Abstract
Objectives
The primary goal is to address the challenges in brain tumor segmentation (BraTS), such as limited accuracy and high computational costs, by developing a more precise and efficient segmentation technique. The study aims to improve the diagnosis and treatment planning of brain tumors by enabling clinicians to accurately localize and assess tumor regions from multimodal magnetic resonance imaging (MRI) scans.
Methods
We proposed self-attention U-Net (SAU-Net), a novel model that integrated self-attention mechanisms with the U-Net convolutional architecture. This design allowed the model to preserve spatial context while selectively concentrating on pertinent features, thereby enhancing tumor (ET) boundary delineation and overall segmentation accuracy. Extensive experiments were conducted on the BraTS 2018 and BraTS 2020 datasets using thorough cross-validation and testing protocols. The performance of SAU-Net was evaluated and compared against other attention-based U-Net models, including adaptive attention U-Net, multi-head attention U-Net, and group query attention U-Net.
Results
On the BraTS 2018 dataset, SAU-Net achieved Dice scores of 98.16% (whole tumor (WT)), 98.87% (tumor core (TC)), and 98.23% (ET), with an average Dice score of 98.23%. For the BraTS 2020 dataset, the model recorded Dice scores of 98.99% (WT), 98.70% (TC), and 99.18% (ET), with an average Dice score of 98.62%. In addition to superior segmentation performance, the model demonstrated reduced computational complexity in both training and prediction times, along with optimized memory usage.
Conclusion
SAU-Net is a highly effective and computationally efficient model for BraTS. Its superior performance, as evidenced by the high Dice scores on two benchmark datasets, combined with its reduced computational requirements, underscores its potential for practical and impactful clinical applications.
Keywords
Introduction
Brain tumor segmentation (BraTS) is crucial in medical image analysis, as it provides vital information for the identification, monitoring, and treatment of brain tumors. 1 Accurately delineating tumor regions in medical images, particularly in neuroimaging, is essential for effective diagnosis, treatment planning, and patient monitoring. 2 The task involves segmenting brain tumors in images such as magnetic resonance imaging (MRI) and multimodal MRI scans, enabling clinicians to pinpoint regions of interest and make informed decisions on intervention strategies. 3 Recent advances in deep learning (DL) have shown strong potential for extracting clinically relevant biomarkers and gene-level signatures from medical imaging data, further highlighting the importance of accurate automated segmentation in oncology and neuro-oncology.4,5 Brain tumors can grow undetected to substantial sizes and exhibit varying shapes and sizes, making segmentation a challenging yet vital task. 6 Tumors are clusters of abnormal cells that proliferate within the brain and, if left untreated, can be life-threatening. These tumors are categorized into malignant (cancerous) and benign (non-cancerous) forms, with gliomas being the most common type of malignant brain tumor among adults. 7 Gliomas are further classified into low-grade gliomas, which are slow-growing, and high-grade gliomas, which are aggressive and fast-growing. 8 The segmentation of glioma regions often involves identifying three key subregions: the tumor core (TC), the enhancing core (EC), and the whole tumor (WT). 9 Gliomas account for a significant proportion of brain cancer cases diagnosed worldwide. 9 Approximately 17,200 people die each year from malignant brain tumors, while more than 90,000 patients receive an initial brain tumor diagnosis. Epidemiological analyses indicate that 25 individuals out of every 100,000 are affected by brain tumors, with approximately 33% classified as malignant. 10
Medical imaging, particularly in neurology, places strong emphasis on brain segmentation due to its critical role in diagnosing and managing neurological disorders, including brain tumors. 11 Accurate segmentation supports diagnostic and therapeutic strategies, surgical planning, disease progression tracking, and personalized treatment development. 12 However, manual segmentation remains labor-intensive, time-consuming, and prone to human error, limiting its scalability and precision, especially for large datasets. 13 Traditional segmentation tools, while useful, often fail to capture complex tumor boundary characteristics, resulting in inconsistent outcomes across patients and tumor types.
In contrast, DL-based approaches, particularly those based on the U-Net architecture, have transformed BraTS by automating segmentation and achieving high accuracy through learning from large-scale datasets. 14 U-Net and its variants utilize convolutional neural networks (CNNs) to extract complex patterns from MRI images, significantly improving tumor boundary delineation accuracy. 15 Prior studies demonstrated that architectures such as Deep U-Net and diffusion-based models outperformed manual techniques and enhanced clinical decision-making.16,17 Despite these advances, existing methods continued to face limitations. Tabassum et al. 18 showed that advanced meta-transfer learning strategies improved U-Net performance for tumor segmentation, particularly for underrepresented tumor types such as meningioma and metastasis. Saeed et al. 19 proposed RMU-Net, which achieved high accuracy on the BraTS 2018 dataset but struggled with heterogeneous tumor subregions. Similarly, Ali et al. 20 introduced an ensemble of U-Net and three-dimensional (3D) CNN models, although the approach was constrained by high-computational complexity, limiting clinical applicability. Tataei et al. 21 employed CNN engineering with ResNet-50, yet the fixed convolutional layers did not fully capture intricate tumor boundary variations. Moreover, commonly used evaluation metrics such as the Dice similarity coefficient (DSC) were not always sufficient to capture segmentation quality comprehensively, motivating the need for more robust assessment methodologies.
Recent developments in DL addressed these challenges through the incorporation of attention mechanisms into segmentation models. Attention-based architectures enabled networks to selectively focus on salient image features, improving segmentation accuracy by prioritizing relevant spatial information. These mechanisms also facilitated improved detection of complex tumor structures while reducing computational overhead, supporting practical clinical deployment.
While previous studies have provided extensive background on brain tumor segmentation, the key novelty of this study lies in introducing a streamlined self-attention mechanism integrated directly into the U-Net architecture. Unlike existing attention-based U-Net models, self-attention U-Net (SAU-Net) was designed to enhance feature selectivity while minimizing computational complexity, thereby enabling more accurate delineation of heterogeneous tumor boundaries.
To overcome the limitations of existing segmentation techniques, this study proposed SAU-Net, a novel architecture that incorporated self-attention mechanisms into the U-Net framework. By selectively emphasizing diagnostically relevant tumor features while preserving spatial coherence, SAU-Net achieved more precise segmentation of diverse brain tumor subregions. Experimental evaluations on the BraTS 2018 and BraTS 2020 datasets demonstrated that the proposed model outperformed state-of-the-art (SOA) methods in terms of DSC, sensitivity, and specificity. Furthermore, SAU-Net exhibited reduced memory consumption and computational complexity, making it well-suited for practical clinical applications.
Contributions
This paper makes the following significant contributions:
Research questions
What architectural advancements in SAU-Net enable higher segmentation accuracy for brain tumors compared to traditional U-Net and modern DL techniques? How does SAU-Net’s performance in segmenting WT, TC, and ET from MRI data compare with SOA models on heterogeneous tumor datasets? How does the integration of self-attention mechanisms contribute to reduced computational complexity while maintaining high segmentation accuracy in clinical applications?
Hypothesis
This study hypothesized that the proposed SAU-Net would outperform existing SOA BraTS techniques, including conventional U-Net models. It was anticipated that integrating self-attention mechanisms into a convolutional U-Net framework would enhance the discrimination of complex tumor subregions, such as the ET, TC, and WT. The model was expected to achieve improved performance across key evaluation metrics, including DSC, sensitivity, and specificity. Additionally, the attention mechanism was hypothesized to reduce computational complexity, thereby enabling efficient and accurate clinical deployment.
This paper is organized as follows: Section “Related works” reviews the relevant literature. Section “Methodology” details the proposed methodology. Section “Performance analysis” presents and discusses the experimental results along with the performance analysis. Finally, Section “Discussion” compares the proposed approach with existing works, and Section “Conclusion” summarizes the study and outlines the key findings.
Related works
The identification and treatment planning of brain tumors depend heavily on accurate segmentation using multimodal MRI data. Numerous DL frameworks have been developed to address this challenging problem by leveraging the rich information contained in MRI images. This section reviews recent advancements in BraTS, with particular emphasis on studies conducted using the BraTS 2018 and BraTS 2020 datasets, highlighting key methodologies and performance outcomes.
Related works on BraTS 2018
Saeed et al. 19 proposed a hybrid model, RMU-Net, which combined MobileNetV2 and U-Net architectures for end-to-end brain tumor segmentation. Using the BraTS 2018 dataset, the model achieved Dice coefficients of 90.80, 86.75, and 79.36 for different tumor subregions. Ali et al. 20 presented an ensemble approach integrating U-Net and 3D CNN models, which yielded Dice scores of 0.750, 0.906, and 0.846 for the ET, WT, and TC, respectively, on the BraTS 2018 test set. Tataei et al. 21 employed CNN-based feature engineering with a ResNet-50 backbone and obtained competitive Dice scores across multiple tumor regions using the BraTS 2018 dataset. Gull et al. 22 developed a CNN-based framework for both tumor segmentation and classification, achieving average accuracies of 96.50% for segmentation and 96.49% for classification.
Ullah et al. 23 demonstrated the effectiveness of a 3D U-Net model for automatic tumor segmentation and MRI image enhancement. Their method produced mean Dice scores of 0.70 for the ET, 0.86 for the TC, and 0.91 for the WT, while testing yielded Dice scores of 0.83 for the ET, 0.90 for the TC, and 0.71 for the WT. MBANet, a multi-branch attention-based 3D CNN, was also evaluated on the BraTS 2018 dataset and achieved Dice scores of 78.21, 89.79, and 83.04 for the ET, WT, and TC, respectively.
Zia et al. 24 proposed a context-aware attentional residual dropout U-Net (ARDUNet), which achieved Dice scores of approximately 0.90 across tumor subregions on the BraTS 2018 dataset, including 0.92 for the ET. Sun et al. 25 developed a three-dimensional fully convolutional network that obtained DSCs of 0.90, 0.79, and 0.77 for the WT, TC, and ET segmentation, respectively. Pedada et al. 9 introduced a residual U-Net architecture and reported a segmentation accuracy of 92.20% on the BraTS 2018 challenge dataset, with enhanced classification performance for the TC, EC, and WT regions.
Related works on BraTS 2020
Isensee et al. 26 utilized the nnU-Net framework for the BraTS 2020 challenge and demonstrated that the baseline configuration achieved strong segmentation performance without manual architectural tuning. Through extensive pipeline optimization, including aggressive data augmentation, region-based training, and post-processing strategies, their approach won the BraTS 2020 challenge, achieving Dice scores of 82.03% for the ET, 85.06% for the TC, and 88.95% for the WT.
Kataria et al. 27 introduced HybriCSF, a hybrid segmentation framework combining CNNs, support vector machines, and fuzzy C-means clustering. Evaluation on the BraTS 2020 dataset showed improved segmentation performance, with Dice scores of 0.63 for the ET, 0.87 for the WT, and 0.81 for the TC. Magadza et al. 28 extended the nnU-Net architecture by replacing standard convolutional blocks with depthwise-separable bottleneck units and incorporating shuffle attention in skip connections. Their optimized network achieved Dice scores of 84.8% for the TC, 91.2% for the WT, and 79.2% for the ET.
Susanto et al. 29 proposed a spatial transformation-based data augmentation pipeline that improved segmentation accuracy on the BraTS 2020 dataset, yielding Dice scores of 87.10 for the ET, 86.89 for the TC, and 90.91 for the WT. Zhang et al. 30 developed a multi-encoder 3D MRI segmentation framework and introduced a categorical Dice loss function to address voxel imbalance. Their method achieved Dice scores of 70.24% for the WT, 88.26% for the TC, and 73.86% for the ET in the BraTS 2020 challenge.
Angona et al. 31 proposed a hybrid 3D ResAttU-Net-Swin model integrating residual learning, attention-based skip connections, and a Swin Transformer. When evaluated on the BraTS 2020 dataset, the model achieved Dice scores of 92.50% for the WT, 89.70% for the TC, and 82.60% for the ET, resulting in an average DSC of 88.27%. Li et al. 32 introduced DPF-Unet, a dual-path 3D segmentation framework that combined CNN-based local feature extraction with transformer-based global modeling. Their approach achieved Dice scores of 91.74% for the WT, 89.44% for the TC, and 84.88% for the ET on the BraTS 2020 dataset.
Overall, these studies demonstrated the steady progress of DL-based methods, particularly U-Net-derived architectures, in improving brain tumor segmentation accuracy and robustness across benchmark datasets.
Methodology
During this investigation, we developed a new SAU-Net for BraTS using MRI images. The proposed methodology followed a clear and sequential process. First, the MRI image data were collected and preprocessed. Next, the SAU-Net architecture was constructed as a modified U-Net incorporating self-attention mechanisms. Finally, the model was trained on the prepared datasets and subsequently used to predict and segment brain tumors in unseen MRI images. The detailed architecture of the proposed SAU-Net is illustrated in Figure 1.

Overview of the proposed SAU-Net architecture for brain tumor segmentation using MRI images. The pipeline consists of three main components: (1) Image preprocessing, including image resizing, wavelet denoising, volume slicing, image scaling, and one-hot encoding; (2) the SAU-Net encoder–decoder architecture, where Blocks 1–4 form the encoder, Block 5 integrates a lightweight self-attention mechanism at the bottleneck, and Blocks 6–9 form the decoder with skip connections; and (3) the segmentation output for WT, TC, and ET. SAU-Net: self-attention U-Net; MRI: magnetic resonance imaging; WT: whole tumor; TC: tumor core; ET: enhancing tumor.
Brain multimodal MRI data collection
The BraTS 2018 and 2020 challenge datasets served as the primary benchmark datasets used in this study. All attention-based models were developed, trained, and evaluated using these datasets.
The BraTS 2018 dataset 33 is a comprehensive resource for segmenting both high-grade and low-grade gliomas. It includes multi-institutional MRI scans with expert-annotated tumor regions. Each patient scan contained four MRI modalities: T1-weighted (T1), T1-weighted with contrast enhancement (T1CE), T2-weighted (T2), and fluid-attenuated inversion recovery (FLAIR). These modalities enabled the identification of the ET, TC, and WT, including edema. The dataset comprised 285 samples, distributed into 205 training cases, 43 validation cases, and 37 testing cases. All images were preprocessed to maintain a consistent spatial resolution, and skull stripping was applied to isolate brain tissue.
The BraTS 2020 dataset 34 extended previous releases by providing a larger and more diverse collection of MRI scans. It contained 368 samples, partitioned into 265 training cases, 65 validation cases, and 47 testing cases. Similar to BraTS 2018, each subject included four MRI modalities (T1, T1CE, T2, and FLAIR), enabling accurate segmentation of glioma subregions. BraTS 2020 further incorporated improved annotation quality and a broader range of clinical cases while preserving its focus on glioma subregion segmentation.
Table 1 summarizes the training, validation, and testing splits for both datasets. Representative sample images from the two datasets, illustrating the different imaging modalities used for BraTS, are presented in Figure 2. Specifically, Figure 2(a) shows samples from BraTS 2018, whereas Figure 2(b) shows samples from BraTS 2020.
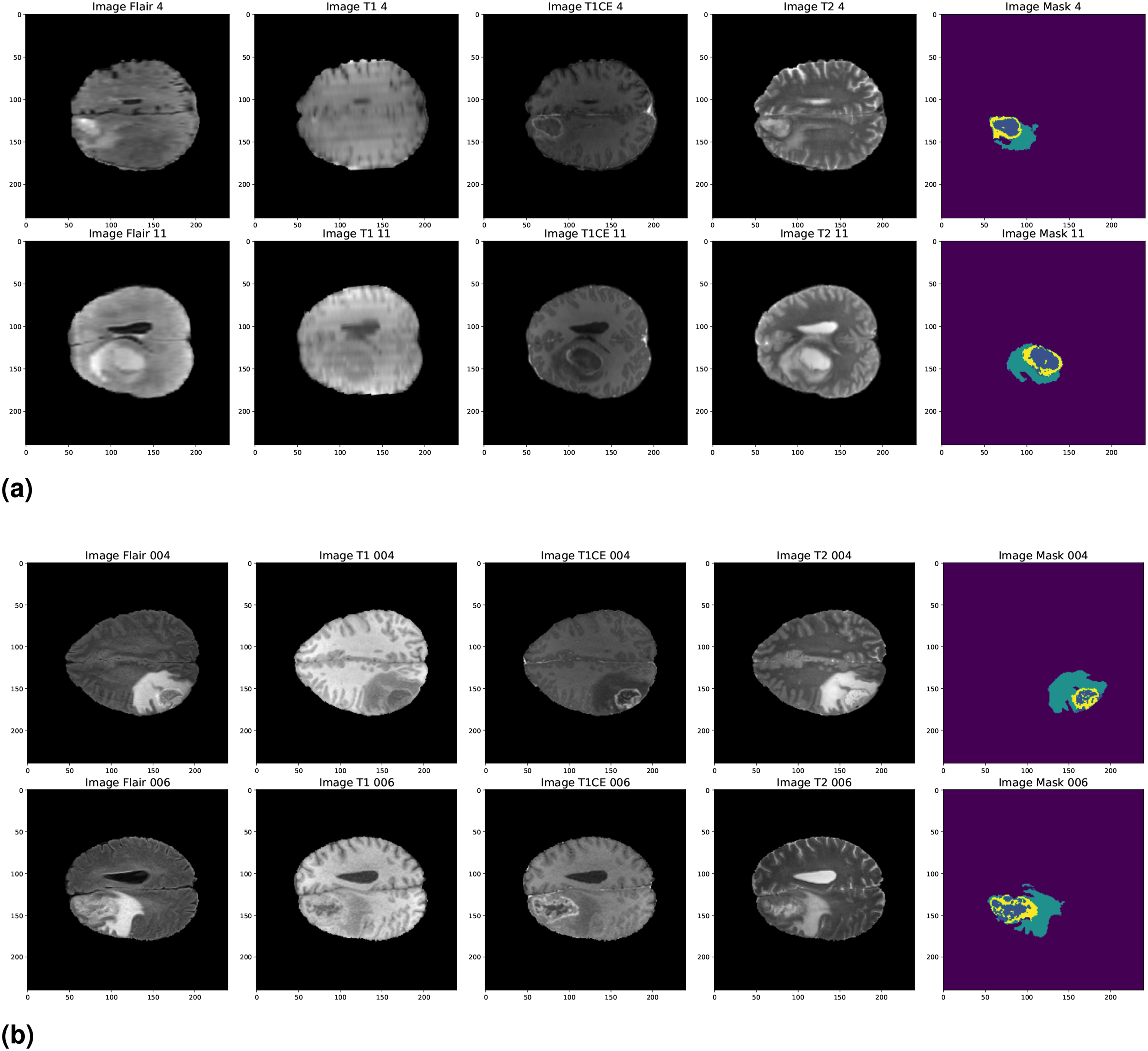
Brain tumor segmentation (BraTS) 2018 and BraTS 2020 dataset sample images. (a) BraTS 2018 and (b) BraTS 2020.
Dataset distribution for brain tumor segmentation (BraTS) 2018 and BraTS 2020 datasets.

Preprocessing
A comprehensive preprocessing workflow was employed to maximize data quality and model performance in preparing the BraTS MRI images for tumor segmentation. This systematic procedure improved image uniformity and enhanced segmentation accuracy.
The preprocessing pipeline consisted of the following key steps:
By implementing this preprocessing pipeline, a consistent and reliable dataset was established for brain tumor segmentation, thereby contributing to improved robustness and accuracy of the segmentation outcomes.
Volume slicing
To prepare the three-dimensional MRI volumes for analysis, an adaptive slicing strategy was employed to extract representative cross-sectional slices. Unlike conventional methods that relied on a fixed slice range, this approach dynamically and uniformly selected a predefined number of slices from each volume, irrespective of its depth. This ensured comprehensive coverage of anatomical structures and enabled the segmentation model to consistently capture salient spatial patterns throughout the brain.
The adaptive slicing strategy also eliminated the need for computationally intensive experimentation with multiple slice resolutions and orientations, which often varied across datasets. By ensuring that critical anatomical regions were consistently included, the method enhanced segmentation performance while substantially reducing data preparation time and computational effort.
Furthermore, the adaptive approach prioritized slices intersecting regions with higher tumor probability, increasing the likelihood that each selected slice contained clinically meaningful pathological information. Consequently, the segmentation model was consistently exposed to tumor-related features across varying brain depths, improving reproducibility and reducing the risk of omitting critical tumor regions due to arbitrary slice selection.
Attention-based U-Net architecture
An attention-based U-Net architecture was employed in this study to efficiently identify and segment tumors in MRI images. This design incorporated several essential components, each contributing to improved segmentation performance. The proposed model was informed by a range of attention-based U-Net variants, including adaptive attention U-Net (ADAU-Net), SAU-Net, multi-head attention U-Net (MHAU-Net), and group query attention U-Net (GQAU-Net), each developed to address specific challenges in medical image segmentation. The key mechanisms underlying these attention models are described below:
In the proposed architecture, a self-attention module was incorporated to enhance feature representation and capture long-range spatial correlations within the input data. This design choice was motivated by the need to improve discriminative capability by selectively emphasizing diagnostically relevant features while suppressing noise and irrelevant anatomical structures. Unlike conventional attention mechanisms that relied on multi-head formulations and introduced additional computational complexity, the implemented self-attention module employed three linear transformations—query, key, and value—to compute attention scores directly within the feature space. This formulation enabled efficient computation of attention weights and dynamic feature reweighting without requiring multiple attention heads.
Furthermore, the proposed approach preserved the spatial integrity of feature maps, allowing the attention mechanism to adaptively enhance segmentation performance by focusing on the most informative image regions. Empirical evaluation demonstrated improved identification of critical tumor areas, particularly in challenging brain tumor segmentation scenarios. Overall, integrating self-attention within the U-Net framework facilitated a more nuanced understanding of spatial dependencies in the data, which was essential for achieving precise and reliable tumor segmentation.
Mechanisms of attention enhancement
The proposed SAU-Net architecture incorporated a streamlined self-attention mechanism designed to enhance feature representation and emphasize diagnostically relevant spatial regions in the input data. To clarify the integration of this module within the U-Net framework, a concise description is provided below, followed by Algorithm 1, which outlines the forward pass of SAU-Net and illustrates how the attention block operates at the bottleneck stage before being propagated through the decoder via skip connections.
The self-attention module computed attention scores by applying linear transformations to the input feature maps to generate query (
The computed attention scores facilitated long-range dependency modeling and contextual feature aggregation. When reintegrated into the U-Net architecture, the attention-weighted features enhanced the decoder’s capacity to reconstruct accurate segmentation maps. Unlike multi-head attention mechanisms, which often introduce additional architectural and computational complexity, SAU-Net employed a lightweight single-head self-attention formulation. This design preserved computational efficiency while maintaining strong representational capability, making the model well-suited for medical imaging applications such as brain tumor segmentation.
Overall, the incorporation of the self-attention mechanism significantly contributed to the segmentation performance of SAU-Net by enabling the network to adaptively focus on the most informative regions of the input data, resulting in improved accuracy and robustness.
Forward pass of SAU-Net (Blocks 1--9) with integrated self-attention.

Brain tumor segmentation (BraTS)
In this study, SAU-Net was employed to segment MRI brain images into three clinically significant tumor subregions. This segmentation strategy provided a comprehensive characterization of tumor pathology and morphology, which was essential for accurate clinical analysis and treatment planning. The three segmented subregions included the WT, the TC, and the ET.
The WT encompassed the entire tumor region, including surrounding edema, thereby offering a complete representation of overall tumor burden. The TC represented the central, actively growing component of the tumor and was critical for assessing tumor aggressiveness. The ET highlighted regions exhibiting contrast enhancement, which were essential for distinguishing viable tumor tissue from non-tumorous regions.
By providing a detailed delineation of these tumor subregions, the proposed segmentation approach enabled more precise treatment planning and facilitated effective monitoring of tumor progression and therapeutic response.
Results analysis
In this study, we employed a range of attention-based U-Net architectures to address the task of brain tumor segmentation. The proposed SAU-Net was rigorously evaluated on the BraTS 2018 and BraTS 2020 datasets, and its performance was systematically compared with existing approaches, demonstrating notable advantages in segmentation accuracy and robustness.
Evaluation setup
The experimental setup was designed to ensure robust and reproducible evaluation. All experiments were conducted in a high-performance computing environment equipped with an NVIDIA Tesla P100 GPU (30 GB memory), 30 GB of system RAM, and 70 GB of dedicated disk space. For model implementation and analysis, widely adopted frameworks and libraries were utilized, including TensorFlow, Keras, Matplotlib, Pandas, and NumPy. The key performance metrics used to assess the segmentation capability of attention U-Net variants are summarized in Table 2.
Performance metrics and formulas.

IoU: intersection of union; TP: true positive; TN: true negative; FN: false negative; FP: false positive.
Performance analysis
The performance of the proposed SAU-Net model was evaluated on the BraTS 2018 and BraTS 2020 datasets through a comprehensive segmentation analysis. These benchmark datasets enabled a systematic assessment of the model’s ability to distinguish different tumor subregions under diverse imaging conditions. Segmentation performance was quantified using standard evaluation metrics, including the Dice coefficient, Jaccard index, sensitivity, and specificity.
Table 3 shows that SAU-Net consistently outperformed the comparative models across all tumor classes. On the BraTS 2018 dataset, SAU-Net achieved a Dice coefficient of 98.35% for the ET, exceeding the performance of ADAU-Net (98.30%) and MHAU-Net (98.23%). The model also demonstrated superior accuracy for the WT and TC categories, with Dice coefficients of 98.16% and 98.87%, respectively. This improved performance was attributed to the effective integration of self-attention mechanisms, which enabled the model to emphasize diagnostically relevant features within MRI images. Such feature selectivity was particularly important in medical imaging, where subtle variations in tumor characteristics can substantially influence segmentation accuracy.
Performance analysis of BraTS on MRI image.
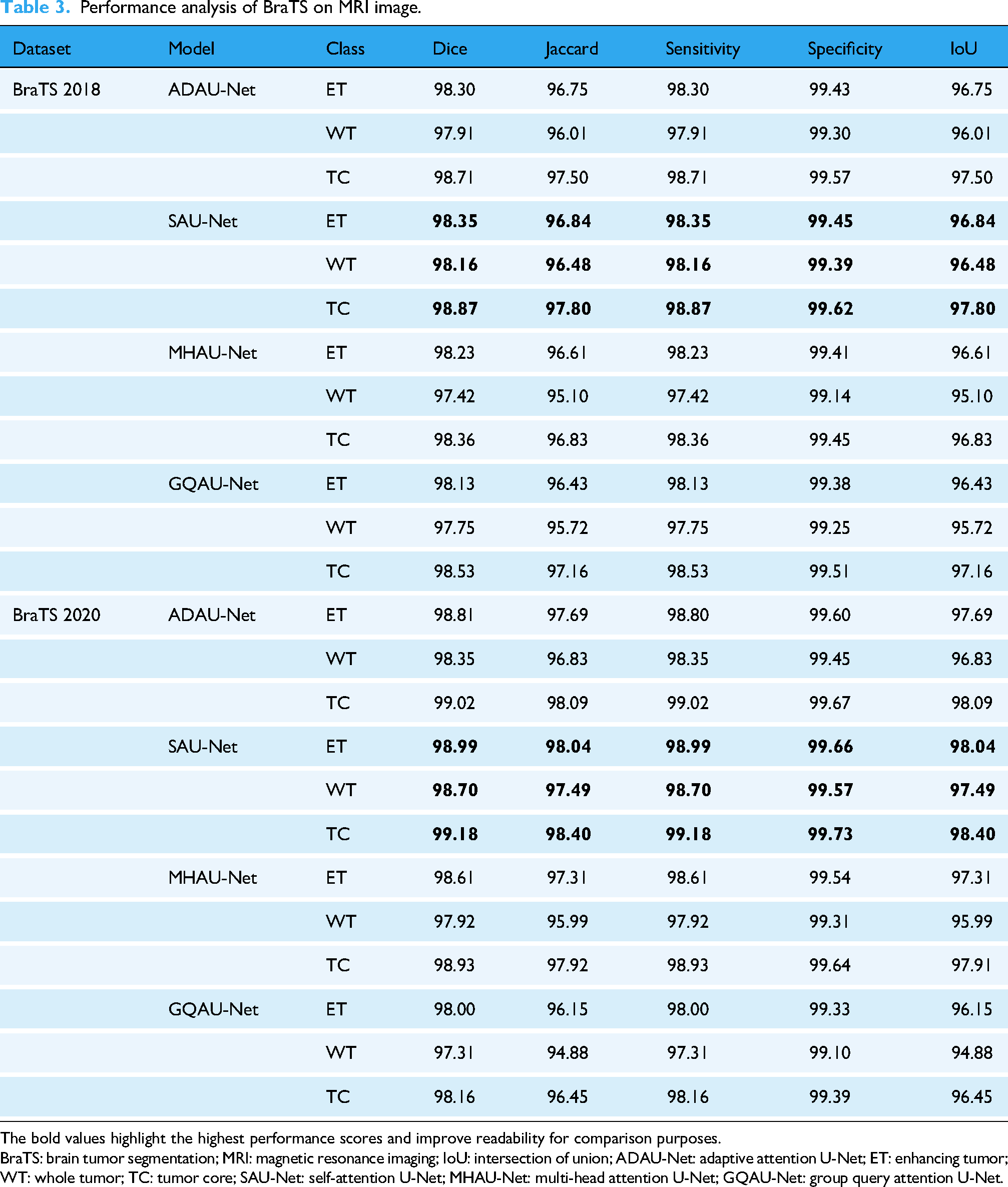
The bold values highlight the highest performance scores and improve readability for comparison purposes.
BraTS: brain tumor segmentation; MRI: magnetic resonance imaging; IoU: intersection of union; ADAU-Net: adaptive attention U-Net; ET: enhancing tumor; WT: whole tumor; TC: tumor core; SAU-Net: self-attention U-Net; MHAU-Net: multi-head attention U-Net; GQAU-Net: group query attention U-Net.
On the BraTS 2020 dataset, SAU-Net maintained strong performance, achieving Dice coefficients of 98.99% for the ET and 99.18% for the TC. These results demonstrated the robustness of the model and its ability to generalize across datasets with varying levels of complexity. In addition, the sensitivity and specificity metrics further confirmed the reliability of the proposed approach. Specifically, SAU-Net achieved sensitivity values of 98.99% for the ET, 98.70% for the WT, and 99.18% for the TC, along with corresponding specificity values of 99.66%, 99.57%, and 99.73%, respectively. Overall, these findings indicated that SAU-Net provided accurate and consistent tumor segmentation performance, supporting its suitability for automated brain tumor analysis.
The analysis of average performance metrics provided a concise and comparative perspective on the effectiveness of the proposed SAU-Net model relative to other SOA BraTS approaches. By averaging performance across the BraTS 2018 and BraTS 2020 datasets, this evaluation enabled a clearer understanding of the overall strengths and limitations of the SAU-Net architecture. The analysis also emphasized the role of individual performance metrics in assessing model reliability and practical applicability.
As summarized in Table 4, SAU-Net achieved the highest average Dice coefficients on both the BraTS 2018 and BraTS 2020 datasets, with values of 98.23% and 98.62%, respectively. These results indicated a strong capability to accurately delineate tumor boundaries. The Dice coefficient, which quantifies the overlap between ground truth annotations and predicted segmentations, remains a critical metric in medical image analysis. The high Dice scores demonstrated that SAU-Net consistently identified tumor regions with high precision, thereby reducing the likelihood of missing clinically significant areas during segmentation. In addition, SAU-Net attained Jaccard index values of 96.63% for BraTS 2018 and 97.34% for BraTS 2020, reflecting strong agreement with the ground truth annotations.
Average performance analysis of BraTS on MRI image.
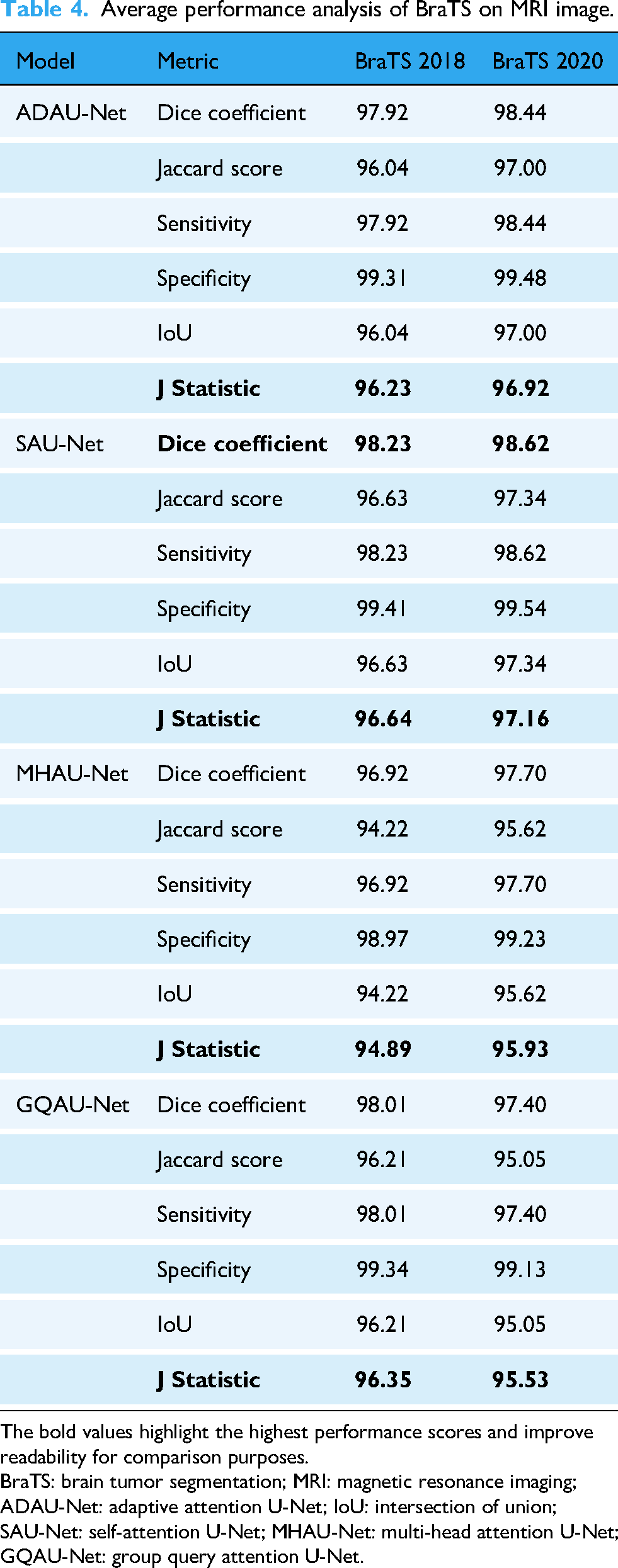
The bold values highlight the highest performance scores and improve readability for comparison purposes.
BraTS: brain tumor segmentation; MRI: magnetic resonance imaging; ADAU-Net: adaptive attention U-Net; IoU: intersection of union; SAU-Net: self-attention U-Net; MHAU-Net: multi-head attention U-Net; GQAU-Net: group query attention U-Net.
Furthermore, sensitivity and specificity metrics reinforced the robustness of the proposed model. SAU-Net recorded average sensitivity values of 98.23% on BraTS 2018 and 98.62% on BraTS 2020, indicating effective detection of true positive tumor cases. The corresponding average specificity values were 99.41% and 99.54%, respectively, demonstrating the model’s ability to accurately identify non-tumorous regions and minimize false positive predictions. Such balanced performance is particularly important in clinical contexts, where misclassification may have significant consequences.
In comparison, although alternative models such as MHAU-Net and GQAU-Net demonstrated competitive performance, their evaluation metrics remained consistently lower than those achieved by SAU-Net. For instance, MHAU-Net reported Dice coefficients of 96.92% for BraTS 2018 and 97.70% for BraTS 2020, whereas GQAU-Net achieved values of 98.01% and 97.40%, respectively. Overall, this comparative analysis confirmed that SAU-Net outperformed competing methods across key evaluation metrics, highlighting its strong potential for reliable deployment in automated brain tumor segmentation tasks.
The tradeoff between sensitivity (true positive rate) and specificity (true negative rate) was examined using the Youden index on BraTS datasets. The values calculated for the different models, as summarized in Table 4, demonstrated each model’s ability to balance false positive and false negative rates while accurately identifying tumor regions. These results supported the statistical significance of the findings and indicated that the proposed SAU-Net model achieved superior segmentation effectiveness compared to the other evaluated approaches. This analysis enabled a quantitative comparison of the models within a clinically relevant framework and underscored the importance of the Youden index as a robust statistical measure in performance evaluation.
SAU-Net incorporated self-attention mechanisms that allowed the model to emphasize diagnostically relevant regions of the input images, thereby capturing fine-grained details of irregular structures and poorly defined tumor boundaries. This capability enhanced sensitivity to subtle variations in tumor morphology and facilitated more accurate segmentation of heterogeneous tumor subregions, including WT, TC, and ET. In addition, extensive experiments were conducted to validate the performance of SAU-Net on multiple benchmark datasets, including BraTS 2018 and BraTS 2020, as reported in Tables 3 and 4. The results demonstrated the robustness of SAU-Net in addressing challenging segmentation scenarios and confirmed its superior performance in terms of Dice coefficient, sensitivity, and specificity relative to existing models.
Performance visualization
Figure 3 illustrates the effectiveness of the proposed SAU-Net model in segmenting brain tumors from MRI images. The close correspondence between the ground truth annotations and the predicted segmentation regions demonstrated the robustness of the proposed approach.

The prediction of BraTS using SAU-Net model. BraTS: brain tumor segmentation; SAU-Net: self-attention U-Net.
The observed performance improvements were attributed to the architectural design of SAU-Net, which incorporated enhanced contextual awareness through the integration of attention mechanisms and context aggregation units. This design enabled the model to capture complex spatial relationships and subtle feature variations that were critical for accurate tumor segmentation. Consequently, tumor boundaries were delineated more precisely, and small or irregular anomalies were detected more reliably, supporting accurate segmentation outcomes.
Benefits and limitations of SAU-Net
SAU-Net offers several practical advantages over traditional U-Net architectures. The lightweight attention block positioned at the bottleneck enables the network to capture broader contextual information while emphasizing clinically relevant tumor regions. This design leads to more precise segmentation of the WT, TC, and ET, while maintaining a relatively low model complexity and computational cost. In addition, SAU-Net retains the conventional encoder–decoder structure, which facilitates its integration into clinical workflows and allows straightforward fine-tuning for other MRI-based segmentation tasks.
Despite these advantages, the model also presents certain limitations. Because SAU-Net was trained and evaluated primarily on BraTS multimodal MRI datasets, its performance on other imaging modalities or data acquired from different institutions may vary unless appropriate domain adaptation techniques are applied. Although the single-head attention mechanism is computationally lighter than multi-head attention variants, it still introduces additional overhead compared to a standard U-Net, which may pose challenges in resource-constrained environments. Furthermore, the approach relies on high-quality manual annotations for supervised training, which are time-consuming and costly to obtain at a large scale.
Discussion
The comparative analysis presented in Table 5 showed that the proposed SAU-Net model achieved superior performance in BraTS, particularly on the BraTS 2018 dataset. The model consistently outperformed other SOA methods in segmenting the three primary tumor subregions—WT, TC, and ET—as measured by the Dice coefficient. Specifically, SAU-Net surpassed competing models such as HTTU-Net (91.50), achieving a Dice coefficient of 98.16 for the WT. This performance can be attributed to the integrated self-attention mechanisms, which enable the model to preserve critical spatial context while selectively emphasizing diagnostically relevant features. Such capability is essential for effectively handling the heterogeneous and complex nature of brain tumors.
Performance analysis (Dice coefficient) of our proposed SAU-Net model with state-of-the-art methods works.

SAU-Net: self-attention U-Net; 3D: three-dimensional; BraTS: brain tumor segmentation; ARDUNet: attentional residual dropout U-Net; CNN: convolutional neural network; WT: whole tumor; TC: tumor core; ET: enhancing tumor.
Similarly, SAU-Net achieved a Dice coefficient of 98.87 for the TC, outperforming existing approaches and underscoring its clinical relevance, as accurate delineation of the TC is crucial for treatment planning. The model also demonstrated strong performance in segmenting the ET, attaining a Dice coefficient of 98.23 and exceeding the results of methods such as 3D-ARDUNet (92.00). This ability to capture subtle variations in ET regions is particularly important for monitoring treatment response and disease progression.
Furthermore, the high segmentation accuracy was supported by the proposed preprocessing pipeline, which incorporated adaptive volume slicing and wavelet denoising to improve image quality and consistency. The carefully designed architecture, combining attention mechanisms with contextual feature aggregation, allows SAU-Net to effectively address challenges such as irregular tumor shapes and poorly defined boundaries, thereby contributing to its robust segmentation performance.
Table 6 presents a comparative sensitivity analysis of the proposed SAU-Net model on the BraTS 2018 dataset against other SOA segmentation approaches. Sensitivity, a critical metric for accurately identifying true positive tumor regions, highlighted notable performance differences among the evaluated models. The 3D-UNet achieved sensitivity values of 91.83% for the WT, 82.05% for the TC, and 84.36% for the ET, indicating relatively limited effectiveness, particularly in TC detection.
Performance analysis (sensitivity) of our proposed SAU-Net model with state-of-the-art methods works.

SAU-Net: self-attention U-Net; WT: whole tumor; TC: tumor core; ET: enhancing tumor; 3D: three-dimensional; BraTS: brain tumor segmentation.
HTTU-Net demonstrated improved sensitivity performance, achieving scores of 94.10% for the WT, 97.20% for the TC, and 94.30% for the ET, thereby showing enhanced capability in identifying TC regions. In contrast, the proposed SAU-Net model achieved substantially higher sensitivity values of 98.16% for the WT, 98.87% for the TC, and 98.23% for the ET, reflecting its strong ability to accurately detect all tumor subregions. This marked improvement can be attributed to the effective integration of self-attention mechanisms within SAU-Net, which enhances feature discrimination and contributes to more reliable BraTS performance. Consequently, the improved sensitivity supports more precise tumor delineation and has meaningful implications for clinical decision-making.
Table 7 presents a comparative performance analysis of the proposed SAU-Net model on the BraTS 2020 dataset against several SOA brain tumor segmentation approaches based on the Dice coefficient. The Dice coefficient, which quantifies the overlap between predicted tumor regions and ground truth annotations, serves as a critical metric for evaluating segmentation accuracy. The nnU-Net model proposed by Isensee et al. 26 achieved Dice coefficients of 88.95% for the WT, 85.06% for the TC, and 82.03% for the ET, indicating solid overall performance while revealing limitations in accurately segmenting ET regions.
Performance analysis (Dice coefficient) of our proposed SAU-Net model with SOA works.

SAU-Net: self-attention U-Net; SOA: state-of-the-art; WT: whole tumor; TC: tumor core; ET: enhancing tumor; BraTS: brain tumor segmentation.
The HybriCSF model introduced by Kataria et al. 27 yielded comparatively lower Dice scores, achieving 87% for the WT and only 63% for the ET, highlighting challenges in ET segmentation. Magadza et al. 28 reported a Dice coefficient of 91.2% for the WT, demonstrating improvement over several earlier approaches, whereas the spatial transformer-based model proposed by Susanto et al. 29 achieved a Dice score of 90.91% for the WT, reflecting reliable segmentation capability.
In comparison, the 3D ResAttU-Net-Swin model proposed by Angona et al. 31 achieved Dice coefficients of 92.50% for the WT, 89.70% for the TC, and 82.60% for the ET on the BraTS 2020 dataset, indicating competitive performance but comparatively lower accuracy in ET segmentation. Similarly, the DPF-Unet model introduced by Li et al. reported Dice scores of 91.74% for the WT, 89.44% for the TC, and 84.88% for the ET, reflecting strong overall segmentation performance. However, both approaches were consistently outperformed by the proposed SAU-Net across all tumor subregions.
The SAU-Net model achieved Dice coefficients of 98.99% for the WT, 98.70% for the TC, and 99.18% for the ET, surpassing all compared methods. These results demonstrate the effectiveness of the self-attention mechanisms integrated into the SAU-Net architecture and position it as a leading approach for brain tumor segmentation. Furthermore, the performance analysis indicates that SAU-Net provides robust and reliable tumor delineation, particularly in accurately identifying complex tumor regions, which is essential for supporting informed clinical decision-making and improving patient outcomes.
The proposed SAU-Net model demonstrated substantially improved performance in segmenting ETs compared to previous studies, as shown in Tables 5 and 7. Historically, ET segmentation has been particularly challenging due to the heterogeneous and irregular nature of enhancing tumor regions, which has often resulted in lower Dice coefficients for earlier models. In contrast, SAU-Net incorporated advanced attention mechanisms that enabled the model to more effectively focus on the complex characteristics of ET regions. This targeted attention enhanced segmentation accuracy by strengthening the model’s ability to distinguish tumor tissue from surrounding non-tumorous areas.
On the BraTS 2018 dataset, SAU-Net achieved a Dice coefficient of 98.23% for ET segmentation, significantly outperforming established methods such as HTTU-Net (88.70%) and RMU-Net (79.36%). Similarly, on the BraTS 2020 dataset, SAU-Net attained a Dice coefficient of 99.18% for the ET, surpassing previously reported SOA results, including nnU-Net (82.03%) and HybriCSF (63%). The superior ET segmentation performance of SAU-Net can be attributed to the effective integration of attention mechanisms and robust feature extraction within its architecture. This design enabled the model to adaptively learn and emphasize the most informative features for ET identification, resulting in consistently enhanced segmentation outcomes relative to contemporary approaches.
Complexity analysis
This section evaluated the computational efficiency of different BraTS segmentation models applied to MRI images, with a focus on build time, prediction time, and memory usage. The analysis included four attention-based U-Net variants—ADAU-Net, SAU-Net, MHAU-Net, and GQAU-Net—and was conducted using the BraTS 2018 and BraTS 2020 datasets.
Table 8 summarizes the computational complexity metrics for each model across both datasets. On the BraTS 2018 dataset, ADAU-Net exhibited the longest build time at 225.25 s, whereas SAU-Net achieved a more efficient build time of 147.74 s. MHAU-Net and GQAU-Net followed with build times of 157.62 s and 152.00 s, respectively. Regarding prediction time, SAU-Net outperformed all other models, requiring only 54.85 s, while ADAU-Net recorded the longest prediction time at 70.20 s. Memory usage remained relatively consistent across models, with ADAU-Net consuming 11,253.04 MB and SAU-Net requiring slightly less at 11,145.25 MB. In contrast, MHAU-Net and GQAU-Net exhibited higher memory consumption, using 11,986.01 MB and 12,606.97 MB, respectively.
Complexity analysis of BraTS model on MRI image.
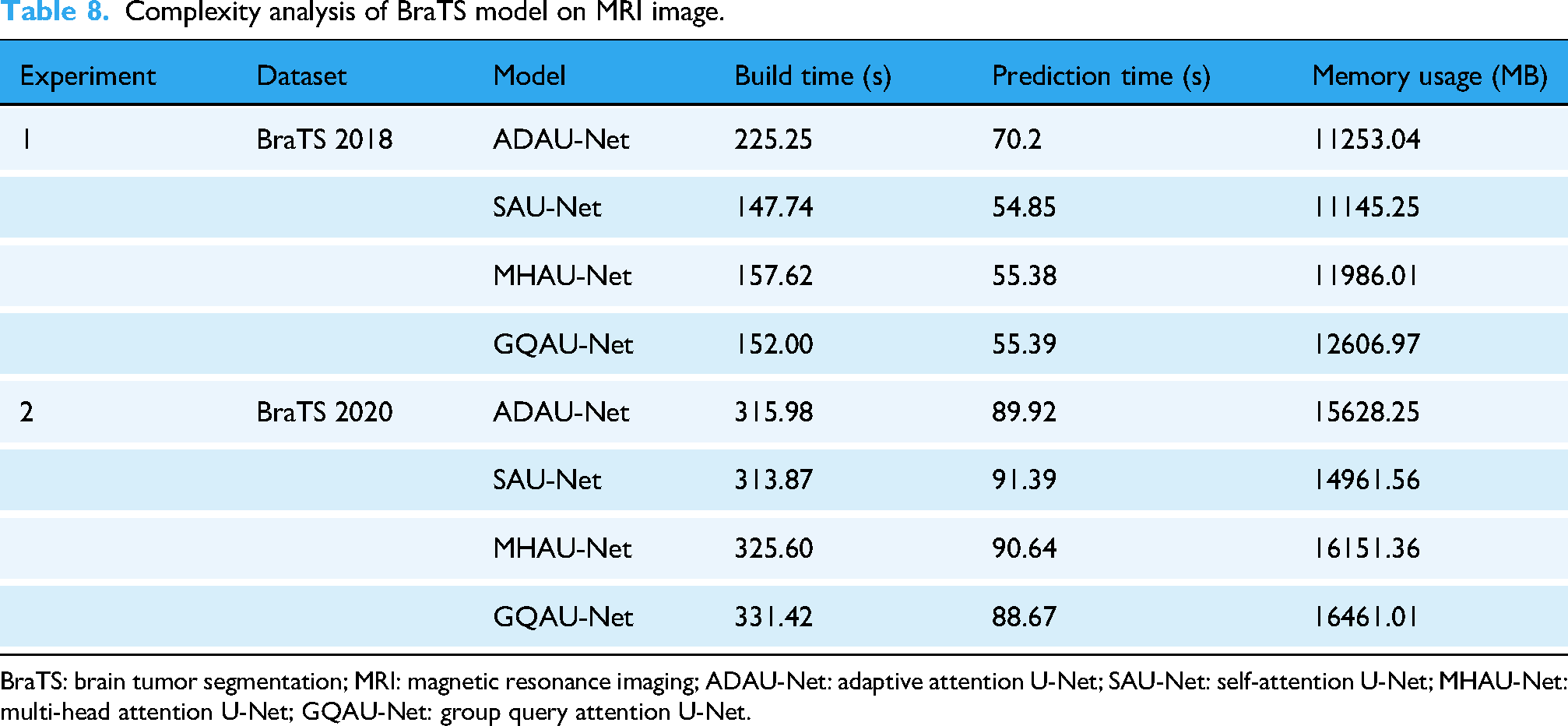
BraTS: brain tumor segmentation; MRI: magnetic resonance imaging; ADAU-Net: adaptive attention U-Net; SAU-Net: self-attention U-Net; MHAU-Net: multi-head attention U-Net; GQAU-Net: group query attention U-Net.
A similar trend was observed on the BraTS 2020 dataset. ADAU-Net again recorded the longest build time at 315.98 s, while SAU-Net maintained competitive efficiency with a build time of 313.87 s. MHAU-Net and GQAU-Net required 325.60 s and 331.42 s, respectively. Prediction times increased for all models due to higher dataset complexity; SAU-Net required 91.39 s, whereas ADAU-Net required 89.92 s.
The memory usage analysis further revealed notable differences among the evaluated architectures. For BraTS 2018, SAU-Net demonstrated the lowest memory usage at 11,145.25 MB, followed closely by ADAU-Net at 11,253.04 MB. In contrast, MHAU-Net and GQAU-Net required substantially higher memory. For BraTS 2020, memory consumption increased across all models, with ADAU-Net using 15,628.25 MB and SAU-Net requiring 14,961.56 MB. MHAU-Net and GQAU-Net showed the highest memory demands at 16,151.36 MB and 16,461.01 MB, respectively. These findings indicated that although memory requirements increased with higher-resolution datasets, SAU-Net consistently maintained lower memory usage relative to the other attention-based models.
In terms of training complexity, SAU-Net demonstrated favorable efficiency when compared with existing approaches such as RMU-Net and HTTU-Net. The analysis showed that SAU-Net required 313.87 s for training, which was substantially lower than HTTU-Net (approximately 36,000 s) and RMU-Net (approximately 10,020 s). This comparison highlighted SAU-Net as a computationally efficient alternative while preserving strong segmentation performance.
Overall, the complexity analysis demonstrated that although all evaluated models exhibited considerable memory usage, SAU-Net consistently achieved lower build and prediction times compared to its counterparts. These results emphasize the importance of balancing segmentation performance with computational efficiency, particularly for clinical applications where rapid and resource-efficient processing is critical. The findings provide valuable insights into the operational characteristics of the evaluated models and inform future research and deployment decisions in brain tumor segmentation.
Model complexity versus accuracy
To further analyze the computational efficiency of the proposed SAU-Net model, segmentation accuracy was examined in relation to the complexity metrics reported earlier, including build time and memory usage (Table 8). Table 9 summarizes this relationship. SAU-Net exhibited the most favorable tradeoff between accuracy and computational complexity. Although its memory usage was slightly higher than that of ADAU-Net, SAU-Net achieved the highest Dice accuracy on both the BraTS 2018 and BraTS 2020 datasets while maintaining among the lowest build and prediction times.
Comparison of model complexity and segmentation accuracy for BraTS 2018 and 2020.

BraTS: brain tumor segmentation; SAU-Net: self-attention U-Net; ADAU-Net: adaptive attention U-Net; MHAU-Net: multi-head attention U-Net; GQAU-Net: group query attention U-Net.
In contrast, MHAU-Net and GQAU-Net incurred higher computational costs while delivering comparatively lower segmentation performance. These findings demonstrate that SAU-Net provides an effective balance between segmentation accuracy and resource consumption, making it well-suited for clinical environments where both computational efficiency and high precision are critical.
Research question validation
RQ1: Effectiveness of SAU-Net compared to U-Net variants
The analysis presented in Table 3 confirmed that the proposed SAU-Net model significantly outperformed traditional U-Net variants, including ADAU-Net, MHAU-Net, and GQAU-Net. SAU-Net consistently achieved higher values across all key evaluation metrics, such as the Dice coefficient, Jaccard index, sensitivity, and specificity. For instance, SAU-Net achieved Dice coefficients of 98.23% on the BraTS 2018 dataset and 98.62% on the BraTS 2020 dataset, exceeding the corresponding scores of ADAU-Net (97.92% and 98.44%), MHAU-Net (96.92% and 97.70%), and GQAU-Net (98.00% and 97.40%). These results indicated that incorporating self-attention mechanisms into the U-Net architecture substantially improved segmentation accuracy. Consequently, the first research question was successfully validated.
RQ2: Comparison with SOA models
The comparative analyses presented in Tables 5 and 6 demonstrated that SAU-Net outperformed SOA segmentation methods in accurately delineating tumor subregions, including the WT, TC, and ET. On the BraTS 2018 dataset, SAU-Net achieved Dice coefficients of 98.16% for the WT, 98.87% for the TC, and 98.23% for the ET, which were substantially higher than those reported for leading approaches such as HTTU-Net and RMU-Net. Similarly, on the BraTS 2020 dataset, SAU-Net attained Dice coefficients of 98.99% for the WT, 98.70% for the TC, and 99.18% for the ET, significantly surpassing models including nnU-Net and HybriCSF. These consistent improvements across both datasets confirmed that SAU-Net effectively and accurately segmented critical tumor subregions, thereby validating the second research question.
RQ3: Impact of self-attention integration
The complexity analysis summarized in Table 8 demonstrated that SAU-Net achieved a favorable balance between segmentation accuracy and computational efficiency. On the BraTS 2018 dataset, SAU-Net recorded the most efficient build time (147.74 s) and prediction time (54.85 s) among the evaluated models, including ADAU-Net (225.25 s build time, 70.20 s prediction time), MHAU-Net (157.62 s build time, 55.38 s prediction time), and GQAU-Net (152.00 s build time, 55.39 s prediction time). Although SAU-Net exhibited memory usage comparable to other models, its superior processing speed combined with high segmentation performance indicated that the integrated self-attention mechanisms effectively enhanced computational efficiency without sacrificing accuracy. These findings therefore validated the third research question.
Hypothesis validation
The experimental findings provided compelling evidence that the proposed SAU-Net outperformed conventional U-Net models as well as other SOA techniques in BraTS. The integration of self-attention mechanisms within the U-Net architecture resulted in substantial improvements in segmentation accuracy, as demonstrated in Tables 3, 5, and 7. On the BraTS 2020 dataset, SAU-Net significantly surpassed existing models such as nnU-Net and HybriCSF, achieving Dice coefficients of 98.99% for the WT, 98.70% for the TC, and 99.18% for the ET.
Similarly, on the BraTS 2018 dataset, SAU-Net demonstrated consistently superior performance, attaining Dice coefficients of 98.16% for the WT, 98.87% for the TC, and 98.23% for the ET. The observed improvements across all evaluation metrics, including Dice coefficient, sensitivity, and specificity, were primarily attributed to the self-attention mechanism’s ability to emphasize diagnostically relevant features while preserving essential spatial context.
Furthermore, the complexity analysis presented in Table 8 supported the second component of the proposed hypothesis concerning computational efficiency. SAU-Net achieved lower build time (147.74 s) and prediction time (54.85 s) compared with alternative models such as ADAU-Net. This combination of high segmentation accuracy and reduced computational complexity confirmed the suitability of SAU-Net for clinical applications requiring both efficiency and precision.
Scientific implications for clinical practice
SAU-Net has important implications for clinical practice, particularly in the field of neuro-oncology. Accurate BraTS is critical for surgical planning, diagnosis, and therapeutic monitoring. By incorporating self-attention mechanisms, SAU-Net enables more precise delineation of tumor boundaries, especially in challenging regions such as the TC and ET. This increased precision can enhance radiologists’ confidence in tumor identification, leading to more informed and accurate treatment decisions.
Moreover, the computational efficiency of SAU-Net supports its potential integration into real-time clinical workflows, including resource-constrained healthcare environments. The utilization of multimodal MRI data provides anatomical insights, which further facilitate personalized treatment strategies and support preoperative and postoperative planning. By reducing manual intervention and accelerating the segmentation process, SAU-Net has the potential to improve patient outcomes and streamline clinical decision-making in brain tumor management, offering a practical and scalable solution for real-world deployment.
Conclusion
This study evaluated the performance of the proposed SAU-Net model for brain tumor segmentation using the BraTS 2018 and BraTS 2020 datasets and demonstrated that it consistently outperformed existing approaches. By effectively integrating context aggregation with advanced attention mechanisms, the model accurately identified tumor regions across diverse and complex cases.
On the BraTS 2018 dataset, SAU-Net achieved high sensitivity and specificity values of 98.87% and 99.05%, respectively, along with a Jaccard index of 96.31% and an average Dice coefficient of 98.16%. On the more challenging BraTS 2020 dataset, the model maintained strong performance, achieving a Jaccard index of 97.04% and an average Dice coefficient of 98.99%. These results indicated that SAU-Net provided superior tumor identification and segmentation accuracy compared to existing SOA methods.
The observed performance improvements suggest that SAU-Net can significantly enhance clinical diagnosis and treatment planning in neuro-oncology, thereby contributing to improved patient outcomes.
Despite these promising results, certain limitations remain, including the lack of investigation into advanced feature fusion and hybrid segmentation strategies. Future research will focus on developing hybrid segmentation frameworks and exploring feature fusion techniques to further enhance segmentation accuracy and model generalization.
Footnotes
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Author contributions
Md. Alamin Talukder: Conceptualization, data curation, methodology, software, resource, visualization, formal analysis, supervision, validation, and writing—original draft, review, and editing. Mehnaz Tabassum: Methodology, software, resource, visualization, validation, investigation, and writing—review and editing. Majdi Khalid: Methodology, software, resource, visualization, validation, and investigation.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of data and materials
The data that support the findings of this study are openly available in the BraTS 2018: https://www.med.upenn.edu/sbia/brats2018/data.html and BraTS 2020: ![]()
Guarantor
Md. Alamin Talukder.
