Abstract
Objectives
Hepatic failure is a common and severe condition among intensive care unit (ICU) patients. Its complication with acute respiratory distress syndrome (ARDS) is consistently associated with poor clinical outcomes and a significant disease burden. Early identification of high-risk patients is essential for improving clinical outcomes. This study aimed to develop and validate a machine learning (ML) model to predict 28-day mortality in ICU patients with hepatic failure complicated by ARDS.
Methods
Data were extracted from the Medical Information Mart for Intensive Care IV database, focusing on patients with hepatic failure complicated by ARDS. The cohort was randomly divided into an 80% training set and a 20% validation set. Six ML algorithms were applied to analyze clinical characteristics. Shapley Additive Explanations (SHAP) were used to interpret the optimal model.
Results
A total of 884 patients with hepatic failure and concurrent ARDS were included, with a 28-day mortality rate of 47.4%. Random forest models demonstrated superior performance, achieving an area under the curve of 0.823 (95% confidence interval: 0763–0.883) in the validation set. SHAP analysis identified eight clinically significant predictors of mortality, ranked by importance: age, neutrophil count, pulse transit time, direct bilirubin, heart rate, fibrinogen, serum sodium concentration, and prothrombin time. SHAP enhanced model interpretability, supporting clinical decision-making and potentially improving patient outcomes.
Conclusions
ML approaches exhibited promising performance in predicting 28-day mortality among hepatic failure patients complicated by ARDS. These models may aid in guiding treatment decisions for patients with hepatic failure patients.
Keywords
Introduction
Hepatic failure, particularly when complicated by acute respiratory distress syndrome (ARDS), poses a significant challenge in intensive care units (ICUs) because of high short-term mortality resulting from multiorgan dysfunction and limited therapeutic options. Patients with acute-on-chronic liver failure (ACLF) or acute liver failure (ALF) frequently develop respiratory complications, including ARDS or pneumonia, due to systemic inflammation, metabolic disturbances, and impaired immune responses.1–3 Previous studies have primarily focused on developing machine learning (ML) models for overall mortality in ACLF or ALF patients.4,5 The 28-day mortality rate for these patients remains persistently high, emphasizing the need for early prognostic biomarkers and targeted interventions. 6 Recent evidence has further elucidated the distinct pathophysiology and heightened risk profile of ARDS in this vulnerable population.5,7,8 Given that ARDS is a fatal and common terminal complication in ACLF patients, analogous to its role in sepsis, it is clinically important to develop a model specifically for predicting this outcome to aid in early risk assessment. 9
Model for end-stage liver disease (MELD) and its sodium-adjusted counterpart, MELD-Na, are commonly used tools for assessing liver disease severity.10–12 While these tools offer utility in general populations, they lack specificity for critically ill patients with respiratory complications. ARDS in hepatic patients is often worsened by factors such as hepatic encephalopathy-induced aspiration, fluid overload, and chronic steroid-induced immunosuppression.13,14 The interplay between liver dysfunction and respiratory compromise creates a self-perpetuating cycle that complicates ICU management. In ARDS patients, advanced age, concurrent shock, and thrombocytopenia are independent predictors of 28-day mortality, mirroring the multiorgan failure observed in hepatic-respiratory syndromes.15–17 It is therefore vital for clinicians to develop and apply a prognostic risk score model to enable risk assessment at ICU admission.
ML, as a branch of artificial intelligence, is capable of handling high-dimensional data and capturing complex, nonlinear relationships without relying on assumptions of data distribution. 18 However, most models focus on hepatic failure-related complications in the general population, with few tailored to predict outcomes in hepatic failure patients experiencing concurrent organ failure in ICU settings.19,20 To address this gap, this study aimed to develop an ML-based model to predict 28-day mortality risk in patients with hepatic failure complicated by ARDS, incorporating SHapley Additive exPlanations (SHAP) to enhance model interpretability. This approach is intended to support early risk assessment by clinicians and ultimately improve patient outcomes.
Materials and methods
Data source
Patient data for hepatic failure complicated by ARDS were extracted from the Medical Information Mart for Intensive Care IV (MIMIC-IV) database. This publicly available resource contains de-identified clinical data from critically ill patients admitted to the ICU at Beth Israel Deaconess Medical Center, USA, between 2008 and 2022. Access to the MIMIC-IV database requires completion of an official certification course hosted by the MIT Laboratory for Computational Physiology, ensuring ethical and responsible data use. The certification was completed by the author Wang Yi (certification ID: 14434877), who is authorized to access and extract data from the database. Data extraction for this study was approved by the Institutional Review Board (IRB) of the Massachusetts Institute of Technology (MIT) (Certification No. 14434877, Named:Wang Yi). The MIMIC-IV database is a publicly available dataset comprising de-identified patient information; therefore, no additional ethical approval is required for its use.
Participants
This study focused on patients with hepatic failure who were admitted for the first time to a general ICU for critically ill patients. It was designed to predict mortality risk based exclusively on data obtained within the first 24 h of ICU admission. This initial “snapshot” captures patients’ baseline disease severity prior to the confounding influence of extensive ICU interventions. Diagnoses were classified according to the International Classification of Diseases, Tenth Revision (ICD-10), and included alcoholic hepatic failure (with and without coma), acute and subacute hepatic failure (with and without coma), chronic hepatic failure (with and without coma), and unspecified hepatic failure (with and without coma). Inclusion criteria were: (1) age ≥18 years; (2) PaO2/FiO2 ≤ 300 mmHg (indicative of ARDS); and (3) ICU stay ≥24 h. Patients with more than 30% missing data were excluded. For patients with multiple ICU admissions, only data from the first admission were analyzed. Clinical information was collected within the first 24 h of ICU admission. To enhance the specificity of ARDS identification, patients with diagnoses of heart failure, aspiration pneumonitis, or transfusion-related acute lung injury were excluded using the corresponding ICD-9 codes.
The primary outcome was 28-day mortality following ICU admission for hepatic failure with ARDS. Because of the retrospective nature of the data, this study predicts crude mortality rather than ARDS-specific mortality. A total of 48 variables, including demographic characteristics, vital signs, laboratory values, and outcomes, were included. To ensure data representativeness, the dataset was split into training and testing sets in an 8:2 ratio.
Data extraction and outcomes
Data were extracted using Structured Query Language (SQL) in PostgreSQL from two PostgreSQL databases for patients admitted to the ICU within the first 24 h. The variables extracted included demographic data, vital signs, laboratory measurements, and outcomes.
Feature selection and data preprocessing
Variables with more than 30% missing data were excluded, while those with ≤30% missing values were imputed using a multiple imputation algorithm to minimize bias. Patients were categorized by 28-day ICU survival status.
After evaluating feature significance through univariate analysis, the dataset was randomly split into 80% training and 20% testing subsets, stratified by 28-day ICU survival status. For predictor selection in the training dataset, logistic regression (LR) with least absolute shrinkage and selection operator (LASSO) and Boruta analysis (Figure S1 in the Supplemental materials) were applied. The regularization parameter λ was determined via 10-fold cross-validation. Because LASSO is sensitive to multicollinearity, correlation analysis was performed to identify highly correlated features (r > 0.7). Ultimately, features were selected based on the intersection of the two algorithms.
Model development and explanation
Six ML algorithms—light gradient boosting machine (LightGBM), support vector machine (SVM), random forest (RF), extreme gradient boosting (XGBoost), LR, and adaptive boosting (AdaBoost)—were used to predict 28-day mortality. Evaluation metrics included area under the receiver operating characteristic curve (AUROC), accuracy, sensitivity, specificity, and F1-score, all assessed using the testing dataset. Net clinical benefit was evaluated using decision curve analysis. Calibration curves were generated to assess model calibration, and the best-performing model was selected based on these metrics. All models were trained using the Tidymodels framework to ensure a consistent and unbiased preprocessing pipeline. The hyperparameter optimization process for each model is detailed in the Supplemental materials (Figures S2 to S7).
SHAP, developed by Lundberg and Lee, provides a robust method for interpreting predictions from complex ML models. 19 SHAP summary and dependence plots were applied to identify key predictors and explore their relationships with mortality risk.
Statistical methods
Univariate analysis identified clinically significant features (p < 0.05). The tidymodel R package was used for ML. All continuous variables were incorporated into the model in their original, nondichotomized form to preserve information and maximize the statistical power of the predictive models. The SHAP method was employed to generate summary plots illustrating the contribution of each feature to predictive outcomes. SHAP evaluations for selected cases further illustrated the magnitude of feature contributions to individual predictions, enhancing model transparency. All statistical analyses were performed using R software (version 4.3.3), and two-tailed p-values <0.05 were considered statistically significant.
Results
Population demographics
Data from 2184 patients with hepatic failure were initially extracted from the MIMIC-IV database between 2008 and 2022. After applying inclusion and exclusion criteria (Figure 1), 884 patients with hepatic failure complicated by ARDS were included in the final analysis. The cohort comprised 367 females (41.5%) and 517 males (58.4%), with a median age of 58 years (interquartile range: 48–67). At 28 days post-ICU admission, 465 patients survived, while 419 died, resulting in a 28-day mortality rate of 47.4%, underscoring the high risk of death in this cohort. Differences between survivors and nonsurvivors are summarized in Table 1. The MELD 3.0 score was incorporated into the analysis and demonstrated a significant difference between survivors and nonsurvivors.

The workflow of the study. MIMIC-IV: medical information mart for intensive care IV; LASSO: least absolute shrinkage and selection operator; LR: logistic regression; RF: Random Forest; SVM: support vector machine; XGBoost: extreme gradient boosting; LightGBM: light gradient boosting machine; AdaBoost: adaptive boosting.
All predictor variables for patients with hepatic failure complicated by ARDS in the MIMIC-IV database set (n = 884).
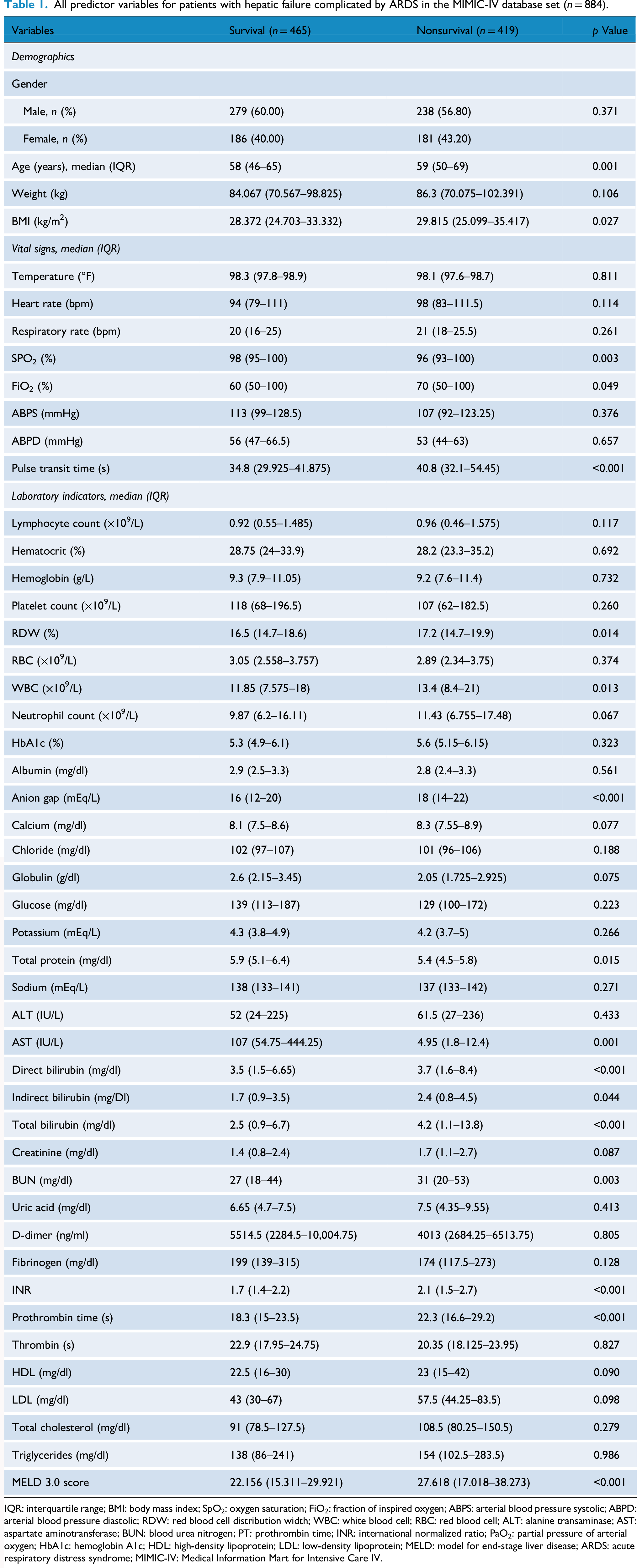
IQR: interquartile range; BMI: body mass index; SpO2: oxygen saturation; FiO2: fraction of inspired oxygen; ABPS: arterial blood pressure systolic; ABPD: arterial blood pressure diastolic; RDW: red blood cell distribution width; WBC: white blood cell; RBC: red blood cell; ALT: alanine transaminase; AST: aspartate aminotransferase; BUN: blood urea nitrogen; PT: prothrombin time; INR: international normalized ratio; PaO2: partial pressure of arterial oxygen; HbA1c: hemoglobin A1c; HDL: high-density lipoprotein; LDL: low-density lipoprotein; MELD: model for end-stage liver disease; ARDS: acute respiratory distress syndrome; MIMIC-IV: Medical Information Mart for Intensive Care IV.
Feature selection
Subsequently, LASSO and Boruta analyses were applied to identify key variables predictive of 28-day mortality in patients with hepatic failure complicated by ARDS. Using the LASSO-bootstrap method, 16 of 38 variables were retained as relevant features (Figure 2(a) and (b)). Eleven features were identified by the Boruta algorithm, six of which overlapped with the LASSO results: age, neutrophil count, pulse transit time, direct bilirubin, heart rate, fibrinogen, serum sodium, and prothrombin time (PT) (Figure 2(c)).

LASSO regression-based variable screening. (a) Variation characteristics of variable coefficients. (b) The process of selecting the optimal value of the parameter λ which indicate the penalty in the LASSO regression model is carried out by the cross-validation method. (c) Key variables identified by Boruta algorithm. LASSO: least absolute shrinkage and selection operator.
Model evaluation and comparison
Six ML algorithms—including LightGBM, SVM, RF, XGBoost, AdaBoost, and LR—were developed using the training data. Model performance was evaluated using AUROC, accuracy, specificity, sensitivity, and F1-score metrics on the testing set. In this evaluation, the RF model demonstrated the highest AUROC (0.823, 95% confidence interval (CI): 0.763–0.883), followed by LightGBM (0.736, 95% CI: 0.663–0.809), XGBoost (0.712, 95% CI: 0.637–0.788), SVM (0.683, 95% CI: 0.605–0.762), AdaBoost (0.671, 95% CI: 0.593–0.748), LR (0.659, 95% CI: 0.578–0.739), and MELD (0.532, 95% CI: 0.443–0.621). The predictive performance of the ML models was benchmarked against the MELD 3.0 score, which showed superior AUROC performance. Following rigorous hyperparameter tuning, RF emerged as the top-performing model. The performance of all comparative models improved substantially after this optimization process (Table 2). Calibration curves indicated a strong agreement between predicted and observed outcomes for both RF and LR models (Figure 3(b)). RF offered the greatest net clinical benefit across various threshold probabilities, establishing it as the optimal model for predicting 28-day mortality risk in this patient cohort (Figure 3(c)).

(a) Receiver operating characteristic curves for the ML models. (b) Calibration curve for the ML models. (c) Decision curve analysis of all the model and MELD 3.0 score. (d) Clinical impact curve of random forest model. ML: machine learning; MELD: model for end-stage liver disease.
Performance of ML models.

ML: machine learning; SVM: support vector machine; XGBoost: extreme gradient boosting; LightGBM: light gradient boosting Machine; AdaBoost: adaptive boosting.
Feature importance and interpretation
Given the superior performance of the RF model, a detailed feature interpretation was conducted. The SHAP summary plot illustrates the direction and magnitude of each variable's contribution to the model (Figure 4(a)). For example, higher fibrinogen levels were associated with reduced mortality risk, while advanced age and elevated direct bilirubin were linked to increased mortality. The variables were ranked by mean absolute SHAP value, highlighting their overall impact (Figure 4(b)). There were significant correlations between the SHAP value of the variables (Figure 4(c)).

Visually interpretable deep learning models using SHAP in the validation cohort. (a) SHAP force plot. Positive SHAP values indicate features that increase the prediction score, while negative values correspond to features that decrease it. (b) SHAP summary point. Each point represents a SHAP value for a feature and an instance; (c) Correlation heatmap plot of the variables with their SHAP value. Colors represent the correlation coefficient. SHAP: Shapley additive explanations.
Discussion
In this study, we developed and validated an interpretable ML model using the RF algorithm to predict 28-day mortality in ICU patients with hepatic failure and ARDS. The model incorporated eight features—age, neutrophil count, pulse transit time, direct bilirubin, heart rate, fibrinogen, sodium, and PT—and demonstrated excellent predictive performance with an AUROC of 0.823. Our findings show that the ML model significantly outperforms the established clinical standard, MELD 3.0, in predicting 28-day mortality. Consistent with previous studies, we suggest that the ML approach provides substantial incremental value for risk stratification. 21 A potential clinical application of this model is its integration into the electronic health record system to generate real-time risk scores. Upon ICU admission, the model could automatically calculate a patient's risk, flagging those at high risk for clinicians to review and consider for more intensive monitoring or preemptive therapy.
The RF algorithm aggregates predictions from multiple decision trees, reducing overfitting risk and improving model stability and generalization. 20 The finding that RF outperformed LightGBM may be attributable to the specific characteristics of our dataset, which is relatively modest in size and dimensionality. The selection of the Brier score as the primary tuning objective likely contributed to the robust performance of the final model by favoring well-calibrated predictions and mitigating overfitting. RF's inherent robustness and bagging ensemble method likely afforded it greater resistance to overfitting and enhanced generalizability. This underscores that there is no universally optimal model, and the best choice is contingent upon the data structure at hand. Importantly, SHAP analysis was employed to interpret the model, enhancing its transparency. This analysis revealed that elevated age and bilirubin levels were associated with increased mortality, while higher fibrinogen levels were protective.22,23 These findings are consistent with existing literature and support the biological plausibility of the identified predictors. Patients with hepatic failure frequently exhibit coagulopathy and systemic inflammation, contributing to poor outcomes. Fibrinogen and platelet counts, for instance, may provide better prognostic information than international normalized ratio alone.24–26 Additionally, hyponatremia is a common electrolyte disturbance, although its relationship with mortality remains poorly defined.27,28 We also found the prolonged PT and neutrophil counts in the final model. In ARDS, systemic inflammation further consumes coagulation factors, leading to the formation of intra-alveolar microthrombi and aggravating lung injury, and prolonged PT, reflecting impaired synthesis of coagulation factors, signifies an imbalance between procoagulant and anticoagulant and is associated with poor prognosis. 29 Neutrophils are the central in ARDS-related organ injury with their excessive activation, and infiltrate the organs and directly damage the vascular endothelial barriers by releasing neutrophil elastase, reactive oxygen species, and neutrophil extracellular traps, leading to organ failure. 30 According to the previous study, elevated blood glucose, related to the stress response and inflammatory cytokines, reflected the hyperglycemic environment which could exacerbate endothelial dysfunction and oxidative stress, and perpetuates the inflammatory state by promoting a pro-inflammatory phenotype, thereby exacerbating ARDS. 31 Similar with Song et al., our study also found the elevated direct bilirubin, a direct marker of impaired hepatic excretory function, associated with both hepatic insufficiency and ARDS due to potential aggravating endothelial injury as endogenous toxins. 32 According to Wu et al., we also reveal the elevated hematocrit at the early stage, as a marker of severe capillary leakage and hypovolemia, could be potentially linked to poor prognosis. 33 Our study helps clarify these associations. This model could serve as an early warning tool, enabling clinicians to identify high-risk patients at admission and initiate prompt, targeted interventions. 34
Several limitations should be acknowledged. First, the model was developed using the single-center MIMIC-IV database, which may limit generalizability. External validation using multicenter datasets is necessary to confirm model robustness. Second, the model was constructed using static variables collected at ICU admission. Dynamic changes over time were not considered and may substantially influence prognosis. Third, survival heterogeneity among subtypes of hepatic failure suggests that subgroup analyses could provide additional insight. However, our sample size limited such analyses and may have contributed to overfitting. Fourth, our model is intended as an early warning tool, identifying high-risk patients at admission. We did not include the dynamic evolution of illness or the effects of subsequent treatments, which significantly influence patient trajectory. Analyzing these time-dependent interactions represents a distinct clinical question that requires advanced causal inference models, such as structural nested or marginal structural models, in future studies. Fifth, the retrospective use of the MIMIC database precludes strict adherence to the Berlin Definition for ARDS, as it lacks systematic chest imaging and standardized cardiac function assessments required to definitively exclude cardiogenic pulmonary edema. Although we excluded patients with common alternative causes of hypoxemia using ICD-9 codes, this remains a fundamental constraint of large-scale critical care databases. Therefore, our findings require validation in prospective, multicenter studies with complete clinical and radiographic data. Finally, while the RF model demonstrated robust predictive capacity, more advanced deep learning methods might achieve superior performance in the context of precision medicine.
Conclusions
This study demonstrates that the interpretable RF prediction model holds significant potential in assessing the 28-day mortality risk for patients with hepatic failure complicated by ARDS. By incorporating key clinical predictors, the model provides a reliable tool to assist clinicians in early risk stratification. The interpretable nature of the model offers valuable insights into the contribution of each variable to the mortality prediction, thereby enhancing clinical decision-making. This approach represents a step forward in facilitating timely interventions and optimizing treatment strategies for high-risk patients.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076251408533 - Supplemental material for Machine learning prediction model for 28-day mortality among hepatic failure patients complicated by acute respiratory distress syndrome
Supplemental material, sj-docx-1-dhj-10.1177_20552076251408533 for Machine learning prediction model for 28-day mortality among hepatic failure patients complicated by acute respiratory distress syndrome by Wang Yi, Liu Song, Yang Lv and Zhishui Chen in DIGITAL HEALTH
Supplemental Material
sj-pdf-2-dhj-10.1177_20552076251408533 - Supplemental material for Machine learning prediction model for 28-day mortality among hepatic failure patients complicated by acute respiratory distress syndrome
Supplemental material, sj-pdf-2-dhj-10.1177_20552076251408533 for Machine learning prediction model for 28-day mortality among hepatic failure patients complicated by acute respiratory distress syndrome by Wang Yi, Liu Song, Yang Lv and Zhishui Chen in DIGITAL HEALTH
Footnotes
Ethical considerations
The MIMIC-IV database is a publicly available dataset comprising de-identified patient information; therefore, no additional ethical approval is required for its use. Data extraction for this study was approved by the Institutional Review Board (IRB) of the Massachusetts Institute of Technology (MIT) (Certification No. 14434877, Named:Wang Yi).
Author contributions
Conceptualization, Wang Yi, Yang Lv, and Zhishui Chen; methodology, Wang Yi, Liu Song and Yang Lv; software, Wang Yi, Liu Song and Yang Lv; data curation, Wang Yi and Yang Lv; writing-original draft preparation, Wang Yi. All authors have read and agreed to the published version of the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by National Natural Science Foundation of China 82371794 (ZC).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of data and materials
The datasets generated and/or analyzed during the current study can be found in MIMIC-IV (https://physionet.org/content/mimiciv/3.1/). The code has been posted to public Github, and is available via ![]() .
.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
