Abstract
Objective
The precise segmentation of kidneys from a 2D ultrasound (US) image is crucial for diagnosing and monitoring kidney diseases. However, achieving detailed segmentation is difficult due to US images’ low signal-to-noise ratio and low-contrast object boundaries.
Methods
This paper presents an approach called deep supervised attention with multi-loss functions (MLAU-Net) for US segmentation. The MLAU-Net model combines the benefits of attention mechanisms and deep supervision to improve segmentation accuracy. The attention mechanism allows the model to selectively focus on relevant regions of the kidney and ignore irrelevant background information, while the deep supervision captures the high-dimensional structure of the kidney in US images.
Results
We conducted experiments on two datasets to evaluate the MLAU-Net model's performance. The Wuerzburg Dynamic Kidney Ultrasound (WD-KUS) dataset with annotation contained kidney US images from 176 patients split into training and testing sets totaling 44,880. The Open Kidney Dataset’s second dataset has over 500 B-mode abdominal US images. The proposed approach achieved the highest dice, accuracy, specificity, Hausdorff distance (HD95), recall, and Average Symmetric Surface Distance (ASSD) scores of 90.2%, 98.26%, 98.93%, 8.90 mm, 91.78%, and 2.87 mm, respectively, upon testing and comparison with state-of-the-art U-Net series segmentation frameworks, which demonstrates the potential clinical value of our work.
Conclusion
The proposed MLAU-Net model has the potential to be applied to other medical image segmentation tasks that face similar challenges of low signal-to-noise ratios and low-contrast object boundaries.
Introduction
Ultrasound (US) is one of the mainstays of medical diagnostics for its broad applicability and efficacy. It allows for both a qualitative and quantitative evaluation, offering real-time insights without ionizing radiation. Despite its advantages, US imaging encounters challenges such as artifact presence, noise interference, and subjective interpretation, distinguishing it from modalities like X-ray, computed tomography (CT), and magnetic resonance imaging (MRI). 1 A valid strategy to deal with many of these limitations is the use of computer-aided diagnosis (CAD), which greatly enhances the diagnostic accuracy. Effective image segmentation significantly, enhances the analysis of US images, ensuring precise delineation of anatomical structures for thorough interpretation and clinical diagnosis. Although several techniques have been developed for segmenting US images, their segmentation capability is still inadequate when dealing with relatively complex images. 2 US, one of the most commonly used imaging modalities, is a ubiquitous and effective screening and diagnostic tool for physicians and radiologists. In particular, US imaging is widely used in prenatal screening worldwide due to its safety, low cost, non-invasiveness, real-time imaging, convenience, and user experience. 3
Recently, there has been a growing interest in automated medical image analysis methods, including kidney ultrasound (KUS) segmentation. Segmentation of KUS images is a difficult task due to the complexity of the kidney and its structure and speckle noise in US images.4,5 In contrast, accurate segmentation of the kidney region is essential for various clinical applications such as disease diagnosis, surgical planning, and treatment monitoring. 6 Traditionally, KUS image segmentation involved manual contouring of the kidney, which is time-consuming and subject to user variability.7,8 To address these limitations, several semi-automatic segmentation methods have been proposed.9–11 Conversely, these methods rely heavily on manual initialization and may produce errors due to unclear boundaries and uneven intensity distribution. 12 Consequently, there is a growing need for automatic and robust KUS segmentation techniques to improve segmentation accuracy and efficiency.
Interest in medical image segmentation (MIS) using deep learning (DL) techniques has grown significantly, and various convolutional neural network (CNN) architectures such as U-Net, 13 FCN, 14 CPFNet, 15 Deeplabv3, 16 and SegNet have been proposed. 17 These approaches have greatly improved segmentation accuracy and reliability compared to traditional segmentation networks. The deep neural networks are used to learn high-level representations of medical images and capture the spatial relationships between complex anatomical structures. The stimulation method includes depth supervised attention (DSA) and multiple loss functions (MLFs). The use of various loss functions and an attention module enhances the flexibility and precision of segmentation results. The observation mechanism allows the network to selectively highlight relevant image regions while using MLFs, allowing the model to cover different aspects of the segmentation task and improve the model and overall performance. For example, Chen et al. proposed a similar approach for US kidney segmentation using a modified U-Net architecture with a deep controlled attention mechanism and multiloss features. 18 They reported a significant improvement in segmentation accuracy compared to existing methods, showing the potential of this approach for KUS segmentation problems. Moreover, Feng et al. presented a CNN-based approach that integrates multi-level contextual information and multitask learning for re-segmenting US images. Their segmentation methods were more accurately applied than conventional ones. In addition, a DL-based framework using a bidirectional network with multilevel feature fusion was proposed for kidney segmentation in 3D-US volumes, which showed strong performance in segmenting kidneys of different shapes and sizes. 19
Furthermore, attention mechanisms have been commonly employed in MIS tasks. Attention-guided U-Net for brain tumor segmentation based on MRI images has improved accuracy by incorporating attention mechanisms to guide the network and focus tumor regions. 20 In addition, a self-supervised attention-guided network has been introduced for cardiac image segmentation, which uses self-supervised learning and attention mechanisms to improve segmentation accuracy, especially when labeled data is limited. 21 Multi-loss features have also shown their ability to increase the performance of DL networks in MIS tasks. The accuracy of pancreas segmentation from CT images was improved by incorporating a multiloss attention-guided network with dice loss and focal loss functions. 22 Moreover, a multi-task DL approach with MLFs was shown to improve the segmentation accuracy of liver tumors from CT images. 23 However, kidney segmentation in 2D US images is still a problem in most cases, as the signal-to-noise ratio is low and the contours of the object are poorly contrasted. In most recently proposed CNNs for segmentation, 24 they do not allow capturing the whole high-dimensional structure of a 2D kidney from US entirely, as loss functions based on weak supervision tend to be inefficient to integrate spatial relationships between neighboring pixels; such approaches fail to produce regularly shaped segmentation masks.
To improve the performance of MIS and reduce the complexity of the network structure, we proposed a model called deep tracking MLAU-Net for KUS image segmentation. The model consists of 2D U-Net with two main regulatory components: attention and depth tracking. The main contributions of our work are threefold. (a) The proposed MLAU-Net introduces new depth enhancements and attention gates that enrich the model and allow it to focus on essential image features while ignoring noise. (b) The model includes extensive monitoring as an adjustment method and encompasses target functions. Across the decoder layers, these target functions deal with small data and deeper networks. The process involves carefully transitioning from deeper blocks to the original segmentation size, ensuring that the model effectively captures high-dimensional structures. (c) A preprocessing pipeline, including custom normalization and efficient data augmentation during training, ensures model reliability and efficiency using various US images. These advances pertain to enhancing the accuracy and efficiency of medical image analysis in CAD. The remaining sections of this work are organized in a uniform manner. In the “Related work” section, various related studies are reviewed and the characteristics and challenges associated with existing approaches are described, while the proposed MLAU-Net approach for KUS image segmentation is detailed in the “Materials and methods” section. The performance of the various measures is reviewed in the “The Experiment” section; finally, the conclusions of the proposed approach are presented in the “Experimental results and discussion” section.
Related work
In recent years, considerable research has been conducted on KUS segmentation using DL methods. Several studies have proposed different methods to achieve accurate and efficient kidney segmentation in US images. U-Net, 13 a pioneering encoder–decoder CNN-based framework, has demonstrated exceptional image segmentation, leading to the development of many U-shaped variants. Weighted Res-UNet 25 uses a weighted attention mechanism to segment small regions, while U-Net++ 26 introduces a U-shaped layout with nested dense bypasses to reduce the semantic gap. Dense-UNet uses a densely connected structure to provide optimal separation between intra-output and inter-institution scans. 27 The integration of low- and high-level details is achieved through full-scale skip connections in U-Net3+. ENS-UNet provides a U-shaped architecture with minimal pre- and post-processing requirements, 28 while C-UNet includes inception-like convolutional blocks, recurrent convolutional blocks, and extended convolutional layers to segment skin lesions. 29 Image segmentation tasks often utilize CNN-based techniques that leverage their potent feature extraction capabilities by concentrating on adjacent pixels.30,31
An attention-based U-Network, which includes attention mechanisms to improve segmentation accuracy, has been proposed for kidney segmentation in US images. 32 This innovative method is complemented by an efficient kidney segmentation approach that uses an attention-based dual network that efficiently captures contextual information to enable precise segmentation. 23 Additionally, a self-guided attention model adaptable for kidney segmentation using US images has emerged, employing self-supervised learning to enhance network performance. 33 Moreover, a multidimensional attention-guided U-Network tailored for renal tumor segmentation from CT images has been introduced, enhancing segmentation accuracy, particularly for tumor regions. 34 Bidirectional attention-guided U-Net for kidney segmentation from multicontrast CT images has demonstrated the efficacy of bidirectional and attentional mechanisms for accurate segmentation.35,36 Similarly, a comparable attention module, incorporating two convolutional layers followed by softmax, was integrated into the U-Net hierarchical pooling framework for left atrial segmentation. 37 Recent advancements include the incorporation of additional attention gate modules into the bypass interfaces of the U-Net decoding path, enhancing the model’s ability to capture additional information from the encoder.38,39 Table 1 summarizes previous studies on KUS segmentation using deep-learning approaches. These studies had different goals and specific limitations. Their objectives ranged from applying multitasking CNNs to the precise kidney segmentation of US.
A summary of the objectives and limitations of previous studies using deep learning methods for human kidney ultrasound segmentation.

Materials and methods
The main objective of this work was to propose an MLAU-Net model designed for accurate US kidney segmentation of healthy organs to assist the radio-oncologist. This section provides a detailed overview of the proposed segmentation methodology. Figure 1 depicts the framework tailored for KUS segmentation.

Proposed MLAU-Net pipeline for KUS segmentation.
KUS dataset
One of the most challenging aspects of deep learning (DL) approaches is developing datasets that require many manually labeled images to train a neural network effectively. The WD-KUS dataset was created via our collaborative research with Guangzhou Medical University and its First Affiliated Hospital. This dataset comprises 44,880 KUS images obtained from patients with a clinical indication for US investigations of their kidneys using TELEMED SmartUs EXT-1 M/3 M. In accordance with privacy measures, all personally identifiable information (PII) was carefully removed during the dataset collection process. The study was conducted in accordance with the Institutional Ethics Committee (IEC) at the First Affiliated Hospital of Guangzhou Medical University.
Deep supervised multi-loss attention 2D U-Net framework (MLAU-Net)
The model uses a single-channel 2D renal US image as input, and the output is a 2D image of identical dimensions depicting the kidney segmentation map. To achieve these segmentation goals, the MLAU-Net approach is proposed with specific modifications, including feature depth enhancement and an attention gate, which enhance focus on important image features while reducing noise. In addition, the model includes deep supervision as a regularization technique that embeds objective functions in the decoder layers to address the challenges of small datasets and deeper networks. This approach requires carefully transitioning from deeper blocks to the original segmentation size, ensuring the efficient capture of high-dimensional structures. In Figure 2, MLAU-Net framework for KUS segmentation, emphasizing the integration of attention mechanisms and MLFs. Attention gates selectively focus on relevant features, while various loss functions, including dice and focus loss, optimize training for accurate segmentation.

MLAU-Net framework for KUS segmentation incorporated with deep supervision and an attention gate.
Depth and the number of initial filters
The model designs seven layers and 16 starting filters instead of the conventional five layers and 64 starting filters. Increasing the depth of the model ensures that the deeper layers capture more complex and abstract information essential for encoding latent details needed for effective feature representation. Concrete concerns about available computational resources recommend using a five-layer model with 16 initial filters for the most efficiency, even though the seven-layer structure could boost performance metrics. Based on the flexible and parametric structure of the model, it is possible to easily modify the number of layers, providing flexibility in balancing computational restrictions with performance. A comprehensive description of the architecture of the proposed MLAU-Net model is given in Table 2. The framework's multilevel attention (MLA) processes enhance its overall performance and capacity to hold intricate features. The essential features, including layer type, filter size, and activation functions, are detailed in each row, corresponding to a specific layer or module in the model. This detailed model summary is invaluable for understanding the proposed MLAU-Net architecture's structural complexity.
Summary of the proposed MLAU-Net model.

Attention mechanism in MLAU-Net
The introduction of attention gates into the U-Net algorithm is done at the level of the concatenation of the skip connections with the up-sampled signal coming from deeper layers in the decoder module. The intricate details of attention gates, elucidating the process, are visualized in Figure 3, which comprises a visual representation of how an attention gate enhances the model's focus on relevant KUS features. 43

The attention gate works by adjusting the input functions (
Here, 


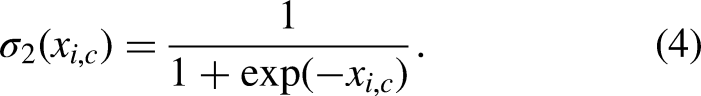


Deep supervision
Deep supervision involves incorporating companion objective functions into the final three hidden layers in the decoder of the network. The final loss, as illustrated in Figure 5 featuring the proposed MLAU-Net, incorporates dice loss

Calculation of the loss in the proposed approach.
Learning rate decay
Learning rate decay is a primarily utilized technique to improve performance in DL models. This involves periodically reducing the learning of the optimizer when the validation metric plateaus for a certain number of time steps (e.g., 20). This adjustment helps the model approach the actual minimum of the loss function, which can improve overall performance. Implementing a learning rate decay policy is often valuable for optimizing DL frameworks.
The experiment
The input provided to the network is the US 2D image containing the renal US data, and the output will be the 2D kidney segmented image. The following pipeline is created to train and test image classification models on the WD-KUS dataset depending on the different types of data augmentation needed. It also includes using the AdamW optimizer and two loss function training sets with different weights to make the model work appropriately.
Preprocessing
Normalization methods
In the proposed MLAU-Net, we employed min-max normalization to scale the data between 0 and 1, ensuring consistency across all input images. This approach was chosen due to the absence of significant outliers in our US image dataset. Specifically, we applied the following formula to normalize the data:
Resizing
Finally, every image is resized to a standard size of [128, 128] pixels since all the input/output images must have the same size to be fed into the DL model. The values chosen are explained in terms of powers of 2. Since we are implementing a U-Net architecture that gradually down samples the volumes by a factor of 2, using a size that is a power of 2 will avoid size errors in the inner layers of the model. This value is a parameter that should be low enough (limited by the computational resources) to be able to perform the training and high enough to have an amicable spatial resolution. This value can be optimized as a parameter and increased for future finer outputs.
Data augmentation
Regarding data augmentation, the most common techniques for medical images are as follows:
Affine transformations:
Rotations: Slight rotations in a range (−10°, 10°) will considerably increase the number of cases and make the model more robust. Scalings: Slight scaling are applied to the input volume that zooms the patient's body by 10%.
Training for several epochs on a limited dataset requires us to take specific precautions against overfitting. To combat overfitting, we use many different data augmentation methods. Through the training cycle, the following data augmentation methods are applied to the data on the fly: random rotations, random scaling, random skewing, gamma correction augmentation, and mirroring. These methods augment the original training dataset and improve the model on new data that may look different. Annotated data additions are fed into the data loader of the DL model and applied with predefined probabilities. In other words, at each step of each epoch, each data sample is transformed according to defined transformations and probabilities. The training speed is also not compromised because monetary transformations allow us to complete these steps efficiently. Other transformations for data augmentation, such as noise addition, flipping, or other distortion transformations, should not be included since, in the medical image domain, it is essential to preserve the original relationships and orientations for the model to learn contextual anatomical information. Figure 4 represents the results of data augmentation on a single image.

Data augmentation effects on a single image. Various transformations, including random rotations, scaling, skewing, gamma correction, and mirroring, applied during on-the-fly augmentation contribute to the enhanced training dataset.
Dataset splitting
Detailed overview of dataset splitting for kidney ultrasound (WD-KUD) images. This table outlines the division of patients’ kidney ultrasound images into training and test sets, including the number of patients, total images, ground truth annotations, and allocation for training and validation subsets.

Implementation and evaluation methods
Loss function and model training
All experiments were executed in Pytorch, and the GPU was an RTX 4090. The DL process uses the WD-KUS and Open Kidney Dataset’s 47 dataset for training networks to classify renal US images. The final results of the trained models are tested with an independent test set. In the training phase, training frames are applied to various random data augmentation strategies such as auto-brightness, contrast, gamma, Gaussian blur, Gaussian noise, arbitrary rotation, elastic deformation, random clipping, and scaling. The AdamW optimizer, with 100 epochs, a batch size of 16, and a weight decay of 0.001, is utilized to optimize the networks and reduce overfitting. The final loss used in the model is calculated as shown in Figure 5.
Two loss functions, dice loss L
The weighted dice loss was selected based on a quantitative evaluation of loss functions. The comprehensive loss function for the network can be expressed as presented in equation (9).
Focal loss (
We chose various standard evaluation metrics to assess the segmentation performance of our network. These metrics include dice score, average symmetric surface distance (ASSD in mm), accuracy, specificity, recall, and 95th percentile of Hausdorff distance (HD95 in mm). The formulations for these metrics are provided in equations (11)–(18), as detailed in Ref. 48 The metrics used for evaluation are based on true positives (TPs). The segmentation model correctly identifies these pixels as belonging to the target class (e.g., lesion, organ, etc.). True negatives (TNs this term isn't commonly used because the focus is on identifying positive (target) regions rather than classifying the entire image as negative), false positives (FPs), and false negatives (FNs). The metrics determined should range from 0 to 1 or 0 to 100%, and the performance of the proposed model increases when the calculated metrics’ values are high.



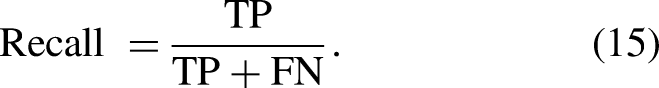



Experimental results and discussion
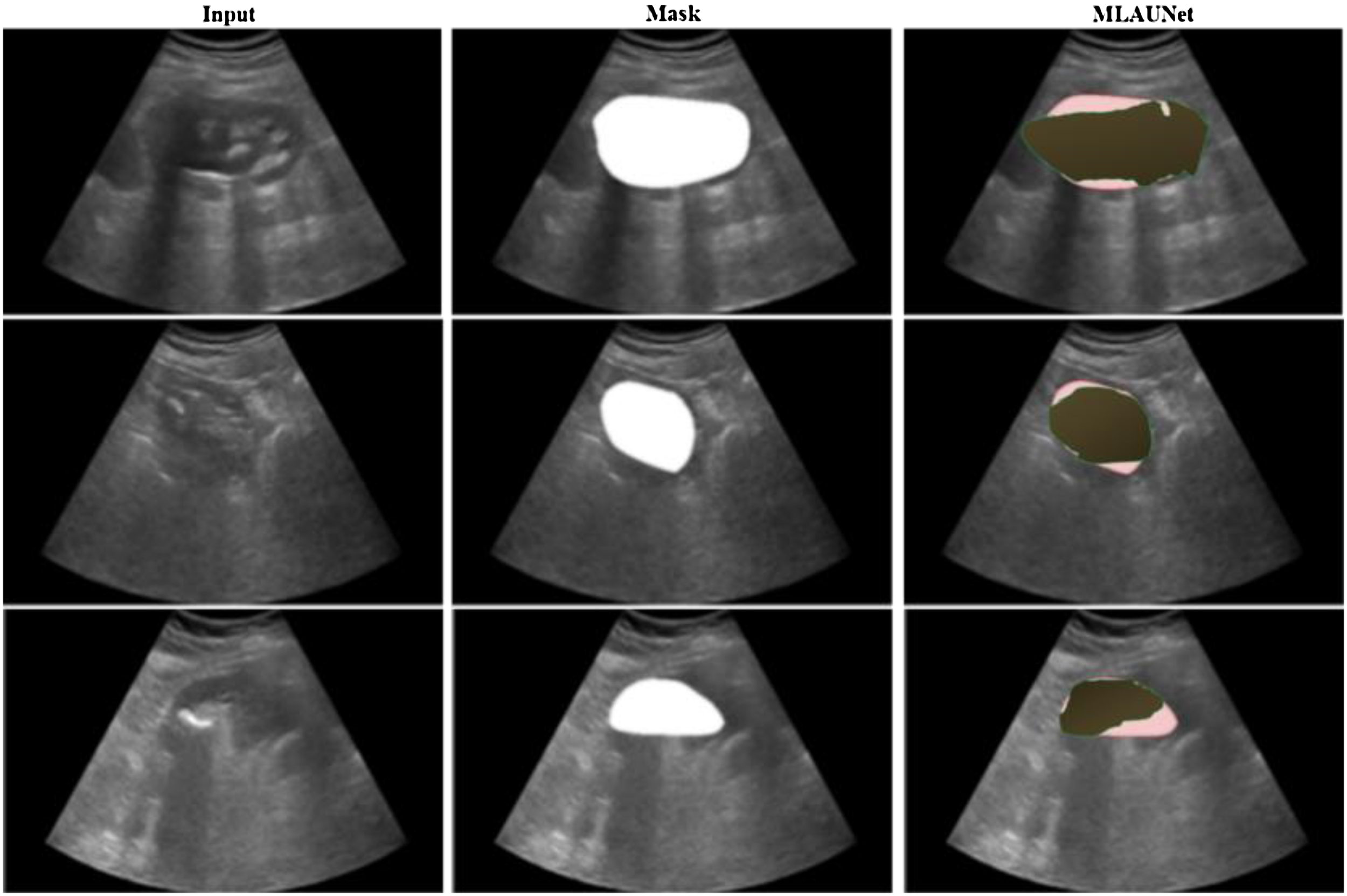
In this study, we adopted six frequently employed metrics to quantitatively compare various methods for KUS image segmentation performance. The six evaluation indicators include accuracy, dice, HD95, recall, specificity, and ASSD. Accuracy, dice, recall, and specificity vary from 0 to 1. A higher score indicates superior method quality, whereas lower ASSD scores correspond to improved segmentation results for the method. Figure 6 shows results in images featuring both the input images and the segmented output masks obtained using the proposed MLAU-Net framework. In this representation, the red color indicates the label, while the green color shows the prediction. Figure 6 show that the images the proposed framework predicted closely resemble the original mask.

Visual analysis of MLAU-Net results; red denotes the label, and green represents the model's predictions.
Qualitative analysis
In particular, Figures 7 and 8 present a qualitative analysis of the WD-KUS dataset and Open Kidney Dataset 47 using the proposed MLAU-Net model, contrasting it with seven other state-of-the-art approaches. For visual comprehension, segmented maps of sample images from the test dataset are depicted for both the proposed models and the competing models in Figure 7.

Qualitative analysis of MLAU-Net and different baseline methods on the WD-KUD dataset. Red indicates the label, and green represents the model's predictions. Column 1 displays the input image, Column 2 shows the mask, Columns 3–9 exhibit the outputs of baseline models, and the final column depicts the prediction of the proposed model.

The open kidney dataset was used to perform a qualitative analysis of MLAU-Net and various baseline methods. This figure is represented by the red color, which symbolizes the label, and the green color denotes the model's predictions. Column 1 depicts the input image, column 2 shows the ground truth mask, and columns 3–9 illustrate results of different baseline models. The last one gives an output for MLAU-Net predicted thus.
The suggested framework consistently outperforms the competing models, even in scenarios involving missing boundaries and shape variability. The boundary of the target area appears indistinct in U-Net and its variants. In contrast, the proposed approach demonstrates resilience to noise and other factors in US images, resulting in segmentations that closely align with the mask.
Quantitative analysis
Numerous kidney segmentation techniques have recently been developed and implemented across various studies. To assess the performance of these methods in comparison to our proposed approach, we conducted a thorough quantitative analysis using metrics such as dice, specificity, accuracy, HD95, recall, and ASSD. 48 The results of this comprehensive evaluation are presented in Table 4, which offers a quantitative comparative analysis of MLAU-Net and seven state-of-the-art segmentation approaches. All the ablation experiments were conducted in a consistent environment using our collected WD-KUS dataset, and in Table 5, a comprehensive assessment of performance metrics was conducted on an Open Kidney Dataset in a consistent experimental environment. The table showcases the results of ablation experiments, highlighting the efficacy of MLAU-Net against established segmentation methods.
Quantitative analysis comparing MLAU-Net with seven state-of-the-art segmentation approaches. A comprehensive assessment of performance metrics was conducted on our collected WD-KUS dataset in a consistent experimental environment. The table showcases the results of ablation experiments, highlighting the efficacy of MLAU-Net against established segmentation methods.

Quantitative study of MLAU-Net against SOTA techniques using an open kidney dataset.

For quantitative analysis, comparative experiments were conducted with seven widely employed segmentation methods, specifically AttU-Net, 43 Seg-Net, 17 FCNN, 51 U-Net, 13 SDFNet, 50 SwinUnet, 52 and LinkNet. 49 To ensure a fair and unbiased comparison, each method underwent complete training, and its segmentation results were not subjected to any post-processing. The segmentation accuracy of U-Net, SegNet, SwinNet, and SDFNet does not match that of FCNN, but their results exhibit superior visual effects. Conversely, FCNN, Linknet, and AttUnet display less favorable visual outcomes, characterized by noticeably jagged contours, suggesting a weakness in the learning ability of these methods, particularly along the kidney's edge. Two conjectures arise: the methods struggle to extract finer kidney features comprehensively, and substantial loss of kidney information occurs during deconvolution. Through a comprehensive analysis of evaluation matrices, significance tests, and segmentation results across various networks, our proposed MLAU-Net demonstrates a significant competitive advantage. Notably, it reduces false and missed detection rates on the WD-KUS and datasets. Computational performance trade-offs are illustrated in Figure 9 by comparing accuracy against the number of parameters for various baseline models and the suggested MLAU-Net on the WD-KUS dataset.

The proposed model effectively balances performance efficiency and computational cost, showcasing the lowest number of parameters among all baseline approaches. Presetting these parameters enables users to tailor computational resources and choose the appropriate encoder–decoder for feature extraction, depending on their specific requirements in MIS.
To enhance the assessment of segmentation methods on KUS images, we generated curves for accuracy, dice, recall, and specificity, as depicted in Figure 10. The visual representation emphasizes the superior performance of our approach to others, underscoring its aptness for WD-KUS image segmentation.

Proposed approach performance evaluation curves for accuracy, dice, recall, and specificity.
In addition to assessing traditional metrics, we conducted a comprehensive performance comparison between different models using HD95 and ASSD, as shown in Figure 11. The evaluated networks included AttU-Net, LinkNet, SDFNet, FCNN, U-Net, SegNet, SwinUnet, and the proposed MLAU-Net. Significantly, the suggested framework outperformed all others across HD95 and ASSD metrics, affirming its exceptional segmentation accuracy and dice for both datsets.
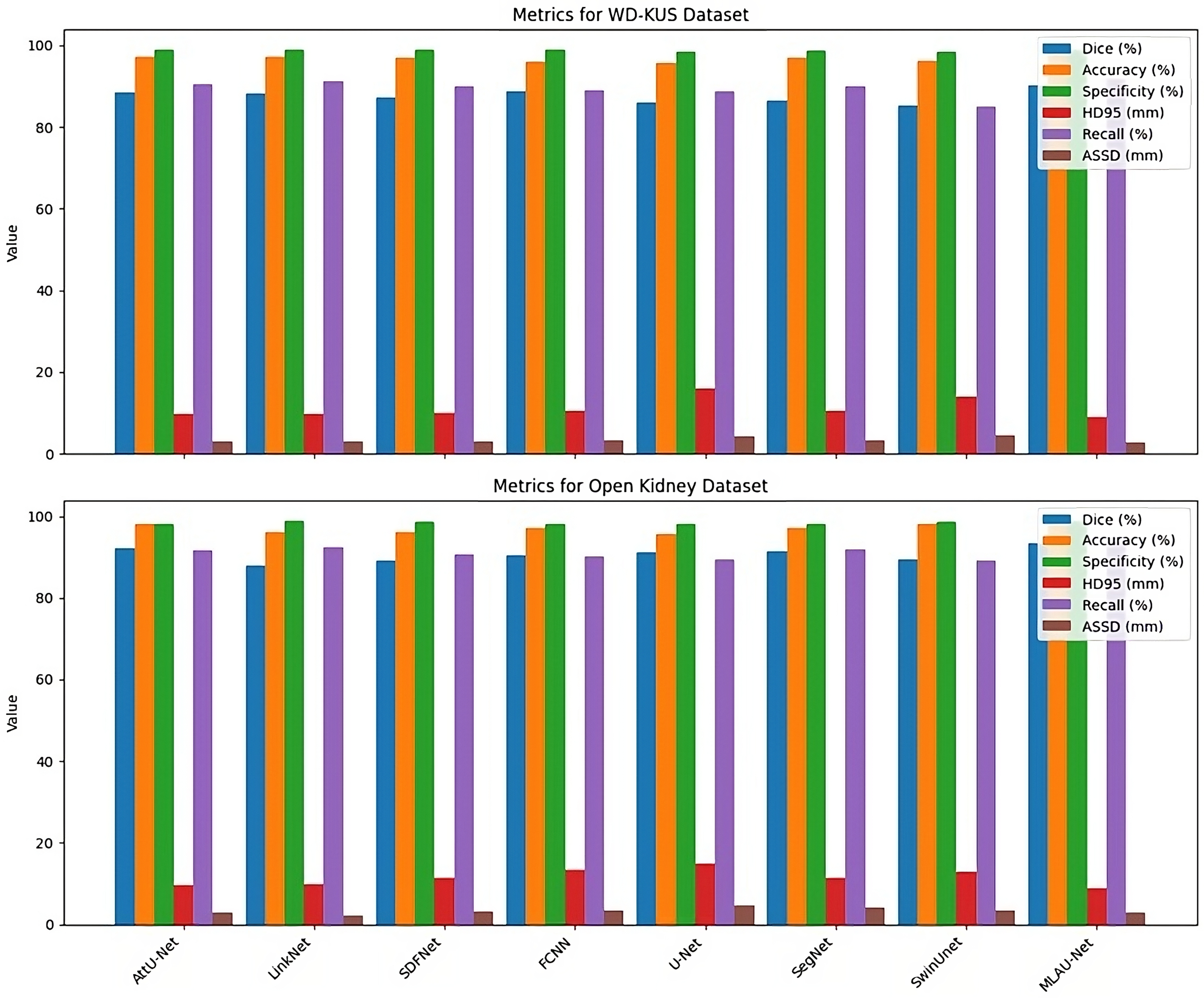
Quantitative analysis of models using segmentation metrics for different SOTA models.
Through qualitative and quantitative analyses of our proposed framework, we demonstrate the efficacy of each designed component. Comparisons with state-of-the-art segmentation methodologies reveal that our suggested network consistently outperforms competitors across six widely used evaluation indicators for two different datasets, as displayed in Figure 12.

Performance comparison of MLAU-Net across two datasets.
Despite instances of false and missed detection in the segmentation outcomes, our method demonstrates impressive performance compared to alternative approaches. Our method exhibits enhanced robustness to WD-KUS images, showing resilience against various influencing factors. In conclusion, the segmentation approach presented in this study effectively addresses the challenges associated with the automatic segmentation of the WD-KUS dataset, marking a substantial advancement in this domain.
Ablation experiments
In order to assess the effect of each component in the proposed MLAU-Net framework, we performed a detailed ablation study on three major modules: hybrid loss, deep supervision and attention mechanism. K-fold cross-validation with
In the second experiment, we eliminated deep supervision, leaving us with lower performance measurements that highlighted its contribution to reducing FNs and improving the segmentation accuracy. The last one turned off an attention mechanism, which brought about evident worsening in HD95, especially ASSD, showing how it enhanced boundary delineation and reduced segmentation errors. Table 6 presents average figures (mean ± SD) for some essential performance metrics obtained via the
Ablation study of MLAU-Net with K-fold cross-validation results for different modules.

Computational complexity analysis
The comparative analysis proposed study compared the computational complexity levels for MLAU-Net with other state-of-art models in terms of parameters and floating-point operations per second. This comparison is important as it determines how well a model performs in relation to its computational efficiency. The segmentation models that we evaluated were AttU-Net, Seg-Net, FCNN, U-net, SDFNet, SwinUnet, and LinkNet. We trained all these methods from scratch and fairly assessed their segmentations without post-processing. As shown in Figures 7–10 and Tables 4–5, our results demonstrate that MLAU-Net achieves better segmentation performances with less parameter numbers and lower computational budget. Table 4 highlights that MLAU-Net outperforms U-Net and FCNN on the WD-KUS dataset, with a Dice Score of 90.21%, while maintaining fewer parameters (Column 3) than other models. Similarly, Table 5 presents the results on the Open Kidney Dataset, where MLAU-Net also achieves a Dice Score of 93.43%, surpassing state-of-the-art models in accuracy and computational efficiency. This proves the effectiveness of MLAU-Net, significantly reducing false alarms and missed segmentations on both the WD-KUS dataset and public datasets. Quantitative comparisons are made in Table 4 (WD-KUS) and Table 5 (Open Kidney Dataset), while Figure 7 provides qualitative analysis for the WD-KUS dataset and Figure 8 for the Open Kidney Dataset. The performance evaluation curves in Figure 9 further confirm MLAU-Net's ability to balance accuracy and computational cost, reinforcing its robustness in segmentation tasks across multiple datasets.
Analysis of segmentation challenges in MLAU-Net
While MLAU-Net achieves overall significant performance, here are certain cases when the situation is different. This includes images with very low signal-to-noise ratio and those that differ significantly from what was used for training. In such situations, the attention mechanism that is responsible for highlighting crucial details might find it difficult to distinguish between noise and subtle renal contours. Furthermore, the deep supervision part may not completely overcome poor contrast, thereby making it difficult to draw exact boundaries. In future research, more advanced techniques of denoising or adversarial training can be studied in order to make handling of such images more robust.
Dealing with small or nonuniform kidneys is another problem. The model could encourage larger kidneys over others because of its inherent class imbalance in the data. Research into more advanced data augmentation methods and loss functions that are class-weighted might help improve the ability of neural networks to interpret kidney shapes and sizes.
Moreover, anatomical variations or artifacts like renal cysts, stones, or US beam distortions may cause uncertainties, leading to less effective segmentation results. The proposed approach often struggles to distinguish the target from the background in these cases accurately. For example, stone shadows may obscure important structures, distortions or artifacts can make shapes unclear, and irregular anatomical structures complicate the segmentation process. Addressing these issues may require incorporating domain-specific knowledge and multi-modal information. In Figure 13, we visualize some challenging cases where MLAU-Net fails to accurately separate the target from the background due to these factors.
Segmentation challenges in different scenarios: (Row 1) Stone shadow obscuring the target, (Row 2) Unclear shapes caused by distortions or artifacts, and (Row 3) Irregular kidney shape due to anatomical variations
Conclusion
Automatic segmentation of human KUS images is essential in helping urologists diagnose and treat kidney diseases in clinical practice. Nevertheless, factors such as the image quality, kidney morphology, and heterogeneous structures present challenges for accurate and automatic segmentation. In this study, we introduced MLAU-Net, a novel framework that leverages well-controlled attention mechanisms and a hybrid loss strategy to enhance the segmentation of low-resolution renal US images. Key components of MLAU-Net, including attention gates, deep supervision, and a meticulous preprocessing pipeline, significantly improve existing methods. Our results demonstrate that MLAU-Net excels in producing accurate segmentations thanks to its advanced features. Including attention, gates ensure that the model focuses on critical regions, while deep supervision aids in refining segmentation outputs. This makes MLAU-Net a valuable tool for precise medical image analysis and diagnosis, addressing the inherent challenges of low-resolution renal US imaging. In addition, through performing more evaluation on MLAU-Net using Open Kidney Dataset which is an open access dataset used to further confirm its ability. According to results, regardless of various metrics such as dice coefficient, accuracy, specificity, HD95, recall or ASSD this technique outperforms all modern techniques. This extensive evaluation underscores the robustness and generalizability of MLAU-Net in different clinical scenarios. The proposed methodology, incorporating domain information integration and weighted feature fusion, yielded superior results, particularly for segmenting malignant masses. MLAU-Net demonstrates high accuracy and efficiency in KUS image processing and holds promise for broader applications in other MIS tasks. In the future, we aim to refine MLAU-Net by incorporating specific pig KUS data and CT scan results as more data becomes available. This continuous improvement will increase its applicability and effectiveness, overcoming difficulties of low signal-to-noise ratios and weakly contrasted boundaries between objects. Developing MLAU-Net further presently aims at producing an all-encompassing solution towards MIS thereby facilitating improved clinical practice diagnosis and treatment planning.
Footnotes
List of abbreviations
Acknowledgment
This study was supported by the Guangzhou Science and Technology Project (202201020535), Guangzhou Medical University (2024SRP077), the National Natural Science Foundation of China (82100805), Guangzhou Science and Technology Planning Project (202102021129), and the Educational Commission of Guangdong Province (2022ZDJS113). The authors also acknowledge the support from Wuerzburg Dynamics Inc. for the Weiding Joint Laboratory of Medical Artificial Intelligence at Shenzhen Technology University.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The authors state that they have no known competing financial interests or close personal ties that could have influenced the research work presented in this paper.
Ethical approval
This study was conducted in accordance with the ethical guidelines and approval of the Institutional Ethics Committee (IEC) at the First Affiliated Hospital of Guangzhou Medical University (No. ES-2024–046-02).
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China, The Educational Commission of Guangdong Province of China, the School-Enterprise Cooperation Fund provided by Wuerzburg Dynamics Inc. to the Weiding Joint Laboratory of Medical Artificial Intelligence, Shenzhen Technology University, Guangzhou Science and Technology Project, Guangzhou Science and Technology Planning Project, Guangzhou Medical University Scientific Research Capacity Improvement Program (grant number: 82100805, 2022ZDJS113, 202201020535, 202102021129, and 2024SRP077).
Guarantor
Rashid Khan and Bingding Huang.
Informed consent
All study participants provided written informed consent or had their legally authorized representatives do so prior to the initiation of the study. The study design and informed consent procedures were reviewed and approved by the Institutional Ethics Committee at the First Affiliated Hospital of Guangzhou Medical University.
