Abstract
Objective
This study aims to investigate the development of automated International Classification of Diseases (ICD) coding models using the Medical Information Mart for Intensive Care (MIMIC) dataset. This work integrates computer science and clinical perspectives to evaluate progress, identify challenges, and provide insights for future ICD coding automation.
Methods
We conducted a systematic review following the Preferred Reporting Items for Systematic Reviews and Meta-Analyses guidelines. We selected 73 studies between 2014 and 2024 and extracted key information about data preprocessing, knowledge integration, model architectures, evaluation strategies, and explainability.
Results
In the reviewed papers, 69.57% (48 papers) focused on utilizing medical knowledge, primarily through knowledge graphs. The methods have evolved from traditional machine learning techniques to more advanced approaches, such as deep learning, knowledge reasoning, information retrieval, and generative models. Since 2019, F1-micro scores have consistently improved: Studies using the MIMIC-III full dataset have shown a 6.4% increase, while those using the MIMIC-III top-50 dataset have experienced a 10.2% improvement. Furthermore, 60.27% (44 papers) implemented strategies to enhance explainability, which included attention visualization and analysis.
Conclusion
Automated ICD coding tasks have improved, but ongoing challenges remain. These challenges include a lack of diverse data, inadequate use of medical knowledge, complex algorithms, and insufficient validation by clinical coders. Those issues obstruct the model implementation. Future research should focus on integrating a wider range of multimodal data, enhancing the application of medical knowledge, and improving the explainability of models.
Keywords
Introduction
The International Classification of Diseases (ICD) is crucial for efficiently storing, retrieving, and analyzing health data. It supports payment systems, service planning, and the administration of quality and safety in healthcare settings. 1 The critical role of ICD in optimizing healthcare delivery underscores the commitment of hospital administrators to enhancing the quality of ICD data.
However, manual ICD coding presents significant challenges. Coders must meticulously review medical records, verify contexts, consult with physicians, and follow frequently updated coding guidelines.2,3 This complex task requires extensive skills in clinical medicine, health statistics, and disease classification. Moreover, manual coding is prone to errors, with accuracy rates ranging from 50% to 98%, and a median accuracy of 80%.4,5 It is also time-consuming; a study by the American Health Information Management Association (AHIMA) on coding productivity 6 showed that coders processed about 24 inpatient records per day, spending roughly 20 min per record using the ICD-9 Clinical Modification (ICD-9-CM) system. In NHS Scotland, clinical coders typically handle around 60 cases per day. 7 With the shift to the more detailed ICD-10 Clinical Modification/Procedure Coding System (ICD-10-CM/PCS) the average time per inpatient record increased to approximately 38 min. 6
Given these challenges, the healthcare industry needs tools that enhance and streamline the coding process. Automated clinical coding, a subset of Computer-Assisted Coding, utilizes artificial intelligence techniques, such as natural language processing and machine learning, to improve efficiency and data quality. 8
Previous reviews
9
on automated clinical coding have primarily analyzed this task from a computer science perspective. ICD coding intersects multiple disciplines, requiring a strong understanding of medical knowledge and interdisciplinary teamwork to comprehend the task and effectively improve the model method. This review aims to provide a thorough, clinically based evaluation of automated ICD coding models built using the Medical Information Mart for Intensive Care (MIMIC) dataset. This review advances the field in several key areas:
It highlights the clinical perspective of ICD coding by emphasizing practical classification needs, data characteristics, and the importance of domain-specific medical knowledge, as discussed in the “Introduction” section. It thoroughly examines the development of ICD coding models across essential aspects such as data handling, knowledge integration, model frameworks, paradigms, evaluation metrics, and explainability, as detailed in the “Result” section. It evaluates model performance across studies using the MIMIC dataset and a consistent data-splitting strategy, enabling fair comparison of evaluation metrics over publication years and paradigms, as shown in “Evaluation metrics and results” section. From a practical standpoint, it evaluates whether current models fulfill real-world clinical needs and provides clear, actionable recommendations for their improvement and implementation in healthcare, as discussed in “Discussion” section.
Characteristics of the ICD system
Continuous updating
The ICD, established by the World Health Organization (WHO), is a classification of diseases that can be defined as a system of categories to which diagnoses of diseases and other health issues are assigned according to established criteria. 10 Introduced in 1976, ICD-9 included nearly 5000 categories, while ICD-10, introduced in 1995, expanded to about 8000. The ICD-11 further increased granularity to over 50,000 categories to meet the growing demands of morbidity data analysis. In practice, coding often requires greater specificity, extending to detailed levels in Clinical Modifications for billing and reimbursement purposes.11,12
The latest update, ICD-11 for Mortality and Morbidity Statistics (ICD-11-MMS), was rolled out in early 2022. This version introduced significant changes, such as a new chapter structure, new diagnostic categories, and revised diagnostic criteria. A key innovation in ICD-11 is the “Foundation Component,” a semantic network that builds a comprehensive polyhierarchy of medical concepts. 13
Structure and principles
The ICD taxonomy is organized into a hierarchical structure with parent–child nodes, showing clear levels of inheritance and distinct layers. The ICD system does not aim to mirror the real world directly but instead categorizes diseases based on necessary and sufficient conditions to establish mutually exclusive disease category classifications. 14
The principles of ICD classification are manually defined. For example, pregnancy and perinatal conditions are prioritized and categorized into the “special groups” chapters. Sibling nodes at the same level in the ICD taxonomy are organized by axes, which include etiology, pathology, anatomical location, and clinical manifestations. Conditions that cannot be classified according to these axes are categorized as “other” conditions, including rare conditions and ‘unspecified’ cases. Additionally, the system supports co-occurrence codes, which indicate combinations of conditions such as tumor morphology, pathogens, or causes of injury. 10
Insufficient definitions
The definitions provided in Volume 1 of the ICD Regulations regarding nomenclature offer those working with statistics a clear explanation of what is included and excluded in the categories, subcategories, and tabulation list items in statistical tables. However, these definitions often lack enough semantic depth and context, making classification difficult and confusing. Therefore, there is a need for additional external knowledge and guidelines.
To accommodate local practices and meet country-specific reporting requirements, many nations and regions have developed modified versions of the ICD system. 15 For example, the United States uses ICD-10-CM, Canada employs ICD-10 Canadian Modification (ICD-10-CA), Germany relies on ICD-10 German Modification (ICD-10-GM), and Australia adopts ICD-10 Australian Modification (ICD-10-AM). In China, the ICD-10-CM has been under development and in use since 2017 to meet local categorization needs.
Characteristics of the automated ICD coding task
Input: Clinical notes
Early research in automated coding mainly concentrated on classifying term excerpts by extracting “diagnosis descriptions".16,17 However, the brevity of these descriptions often resulted in a lack of comprehensive contextual information, leading to suboptimal outcomes. With advancements in NLP technologies, models have been trained on large public Electronic Health Records (EHRs) datasets. These include the MIMIC, 18 UKLarge and UKSmall, 19 CLEF dataset, 20 and CodiEsp dataset, 21 among others. The MIMIC dataset is the most widely used published research, supporting numerous state-of-the-art (SOTA) advancements in automated ICD coding. Consequently, studies utilizing this dataset are particularly significant for comparing and furthering field research.
Clinical notes consist of free text with a large amount of professional medical terminology and noisy elements such as non-standard synonyms and misspellings. These documents are usually extensive, covering a wide range of clinical information, including health profiles, laboratory test results, radiology reports, operative notes, and medication records. As a result, they tend to be lengthy, with an average of 1609 words in MIMIC-III and 1151 words in MIMIC-II.
Output: ICD codes
All public datasets exhibit the following common characteristics in the distribution of ICD data: Firstly, the label and feature spaces in ICD classification are extremely large. For instance, the ICD-9-CM and ICD-10-CM coding systems contain over 14,000 and 70,000 codes, respectively. This extensive label space complicates the prediction process, causing many models to mainly focus on classifying the top 50 or 100 codes. Additionally, the distribution of ICD coding shows a long-tail pattern. A few codes, like those related to respiratory infections and coughs, appear very frequently in Electronic Medical Records (EMRs), while most codes are rarely used. 22
We analyzed the distribution of diagnosis and procedure codes in the MIMIC-III dataset. The results are shown in Figures 1 and 2, respectively. In the MIMIC-III dataset, 21.8% of the diagnostic codes appear only once, and about 4201 labels appear between one and ten times. Similarly, 20.1% of the procedure codes appear only once, and about 1200 labels appear between one and ten times. More seriously, over 50% of the diagnostic and procedure codes, approximately 17,000, never appear in the dataset. These characteristics make Extreme Multi-label Text Classification (XMTC) techniques highly suitable for managing the large-scale label spaces in ICD datasets, as research has demonstrated their effectiveness.23,24

Distribution of diagnosis labels in the Medical Information Mart for Intensive Care III (MIMIC-III) dataset.

Distribution of procedure codes in the Medical Information Mart for Intensive Care III (MIMIC-III) dataset.
Knowledge-based model
Task models for ICD classification require not only the ICD standards and disease information but also patient characteristics, anatomical sites, etiology, pathology, diagnostics, prognosis, and the classification rules based on these factors. Although biomedical knowledge sources like ICD ontologies, Systematized Nomenclature of Medicine Clinical Terms (SNOMED CT), and coding standards offer expert-curated insights, reliable methods to represent and integrate this knowledge have limited their practical usefulness. Developing computational techniques to represent and interpret these standards is essential for accurately guiding models to assign ICD codes in real-world applications. 25
Methods
Eligibility criteria
This study presents a systematic review of automated ICD coding models developed using the MIMIC dataset. This review followed the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA). 26 The inclusion and exclusion criteria in this review are detailed in Appendix 1.
Information source
The studies were sourced from a range of high-quality academic platforms, including PubMed, ScienceDirect, IEEE Xplore, arXiv, and SpringerLink. Additional sources included the Association for Computing Machinery (ACM) Digital Library, as well as conference proceedings from the Association for Computational Linguistics (ACL) Anthology, the Association for the Advancement of Artificial Intelligence (AAAI) Conference on Artificial Intelligence, and the Conference of the North American Chapter of the Association for Computational Linguistics (NAACL). The review protocol was prospectively registered in the International Prospective Register of Systematic Reviews (PROSPERO), hosted by the National Institute for Health Research (NIHR), to avoid duplication and enhance credibility. The registration was completed before the formal literature search, with the registration number: [CRD42023457388]. The protocol specifies the review objectives, inclusion and exclusion criteria, data sources, and analytical tools to guarantee that the study selection and synthesis follow a planned, systematic methodology.
Search strategy
To construct the search query, keywords within each conceptual group were combined using the OR operator, while different conceptual groups were linked using the AND operator. The specific keywords used for the search are listed in Appendix 2.
Selection process
The retrieved publications were stored in Zotero 6 (Corporation for Digital Scholarship) reference management software. Duplicates were identified and removed using the software's tools, supplemented by manual deletions. After deduplication, 3098 records remained. These were screened by title and abstract to assess their relevance, narrowing the selection to 262 publications. In the second stage, full texts were retrieved automatically or via library access. The third stage involved a detailed review of each full text, focusing on the methodology and experimental sections to ensure the inclusion criteria were met. As a result, 162 papers were excluded, and 73 papers published between 2014 and 2024 were included in the final review. All 73 papers explicitly used the MIMIC dataset to develop and evaluate automated ICD coding models. The detailed study selection pathway is illustrated in Figure 3 (PRISMA flow diagram).

Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) flow diagram for the review.
Data collection, extraction, and synthesis methods
The key variables were collected on the following aspects: Data handling and preprocessing (specific methods and categories), knowledge integration (detailed sources and categories), modeling paradigms (specific approaches and categories), evaluation metrics (e.g., F1 score, Area Under the Curve (AUC), and Precision), and model explainability (specific interpretability mechanisms).
After data collection, organization, and verification, this review classifies and summarizes the included studies by categories and conducts descriptive statistical analyses. For model evaluation, it performs comparative analyses based on publication year and modeling paradigm. Studies that employ the same dataset division method as Mullenbach et al. 27 are included in the comparative analysis of evaluation metrics. If a study reports multiple outcomes, the highest-performing result is selected. The analysis examines two perspectives: Micro-F1 and a composite metric. The composite metric for each model is calculated using a weighted sum of several evaluation metrics: AUC-macro (0.20), AUC-micro (0.15), F1-macro (0.20), F1-micro (0.15), P@5 (0.10), P@8 (0.10), and P@15 (0.10). For missing metric values, a penalty is applied by replacing the missing value with the lowest observed value for that metric across all records. All statistical analyses and visualizations are performed using Python.
Quality assessment and risk of bias
Two independent reviewers meticulously collected data for this review to ensure accuracy and reliability. The reviewers directly contacted study investigators to clarify data points and verify methodological details. Regular consensus meetings were held to resolve discrepancies and ensure consistency in data interpretation.
A quality assessment was conducted to verify that the selected studies aligned with the review's objectives. A checklist of ten closed-ended questions was created, and each publication was required to score at least 7 out of 10 to be included in the final analysis. The checklist items are listed below.
Q1. Is the research objective clearly stated and well-defined? Q2. Is the methodology clearly described? Q3. Is the MIMIC dataset identified and appropriately used? Q4. Are the data preprocessing or input preparation steps clearly described and justified? Q5. Are the input representations (e.g., features, embeddings, or encodings) clearly described? Q6. Are the feature extraction and engineering methods clearly described? Q7. Are the classifiers used in the study clearly described? Q8. Does the study provide a clear and structured comparison with existing baseline models? Q9. Is the system's performance evaluated, and are the results properly interpreted and discussed? Q10. Does the conclusion reflect the research findings?
Results
Overview of included papers
Since their introduction in the 1990s, 28 automated ICD coding methods have significantly improved. The 73 papers included in the review range from 2014 to 2024. Table 1 shows the publication year, paper title, and model name for each study. Of these, 43 achieved SOTA results. Studies marked with an asterisk (*) are gray literature, such as preprints. Risk of bias was evaluated with a 10-item checklist, and all included studies scored at least 7 points. Detailed scores are available in Appendix 3.
Summary of the included papers.

ICD: International Classification of Diseases; GCCN: graph convolutional network; EHR: electronic health records; SOTA: state-of-the-art; SVM: support vector machine; CNN: convolutional neural network; UMLS: Unified Medical Language System; BERT: bidirectional encoder representations from transformer.
— indicates that the study used a model but did not specify its name.
* indicates that the study is a preprint paper.
Data handling and preprocessing
Dataset division strategies
Data division is an essential methodological element in automated clinical coding research. Among the 73 reviewed studies, 91.78% (67 papers) explicitly detail how the MIMIC dataset was divided into training, validation, and test sets. Of these, 58.21% (39 papers) follow the split strategy proposed by Mullenbach et al. 27 This MIMIC-III full codes dataset is divided into 47,719/1632/3372 (train/validation/test), while the top-50 codes dataset is divided into 8067/1574/1730. Papers that used the MIMIC-II full dataset employed a 20,533/2282 (train/test) split.
However, some studies only report proportional splits or omit data partitioning details. As the field matures, clear and consistent reporting of dataset division strategies remains crucial for ensuring reproducibility and enabling reliable comparisons across studies. Appendix 4 provides detailed data split information for each included publication.
Text preprocessing techniques
Clinical narratives are often noisy, sparse, and contain many misspellings, non-standard synonyms, and grammatical errors. Various pre-processing methods have been developed to tackle these challenges, 100 including tokenization, converting to lowercase, removing stop words, sentence segmentation, expanding abbreviations, spelling correction, and lemmatization. Most research on text representation has primarily used models like Word to Vector (Word2Vec), Convolutional Neural Network (CNN), and Bidirectional Long Short-Term Memory (Bi-LSTM). However, recent developments in graph models and large language models have led to more advanced representations used in this test, such as Clinical Bidirectional Encoder Representations from Transformers (BERTs),87,89 PubMedBERT, 62 Bigbird, 74 and ClinicalLongformer. 97
Besides basic text representation, many studies have highlighted the importance of including hierarchical structures in clinical text representations. The HA-GRU model 31 used paragraphs as an additional representation layer for input text, considering the strong hierarchical structure of discharge summaries. Similarly, the EnHANs 35 and DeepLabeler 41 models established word, sentence, or document-level representation layers, creating a hierarchical representation of clinical narratives.
Given the length and complexity of clinical notes, several approaches have been proposed to divide the text into chunks, thus improving model training and prediction efficiency. Models like Hierarchical BERT, 70 HiLAT, 79 and SCB-T 89 models create text chunks based on token length, while more advanced methods segment clinical notes based on semantic content or semi-structured medical information. For instance, The CM model 94 introduces the DF-IAPF algorithm, which automatically segments clinical notes based on their semi-structured characteristics, thereby reducing data variability. Additionally, the LAHST model 96 organizes and segments input data using clinical note timestamps to preserve temporal order in the data.
To further address the challenges posed by complex medical texts, more advanced data augmentation techniques have been introduced to improve model performance. The CAML model 80 employs tools like SemEHR 101 and MedCAT 102 for data augmentation and synthesis; The LAHST model 96 utilizes the Extended Context Algorithm (ECA) to enrich datasets by providing more context, thereby improving model predictions.
Knowledge integration sources
The task of automated clinical coding is inherently knowledge-guided; in the analyzed articles, 69.57% (48 papers) incorporated knowledge in automated clinical coding. Following Hu et al., 103 we divide the knowledge designed for ICD coding tasks into three types: Text knowledge, knowledge graph, and rule knowledge. The sources, classification, and application methods of each type are detailed in Appendix 5. Table 2 provides statistics on the types of knowledge integrated into different studies. The statistical results clearly show that research on knowledge integration has steadily increased since 2019. The specific applications of each type of knowledge are described as follows.
Statistics on types of knowledge integrated in the included papers.

Knowledge graphs
Entity Knowledge
The literature focusing solely on using Entity Knowledge resources accounts for 35.82% and remains a consistently popular approach. This method depends entirely on the textual descriptive information of ICD codes, which includes labels, definitions, synonyms, and terminology. Common practices include using the ICD label information released by the WHO27,34,56,66,6787–89,94 and leveraging other terminology libraries, such as the Unified Medical Language System (UMLS),43,51,73,92,93,95,99 Medical Subject Headings,42,97 and PyMedTermino. 80 Some studies employ entity linking75,80 techniques to enrich the descriptions of ICD codes. Early research used models such as Word2Vec, CNN, and Bi-LSTM to generate ICD description representations, while recent studies have utilized pre-trained models like ClinicalBERT,87,89 PubMedBERT, 85 and RoBERTa,88,96 etc., to obtain ICD description embeddings.
Triplet Knowledge
The utilization of the Triplet Knowledge resource accounts for 38.81% within this study. Early research primarily focused on hierarchical relationships, capturing only “parent-child” connections and failing to represent more complex interactions, such as mutual exclusion or weak links between different code families. SCB-T model 89 developed the Hierarchical Information Transmission module using Gated Recurrent Unit (GRU) cell algorithms to capture and utilize hierarchical relationships between ICD codes, enhancing prediction accuracy, particularly for rare codes. However, this performance improvement involves higher model complexity and more computation.
With technological advancements and a deeper understanding of the task, recent studies have expanded to incorporate co-occurrence relations derived from datasets such as MIMIC,57,61,93 ICD ontologies,48,49 and UMLS.92,99 Wang et al. 98 investigated external auxiliary knowledge from EHR data, including Diagnosis-Related Group (DRG) and Current Procedural Terminology (CPT) codes, by combining it with co-occurrence relations of ICD labels. While these relations show whether two codes appear together in the training data, they do not specify the type or nature of the relationships. The CGNEHR model 99 indicates that integrating external knowledge graphs, like UMLS, does not align with the specific coding rules of ICD systems, resulting in decreased performance compared to baseline models.
Text-based knowledge
This category of methods focuses on enhancing the representation learning of ICD-related knowledge by leveraging text data sources such as medical domain texts (e.g., Wikipedia documents about diagnoses),30,39,47,60,77 and de-identified medical records from datasets like the US Veterans Health Administration Corporate Data Warehouse. 84 Other sources include the Partners HealthCare Biobank, 47 Biomedical Semantic Indexing and Question Answering, Hallmarks of Cancers biomedical literature, 45 and MIMIC data.58,77,79,97 With advancements in large model technology, these approaches have become increasingly popular since 2021, accounting for 23.88% of the methods used. Most studies integrate medical text knowledge by directly employing pre-trained language models (PLMs), conducting separate pre-training stages, or incorporating such knowledge into encoder architectures.
Rule-based knowledge
ICD coding rules are mainly based on standardized processes and guidelines, which are often formalized as sets of rules and terminologies used within the healthcare system. For example, the mapping from SNOMED CT to ICD-10 mapping rules primarily involve gender, patient age, acquired versus congenital conditions, poisonings, external causes, dagger, and asterisk. 104 In research literature, the TreeMAN 81 model attempts to extract information such as patient physiological indicators, gender, admission type, and treatment events from the MIMIC dataset and trains decision trees based on these characteristics of structured data.
Gaps in medical knowledge utilization
Based on extensive experience and intuition in the medical field, we believe the following knowledge sources can be highly effective for this task: SNOMED CT contains a vast collection of concepts, relationships, and descriptions. By leveraging the sufficiency and necessity conditions defined in this terminology system and utilizing its existing OWL representation, the accuracy of ICD coding can be significantly enhanced. 105 Another valuable resource is the SNOMED CT to ICD-10-CM mapping, 104 which includes manually edited logical rules and mapping examples. These mappings capture the expertise of medical coding professionals, enabling the model to learn from human expertise and enhance coding accuracy. 106 Furthermore, ontology representations of ICD-10 or ICD-11 can provide valuable semantic knowledge for the coding task. The principles, such as inclusion, exclusion, and “code also,” encapsulate complex relationships and dependencies between ICD codes. 13
Model frameworks
The typical architecture of ICD coding models includes four core layers: Input layer, representation layer, feature combination layer, and output layer. Larger models often have parameter counts over 100 million, whereas smaller models range from 10 to 30 million parameters. Further details about each study's model architecture, representation, feature layers, output strategies, training techniques, and parameter sizes can be found in Appendix 6.
i. Input layer
The input layer handles diverse data sources, including clinical narratives, multimodal clinical data, and structured domain knowledge (e.g., ICD ontologies). Input information is tokenized using various techniques tailored to the specific language expressions.
ii. Representation layer
This layer converts tokens into vectorized representations. Techniques include traditional embeddings such as Word2Vec
107
and FastText ,
108
Statistical methods such as Term Frequency-Inverse Document Frequency (TF-IDF) and Bag-of-Words, along with deep learning-based dynamic representations like RNNs (LSTM, GRU) and CNNs. Recent studies increasingly incorporate contextualized embeddings from pre-trained models like BERT and its variants to better capture semantic nuances.
iii. Feature combination layer
The feature combination layer creates more advanced architectures to capture multi-level and richer features. It uses methods like attention mechanisms, multi-scale feature extraction, and complex structures such as RNNs, CNNs, or Transformers. The aim is to ensure that the extracted features match the structural characteristics of clinical data and support the decision-making processes of coders.
iv. Output layer
The output layer converts learned features into ICD codes using classification, retrieval, and generation. Classification remains the most common method. Retrieval techniques offer greater flexibility by finding the most relevant codes. Generation-based methods dynamically create ICD codes.
While not inherently part of the model architecture, training strategies are crucial for the model's performance. Typically, models are optimized using objective functions like binary cross-entropy loss or ranking loss. Advanced techniques have been developed to tackle specific challenges, for example, the JLAN model 72 includes a Truncation Loss function and a Dynamic Threshold Function to minimize noise impact in ICD coding. The CoRelation model 93 combines cross-entropy loss with a complexity penalty loss to simplify relationship reasoning and enhance the model's efficiency in handling complex ICD code relationships.
Modeling paradigms
With technological advancements, the modeling paradigms for ICD coding tasks have become increasingly diverse. Each study may adopt one or more paradigms. We recorded and organized those modeling paradigms explicitly stated in the reviewed studies (excluding inferred or implied paradigms) and classified them into five main categories: Deep Learning, Knowledge Representation and Reasoning, Information Retrieval, Machine Learning, and Generation. Details of each study's adopted paradigm(s), model architecture, representation and feature layers, output strategies, training methods, and parameter sizes are provided in Appendix 7, while the frequency of paradigms for the included studies is statistically summarized in Table 3.
Statistical summary of modeling paradigms for the included papers.

CNN: convolutional neural network; GCN: graph convolutional network; SVM: support vector machine; LSTM: long short-term memory; BERT: bidirectional encoder representations from transformers; GAN: generative adversarial networks.
We observed significant differences in how the ICD coding task is defined across various studies. Although it is commonly described as a multi-label classification problem, some studies provide different interpretations of the task. For instance, Zeng et al. 42 define it as an indexing task, mapping medical text to predefined ICD code indices. DeYoung et al. 75 and Ziletti et al. 109 treat it as ontology linking or entity normalization, linking text to concepts within the ICD ontology or normalizing it to ICD codes. Prakash et al. 30 and Guo et al. 51 treat it as a disease inference task, identifying the patient's disease based on the provided information text. NMT models 91 treat the task as a translation problem, converting diagnostic descriptions into the corresponding ICD codes. These distinctions indicate that researchers frequently lack a clear understanding of the difference between “disease classification” and “disease diagnosis,” which could undermine the validity and usefulness of the models.
Deep learning
Deep learning is the most commonly used approach in ICD coding models among the studies surveyed, with 131 times. It acts as a flexible computational framework that can be integrated into different methodologies. When developing deep learning models for ICD coding, several important challenges must be considered.
Firstly, considerable research has been done on model architecture. Some studies have developed hierarchical structures to create deep learning architectures, such as JointLAAT, 46 ZAGCNN, 34 HLAN, 69 Two-stage decoding model, 83 and Hierarchical BERT, 70 XR-LAT. 88 These models utilize attention mechanisms, label embeddings, or various trained structures like BERT, taking advantage of the ICD coding system's natural hierarchy to improve overall performance. While hierarchical models reflect human disease classification logic, they pose risks, such as higher-level errors impacting later predictions. Other studies have constructed complex network architectures, such as capsule networks, 67 Multi-CNN, 53 Multi-Scale,38,73 and Transformers,57,68,85 to extract more features. However, these models often have too many parameters, require high computational resources, and are challenging to optimize and debug. Meanwhile, LSTM models73,83 have also achieved SOTA results.
Secondly, attention mechanisms have evolved considerably over time. Initially, single-layer attention was used to pinpoint relevant keywords. This approach later advanced to hierarchical attention for a more organized text representation. More sophisticated mechanisms, including label-wise attention, parent-child label attention, and multi-head self-attention, have enhanced feature extraction by integrating ICD-specific traits and hierarchical structures. For example, the JLAN model 72 employs a joint learning mechanism to combine self-attention and label attention, creating specialized representations for both high- and low-frequency labels. This reflects a shift from simple stacking to interactive designs, integrating task-specific focus with medical domain knowledge.
Thirdly, recent studies have commonly employed PLMs, such as BioBERT, 110 ClinicalBERT, 111 PubMedBERT, 112 and RoBERTa-PM, 113 which utilize extensive scientific domain data to enhance semantic understanding and ICD coding accuracy. Ji et al. 70 suggest that pre-trained models, like BERT, do not necessarily improve the ICD coding results. The Gpsoap model 84 uses the SOAP structure to create pre-training tasks. While these models possess strong semantic capabilities and benefit from large corpora, they are complex and may exhibit limited gains. The XR-LAT model 88 emphasizes the importance of domain-specific context, as it was pre-trained on biomedical data using the BIGBIRD model.
Lastly, various deep learning techniques have been developed to address specific challenges in ICD coding. Few-shot learning methods, like AGM-HT 59 and Mining Inter-Code Relations, 82 target issues related to sparse label prediction. Transfer learning, discussed in Li et al.'s study, 41 utilizes knowledge from related areas, though its effectiveness depends on the correlation between source and target tasks. The ISD 57 model employs a self-distillation mechanism to minimize noise in input texts. Contrastive learning approaches have also been used to improve feature representations, including text-label contrastive learning in the FLASH framework, 86 graph contrastive learning in the CoGraph model, 60 and tree-based contrastive learning in the CM framework. 94 Multi-task learning methods, such as Yang et al., 87 combine ICD with CPT and DRG code classification, highlighting the benefit of handling diagnostic and procedural classifications separately. The KEMTL 92 model views these tasks as a multi-task learning problem, encompassing ICD coding, treatment recommendations, and mortality prediction.
Knowledge representation and reasoning
The knowledge representation and reasoning paradigm was mentioned 28 times in the surveyed studies. Early research focused on utilizing deep learning techniques such as Bi-LSTM, CNN, and Embeddings from Language Models for knowledge representation. With advances in knowledge graph representation and application technologies, many studies have aimed to develop structured and semantic ICD knowledge bases. For instance, GRU36,60,89 and Graph Convolutional Network (GCN)38,48,74,82,99 have been applied to learn coded representations on the ICD graph.
The KEMTL model 92 employs a Graph Attention Network that integrates the UMLS medical knowledge base to construct a heterogeneous textual graph. This approach effectively captures essential information within clinical texts and elucidates the semantic relationships between concepts. In comparison, the GCNEHR model 99 established GCNs based on UMLS for this purpose, but it did not yield significant performance improvements. Furthermore, many studies underutilize ICD knowledge and neglect the broader semantic relationships among medical concepts. The complexity of incorporating knowledge graphs and the necessity for extensive hyperparameter tuning pose additional challenges for practical model training.
Several studies have investigated multimodal methods58,81 that combine structured data with textual information. These approaches have shown that integrating structured data can significantly improve the performance of ICD coding. For instance, the ICDXML model 97 addresses the diverse nature of PLMs and domain knowledge using a Multi-modal Factorized Bilinear operation. This technique effectively incorporates domain knowledge while maintaining the original semantic information.
Other specific representation techniques have been developed to capture ICD code knowledge. For instance, Hyperbolic Geometry 48 learns continuous vector representations reflecting the ICD hierarchy, while the Path Generator models 49 ICD coding as a process of generating paths along the ICD tree. Wang et al. 98 utilize the BM25 algorithm to extract co-occurrence information of ICD labels and employ the Graphormer model to encode the semantic relationships between these labels. The CoRelation model 93 utilizes a Contextualized Code Relation Learning mechanism that dynamically captures the intricate relationships between ICD codes in the processed case context. This model performs exceptionally well in the top-50 and full dataset results, making it one of the best models in Knowledge Representation and Reasoning paradigms.
Information retrieval
Information retrieval techniques for ICD coding have been utilized 14 times in the surveyed studies. These studies focus on transforming clinical texts and terminologies into vector spaces to improve the effectiveness of matching and mapping. In earlier research, Perotte et al. 16 combined TF-IDF features with Support Vector Machines (SVMs) classifiers for ICD coding. The C-MemNNs 30 model utilized stored information to assist in diagnostic retrieval, while Shi et al. 33 and Guo et al. 51 implemented attention mechanisms to align diagnostic descriptions with ICD knowledge. The MSMN model 56 enhanced ICD code representation by leveraging synonyms through a multiple-synonym matching network. The KSI model 39 computed matching scores between clinical notes and external knowledge sources like Wikipedia. The UNITE model 47 utilized word vector techniques to create representations from both EMRs and online knowledge sources.
Ranking methods aim to optimize label ranking in multi-label classification problems. The MADE Reranker 61 was the first to apply re-ranking methods to prioritize the selection of primary diagnoses and procedures. It adjusted coding sequences by estimating probabilities and leveraging label correlations. To tackle challenges such as a large label space and long-tail label distribution, Wang et al. 98 introduced a two-stage retrieval (utilizing auxiliary knowledge and BM25) and re-ranking phase (incorporating contrastive learning and label co-occurrence relationships), achieving a 6.40% improvement in F1-micro scores on the full dataset. Similarly, the FLASH framework 86 presents an innovative retrieval and re-ranking method, resulting in a 5.10% improvement in F1-micro scores on the full dataset. The two-stage retrieval and re-ranking model 98 and the FLASH framework 86 represent significant advancements in the category of retrieval methods.
Machine learning
Machine learning methods have been applied in ICD coding in 12 surveyed studies. Early approaches primarily utilized SVM.16,32,43 The G_Code model 53 generates adversarial examples to enhance sample diversity and improve the robustness of ICD code assignments. The RPGNet model 49 uses adversarial and reinforcement learning to frame ICD coding as a path generation task. The MCDA model 77 employs Latent Dirichlet Allocation (LDA), an unsupervised learning method, to extract medical concepts from clinical notes and Wikipedia. Additionally, some studies combine structured data with Decision Trees44,81 and deep learning architectures to enhance model performance.
Generation
Research on generative paradigms is limited; it has been applied 5 times in studies. The prompt-based paradigm has recently gained attention. The AGM-HT model 59 uses a Wasserstein Generative Adversarial Network with Gradient Penalty to leverage the hierarchical structure of ICD codes, improving zero-shot classification. Yang et al. 78 adopt a prompt-based fine-tuning approach, framing the task as filling in prompts, while another study 84 employs an autoregressive encoder-decoder for generating ICD codes using a cloze-style prompting method.
Evaluation metrics and results
Evaluation metrics
In the study by Mullenbach et al., 27 the model was evaluated using three datasets: MIMIC-III full codes, MIMIC-III top 50 codes, and MIMIC-II full codes. The reported metrics included AUC-macro, AUC-micro, F1-macro, F1-micro, and Precision@5, Precision@8, and Precision@15. These benchmarks have served as important reference points for later research, with many subsequent models comparing themselves to and striving to exceed these benchmark performances. Detailed metric results from the literature are included in Appendix 8.
Performance outcomes
In ICD coding research, most studies focus on ICD-9 coding tasks due to the MIMIC dataset only containing ICD-9 codes, while only a few have extended to manual annotation and handling of ICD-10 classification. For example, Xu et al. 44 and Rajendran et al. 58 manually mapped 32 ICD-9 codes to ICD-10 codes. The HAN GRU model 114 used mapping tables to convert 5935 unique ICD-9-CM codes to ICD-10, which could introduce potential noise or inaccuracies. Additionally, the study by DeYoung et al. 75 involved professionals undertaking tasks like ICD-10-CM coding, ordering, and entity annotation. Given that most studies focus on ICD-9 classification, this review exclusively compares and analyzes the classification outcomes of ICD-9 coding.
We used Mullenbach et al. 27 as the baseline and organized the literature by year to analyze differences in F1-micro, which is the most frequently reported metric, and composite metrics, as shown in Figure 4, Figure 5. The analysis shows an upward trend in the F1-micro metric since 2019, with the most notable improvements observed in the MIMIC-III top-50 dataset. Composite metrics have shown consistent improvement since 2022. In the full dataset, the results from the HLAN 69 and BERT-hier + LAN 70 models fall below the baseline, and in the top-50 dataset, the results from Jin et al. 90 fall below the baseline.
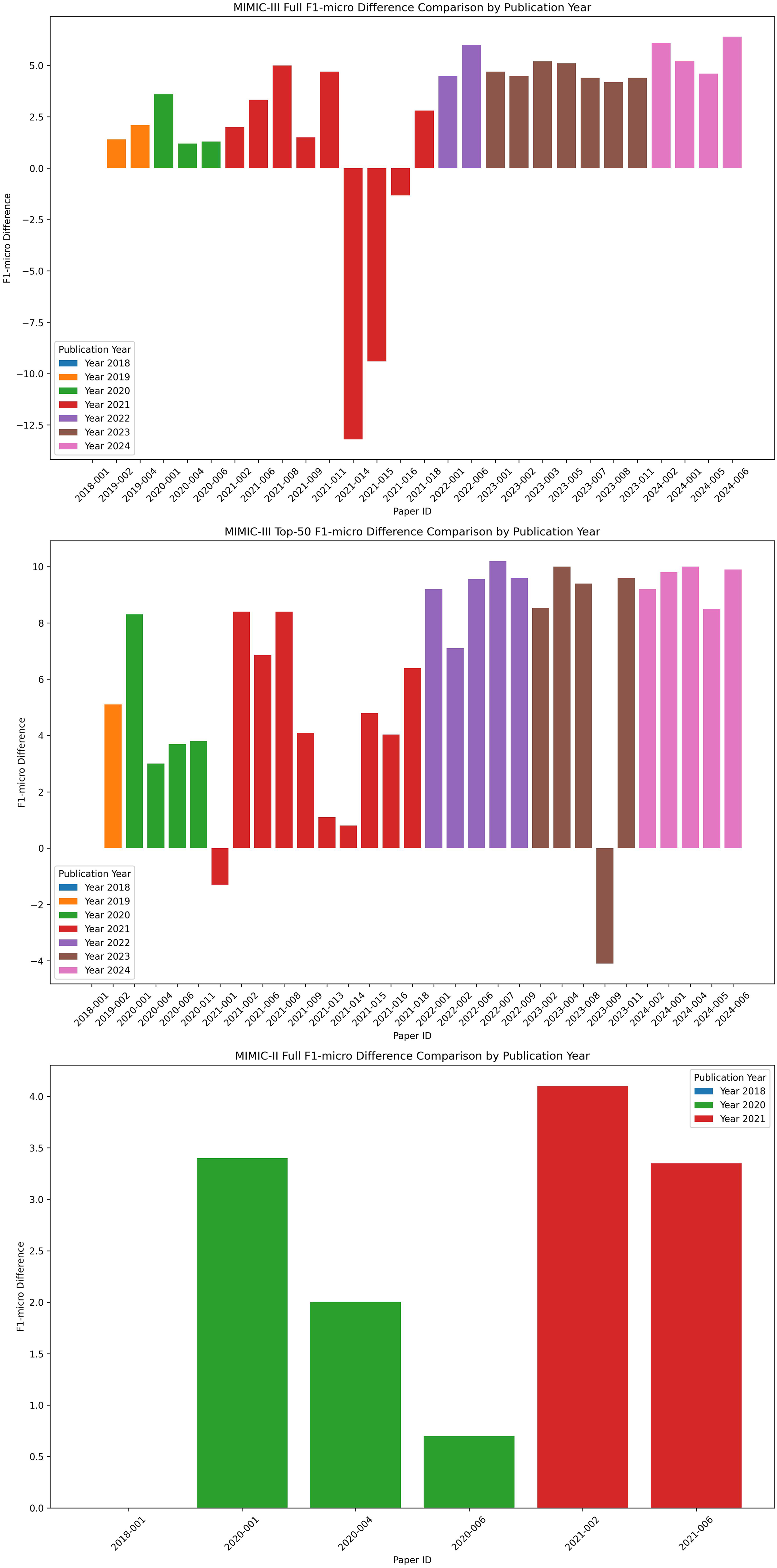
Medical Information Mart for Intensive Care (MIMIC) F1-micro scores difference comparison by publication year.
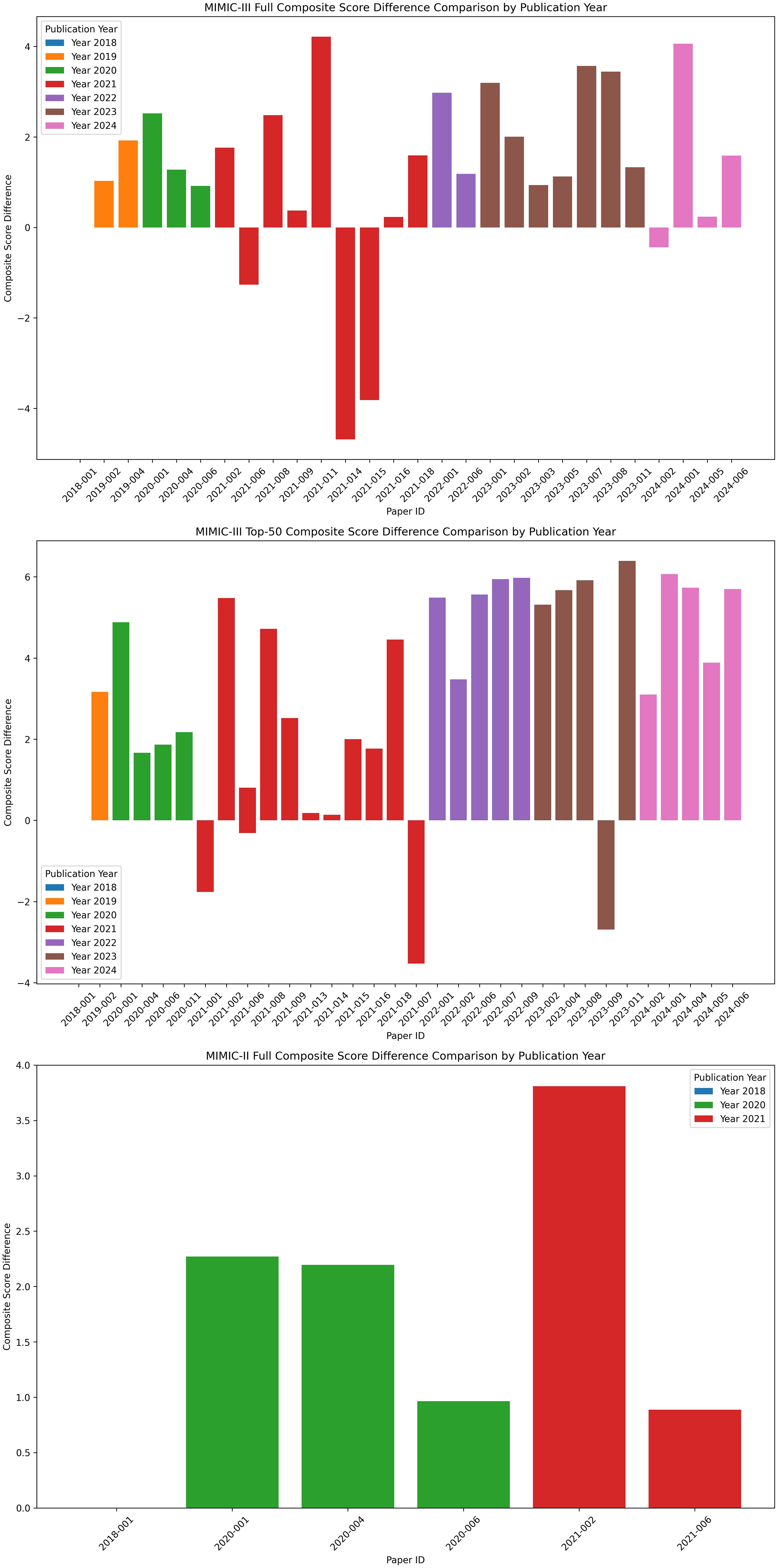
Medical Information Mart for Intensive Care (MIMIC) composite scores difference comparison by publication year.
The literature was categorized by paradigms, and statistics for F1-micro and composite metrics were summarized for each paradigm to compare model performance. The results indicate that knowledge-based and generation methods significantly improve F1-micro scores, especially when using the full dataset. In contrast, deep learning, knowledge-driven, and information retrieval methods perform better in composite metrics. The Violin plot comparisons for each paradigm are presented in Figures 6 and 7.

Violin plot of Medical Information Mart for Intensive Care (MIMIC) F1-micro scores difference by paradigm.

Violin plot of Medical Information Mart for Intensive Care (MIMIC) composite scores difference by paradigm.
Metrics improvement in ICD coding
Currently, few studies have explored the classification tasks of label quantity and order. The ML-Net model 45 develops a label count prediction network, treating label quantity prediction as an N-way classification task and using a multi-task learning approach to simultaneously predict label quantity and ICD codes. Additionally, Tsai et al. 61 were the first to use a reranking method to adjust the order of automatic ICD coding. However, these studies lack evaluations or analyses of label quantity and order predictions.
Perotte et al.16 introduced novel metrics to improve evaluation metrics, including shared path and depth metrics. These metrics utilize the hierarchical structure of ICD codes to assess the relationship between predicted and gold standard codes, thereby facilitating error analysis. Amigo and Delgado 115 proposed the Information Contrast Model for multi-label hierarchical extreme classification. This model compares the informational content of predicted and actual label sets, effectively addressing challenges such as hierarchical similarity and class imbalance.
Model explainability
Case analysis and visualization techniques
Early research on automated ICD coding largely overlooked the importance of interpretability. Among the literature reviewed, 60.27% (44 papers) tried to enhance interpretability, primarily through qualitative visualization and case analysis methods.
In case analysis, Luo et al. 93 examined various coding systems and their interrelationships, while Williamson et al. 95 concentrated on rare codes. Falis et al. 80 introduced the Weak Hierarchical Confusion Matrix method, which allows for a more nuanced evaluation of errors and links algorithmic outcomes to professional expertise. Despite these advancements, many case analyses still fail to provide comprehensive assessments of classification results from the perspective of ICD coding professionals.
Regarding visualization, the HA-GRU model 31 made a groundbreaking contribution in 2017 by employing attention mechanism visualization to tackle the interpretability challenges in ICD classification. Following this innovation, many studies31,40,48,68,69 have adopted similar methods to clarify model predictions, focusing on keywords and sentences related to specific ICD codes. The HiLAT model 79 further improved this understanding by comparing these keywords with established ICD knowledge bases like SNOMED CT and UMLS. Additionally, the HLAN model 69 quantitatively identified the most significant words and sentences for each label and compared its findings across multiple models. However, discussions about professional aspects and the mechanisms of visual interpretability are often overlooked in the model. Most existing research inadequately examines the logical relationships between visualization keywords.
Explainability improvements in ICD coding
Automated ICD coding is a specialized task where interpretability is crucial. However, deep learning methods are often considered “black boxes,” making it difficult to understand their decision-making processes. Recent studies 116 have shown that attention weights do not always effectively explain model decisions, as they can be influenced by various factors like data and model parameters.
To enhance the explainability of ICD coding models, clinical experts should evaluate the outputs of algorithms. Research should establish reasoning pathways similar to those employed by human coders. Knowledge-enhanced PLMs integrating explicit reasoning and dynamic relationship modeling from various knowledge sources can significantly improve performance and interpretability. 96 High-quality medical datasets, such as the MDACE dataset, 117 along with multimodal sources like laboratory test data, medical imaging, and associated reports, can increase the reliability of ICD coding and improve the clarity of coding explanations. Furthermore, building on the studies conducted by Balkir et al. 118 and Darwiche and Ji, 119 the focus on generating “sufficient and necessary explanations” can provide more reliable justifications for predictions.
Discussion
Common errors in medical data, including diagnostic and ICD codes, pose serious challenges for healthcare systems worldwide.120,121 These errors often necessitate extensive human resources for quality management. It is essential to develop automated decision-support tools for ICD coding to address these issues. These tools need to process medical documents that can be structured in various formats, may be lengthy, noisy, and often incomplete. The outputs of these tasks tend to be imbalanced and involve a wide range of labels. Additionally, the classification systems and coding rules are complex and subject to change, which makes effective implementation in real-world medical settings quite challenging. 8
Our study explored the development of automated ICD coding models based on the MIMIC dataset from 2014 to 2024, focusing on both computer science and clinical perspectives. To address the critical dimensions of this evolving research field, we analyzed and compared the results using quantitative classification methods. The key findings and synthesis of the reviewed studies are as follows:
I. Input data
To ensure reproducibility and comparability, algorithms must use consistent and well-documented data division strategies. Paragraph-level representation, text chunking, and structured data extraction can further enhance performance. Currently, studies in automated clinical coding primarily rely on specialized datasets, including MIMIC, Centers for Disease Control, and CodiEsp dataset.
100
This review focuses on widely used published research datasets, specifically the MIMIC datasets. However, these datasets have inherent limitations. The data primarily comes from intensive care unit (ICU) patients, resulting in a biased distribution of diseases, with severe conditions potentially being overrepresented. Furthermore, the dataset lacks diversity, including only a small subset of all possible ICD-9 codes.
122
Williamson et al.
95
noted that the limited data extraction points and inadequate labeling were primary constraints in their research.
II. Knowledge integration
Of the 48 papers reviewed, 69.57% utilized various types of knowledge, including text-based information, knowledge graphs, and rule-based systems, with knowledge graphs being the most commonly used method. The primary sources of knowledge included the ICD ontology, the UMLS ontology, and medical information sourced from platforms like Wikipedia. The CGNEHR system faced performance issues due to the limited alignment between utilizing the UMLS knowledge graph and coding-specific requirements. The integration methods employed included graph algorithms, PLMs, and hierarchical algorithms, highlighting the importance of incorporating medical knowledge to enhance model performance. However, effectively integrating and utilizing this knowledge remains a challenge, as standards, structures, and guidelines related to ICD coding have not been fully utilized.
III. Model frameworks and paradigms
Algorithm development has evolved from traditional machine learning to more advanced paradigms, including deep learning, knowledge representation and reasoning, information retrieval, and generative models. With the development of PLM, the application of knowledge representation and reasoning has increased significantly. Deep learning techniques have been extensively explored, including few-shot learning, transfer learning, self-distillation, and contrastive learning. Developing these methods aims to replicate the reasoning processes used by clinical coders. This involves the innovative construction of complex model architectures, such as multi-scale and multi-head structures. However, these advancements often introduce increased complexity, which may lead to substantial computational costs and pose challenges in real-world applications.
IV. Performance outcomes
Studies that employ the same dataset division method as Mullenbach et al. 27 are included in the comparative analysis of evaluation metrics. Literature reviews indicate a consistent improvement in F1-micro scores since 2019, particularly for the top-50 dataset from MIMIC-III. Additionally, composite metrics have shown enhancements since 2022. For the full MIMIC-III dataset, the Multi-Stage Retrieve and Re-Rank Model 98 achieved a 6.4% increase in F1-micro scores. For the top-50 MIMIC-III dataset, the HiLAT + ClinicalPlusXLNet Model 79 recorded a 10.2% improvement in F1-micro scores. It is important to note that larger models do not always ensure better classification performance, as several smaller models73,77 have also achieved SOTA results.
Methods that employ Knowledge Representation and Reasoning and information retrieval generally achieve better F1-micro scores. In contrast, deep learning and Knowledge Representation and Reasoning perform better on composite metrics. Models based on Information Retrieval paradigms provide significant advantages; this improvement is mainly due to the models’ ability to tackle challenges inherent in ICD coding tasks, such as the extremely large label space and long-tail label distribution. By treating coding as a retrieval task, these models effectively narrow the set of candidate codes, eliminating the need for extensive computations across numerous classes. The integration of domain-specific knowledge not only improves the model's understanding of the semantic alignment between clinical narratives and target codes but also enhances interpretability. Research into Knowledge Representation and Reasoning is particularly promising, especially for addressing low-resource conditions, such as rare-50 coding.
95
V. Model explainability
60.27% (44 papers) focused on enhancing explainability through qualitative visualizations, such as attention mechanism visualizations, and case-based analysis methods. However, these approaches often lack thorough review or validation by medical professionals.
Limitations of this review
This review lacks a comprehensive analysis of deep learning algorithms, particularly regarding parameter settings, algorithm transparency, ablation studies, and computational efficiency. A thorough examination of these elements is crucial for understanding the underlying mechanisms of the models, optimizing their performance, and promoting further algorithm development. Future review studies should strive to evaluate these dimensions more systematically and comprehensively to provide deeper insights and practical guidance. As an interdisciplinary field, automated clinical coding requires integrated medical, computer science, and informatics expertise. Therefore, future research should place a greater emphasis on cross-disciplinary collaboration.
Implications for practice, policy, and future research
Future models should be trained on more comprehensive datasets that accurately reflect real-world clinical scenarios across various medical specialties, institutions, and countries. The ICD coding systems are continuously updated to keep pace with advancements in medical technology and changes in healthcare policies. The MIMIC-II and MIMIC-III datasets, which are limited to ICD-9 coding and primarily focus on data from ICUs, pose challenges for broader applicability. As ICD-9 gradually transitions to ICD-10 and ICD-11, it is crucial to address the differences in label structure, semantic density, and granularity when adapting models to these newer coding systems. 123 The recently released MIMIC-IV dataset, which supports multiple ICD versions (excluding ICD-11), represents a significant step toward bridging this gap. 124 Mapping and converting between different ICD systems will be essential for facilitating knowledge transfer and enabling model reuse across standards, ultimately improving coding accuracy and applicability. Furthermore, integrating multimodal real-world clinical data, such as clinical narratives alongside structured information (such as lab results and vital signs) and diagnostic imaging (such as chest X-rays and reports), can greatly enhance contextual understanding and diagnostic accuracy. 7 This integration is particularly valuable in complex cases involving complications, comorbidities, or unclear documentation. Additionally, it establishes a robust chain of evidence that supports model interpretability.
The review highlighted the significant advantages of knowledge representation and reasoning and information retrieval in models, emphasizing the importance of incorporating broad and deep domain knowledge for future advancements. The ICD coding task is a tightly regulated medical task that demands precise input, accurate outputs, and clear interpretability. It is essential to integrate various forms of medical knowledge, including ICD coding standards and guidelines, clinical terminology, and treatment pathways. The efficiency, accuracy, and consistency of EHR data representation should be based on domain-specific medical knowledge. For instance, adopting standardized frameworks such as Fast Healthcare Interoperability Resources (FHIR) can provide a practical solution for more effectively representing medical information. 125 In the scope of medical knowledge, efforts should focus on verifying the validity of knowledge sources, enhancing the density and richness of information embedded in models, and developing efficient organizational structures for managing medical knowledge.103,126
The primary goal of automated clinical coding is to improve efficiency and alleviate the burden of manual coding. However, several practical challenges need to be addressed carefully. These challenges include aligning with regional healthcare administration and insurance regulatory requirements, designing effective workflows for EHR integration, and ensuring the quality of ICD coding assessments, including diagnostic ordering, accuracy, and completeness. Establishing trust among clinicians is crucial, as it allows physicians to validate and depend on the outputs of models more effectively. Visualizing “reasoning chains,” which illustrate the sources of text evidence and coding rules and guidelines, can significantly enhance this trust. According to Dong et al., 8 automated clinical coding that is human-centered, explainable, intelligent, and robust enough to handle complex real-world scenarios still faces significant challenges. To effectively tackle these challenges, promoting interdisciplinary collaboration between medical informatics and computer science is essential.
Conclusion
This review systematically evaluates automated ICD coding models developed using the MIMIC dataset from 2014 to 2024. The analysis shows that 69.57% (48 papers) of the reviewed studies incorporate various forms of medical knowledge, with knowledge graphs being the most commonly used. Algorithm development has advanced from traditional machine learning to more sophisticated paradigms, such as deep learning, knowledge reasoning, information retrieval, and generative models. Knowledge representation, reasoning, and information retrieval have shown significant improvement. There has been a consistent improvement in F1-micro scores since 2019, and composite metrics have shown enhancements since 2022. There was a 6.4% increase in F1-micro scores for the full MIMIC-III dataset, while the top-50 MIMIC-III dataset recorded a 10.2% improvement. Additionally, 60.27% (44 papers) of the studies include efforts to enhance model explainability, primarily through attention visualization and case-based analysis.
This review highlights several critical limitations in the development of models. For instance, the MIMIC data in these models lacks diversity, and the application of medical domain knowledge has not been fully realized. Additionally, developing these models involves high algorithmic complexity and insufficient validation by clinical coders. These factors reduce their reliability in clinical settings. Future research should prioritize key areas such as incorporating a wider variety of multimodal data sources, more effective integration of medical knowledge, and enhancements in model explainability. There is a significant gap between algorithm development and practical application, requiring multidisciplinary experts’ collaboration and effort.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076251404518 - Supplemental material for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset
Supplemental material, sj-docx-1-dhj-10.1177_20552076251404518 for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset by Ying Zhang, Chen Lyu, Lu Chang, Hong Yang, Bin Ji and Ling-yun Wei in DIGITAL HEALTH
Supplemental Material
sj-docx-2-dhj-10.1177_20552076251404518 - Supplemental material for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset
Supplemental material, sj-docx-2-dhj-10.1177_20552076251404518 for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset by Ying Zhang, Chen Lyu, Lu Chang, Hong Yang, Bin Ji and Ling-yun Wei in DIGITAL HEALTH
Supplemental Material
sj-docx-3-dhj-10.1177_20552076251404518 - Supplemental material for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset
Supplemental material, sj-docx-3-dhj-10.1177_20552076251404518 for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset by Ying Zhang, Chen Lyu, Lu Chang, Hong Yang, Bin Ji and Ling-yun Wei in DIGITAL HEALTH
Supplemental Material
sj-docx-4-dhj-10.1177_20552076251404518 - Supplemental material for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset
Supplemental material, sj-docx-4-dhj-10.1177_20552076251404518 for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset by Ying Zhang, Chen Lyu, Lu Chang, Hong Yang, Bin Ji and Ling-yun Wei in DIGITAL HEALTH
Supplemental Material
sj-docx-5-dhj-10.1177_20552076251404518 - Supplemental material for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset
Supplemental material, sj-docx-5-dhj-10.1177_20552076251404518 for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset by Ying Zhang, Chen Lyu, Lu Chang, Hong Yang, Bin Ji and Ling-yun Wei in DIGITAL HEALTH
Supplemental Material
sj-docx-6-dhj-10.1177_20552076251404518 - Supplemental material for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset
Supplemental material, sj-docx-6-dhj-10.1177_20552076251404518 for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset by Ying Zhang, Chen Lyu, Lu Chang, Hong Yang, Bin Ji and Ling-yun Wei in DIGITAL HEALTH
Supplemental Material
sj-docx-7-dhj-10.1177_20552076251404518 - Supplemental material for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset
Supplemental material, sj-docx-7-dhj-10.1177_20552076251404518 for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset by Ying Zhang, Chen Lyu, Lu Chang, Hong Yang, Bin Ji and Ling-yun Wei in DIGITAL HEALTH
Supplemental Material
sj-docx-8-dhj-10.1177_20552076251404518 - Supplemental material for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset
Supplemental material, sj-docx-8-dhj-10.1177_20552076251404518 for A systematic review of automated International Classification of Diseases coding models using the Medical Information Mart for Intensive Care dataset by Ying Zhang, Chen Lyu, Lu Chang, Hong Yang, Bin Ji and Ling-yun Wei in DIGITAL HEALTH
Footnotes
Acknowledgements
This work was carried out in the Information Department of Guangdong Women and Children Hospital, in collaboration with the research team from the School of Computer Science, Sun Yat-sen University. The authors would like to express their gratitude to all colleagues and collaborators for their valuable contributions to the technical discussions and manuscript preparation.
Author contributions
Ling-Yun Wei: Conceptualization, project administration, resources, supervision, funding acquisition, writing—review & editing. Ying Zhang: Data curation, formal analysis, investigation, methodology, software, funding acquisition. Chen Lyu: Methodology, validation, writing—review & editing. Lu Chang: Investigation. Hong Yang: Visualization, writing—original draft. Bin Ji: Writing—review & editing.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Natural Science Foundation of Guangdong Province (Grant number: 2021A1515110721).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of data and materials
The data that support the findings of this study are available from Guangzhou Healthcare Security Administration, but restrictions apply to the availability of these data, which were used under license for the current study, and so are not publicly available. Data are however available from the corresponding author upon reasonable request and with permission of Guangzhou Healthcare Security Administration.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
