Abstract
Objective
To systematically map the research landscape of synthetic data in healthcare between 2000 and 2024, revealing prevalent topics and tracking their evolution over time and across geographic locations.
Methods
We applied structural topic modeling (STM) to map this landscape, identifying prevalent topics and their evolution over time and geography. PubMed articles from 2000 to 2024 with “synthetic data,” “artificial data,” or “simulated data” in the title/abstract were analyzed. Texts were preprocessed (lowercasing, stopword removal, stemming), and STM was run with year and continent as covariates. The optimal number of topics (K = 10) was selected based on held-out likelihood and interpretability. Topic trends and correlations were analyzed using stacked area charts and network analysis.
Results
Among 7533 articles, a 20-fold growth in publications was observed. North America (48.1%) and Europe (31.8%) dominated early research, while Asia's share rose from 4.7% to 24.1%. Topics grouped into four themes: Biomedical Imaging & Signal Processing (21.1%), Synthetic Data Applications (20.7%), Computational & Statistical Methods (34.3%), and Genomics & Molecular Biology (23.9%). Initially prominent topics such as “Bayesian Modeling” (23.1%–10.8%) and “Statistical Bias & Missing Data” (21.9%–7.1%) declined, while “Synthetic Data Generation” (2.7%–23.0%), and “Disease Modeling and Public Health” (3.5%–14.3%) grew significantly.
Conclusion
Synthetic data research in healthcare is expanding, with shifting regional contributions and evolving topic focus. Realizing its potential requires cross-disciplinary collaboration, bias mitigation, and equitable partnerships.
Introduction
The healthcare sector is increasingly leveraging data-driven approaches to improve patient outcomes, optimize operational efficiency, and advance medical research. However, acquiring comprehensive healthcare data is often constrained by high costs associated with advanced data acquisition techniques, privacy regulations, and ethical considerations regarding patient data collection and sharing.1,2 These factors collectively impede the utilization of comprehensive data for advancing patient care and medical research. To address these challenges, synthetic data offer a promising solution.
Synthetic data refer to artificially generated datasets that mimic the statistical properties of real-world data without exposing sensitive information. 3 It mitigates data scarcity and privacy concerns, enabling a broader scope of research and experimentation without patient data exposure. 4 Moreover, it facilitates the creation of diverse and representative datasets, improves AI model generalizability, and mitigates bias arising from skewed or underrepresented populations. 5 Synthetic data also facilitate innovation while safeguarding patient confidentiality, making it a crucial tool for overcoming data access barriers.
While a number of reviews have examined synthetic data in healthcare,3,5–11 they predominantly focus on data generation methods (Table 1). These existing reviews of synthetic data in healthcare, while valuable, often rely on manual theme identification, introducing potential subjectivity and bias. Structural topic modeling (STM) is an advanced probabilistic unsupervised machine learning topic modeling technique designed to uncover latent themes in textual corpora by incorporating document-level metadata. 12 Compared to traditional systematic reviews or bibliometric analyses, STM provides a scalable, data-driven way to identify themes, quantify their prevalence, and examine how they evolve over time.13,14 In contrast to Latent Dirichlet Allocation (LDA) and related topic models, STM extends the analysis by explicitly modeling covariates, thereby enabling richer insights into research trends and contextual factors. While STM mitigates certain forms of subjective bias, its outputs remain sensitive to preprocessing decisions, parameter choices, and corpus selection, and thus the method cannot be regarded as entirely unbiased. Taking this into account, STM complements rather than replace existing approaches by offering an additional lens to explore thematic structures, and evolution of research priorities in the literature. This is important in informing strategic decision-making by policymakers, funding agencies, and researchers.15,16
Summary of review studies on synthetic data in healthcare.

Despite its growing adoption, the thematic landscape of synthetic data research in healthcare remains underexplored. Here, we apply STM to map the research landscape of synthetic data in healthcare, revealing prevalent topics and tracking their evolution over time and across geographic locations. Specifically, this study aims to address this gap by answering two research questions: (i) What are the dominant topics in synthetic data research? (ii) What are the patterns in the thematic and geographical evolution of synthetic data research over time?
Materials and methods
Study design
This study is a systematic bibliometric review of globally published literature on synthetic data in healthcare, covering the period 1 January 2000 to 31 December 2024.
Data collection
We systematically retrieved relevant publications from PubMed using a search strategy that targeted articles containing terms related to synthetic data in the title or abstract: (Synthetic data"[Title/Abstract] OR “Artificial data"[Title/Abstract] OR “Simulated data"[Title/Abstract] OR “generative data"[Title/Abstract] OR “synthetic patient"[Title/Abstract] OR “synthetic health"[Title/Abstract] OR “in-silico data"[Title/Abstract] OR “simulation data"[Title/Abstract] OR “data simulation"[Title/Abstract]”). The batch_pubmed_download function was used to systematically retrieve articles in XML format while adhering to PubMed API rate limits. 17 A semiautomated screening process was implemented using a rule-based algorithm to filter potentially relevant articles. Specifically, the algorithm filtered articles by detecting the co-occurrence of terms related to synthetic or simulated data and also referenced healthcare or biomedical contexts, including clinical, epidemiological, or genomic applications. Following the automated screening, a random sample of 5% of abstracts from the final set of 7533 identified articles was manually reviewed to verify the accuracy of the rule-based approach, confirming that the majority (99.7%) aligned with the intended scope. We included English articles published between 2000 and 2024 with an abstract.
Data processing
Each downloaded XML file was assigned a unique timestamp-based filename to prevent duplication. The downloaded XML files were processed to extract key bibliographic information, including title, abstract, publication year, and first author affiliations. Metadata variables were extracted, including publication year and country. Countries were categorized into continents (Africa, Asia, Europe, North America, South America, and Oceania). To ensure data integrity, articles were assessed for missing values in key fields, duplicate titles and extracted fields were cleaned to remove HTML/XML artifacts.
Text preprocessing
Titles and abstracts from the dataset were combined to form a text corpus. The corpus was preprocessed using the tm package in R. This process involved conversion of text to lowercase, removal of punctuation, numbers and English stopwords and stripping of extra whitespace. 18 Thereafter, we stemmed all words using the Porter stemming algorithm. 19 A Document-Term Matrix was then created, transformed into a matrix format, and transposed to generate a term matrix.
Topic modeling
Structural topic modeling is an advanced probabilistic unsupervised machine learning topic modeling technique designed to uncover latent themes within a collection of textual documents by incorporating document-level metadata.12–14 Preprocessed text data were formatted for STM analysis. Documents were structured into a list format compatible with the stm package, and the vocabulary was extracted from the term matrix.
To determine the optimal number of topics, a search was performed using searchK(), which evaluates model fit using held-out likelihood. 18 A range of topic numbers (K = 5, 10, 15, 20, 25, 30) was tested, incorporating year and continent as prevalence covariates. The optimal number of topics (K) was selected by identifying the “elbow point” in the likelihood plot, balancing semantic coherence and exclusivity, and evaluating the interpretability of the resulting topics 20 (Figure S1). A final STM model was fitted with K = 10 topics, using year and continent as prevalence covariates. The Expectation–Maximization algorithm was run for 150 iterations, initializing with a LDA approach. 21
The top terms associated with each topic were extracted using the labelTopics() function. Based on interpretability, topics were assigned descriptive labels reflecting their thematic content. Furthermore, to enhance the interpretability of the topics generated by the STM model, word clouds were created for each topic, highlighting the most frequently associated words. 21 The cloud() function from the stm package was used, with word sizes scaled to reflect their relative importance within each topic. To enhance interpretability, topics were then grouped into broader thematic categories according to their primary orientation, distinguishing between domain-specific applications, translational and clinical research contexts, methodological innovations, and biological sciences.
To assess the temporal trends in STM research, we computed the annual count of articles for each continent and standardized the data to ensure a complete set of year-continent combinations. Missing values were replaced with zero to maintain data consistency. We then calculated the annual proportion of articles per continent to facilitate a comparative analysis. We employed 100% stacked area chart to illustrate the temporal distribution of STM research by continent. To examine how research topics evolved over time, we linked topic prevalence data with publication years. We generated 100% stacked area chart to depict changes in topic proportions across years.
We assessed relationships between topics using correlation-based network analysis.18,21 A topic correlation matrix was derived from the STM model, with edges retained for correlations above a predefined threshold (0.1). An adjacency matrix was created to construct a graph-based network, which was visualized using the igraph and ggraph packages. To improve readability, we scaled node sizes based on topic prevalence, colored nodes according to their thematic classification, and weighted edges based on correlation strength. Additionally, a force-directed layout was employed to enhance network interpretability, and labels were added to identify key topics.
Descriptive analysis using percentages and topic modeling were performed using R version 4.4.1 (R Foundation for Statistical Computing, Vienna, Austria).
Ethical approval
Ethical approval was not required for this study as it did not involve human or animal participants.
Results
During the review period, the search criterion yielded 20,011 articles. Of these, 25 articles without abstracts were excluded. An additional 12,437 articles were excluded following rule-based screening resulting in 7533 articles retained for further analysis (Figure 1). The number of publications analyzed exhibited a fluctuating upward trajectory, increasing approximately 20-fold from 43 articles in 2000 to 863 articles in 2024 (Figure 2).
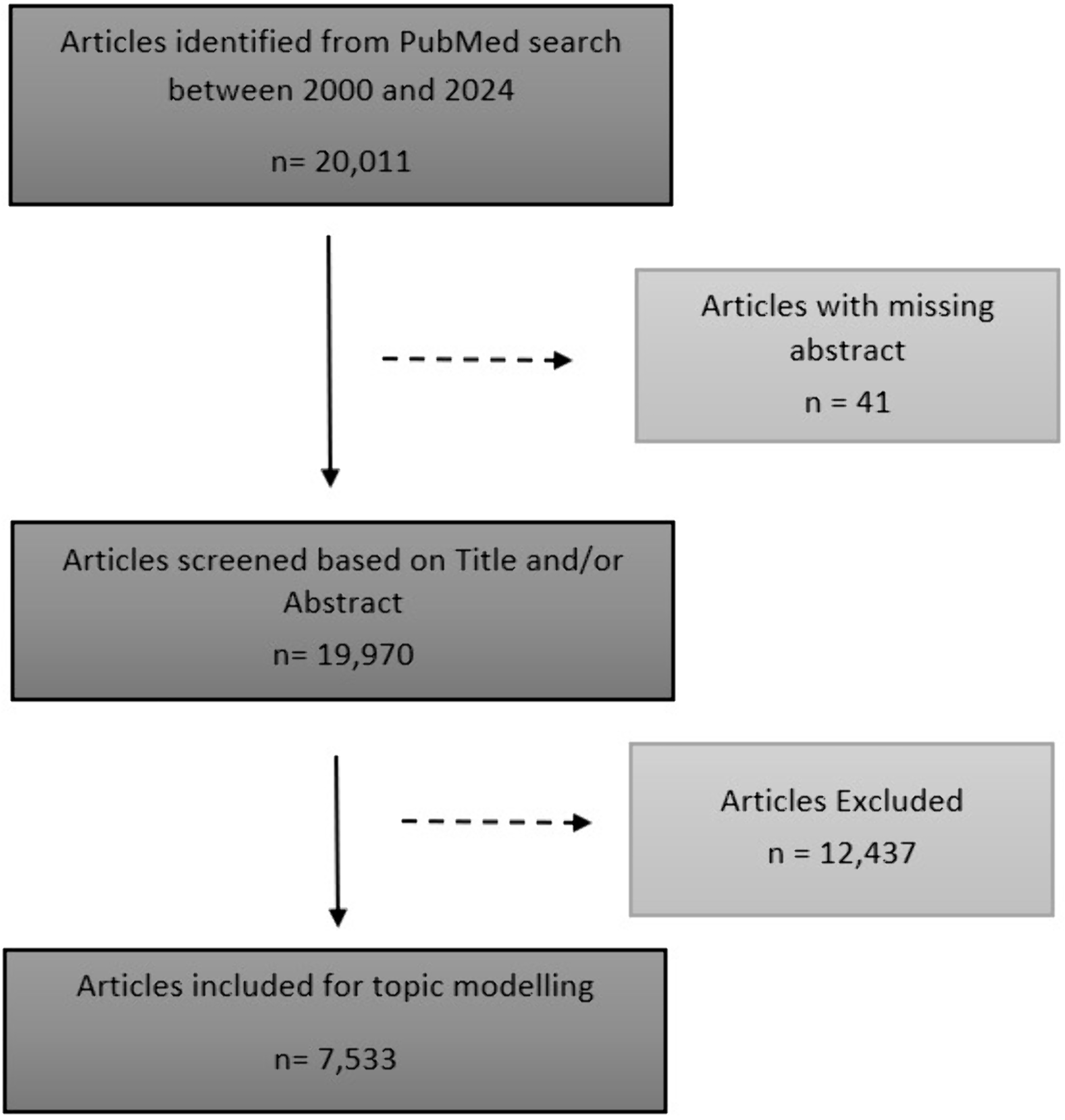
Flow diagram of included studies for synthetic data in healthcare, 2000–2024.
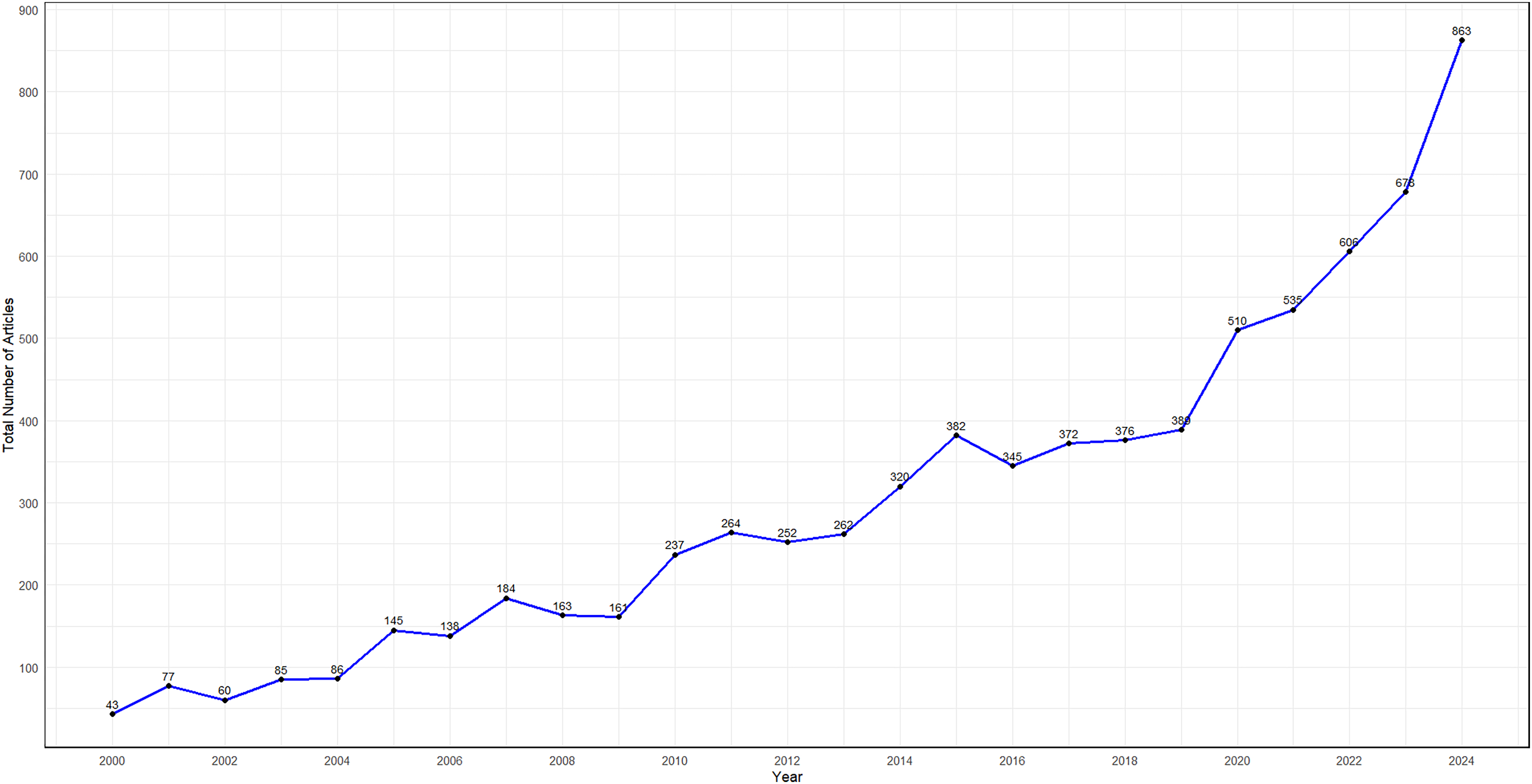
Number of articles related to synthetic data in healthcare between 2000 and 2024.
Geographical patterns
Overall, North America (3625 [48.1%]) and Europe (2396 [31.8%]) were the primary contributors to the research, followed by Asia (1293 [17.2%]). In contrast, Oceania (112 [1.5%]), South America (74 [1.0%]), and Africa (33 [0.4%]) had the lowest contributions. Although North America contributed the most, its share declined from 58.1% in 2000 to 40.8% in 2024. Europe experienced a marginal drop, decreasing from 34.9% to 32.0% over the same period. Conversely, Asia experienced a steady rise, increasing from 4.7% in 2000 to 24.1% in 2024. Oceania and South America demonstrated modest contributions with minor fluctuations between years, ranging from 0.0% to 2.8% and 0.0% to 1.9%, respectively, between 2000 and 2024. Africa's contribution remained low and was minimal before 2018 but has shown steady growth since, peaking in 2019 (∼1.3% of global output) (Figure 3). At the country level, the top five contributors to the research were the United States of America (3388 [45.0%]), China (649 [8.6%]), the United Kingdom (562 [7.5%]), Germany (442 [5.9%]), and France (285 [3.8%]).

Annual proportion of synthetic data research in healthcare by continent (2000–2024).
Topic identification and prevalence
Table 2 presents the identified topics along with key terms for each metric, as well as the thematic areas encompassing the ten topics. Figure 4 presents word clouds visualizing the most frequent words for each topic, ranked by the FREX criterion. Word size corresponds to their relative importance within each topic. Among these, Topic 2: Bayesian Modeling & Inference (15.1%), Topic 6: Statistical Bias & Missing Data (13.0%), Topic 8: Gene Expression & Transcriptomics (12.0%), Topic 9: Genetic Association Studies & Genomics (11.9%), and Topic 10: Medical Image Reconstruction (11.4%) emerged as the most prominent topics in synthetic data research in healthcare. In 2000, the most important topics were Topic 6: Statistical Bias & Missing Data (21.9%), Topic 2: Bayesian Modeling & Inference (17.9%), and Topic 7: Neuroimaging & Signal Processing (14.4%). By 2024, the focus shifted to Topic 4: Synthetic Data Generation (23.0%), Topic 3: Disease Modeling and Public Health (14.34%), and Topic 2: Bayesian Modeling & Inference (10.8%), reflecting an evolving research landscape (Figure 5).

Word cloud figure for the ten topics in synthetic data research in healthcare, 2000–2024.
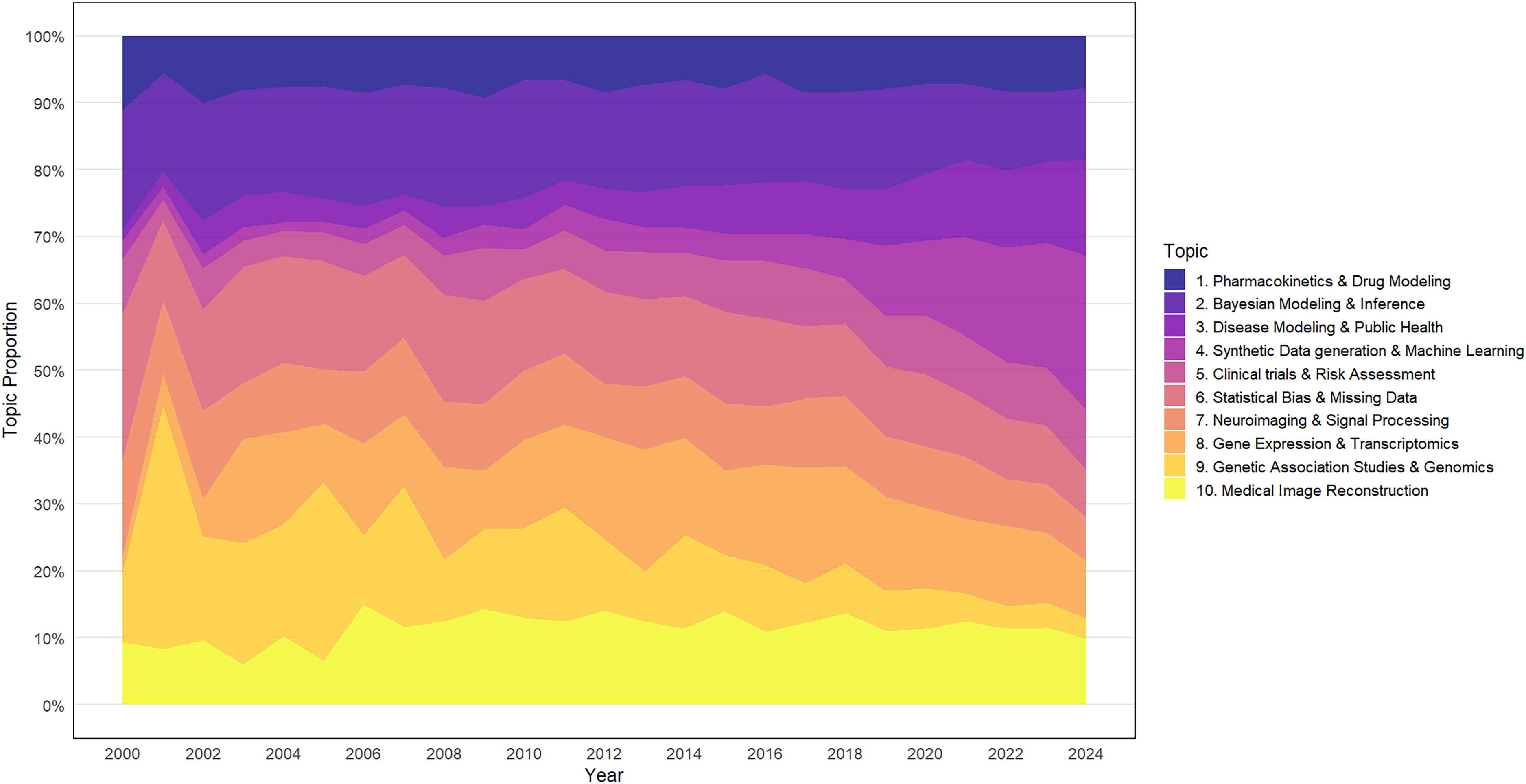
Temporal patterns in synthetic data research in healthcare topics, 2000–2024.
Themes, topic labels, and key terms in synthetic data research for healthcare.

Thematic groups and temporal dynamics
The identified topics were grouped into four thematic areas:
Theme 1: Biomedical imaging and signal processing
This theme encompasses two topics: Topic 7: Neuroimaging & Signal Processing, and Topic 10: Medical Image Reconstruction (Table 2), collectively representing 21.1% of all evaluated publications. Topic 7 accounted for 9.7%
Theme 2: Synthetic data applications in biomedical research
Comprising Topic 1 (Pharmacokinetics & Drug Modeling), Topic 3 (Disease Modeling & Public Health), and Topic 5 (Clinical Trials and Risk Assessment) (Table 2), the Synthetic Data Applications in Biomedical Research theme (Table 2) represented 20.7% of the total publications. Topics 1, 3, and 5 individually contributed 7.8%, 6.3%, and 6.6%, respectively. These topics had differing temporal trajectories: Topic 1 had a marginal decline from 11.0% in 2000 to 7.8% in 2024, while Topic 3 saw growth from 1.7% to 14.3% during the same time frame. However, Topic 5 fluctuated between 3.0% (2001) and 9.0% (2024) (Figure 5).
Theme 3: Computational and statistical methods
Comprising Topics 2 (Bayesian Modeling & Inference), 4 (Synthetic Data Generation), and 6 (Statistical Bias & Missing Data) (Table 2), the Computational & Statistical Methods theme accounted for a significant 34.3% of the research output. With individual contributions of 15.1%, 6.2%, and 13.0%, respectively, these topics exhibited contrasting temporal patterns. While Topic 2 declined from 23.1% to 10.8%, and Topic 6 from 21.9% to 7.1%, Topic 4 had an upward trend, rising from 2.7% in 2000 to become the leading topic in 2024, accounting for 23.0% of research topics (Figure 5).
Theme 4: Genomics and molecular biology
Comprising two topics, 8 (Gene Expression & Transcriptomics) and 9 (Genetic Association Studies), the Genomics and Molecular Biology theme (Table 2) accounted for a 23.9% of the analyzed research. The temporal trends within this theme were varied. Topic 8 was lowest at 2.6% in 2000, rose to a peak of 18.2% in 2013, and then gradually declined to 8.6% by 2024. In contrast, Topic 9 showed an early dominant focus, peaking at 36.5% in 2001, but has since declined sharply to just 3.1% in 2024 (Figure 5).
Topic co-occurrence and correlations
The interconnections between different research topics in synthetic data research in healthcare are shown in the topic network diagram (Figure 6). The network is anchored by a strong Computational & Statistical Methods core, dominated by the close integration of “Bayesian Modeling” (Topic 2) and “Synthetic Data Generation” (Topic 4). This analytical foundation directly supports “Pharmacokinetics & Drug Modeling” (Topic 1), a central application hub in the quantitative domain. In parallel, “Medical Image Reconstruction” (Topic 10) functions as a key translational hub, linking “Neuroimaging” (Topic 7) with both clinical practice and molecular biology, particularly through its strong ties to “Genetic Association Studies & Genomics” (Topic 9). These hubs converge on “Clinical Trials & Risk Assessment” (Topic 5), which serves as the central clinical anchor connecting imaging, genomics (“Gene Expression & Transcriptomics,” Topic 8), and “Disease Modeling & Public Health” (Topic 3). More peripheral but still important are topics such as Topic 8, Topic 3, and Topic 6 (“Statistical Bias & Missing Data”), which reinforce applied and methodological depth. Overall, the landscape is defined by highly central translational hubs (Topics 10, 1, and 5) built upon a robust computational–statistical foundation (Topics 2 and 4), enabling integration across clinical, genomic, and methodological domains.

Topic network in synthetic data research in healthcare, 2000–2024.
Discussion
We mapped 25 years of synthetic data research in healthcare using a STM approach. The research output grew twentyfold, from 43 articles in 2000 to 863 in 2024, reflecting a rising interest in synthetic data applications in healthcare. Furthermore, we observed substantial changes in the geographic distribution of research activity, despite the global North-South disparity, and a dynamic evolution of key research topics. Specifically, initially prominent topics including “Bayesian Modeling,” “Neuroimaging,” and “Statistical Bias & Missing Data” gradually decreased as the research focus shifted to “Synthetic Data Generation” and “Disease Modeling and Public Health” in 2024 with “Bayesian Modeling” still ranking third despite its reduction.
Geographical landscape
Geographically, North America and Europe were initially the dominant contributors, but Asia's contribution significantly increased over time. Overall, Oceania, South America, and Africa contributed minimally (<2% each). The observed geographic patterns reflect both regional dominance and disparity in synthetic data research in healthcare. North America and Europe's historical leadership stems from established institutions, robust funding, strong industry-academia collaborations, and mature regulatory frameworks. Conversely, Asia's rising contribution, particularly from China driven largely by state-led investment in research (2.4% of GDP in 2022) prioritizing AI and healthcare, signals a shifting research landscape. 22 The persistent global North-South disparity23–25 is exacerbated by limited funding, infrastructure deficits, brain drain, and weak regulatory frameworks in Africa, South America, and Southeast Asia.
Thematic landscape and temporal dynamics
The thematic contributions and temporal dynamics in synthetic data research reflect evolving technological priorities, healthcare demands, and methodological advancements. Biomedical Imaging & Signal Processing theme representing approximately a fifth of the research output, once dominant, has seen gradual declines in subtopics like “Neuroimaging.” This trend may stem from the maturation of imaging technologies, where foundational innovations in computed tomography, magnetic resonance imaging, and signal processing have already been integrated into clinical workflows.26,27 This may reduce the urgency for novel breakthroughs. Additionally, the rise of computationally intensive methods in other areas, such as synthetic data, may have diverted the focus from these areas.
The Computational & Statistical Methods theme, the most dominant, representing slightly more than a third of the research output, illustrates a pivotal shift. The surge in “Synthetic Data Generation” overshadowed the declines in “Bayesian Modeling & Inference” and “Statistical Bias & Missing Data.” This shows the growing demand for scalable, AI-driven solutions in research and healthcare. This is especially true in contexts where synthetic data addresses challenges such as data privacy, scarcity of labeled datasets, and the need to train robust machine learning models. 4 Bayesian methods, while robust, may have waned due to their computational complexity and the preference for faster, data-driven approaches in large-scale applications. 28 Moreover, the declining prominence of “Statistical Bias & Missing Data” does not reflect reduced importance, but rather a shift from methodological innovation to maturity and integration into standard research practice.29,30
The Synthetic Data Applications in Biomedical Research theme representing one-fifth of the research output highlights contrasting trajectories. “Disease Modeling & Public Health” had an upward trend reflecting its growing centrality in global research and the need for predictive modeling to inform policy and outbreak management. 31 This growth has been driven by the increasing burden of infectious and chronic diseases, heightened urgency during major outbreaks such as SARS, H1N1, Ebola, and COVID-19. The parallel growth of computational power and large-scale health data has also enabled more sophisticated models for accurate forecasting, intervention planning, and informing public health policy. Conversely, “Clinical Trials & Statistical Inference” declined, possibly due to increasing regulatory scrutiny and the difficulty of translating synthetic data into accepted clinical endpoints. In parallel, the decline of Pharmacokinetics & Drug Modeling likely reflects its transition into a mature and highly standardized discipline within drug development. As pharmacokinetic methods and software became routine, the need for extensive methodological publication decreased, while research focus broadened toward biologics, complex therapies, and precision medicine, often emphasizing delivery systems or integrative approaches. 32 At the same time, the rapid rise of computational fields such as Bayesian modeling, synthetic data generation, and machine learning has absorbed much of the methodological innovation once attributed to the topic, redistributing publications under these broader computational themes. Genomics & Molecular Biology, representing slightly more than one-fifth of the research output, reveals subfield variability. Synthetic genomic data address data scarcity and privacy concerns by enabling researchers to share datasets mimicking real sequences without exposing individual identities. This facilitates studies of rare variants and diseases, modeling evolutionary dynamics, and validating algorithms.33–36 It supports hypothesis testing via evolutionary simulations, allows for controlled benchmarking and bias mitigation, and facilitates large-scale population genomics and high-risk climate adaptation modeling. Furthermore, synthetic data democratize genomic research by providing usable datasets for resource-limited settings and addressing ethical concerns in indigenous communities, promoting equitable access and collaboration in the field. While synthetic genomic data are increasingly utilized, concerns exist regarding its fidelity and ability to capture complex genomic phenomena such as epistasis and epigenetic regulation.37,38 Additionally, establishing statistical validation standards is crucial to ensure the trustworthiness of synthetic genomic datasets. Temporally, “Gene Expression & Transcriptomics” demonstrated growth possibly fueled by reducing sequencing costs and the rise of precision medicine. On the contrary, “Genetic Association Studies” declined, potentially due to saturation in identifying common genetic variants and shifting focus to functional genomics. 39
Topic co-occurrence and correlations
The network structure highlights the growing centrality of computational and statistical methods, showing how advances in Bayesian modeling and synthetic data generation now form the analytical backbone of biomedical research. This methodological foundation is not only driving innovation but also enabling the shift toward multimodal data integration, particularly imaging and genomics, into more holistic models of disease and treatment. The translational role of medical imaging and genomics, converging on clinical trials and risk assessment, emphasizes the importance of aligning methodological sophistication with clinical relevance to directly inform patient care and policy. At the same time, peripheral but strategically important areas such as bias correction and handling of missing data remain critical for ensuring validity and reproducibility in an era of increasingly complex datasets. The observed structure reflects a field in transition, where traditional application areas like pharmacokinetics are becoming embedded within broader computational frameworks, signaling the need for domain expertise to evolve in tandem with advanced data science skills. Collectively, these dynamics point to the importance of cross-disciplinary training and collaboration as essential enablers of the next generation of biomedical research.
Policy implications
Our findings carry several practical and policy implications for fostering innovation, advancing equity, and addressing global health challenges. At the global level, closing the persistent North–South disparity requires targeted investments in infrastructure, training, and local talent retention in underrepresented regions. This should go hand-in-hand with equitable international partnerships that support technology transfer and collaborative research in these settings. Additionally, policymakers in emerging regions may draw lessons from Asia's rapid rise, particularly China's strategic state-led investment in AI and healthcare, as a model for catalyzing local research growth. Strategic global investment is also essential, with resources directed toward emerging areas such as AI-driven data generation, disease modeling, and genomics. In parallel, there is need to ensure that more established or hard-to-regulate domains, such as imaging and clinical trials, are not neglected.
At the research and regulatory level, fostering interdisciplinarity is key. Institutions and funders should encourage cross-cutting research collaborations that connect computational methods with applied fields such as neuroimaging and disease modeling, as well as foundational sciences such as genomics. To support this, policymakers should work on establishing agile regulatory frameworks that uphold fidelity, privacy, and generalizability. This is particularly important for sensitive applications in clinical trials and genomic data to build public trust while enabling responsible innovation. Ethical governance will be equally crucial to ensure equitable access, prevent misuse, and mitigate risks of bias in synthetic data.
At the sectoral level, several themes demand attention. The rise of AI-driven synthetic data generation highlights the need for scalable, privacy-preserving solutions to meet the demands of modern machine learning. Growth in disease modeling highlights synthetic data's potential for pandemic preparedness and public health policy. Similarly, advances in synthetic genomics offer opportunities to democratize research, enable studies of rare variants, and drive precision medicine, provided standards for fidelity and validation are rigorously enforced.
Research gaps
Despite the diversity of themes identified in this analysis of synthetic healthcare data, key gaps remain. These thematic gaps are underpinned by common, cross-disciplinary challenges: technical limitations, ethical considerations, and regulatory uncertainty.
Theme 1: Biomedical imaging and signal processing
Despite advances in neuroimaging and medical image reconstruction, important research gaps remain. In neuroimaging, there is limited progress in developing synthetic data approaches that accurately capture the complexity of brain structure, function, and connectivity, especially for multimodal imaging and rare neurological conditions. 40 Similarly, in medical image reconstruction, while deep learning has enhanced image quality and reduced noise, challenges persist in generalizability across scanners, populations, and acquisition protocols. 41 Furthermore, research is needed to develop standardized digital phantoms and validation frameworks that can be universally adopted to benchmark generative models across different imaging modalities, moving beyond single-institution validation.42,43 Limitations in molecular modeling, high computational costs, and the lack of standardized imaging benchmarks hinder broader adoption.
Theme 2: Synthetic data applications in biomedical research
Synthetic electronic health records (EHRs), clinical trial simulation, and disease modeling, remain underdeveloped, largely due to fragmented data standards, regulatory barriers, and data scarcity. 44 This is particularly pronounced in applications for health economics and outcomes research and for modeling care delivery including areas like telemedicine. Personalized medicine beyond genomics is particularly underrepresented, suggesting a need for specialized tools and cross-institutional data-sharing frameworks.
Theme 3: Computational and statistical methods
Despite rapid methodological advances, key methodological gaps persist. Bayesian approaches in healthcare remain underutilized for handling uncertainty in multimodal and longitudinal clinical data, particularly when data are sparse or incomplete. Similarly, while synthetic data methods are advancing, they often overlook statistical biases and missingness mechanisms inherent in real-world health datasets, limiting their reliability for downstream inference.45,46 A critical gap lies in developing integrated frameworks that combine Bayesian modeling, principled missing data strategies, and fairness-aware synthetic data generation to ensure both validity and equity in clinical research and policy applications. Additionally, methods for generating complex, multimodal data are still nascent and require further development to address the complexity of clinical data. 47
Theme 4: Genomics and molecular biology
The primary gap here is the narrow focus on genomic sequence data itself. There is need for expansion of the field to address the challenge of generating synthetic data for integrated multiomics and for spatial transcriptomics. 48 Furthermore, there is a significant opportunity to extend synthetic data applications beyond basic research toward personalized medicine and drug discovery. These areas face hurdles due to the complexity of molecular modeling and a lack of suitable synthetic data for preclinical and clinical stages.
Limitations
This study's methodology is subject to some limitations. The search strategy, restricted to specific keywords in titles/abstracts and reliance on PubMed, introduces potential selection bias by excluding relevant literature using alternative terminology or indexed in other databases. Although, the semiautomated screening approach used enhanced efficiency and consistency, we cannot rule out that some studies only tangentially related to synthetic data may have been included due to its reliance on predefined keyword rules. This could potentially introduce minor noise and slightly reduce topic precision. Geographic classification based solely on first-author affiliation may misrepresent multinational collaborations and regional contributions. While language bias was minimized via English titles/abstracts, exclusion of non-English publications is still possible. Future analyses should mitigate these issues through expanded search terms and data sources, expert validation, and refined geographic classification methods to enhance robustness and representativeness.
Conclusion
Through STM analysis of 25 years of synthetic data research in healthcare, this study reveals significant growth and evolution of topics. Research output has increased nearly twentyfold, accompanied by a shift in geographic contributions, with Asia's presence growing alongside the historical dominance of North America and Europe, although a persistent Global North-South disparity remains. Thematic foci have evolved, with increased emphasis on “Synthetic Data Generation” and “Disease Modeling and Public Health.” Methodological interconnections are evident. Despite these advances, key gaps exist in areas such as drug discovery, synthetic EHRs, personalized medicine beyond genomics, telemedicine, mental health applications, health economics, and ethical AI. These highlight the need for cross-disciplinary collaborations, bias mitigation strategies, and equitable partnerships to fully realize the potential of synthetic data in healthcare.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076251404530 - Supplemental material for A quarter-century of synthetic data in healthcare: Unveiling trends with structural topic modeling
Supplemental material, sj-docx-1-dhj-10.1177_20552076251404530 for A quarter-century of synthetic data in healthcare: Unveiling trends with structural topic modeling by Billy Ogwel, Vincent H Mzazi, Alex O Awuor, Gabriel Otieno, Sidney Ogolla, Bryan O Nyawanda and Richard Omore in DIGITAL HEALTH
Footnotes
Contributorship
BO conceived the study, BO, VM, BON, AOA, GO, and RO contributed to study design and implementation. BO and BON analyzed and interpreted the data. BO drafted the manuscript and all authors critically reviewed the manuscript for intellectual content and approved the final manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of data and materials
Disclosure
The findings and conclusions in this report are those of the authors and do not necessarily represent the official position of the Kenya Medical Research Institute or partnering institutions.
Use of artificial intelligence (AI) tools
The authors would like to acknowledge the use of AI technology (Gemini and Deepseek) for grammar checking and proofreading of this manuscript.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
