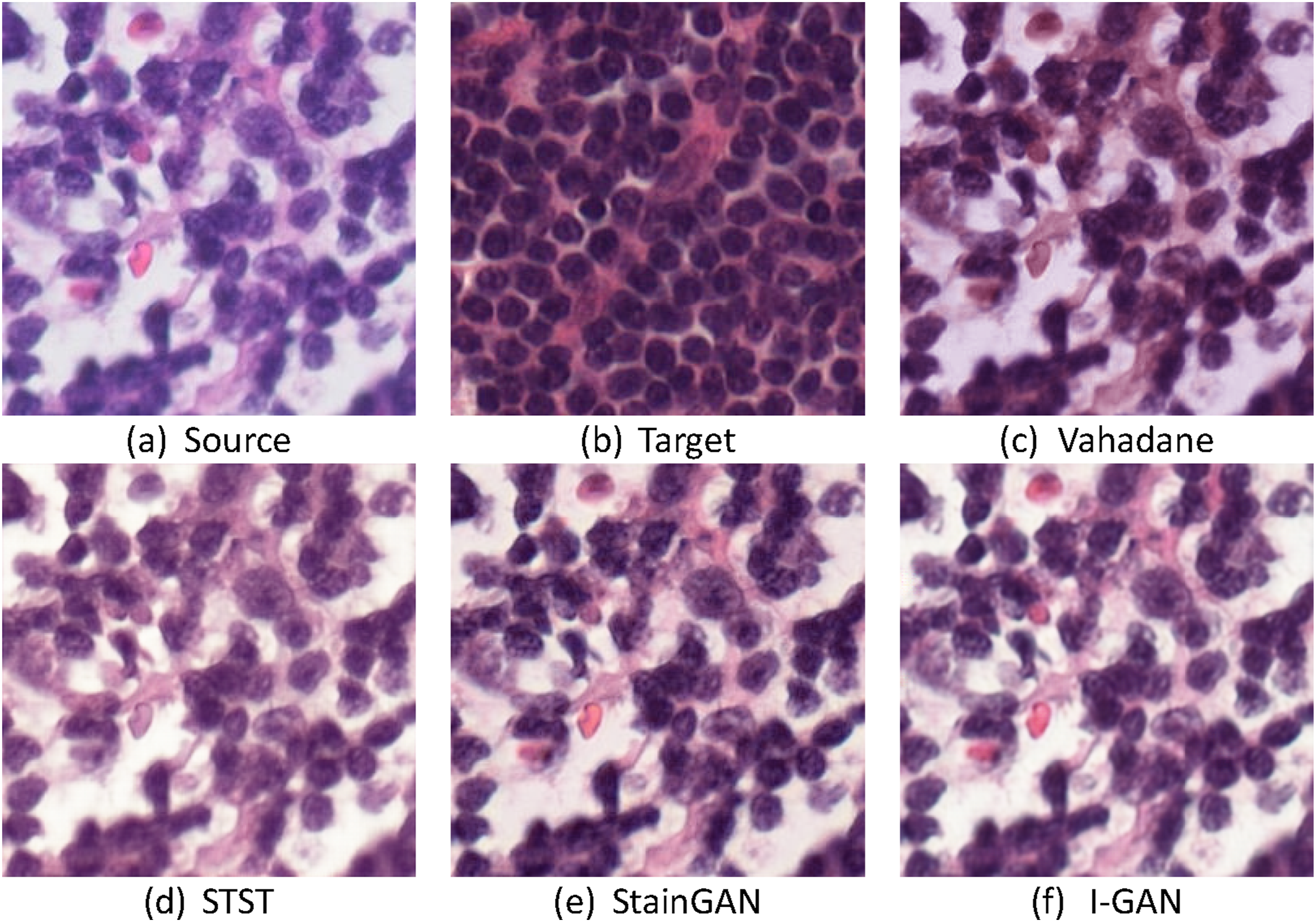
3.7. ICIAR2018 BACH challenge
In addition, the ICIAR2018 BACH Challenge dataset was used in this study to compare the prediction results for the classification of four categories with five different models (shown in Figure 8). The Stain normalization model was trained using the Camelyon17 dataset and its corresponding grayscale image dataset, whereas Vahadane randomly selected an image from the dataset as the target color space. The accuracy, precision, and recall values of the four classification categories were evaluated with five different color-space conversion techniques using the ResNet model classifier. The results was shown in supplemental Table C, which shows the prediction performance after staining normalization to Camelyon17’s Center4. The detailed data are provided in the supplemental Table D. It is clear from supplemental Table C that the proposed I-GAN model significantly outperforms other models in terms of an accuracy of 0.87, a recall of 0.86, and a precision of 0.87 after training and testing. An example of different models for the ICIAR2018 BACH challenge dataset: (a) source image; (b) target image; (c) Vahadane; (d) STST; (e) StainGAN; (f) I-GAN.
4. Discussion
This study proposes a novel I-GAN training and testing framework for stain normalization in digital histopathological images. Unlike conventional StainGAN-based approaches that require explicitly defined source–target stain pairs, the proposed method integrates RGB–grayscale training with identity loss, enabling the model to learn a stable target stain representation while preserving structural information during testing on RGB images.
The experimental results demonstrate that the proposed I-GAN consistently outperforms conventional stain normalization techniques across multiple evaluation metrics, including SSIM, PSNR, and DeltaE-ITP, on the MITOS-ATYPIA 14 dataset. In particular, the higher SSIM values indicate superior preservation of tissue morphology, which is critical for histopathological analysis where nuclear shape and spatial organization play a central diagnostic role. At the same time, the improved PSNR reflects reduced image distortion after stain normalization, while the lower DeltaE-ITP values confirm that the generated images are perceptually closer to the target stain domain.
The ablation study further provides insights into the mechanisms underlying these improvements. When RGB images are used during the testing phase, the model retains stain-specific chromatic cues that are lost in grayscale-only inputs, leading to better separation between hematoxylin-stained nuclei and eosin-stained cytoplasm. This explains why the proposed method achieves high SSIM values without sacrificing color fidelity. Moreover, the incorporation of identity loss encourages the generator to behave as an identity mapping for images already belonging to the target stain domain, effectively learning a fixed-point representation of the target stain style. This mechanism enables the I-GAN to generalize to unseen RGB stain domains without requiring multiple source–target domain pairs.
In addition, replacing the ResNet generator with an Attention U-Net backbone further enhances stain normalization quality. The attention mechanism allows the generator to focus on diagnostically relevant regions, such as nuclei-dense areas, while suppressing background variations and GAN-induced artifacts. This architectural choice results in a favorable balance between structural preservation, low distortion, and accurate color reproduction, making the proposed I-GAN particularly suitable for downstream histopathological image analysis.
The advantages of the proposed stain normalization framework are further reflected in downstream classification tasks. On the Camelyon17 dataset, the I-GAN achieves the highest average precision in binary classification across multiple centers, indicating improved robustness against inter-center and inter-scanner variability. Similarly, on the ICIAR2018 BACH Challenge dataset, the proposed method yields superior accuracy, precision, and recall in four-class classification. These results suggest that preserving structural integrity during stain normalization plays a critical role in improving the generalization performance of deep learning–based classifiers.
5. Limitations and future work
Despite the promising results, this study has several limitations that warrant further investigation. First, the experimental evaluation was conducted on a limited set of publicly available H&E-stained breast cancer datasets, including MITOS-ATYPIA 14, Camelyon17, and ICIAR2018 BACH. Although these datasets covered multiple scanners and centers, the generalizability of the proposed I-GAN framework to other organs, disease types, and staining protocols has not yet been systematically evaluated.
Second, this study focused primarily on single-target stain normalization. While the proposed identity loss–based framework showed robustness in handling multiple unseen stain domains, extending the method to explicitly support multi-target or fully domain-agnostic stain normalization remains an important direction for future work. In addition, the current experiments are limited to hematoxylin and eosin (H&E) staining. The applicability of the proposed approach to immunohistochemistry (IHC) staining protocols and other non-H&E modalities requires further validation.
Future work will therefore focus on expanding the evaluation of the I-GAN framework to larger and more diverse datasets, as well as exploring its extension to multi-target and domain-agnostic stain normalization settings. Investigating the integration of the proposed method with task-aware or end-to-end training strategies for downstream clinical applications is also a promising direction.
6. Conclusion
In this study, we proposed a novel I-GAN training and testing framework for stain normalization of digital histopathological images. By combining RGB-based testing with identity loss, the proposed method effectively preserved structural information while achieving accurate color-domain conversion using a single target stain domain. Comprehensive experiments demonstrated that the proposed approach outperformed conventional stain normalization techniques in terms of SSIM, PSNR, DeltaE-ITP, and downstream classification performance.
The results confirm that the proposed I-GAN framework provides a stable and effective solution for stain normalization in multi-center and multi-scanner settings. For applications requiring GAN-based stain normalization in digital pathology, the proposed training and testing architecture offers a robust and practical choice, with strong potential for improving the reliability of deep learning–based histopathological image analysis.
Supplemental material
Supplemental Material - Improving stain normalization for digital histological image analysis based on the cycle generative adversarial network identity loss model
Supplemental Material for Improving stain normalization for digital histological image analysis based on the cycle generative adversarial network identity loss model by Jung-Ting Chen, Yen-Yin Lin and Tun-Wen Pai in Digital Health.
Footnotes
Author contributions
Conceptualization, T.-W.P.; methodology, J.-T. Chen and T.-W.P.; software, J.-T. C.; validation, J.-T. C., Y.-Y. L., and T.-W.P.; formal analysis, J.-T. Chen and T.-W.P.; investigation, J.-T. Chen and T.-W.P.; resources, Y.-Y. L.; data curation, J.-T. C., Y.-Y. L., and T.-W.P.; writing—original draft preparation, J.-T. C.; writing—review and editing, Y.-Y. L., and T.-W.P.; visualization, J.-T. C.; supervision, Y.-Y. L., and T.-W.P.; project administration, Y.-Y. L., and T.-W.P.; funding acquisition, Y.-Y. L., and T.-W.P. All authors have read and agreed to the published version of the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: We thank the funding supports from National Science and Technology Council, Taiwan (NSTC 113-2221-E-027-109 and NTSC 114-2221-E-027-083) and technical guiding from JelloX Biotech Inc., Taiwan.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The datasets generated during and/or analyzed during the current study are available in the following repositories: MITOS-ATYPIA 14 Challenge (https://mitos-atypia-14.grand-challenge.org/). Camelyon17 Challenge (https://camelyon17.grand-challenge.org/). ICIAR2018 BACH Challenge (![]() ).
).
Supplemental material
Supplemental material for this article is available online.