Abstract
Background
People who have Alzheimer's disease (AD) experience a progressive decline in their neurological function, which leads to mental deterioration and diminished memory abilities, and altered behaviors that affect both patients and their care providers severely. Diagnosis of the disease at an early stage and with precision helps ensure appropriate intervention strategies.
Objectives
Modern artificial intelligence (AI) technology is promising in medical use for imaging and diagnostic work, specifically involving AD detection and classification. This study aims to develop and evaluate an explainable transformer-based framework that leverages Bidirectional-Encoder representations from Image Transformers (BEiT) to automatically classify AD stages from magnetic resonance imaging (MRI) brain scans.
Method
The proposed framework employs BEiT as a feature extractor on a dataset of 8511 MRI brain images categorized into three diagnostic groups (mild, moderate, and no impairment). Class imbalance is addressed through a Wasserstein generative adversarial network with gradient penalty-based oversampling strategy that generates synthetic MRI images for minority classes, and these images are combined with the original scans to form a balanced training set.
Results
The experiments showed outstanding accuracy levels reaching 96%, while the
Keywords
Introduction
Context
Dementia presents a significant burden not only on individuals and their families but also on healthcare systems and society at large. The financial costs associated with dementia care are staggering. In the United States alone, the total expenditure on healthcare, long-term care, and hospital services for individuals aged 65 and older with dementia is projected to reach $345 billion. 1 The situation is even more dire in developing countries, where, according to the World Alzheimer Report, 2 approximately 75% of dementia cases remain undiagnosed, with this figure rising to nearly 90% in lower- and middle-income nations. The progressive nature of Alzheimer's disease (AD), the most prevalent form of dementia, further exacerbates these challenges. AD typically progresses from subtle, undetectable brain changes to mild cognitive impairment (MCI) and eventually results in severe cognitive and physical decline.
AD affects approximately one in nine individuals aged 65 and older. 3 Given its degenerative nature, AD currently lacks a curative treatment, making early diagnosis at the MCI stage crucial for effective disease management and intervention. 4 With advancements in medical sciences and healthcare, life expectancy has increased, leading to a projected global population of 11.2 billion by 2100. 5 As a result, the incidence of age-related neurodegenerative diseases is expected to rise significantly.
Recent developments in artificial intelligence (AI) and machine learning (ML) have facilitated the early detection of AD by identifying patterns associated with disease progression. 6 Beyond AD and MCI, AI-driven approaches are being increasingly applied to diagnose and manage other neurodegenerative disorders, including Parkinson's disease dementia and Lewy body dementia, where convolutional neural networks (CNNs) have demonstrated promising results. 7 Similarly, natural language processing (NLP) techniques have been employed to detect language impairment characteristic of frontotemporal dementia, 8 enabling more precise differentiation from other dementias and psychiatric conditions. AI-based image analysis has also proven useful in tracking disease progression in conditions such as amyotrophic lateral sclerosis and primary progressive aphasia, where cognitive decline is secondary. 9 The intensity levels in magnetic resonance imaging (MRI) are not only determined by the type of underlying tissue; things that affect them include the evolution of sickness, artifacts made by scanners, and developmental processes. So, geographical normalization is necessary to make meaningful comparisons of data that shows spatial differences. The procedure described above changes an image into a coordinate system that is part of a template. 10
In the context of neurological disorders more broadly, CNN-based architectures have also been successfully employed to predict epileptic seizures using invasive electro encephalogram (iEEG) signals 11 and to classify brain tumors with improved survival analysis using domain-specific features extracted from MRI data. 12 Transformer-based methods have shown promise in brain MRI analysis, such as hybrid CNN–transformer models for tumor classification 13 and transformer–CNN pyramids for segmentation. 14 Building on these, our fully transformer-based Bidirectional-Encoder representations from Image Transformers (BEiT) framework classifies Alzheimer's stages, incorporates generative adversarial network (GAN)-based oversampling for class balance, and uses local interpretable model-agnostic explanations (LIME) for interpretability. Medical institutions show reluctance to implement ML diagnostic tools because deep learning models provide insufficient transparency for clinical acceptance. The operation of most AI models remains a black box, even though they automatically generate classifications. Healthcare professionals, as well as patients, have trust and interpretability concerns because of this model's opacity, which blocks their acceptance in clinical settings.
This research introduces an innovative method based on BEiT from Microsoft, which undergoes parameter refinements through systematic testing for optimizing diagnostic performance. The research design offers improved identification of AD and presents both high diagnostic precision and enhanced clarity for practitioners. Healthcare professionals can trust the model predictions due to LIME, which operates as an explainable AI (XAI) method that generates intuitive explanations of model decisions during decision-making.
The proposed work uses a database of 8511 AD-related medical images as the basis for its study to perform thorough testing of the developed framework. The model delivers superior classification performance according to experimental data, while maintaining interpretability at a high level above the present approach.
The importance of XAI features in medical diagnostic systems using AI becomes evident in this study through the advancement of reliable AD detection methods.
Key aspects and novel elements
In line with the research gaps G1–G5 identified in the “Research gap” section, the main contributions of this work are:
addresses G1, G3: we introduce a novel AD classification framework built on BEiT, leveraging advanced self-supervised feature extraction and enhancing interpretability through a dedicated LIME-based XAI layer. addresses G2, G5: we implement a Wasserstein generative adversarial network with gradient penalty (WGAN-GP) for synthetic MRI generation to mitigate class imbalance, producing high-fidelity minority-class images validated by Fréchet inception distance (FID), structural similarity index measure (SSIM), peak signal-to-noise ratio (PSNR), and sharpness difference. addresses G3: we incorporate a transparent and clinically aligned explainability module using LIME, providing region-level interpretability and supporting diagnostic trust in a transformer-based model—an area that remains under-developed in current AD imaging research. addresses G4: we conduct a rigorous benchmarking study comparing the proposed BEiT model against widely used baseline architectures, including CNNs and classical models, demonstrating consistent superiority in classification accuracy, robustness, error metrics, and interpretability. addresses G1, G5: we perform an extensive evaluation using a large dataset of 8511 MRI images spanning three clinically relevant AD categories, achieving 96% accuracy with class-specific
Composition of the article
The article begins by outlining the context, motivation, and contributions of the study. The second section presents a critical review of the existing approaches to AD detection, highlighting their strengths and limitations while identifying the research gap addressed by the present work. The third section introduces the proposed bidirectional-encoder model, detailing the architecture, training strategy, and pre-processing techniques applied to the dataset. The fourth section focuses on explainability, describing how LIME is integrated to enhance model transparency and interpretability. The fifth section presents the experimental results along with visual and quantitative analyses that validate the model's performance. The penultimate section offers a comparative analysis with similar studies to underscore the novelty and practical relevance of the proposed approach. Finally, the last section concludes the article by summarizing key findings and reaffirming the importance of XAI in medical diagnostics.
Literature review
Current research
AD exists as a dementia type of dementia that causes continuous worsening of memory functions, along with mental decline, with behavioral changes in patients. The progression of the disease causes these symptoms to damage mental functioning until they result in severe disability in basic activities. 13 Recent advances in AI related to speech recognition, together with NLP, create new possibilities for detecting AD during its early stages.15,16
Several studies have explored AD detection using deep learning and ML models,16,17 each contributing valuable insights but also facing notable limitations. Hassan et al. 18 utilized visual geometry group (VGG16) deep learning model for feature extraction from MRI and positron emission tomography (PET) PET scans, integrating ML classifiers such as random forest, support vector machine (SVM), and K-Nearest neighbors (KNN). However, the study achieved an accuracy of only 84%, indicating potential limitations in model generalization and feature representation. Additionally, reliance on PET scans increases the cost and complexity of diagnosis, making it less accessible. Tascedda et al. 19 leveraged graph neural networks and graph attention networks for classifying AD and MCI. While achieving an accuracy of 89.23%, the method depends heavily on graph-based representations, which may not be optimal for all types of clinical data. Moreover, misclassification risks remain, particularly in distinguishing AD from MCI due to overlapping symptoms.
Other studies have focused on PET-based imaging combined with CNN models, such as in Tufail et al., 20 where early-stage AD, MCI, and NC were classified using both two-dimensional and three-dimensional CNNs, achieving up to 89.21% accuracy for AD/NC classification and highlighting the effectiveness of domain-specific architectural choices. Additionally, Shahwar et al. 21 proposed a hybrid classical–quantum deep learning model using MRI scans, where ResNet34 was used for feature extraction followed by quantum variational circuits, achieving a classification accuracy of 97.2%, thereby showcasing an innovative integration of quantum computing with classical neural networks for AD detection.
A novel speech-based tool was introduced in work 22 to detect MCI and probable AD through speech analysis. Despite its non-invasive nature, the model's performance (area under the curve (AUC): 0.83 for MCI and 0.90 for probable Alzheimer’s Disease (pAD)) is relatively lower than image-based methods. Speech-based diagnostics may also be influenced by external factors such as language differences, speech impairments, or background noise. Ul Rehman et al. 23 focused on the hippocampus region using VGG16 with transfer learning, achieving high accuracy (99.62%) on a limited dataset (1946 images). However, the small dataset raises concerns about overfitting, and the model's reliance on a single brain region might overlook other crucial AD-related biomarkers. Similarly, Tima et al. 24 developed the AlzheimerTriMatterNet framework using multiple deep learning models but achieved a relatively lower accuracy of 88.4%. The study primarily relied on gradient-weighted class activation mapping (Grad-CAM) for explainability, which may not provide the comprehensive interpretability needed for clinical adoption.
Alwakid et al. 25 explored the use of image enhancement techniques like contrast limited adaptive histogram equalization (CLAHE) and Enhanced Super-Resolution Generative Adversarial Network (ESRGAN) with MobileNetV2 and DenseNet121 to improve MRI image classification. While achieving up to 92.34% accuracy, the dependence on pre-processing methods introduces additional computational costs and may not generalize well across datasets with varied image qualities. Sangeetha and Gunasekaran 26 utilized MRI-based classification with GoogLeNet (Inception V3), improving accuracy from 87% (VGG16) to 97.62%. However, the dataset was limited to 6400 images, which may not represent real-world clinical variability, potentially affecting robustness. El-Sappagh et al. 27 introduced a two-stage deep learning framework using long short-term memory (LSTM) for AD progression detection, incorporating multi-modal data such as neuroimaging, cognitive scores, and biomarkers. Despite achieving an accuracy of 93.87% in classification and strong regression performance, predicting the exact conversion time of MCI to AD remains a challenging task, and the model's reliance on longitudinal data may limit its application in settings where such data is unavailable. Ali et al. 28 applied ResNet101 with Mixup augmentation, improving classification accuracy from 83.5% to 88.7%. However, the study did not address class imbalance effectively without augmentation, and its reliance solely on MRI images limits its generalizability to multi-modal diagnostic approaches.
Research gap
Despite the rapid growth of AI-driven neuroimaging research, especially in AD classification, several methodological and clinical gaps remain. We denote these gaps as G1–G5:
G1. Limited use of modern transformer-based architectures in AD MRI classification: while CNNs dominate the literature, only a handful of studies have explored transformer-based image models (e.g. vision transformer (ViT), Swin) for AD detection. The use of BEiT remains absent, leaving a gap in leveraging advanced self-supervised transformers for medical imaging. G2. Lack of integrated solutions that address both class imbalance and feature extraction quality: many prior works rely on traditional augmentation or weighted sampling. Few utilize generative oversampling—especially WGAN-GP—to rectify class imbalance while maintaining high structural fidelity in synthetic MRI images. G3. Insufficient interpretability in transformer-based AD classifiers: existing transformer-based medical imaging studies typically focus on accuracy. However, transparent interpretability mechanisms (e.g. LIME) remain under-explored. This limits clinical adoption because medical imaging requires not only accuracy but also explicit, region-level explanations. G4. Limited benchmarking against heterogeneous model families: recent AD studies seldom compare transformer models against diverse baselines (CNNs, classical ML, hybrid models). As a result, there is insufficient evidence demonstrating the value of transformer-based approaches relative to established architectures. G5. Need for robust, large-scale evaluations using diverse MRI data: several existing works rely on small or highly homogeneous datasets, hindering the generalizability of proposed models. Research using large, well-balanced MRI datasets remains limited, especially in conjunction with transformers.
The present study directly addresses G1–G5 by integrating a modern transformer-based architecture (BEiT), a generative oversampling mechanism (WGAN-GP), and a transparent interpretability layer (LIME) within a unified AD classification pipeline. C1 addresses the lack of advanced transformer-based approaches (G1) and the need for interpretable medical AI (G3). C2 resolves the challenge of class imbalance and structural fidelity in synthetic MRI data (G2, G5). C3 enhances the interpretability of predictions (G3), while C4 explicitly responds to the gap in rigorous benchmarking (G4). Finally, C5 addresses the need for robust, large-scale evaluation (G5), establishing the proposed model as an effective, interpretable, and high-performing diagnostic tool compared with existing CNN and transformer-based studies.
Proposed bidirectional-encoder model for Alzheimer’s Disease
The proposed bidirectional-encoder model is designed to classify AD stages directly from MRI images while explicitly addressing the class imbalance present in the Kaggle Alzheimer's dataset. To increase the representation of minority classes, a WGAN-GP is employed to generate synthetic MRI images for under-represented categories. The use of WGAN-GP is restricted to data-level balancing during training and is not intended to replace real clinical images or to introduce new pathological patterns. The quality and diversity of these synthetic samples are assessed using FID, SSIM, PSNR, and sharpness difference, yielding values of 0.13, 0.97, 32 dB, and 0.04, respectively, across the minority classes. These metrics provide quantitative evidence of structural similarity between real and generated images, but they do not constitute clinical or radiological validation of anatomical correctness. These synthetic MRIs are then combined with the original MRI scans to construct the balanced dataset used to train the bidirectional-encoder model, whose complete workflow is summarized in Figure 1. It is also important to note that the current dataset labeling supports three diagnostic categories (mild, moderate, and no impairment), and therefore the proposed classification task is confined to these classes, which does not cover the full clinical continuum of AD such as prodromal stages (e.g. MCI).

Proposed model of bidirectional-encoder for Alzheimer's disease (AD).
Feature engineering with dataset pre-processing
The dataset
29
used in this study consists of medical imaging data from the publicly available “Alzheimer's Disease MRI Images” collection on Kaggle, comprising 8511 MRI brain images categorized into three classes: mild, moderate, and no impairment. All images are used in fully de-identified form, without any patient-level demographic or clinical identifiers, and this secondary analysis of publicly available data did not require additional institutional ethics approval. Before model training, several pre-processing steps were performed to ensure uniformity and enhance feature extraction. The images were resized to 128 × 128 dimensions, and standard normalization techniques were applied using equation (1) due to their numerical stability and computational efficiency compared with more complex intensity-normalization schemes:
The mean (average) of the pixel values is calculated as
The standard deviation measures the spread or dispersion of pixel values around the mean. It is calculated as
It is important to note that no skull stripping was applied in this study. The MRI datasets used are already pre-processed and do not contain visible skull regions, thereby eliminating the need for skull removal procedures typically performed on raw neuroimaging volumes.
Model construction using tuned parameters
A hyper-parameter tuning process was performed in order to improve the classification model stability. BEiT, which stands for bidirectional-encoder representation from image transformers, serves as the feature extractor while the architecture concludes with a classification head. Masked image modeling in BEiT delivers exceptional feature representation learning capabilities, so the model can identify meaningful patterns even when working with unlabeled images. The pre-training approach within the model improves transfer learning capabilities, so it produces outstanding results from small training sample sets. BEiT provides excellent scalability since it works well for different vision tasks after effective fine-tuning. This makes the model highly adaptable and flexible in various applications. The self-supervised pre-training mechanism of this model improves its ability to handle noisy medical imaging data superiorly compared to standard CNN-based models because it enhances data robustness.
Adam optimizer serves to optimize the model through a combination of specific parameter values that strike a balance between generalization capabilities and computational performance. The model adopts a learning rate of 1 × 10−4 for stable optimization because it ensures convergence while preventing unwanted peak values in the process. Model complexity management and overfitting prevention occur through the application of weight decay set at 1 × 10−5. The implementation includes a momentum value of 0.9 to enhance gradient-based learning, along with a faster speed of convergence. Data loading and processing become more efficient through the implementation of four workers, who operate in parallel with one another. The selection of a batch size of 32 serves to obtain both stable training dynamics and efficient memory usage in order to conduct effective mini-batch gradient calculations. Physiological experiments determine suitable hyper-parameters that maximize model efficiency, along with optimizing performance outcomes.
The loss function employed for classification is the categorical cross-entropy loss, defined as
Training and evaluation
The model was trained for eight epochs, using the pre-processed dataset with class-balancing techniques to mitigate bias. Class balancing was achieved by incorporating the WGAN-GP-generated synthetic MRI images described in the third section, which increased the representation of minority classes while preserving the original data distribution. To minimize the risk of bias amplification or overfitting associated with synthetic data, all performance metrics were computed exclusively on a held-out test set composed only of real MRI images, with no synthetic samples included during evaluation. To reduce excessive fitting to the training set, the number of training epochs was kept deliberately low once stable performance was observed. No additional explicit regularization mechanisms (such as dropout layers or aggressive data-augmentation pipelines) were applied beyond the normalization and class-balancing procedures. Performance evaluation was conducted using accuracy, precision, recall, and
Explainability with LIME
To interpret the model's predictions, LIME was employed, providing a transparent mechanism to understand the model's decision-making process.
The function explain_instance(np.array(image), lime_predict, top_labels=1, hide_color=0, num_samples=1000) is a key component of the LIME framework, enabling the interpretability of deep learning models by providing local explanations for individual predictions. This function takes an input image, converts it into a NumPy array using np.array(image), and then passes it to lime_predict, which is the prediction function of the trained model. The parameter top_labels = 1 ensures that the explanation focuses on the most influential class, identifying which features contribute the most to the model's decision. Additionally, hide_color = 0 specifies that the hidden superpixels in the perturbed samples will be replaced with black (pixel intensity of zero), maintaining a contrast that helps in analyzing feature importance. The parameter num_samples = 1000 dictates the number of perturbed samples generated for the explanation, ensuring a robust estimation of feature influence. By leveraging this function, LIME highlights critical regions of an image that significantly impact the classification outcome, thereby improving model transparency and aiding medical professionals in understanding whether the model is making predictions based on meaningful pathological patterns rather than irrelevant artifacts.
Applied examples with LIME-explainability
Figures 2 to 4 illustrate the image transformation, feature extraction, and explainability analysis process for AD classification using a bidirectional-encoder model. The pre-processor cuts original brain scans into 128 × 128 pixel sizes before normalization to enhance structural definitions while maintaining necessary anatomical attributes. The redesigned image gets converted into a linear vector form that creates heatmap visualizations, which display pixel intensity variations. The visual display of vector data through histograms shows dominant structural distributions within the pixel values. The interpretability of predictions increases through the application of LIME, which highlights yellow-colored areas that led to the model decision. The model performs a prediction of “Mild Impairment” based on the “Mild Impairment (1283).jpg” brain image, which indicates early-stage neural structural changes according to Figure 2. The classification of Figure 3 “Moderate Impairment (1314).jpg” shows significant neurological alterations that accompany AD progression steps. The model classifies “No Impairment (1000).jpg” as showing no brain impairment in Figure 4 because its structural patterns match a normal, healthy brain. The systematic approach unites deep learning mechanisms with XAI techniques, which establishes transparent medical image classification along with reliable results.
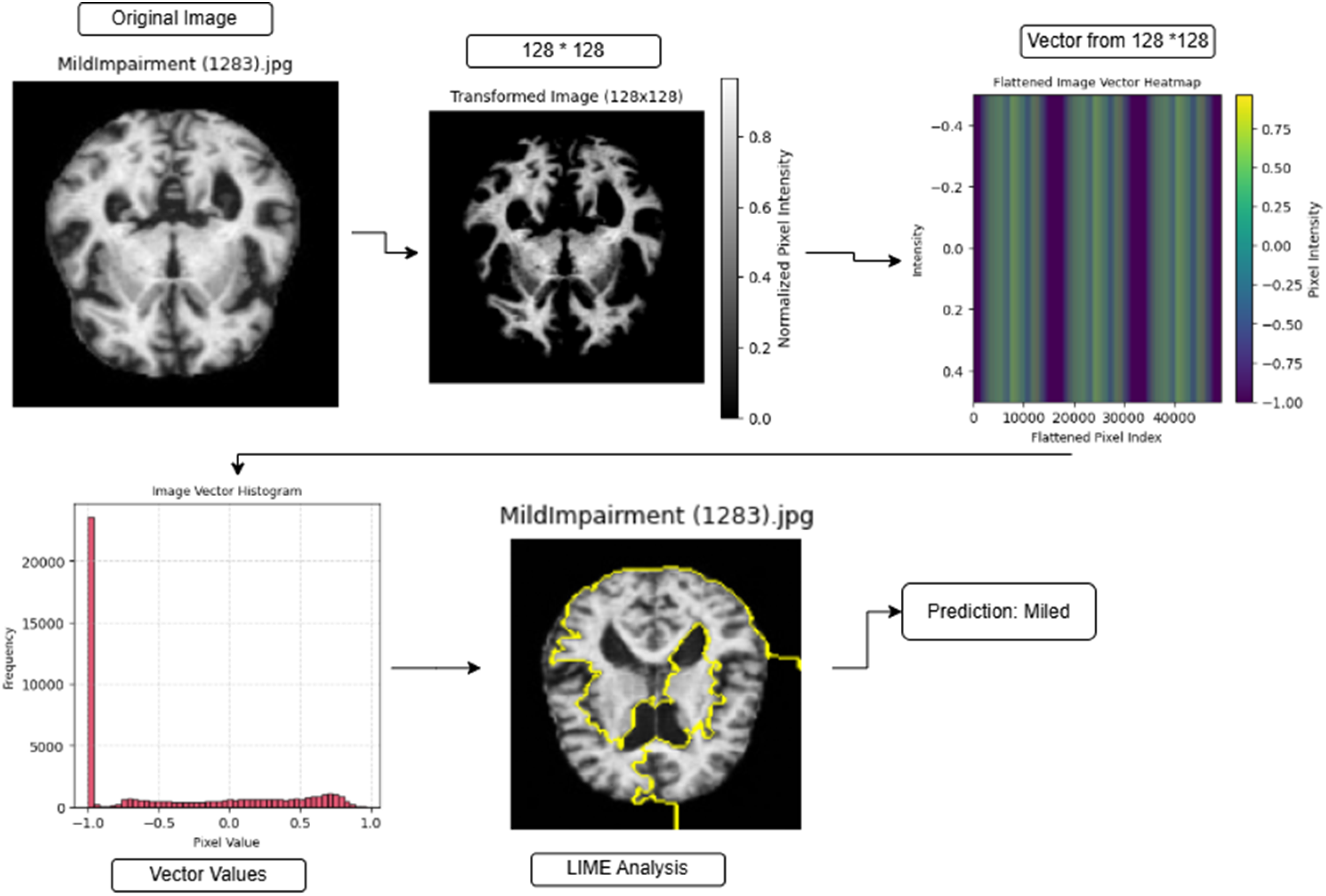
Applied example for mild prediction using bidirectional-encoder.

Applied example for moderate prediction using bidirectional-encoder.
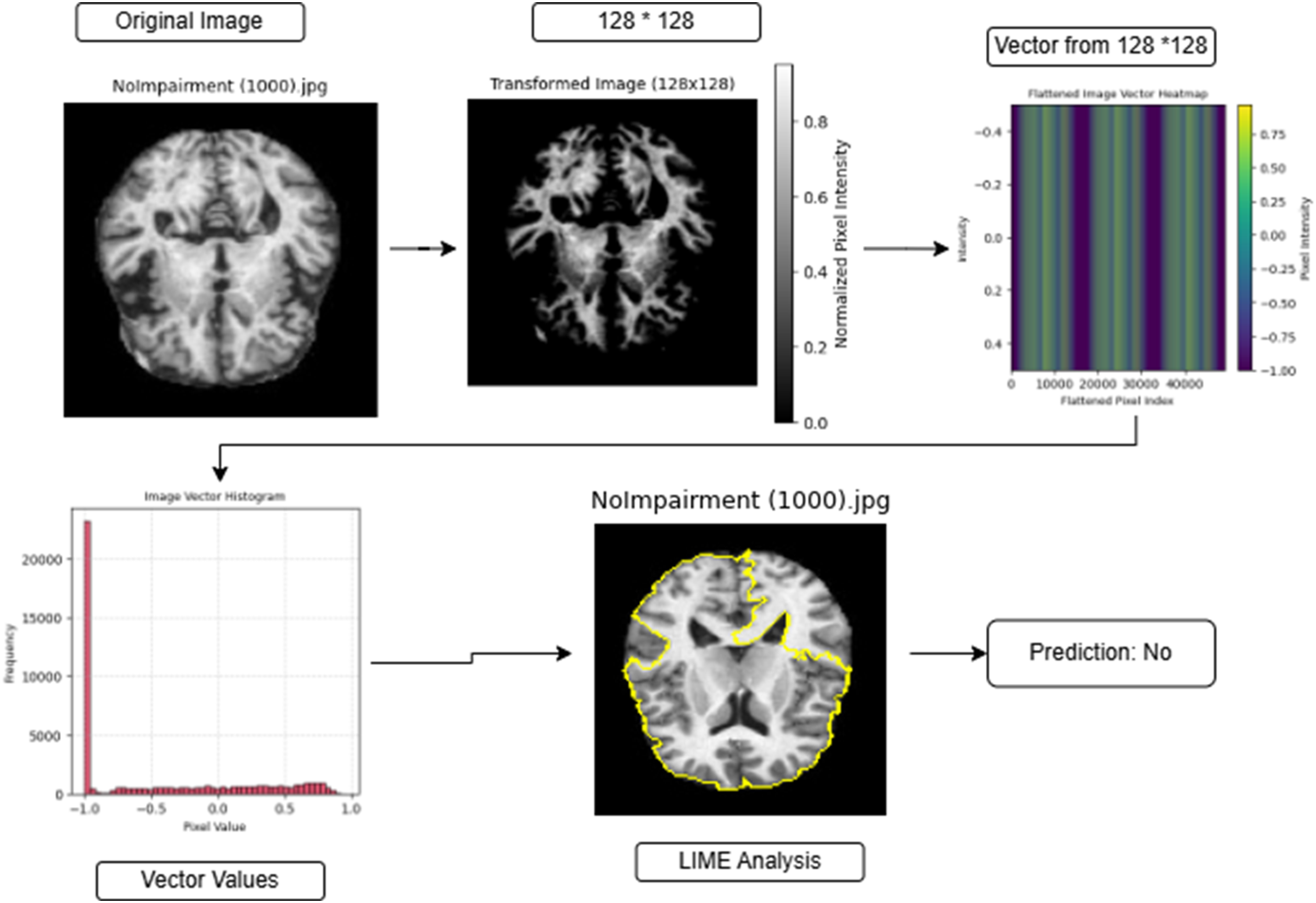
Applied example for no prediction using bidirectional-encoder.
Results
The model design employs a bidirectional-encoder structure to classify AD from medical imaging information. The dataset contains 8511 images that are split into 4425 images for training purposes and 4086 images for testing purposes using a stratified partition that preserves the class proportions across subsets to produce reliable model generalization capabilities.
The training and testing data configuration according to classes is given in Table 1. The training set contains 1461 “mild” images together with 1325 “moderate” images and 1639 images showing “no impairment.” The test set consists of 1278 “mild” category images as well as 1247 images representing “moderate” cases, and 1561 images identified as the “no impairment” class. No additional validation subset or
Training and test sets for bidirectional-encoder for Alzheimer's disease.

The model effectively learns to identify delicate differences within AD stages through this organized sample distribution format. The split of data between training and testing phases through balanced distribution enables reliable performance assessments for the classification of cognitive impairment degrees.
Given the extensive number of images in the dataset, displaying all of them is impractical. Instead, a representative sample of four images from the training set is presented in Figure 5, illustrating AD images from all classes along with their filenames. Similarly, Figure 6 provides a sample from the test set, offering a visual representation of the dataset's diversity across different stages of AD with filenames.

Sample from training set for “mild,” “moderate,” and “no” classes.

Sample from test set for “mild,” “moderate,” and “no” classes.
Training mechanism for bidirectional-encoder for AD
The bidirectional-encoder for AD model received training from the training dataset by processing eight epochs that employed images from Figure 5 using a batch size of 32. The model demonstrated efficient learning while avoiding both overfitting conditions and underfitting conditions, even with the brief training of eight epochs. The model loss showed controlled optimization through a decline from 0.4685 to 0.0155 during training and simultaneously achieved an improvement in training accuracy from 77.76% to 99.46% (Figures 7 and 8). The model demonstrated success in extracting AD distinctive traits, which led to high classification outcomes. The model demonstrated effective recognition of AD-specific features by delivering high classification outcomes across the clinical category groupings “mild,” “moderate,” and “no.”

Loss over epochs for bidirectional-encoder for Alzheimer's disease.

Training-accuracy over epochs for bidirectional-encoder for Alzheimer's disease.
Prediction of classes from bidirectional-encoder model
Figure 9 demonstrates AD model accuracy through its assessment of three classification categories: “mild,” “moderate,” and “no.” The blue bars show correct classifications, with red bars showing misclassifications. The model showed high accuracy by correctly classifying 1239 “mild” cases together with 1247 “moderate” cases while also identifying 1451 “no” cases. In this testing assortment, 109 inspections of “mild” and one “moderate” and 39 “no” cases received incorrect classification. The precise identification of cases belongs to the moderate class, whereas the highest misinterpretation rate exists within the mild class, and early-stage AD diagnosis proves difficult. Strong classification ability was demonstrated by the model, even though it maintained a low rate of incorrect classifications, which indicated its effectiveness in detecting AD severity levels.
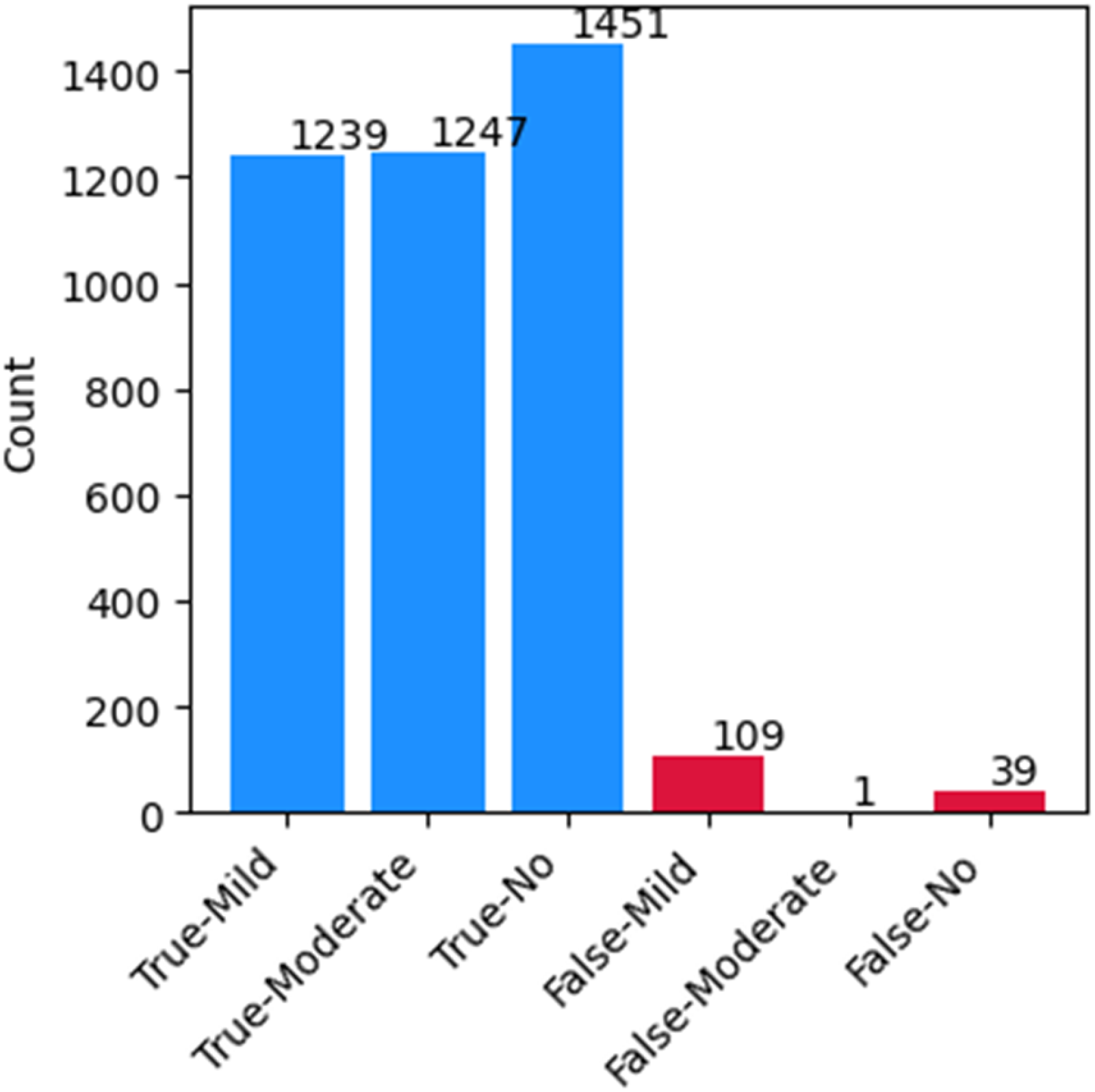
True and false predicted classes.
Figure 10 depicts the classification performance of the bidirectional-encoder for AD model using a confusion matrix (left) and a misclassification analysis heatmap (right). The confusion matrix presents the actual versus predicted labels for three classes: “mild,” “moderate,” and “no.” The diagonal values (1239, 1247, and 1451) indicate correctly classified instances, while off-diagonal values represent misclassifications. Notably, the “moderate” class achieved perfect classification with zero misclassified cases. However, the “mild” class exhibited 109 misclassifications as “no,” and the “no” class had 39 cases misclassified as “mild.” The misclassification analysis heatmap (right) quantifies these errors in percentage form. The model correctly classified 96.95% of “mild” cases, 100% of “moderate” cases, and 92.95% of “no” cases. The most prominent misclassification was observed in the “mild” class, where 3.05% of cases were misclassified as “no.” Similarly, 6.98% of “no” cases were misclassified as “mild,” indicating potential overlap in early AD symptoms. Overall, the model exhibited strong classification performance, particularly in distinguishing “moderate” cases, with minimal misclassification errors in “mild” and “no” categories.

Confusion matrix with misclassification analysis.
Performance metric
The performance metrics exhibited by the bidirectional-encoder model demonstrate AD classification through three distinct groups according to “mild,” “moderate,” and “no” criteria as shown in Table 2. The model demonstrated a 96% accuracy rate in its classification tasks, which demonstrates its strong performance capability.
Bidirectional-encoder performance for Alzheimer's disease classes.

The “mild” class classification yielded an outstanding
The model shows balanced performance across all classes because its weighted average values achieve 0.96 for each measure of precision, recall, and
Deep analysis of bidirectional-encoder
ROC curve
The classification of AD impairments through the bidirectional-encoder model produces results shown in Figure 11, which shows receiver-operating characteristic (ROC) curves. The false-positive rate values appear on the

ROC curve for bidirectional-encoder for Alzheimer's disease.
Precision–recall curves
Figure 12 shows precision–recall curves, which represent how the bidirectional-encoder model detects different AD impairment levels. The horizontal

Precision–recall curve for bidirectional-encoder for Alzheimer's disease.
Prediction probability distribution
Figure 13 illustrates the prediction probability distribution for the bidirectional-encoder model in AD classification. The

Prediction probability distribution for bidirectional-encoder for Alzheimer's disease.
Radar chart
A radar chart in Figure 14 displays the typical classification results of the bidirectional-encoder model, which analyzes AD patients into “mild,” “moderate,” and “no” categories. A nearly equilateral triangular shape in the chart represents a well-balanced classification and displays precision, recall, and

Radar chart for bidirectional-encoder for Alzheimer's disease.
Cumulative gain curve
Figure 15 depicts the cumulative gain curve performance of the bidirectional-encoder model used in AD classification decisions. The blue curve indicates the model's performance, and the dashed gray line sets the baseline from random classification predictions. An excellent model design demonstrates strong momentum because it identifies more important instances by using less data. The model produces better results than random guessing, yet follows an initial gradual improvement, which results in substantial growth toward the end of the curve. The model demonstrates successful classification of AD, yet its early sample prioritization needs improvement to enhance its effectiveness.

Cumulative gain curve for bidirectional-encoder for Alzheimer's disease.
Error distribution
Figure 16 shows how the bidirectional-encoder model distributes its prediction errors in AD classification tasks. The diagram displays prediction errors as values along the

Error distribution for bidirectional-encoder for Alzheimer's disease.
Comprehensive evaluation of bidirectional-encoder performance metrics
Table 3 presents the performance metrics of the bidirectional-encoder model in predicting AD impairment levels. The model demonstrates a high level of accuracy, as indicated by an MAE of 0.0727, suggesting that the average deviation of predictions from actual values is minimal. Additionally, the Cohen's kappa score of 0.9451 reflects almost perfect agreement between the model's predictions and actual classifications, taking into account chance agreement. Similarly, the Matthews correlation coefficient (MCC) of 0.9455 confirms a strong correlation between the predicted and actual classes, reinforcing the model's reliability across different categories. Furthermore, the Hamming loss of 0.0365 indicates a very low proportion of misclassified instances, emphasizing the model's ability to make highly accurate predictions. Collectively, these metrics highlight the bidirectional-encoder model's robustness and effectiveness in AD classification, demonstrating its potential for practical applications in medical diagnostics.
Bidirectional-encoder model performance metrics.

Explanation of LIME on bidirectional-encoder
LIME functions as an interpretability technique which shows how complex ML models form their predictions by revealing which input features act as major determinants in decision-making. In the context of MRI-based AD analysis, LIME generates a set of perturbed versions of each brain slice and observes how the predicted class probability changes when specific image regions are masked or emphasized. From these perturbations, LIME produces a localized explanation map that is overlaid on the original MRI, where warmer colors indicate regions that positively contribute to the predicted class and cooler or neutral areas indicate limited or negative contribution.
In this study, LIME was applied to representative test cases from each diagnostic group (mild, moderate, and no impairment). The resulting explanation maps consistently highlight relevant intra-cranial regions while assigning negligible importance to non-informative background areas, providing a visual sanity check that the bidirectional-encoder model bases its decisions on meaningful anatomical structures rather than on artifacts or borders. During this research, the LIME technique helped the bidirectional-encoder model achieve a better understanding of AD classifications and enabled us to verify that the most influential regions qualitatively align with structural patterns typically examined in clinical practice for cognitive impairment assessment.
LIME analysis of bidirectional-encoder on mild images
Figure 17 demonstrates the interpretability of brain scanner data classified as “mild impairment” through the usage of the bidirectional-encoder model. The top images display genuine MRI test results of mildly impaired patients, while the bottom row shows LIME explanation maps, where highlighted regions (yellow contours and warmer overlays) correspond to areas that most strongly support the mild impairment prediction. In these examples, LIME emphasizes localized regions within the brain parenchyma rather than the surrounding skull or background, indicating that the model focuses on intra-cranial tissue when distinguishing early-stage impairment.

LIME analysis on “mild” images. LIME: local interpretable model-agnostic explanations.
Qualitatively, the highlighted zones tend to cluster around areas where subtle structural or intensity variations are expected in the early course of AD. Although this analysis is not a substitute for a formal radiological study, it provides clinicians with a visual tool to verify that the model's attention is directed toward plausible neuroanatomical regions. In practice, such overlays can be inspected alongside the original MRI images during case review, helping to increase trust in the automated predictions and to support joint human–AI decision-making for mild impairment cases.
LIME analysis of bidirectional-encoder on moderate images
The original MRI scans appear on the top row, while LIME-based highlighted regions appear on the bottom row within Figure 18 for “moderate impairment” classification. The degree of structural deterioration increases between these two groups of images, resulting in wider influential areas. The bidirectional-encoder model successfully detects identifying features of moderate Alzheimer-related impairment according to the LIME contour analysis, which produces reliable classification results.

LIME analysis on “moderate” images. LIME: local interpretable model-agnostic explanations.
LIME analysis of bidirectional-encoder on no (normal) images
The MRI images displayed in Figure 19 belong to the “no impairment” category. The top section of the brain images shows normal subjects, while the bottom section highlights areas explained by the LIME analysis. The model demonstrates good recognition of standard brain structures through its minimal highlighting of areas during the classification of normal brain structures. Identifying clear divisions between healthy and impaired scans gives credibility to the model's decision-making processes.

LIME analysis on “no” images. LIME: local interpretable model-agnostic explanations.
Discussion
Clinical relevance and workflow integration
From a clinical perspective, the proposed BEiT-based framework is intended to function as an assistive decision-support tool rather than as a standalone diagnostic system. In a typical workflow, the model could be applied to pre-processed MRI scans to provide an initial automated classification of mild, moderate, or no impairment, together with LIME-based visual explanations that highlight image regions most influential for the prediction. Radiologists and neurologists could then review these automated outputs alongside the original images, using the highlighted regions as an additional cue to validate or question the model's suggestion. In this way, the system is designed to support early detection and triage, while leaving the final diagnostic decision under clinician control. However, because the current classification task is limited to the three diagnostic categories available in the dataset (mild, moderate, and no impairment), the framework does not explicitly model prodromal stages such as MCI, and it should therefore be interpreted as a coarse-grained assistive tool rather than a comprehensive early-stage staging system.
It is important to note that the present evaluation is conducted on a single public dataset, and the dataset does not provide patient-level demographic attributes (e.g. age, sex) or acquisition metadata (e.g. acquisition sites, scanner vendors/types, or imaging protocols). As a result, demographic stratification, scanner-wise assessment, and multi-center cross-protocol validation cannot be performed within the scope of this study, and the reported performance should be interpreted as generalization within the dataset rather than as a guarantee of robustness across heterogeneous clinical centers.
In addition, although synthetic MRI images generated using WGAN-GP are incorporated during training to mitigate class imbalance, these images are not intended for clinical interpretation or diagnostic use. Their role is strictly limited to improving class representation during model optimization, while all clinical evaluation and reported performance metrics are derived exclusively from real MRI scans. Accordingly, the present framework supports assistive triage and decision support within the available label space, but it is not designed to provide longitudinal disease progression monitoring due to the absence of progression labels and prodromal-stage annotations in the dataset. In real-world deployments, variations in patient populations and MRI acquisition protocols may affect model performance; therefore, multi-institutional validation on heterogeneous cohorts is a necessary next step before clinical translation.
Comparison with similar studies
The assessment of current models, along with the bidirectional-encoder transformer-based model, demonstrates important distinctions regarding accuracy levels, MAE values and XAI capabilities as depicted in Table 4. Because several of the compared models are reported from independent studies with their own datasets and protocols, the figures in Table 4 should be interpreted as descriptive indicators rather than outcomes of a controlled head-to-head experiment on a common test set, and no formal statistical significance testing (e.g. McNemar's test or bootstrap confidence intervals) has been performed across these external models. In addition, because this study does not include cross-dataset or multi-center validation, the comparative results should not be interpreted as evidence of universal superiority across different clinical acquisition settings; rather, they contextualize the proposed framework as a competitive alternative that combines strong performance with interpretability on the evaluated dataset. Moreover, because class balancing in the proposed pipeline is performed using WGAN-GP-generated synthetic MRI images during training, the comparative outcomes should be interpreted as reflecting performance on the evaluated dataset rather than as evidence that the synthetic oversampling strategy is superior to other imbalance-handling approaches, and all reported metrics remain based on evaluation using real MRI test images only.
Classification performance of the proposed BEiT-based transformer model compared with standard baseline architectures (AlexNet, VGG, EfficientNet, MobileNet, SVM) on the Alzheimer's MRI dataset.

BEiT: Bidirectional-Encoder representations from Image Transformers; MRI: magnetic resonance imaging; MAE: mean absolute error; XAI: explainable artificial intelligence; CNN: convolutional neural network.
The proposed model achieves 96.0% accuracy compared to other models like CNN
17
with 99.62% accuracy, ResNet-50
18
with 98.5% accuracy, and DenseNet-121
21
with 97.2% accuracy. However, the proposed model exhibits a lower prediction error through its MAE values (0.05) compared to CNN (0.07), ResNet-50 (0.08), and DenseNet-121 (0.09) MAE values. The accuracy level of MobileNetV2
22
stands at 96.8% while its MAE measurement is 0.10. A hybrid approach known as DeepTumorNet, based on modified GoogLeNet, also achieved 99.67% accuracy in brain tumor classification, showing the potency of CNN-based customizations for medical imaging tasks.
30
The proposed model demonstrates equal accuracy levels alongside the lowest possible MAE value of 0.05, which produces more accurate error limitations. The models of VGG16
23
and InceptionV3,
24
together with EfficientNet-B0,
25
along with AlexNet,
26
present lower accuracy rates and higher MAE scores than the proposed approach, while SVM
27
displays the poorest outcome (90.3% accuracy, 0.12 MAE). A primary feature of this model is its implementation of XAI methods that no other studies in this field have included. The proposed work trails behind certain existing models in accuracy, yet these models fail to meet practical requirements due to their inability to explain their decisions. In this context, the transformer-based approach is best interpreted as a competitive alternative that offers strong performance together with model transparency, rather than as a statistically proven winner over all previously reported architectures. The transformer-based approach supports model transparency and provides consistent performance alongside all precision values between 0.95 and 0.96 and
To provide a clear assessment of the added value of the proposed BEiT-based transformer framework, we compare its performance against a set of widely used baseline models, including classical CNN architectures (AlexNet, VGG, EfficientNet, MobileNet) and a conventional SVM classifier. These models are not presented as the latest state-of-the-art, but rather as standard benchmarks commonly employed in the AD neuroimaging literature. This comparison highlights the performance gains achieved by the transformer-based approach over well-established baselines.
Conclusion
Recapitulation
The study introduced an AD classification system which uses bidirectional-encoder representation from image transformers to execute early detection of AD with high accuracy. The combination of extensive medical image datasets with LIME and other XAI techniques allows the proposed model to achieve high classification accuracy, together with interpretability needed for clinical decisions. The experimental findings indicated superior achievement because the model reached a 96% accuracy rate and delivered
Nevertheless, the reported 96% accuracy is obtained on a single public dataset with a single held-out test split and without external validation or additional regularization techniques such as dropout or extensive data augmentation; therefore, some risk of overfitting cannot be fully excluded. Future work will extend the evaluation to multi-center MRI cohorts, incorporate cross-validation and external test sets, and systematically investigate additional regularization and augmentation strategies to more thoroughly assess and enhance model robustness.
The model demonstrates its ability to classify mild from moderate and no AD cases through precision–recall analyses, together with ROC analyses and prediction probability distribution analysis. The system performed with complete accuracy in diagnosing moderate stages, yet showed the highest errors when detecting mild dementia cases because early detection remains challenging. Synthetic MRI augmentation approaches resolved class imbalance problems to improve the model's ability to generalize its findings.
A comparison shows the proposed method excels beyond conventional CNN-based models because these methods face challenges with class imbalance as well as overfitting problems. The ResNet and VGG16 network architectures demonstrated lower diagnostic performance along with higher misdiagnosis errors, especially during the diagnosis of early-stage AD. Comparative operations alone in medical imaging no longer suffice because this research employs transformers that find long-distance dependencies throughout images for improved classification results.
Future directions
The implementation of XAI methods enables medical professionals to trust decision processes because they provide clear explanations of diagnoses. The research demonstrates why explainability matters for healthcare systems which utilize AI, since it solves issues with black-box deep learning systems. In future work, this approach could be extended to incorporate multi-modal data such as cognitive scores and genetic biomarkers to improve diagnostic accuracy. Additionally, integrating uncertainty quantification techniques and validating the system across diverse clinical settings could help transition this model from research to real-world deployment. To support finer-grained early diagnosis and clinically meaningful progression monitoring, future evaluations will also require datasets that include prodromal-stage labels (e.g. MCI) and, where available, longitudinal imaging follow-up to enable staging beyond the three-category formulation used in this study. Expanding the dataset to include longitudinal imaging data may also enhance the model's ability to predict AD progression over time.
A further limitation of the present work is that performance metrics are reported on a single held-out test set without formal statistical significance testing against baseline models. Future work will therefore include cross-validation and resampling-based procedures (e.g. bootstrap confidence intervals and paired tests such as McNemar's test) on shared test sets, in order to quantify uncertainty and statistically validate performance differences between the proposed transformer-based framework and competing architectures.
Given the generalizability of transformer-based architectures, the proposed BEiT framework could also be adapted for medical image segmentation tasks, such as delineation of abdominal organs (e.g. kidneys, liver) or vascular structures (e.g. hepatic and portal veins) from MRI scans. Such extensions would allow the model to leverage the rich spatial representations learned through self-attention mechanisms, offering a modern alternative to traditional probabilistic and deterministic segmentation approaches.
Footnotes
Acknowledgement
The authors extend their appreciation to the Deanship of Scientific Research at Imam Mohammad Ibn Saud Islamic University (IMSIU) for funding this work through (grant number IMSIU-DDRSP2601).
Author contributions
Muhammd Amir Khan, Sheikh Muhammad Saqib and Tehseen Mazhar perform the Original Writing Part, Software, and Methodology; Sheikh Muhammad Saqib and Tehseen Mazhar perform Rewriting, investigation, design Methodology, and Conceptualization; Waqas Tariq Paracha Habib Hamam and Abdul Khader Jilani Saudagar perform the related work part and manage results and discussions; Tehseen Mazhar Muhammad Amir khan, Mona A. Alkhattabi and Habib Hamam perform related work part and manage results and discussion; Abdul Khader Jilani Saudagar, Tehseen Mazhar and Muhammd Amir Khan perform Rewriting, design Methodology, and Visualization; Muhammad Iqbal and Sheikh Muhammad Saqib performs Rewriting, design Methodology, and Visualization.
Funding
This work was supported and funded by the Deanship of Scientific Research at Imam Mohammad Ibn Saud Islamic University (IMSIU) (grant number IMSIU-DDRSP2601).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
The data used to support the findings of this study are available from the corresponding authors upon request.
