Abstract
Purpose
Sleep staging is critical for assessing sleep quality and diagnosing sleep disorders. However, manual annotation is both time-consuming and labor-intensive, highlighting the need for efficient automated solutions. This study proposes Kolmogorov–Arnold Networks (KAN)-SleepNet, a hybrid deep learning model designed for automated sleep stage classification using single-channel electroencephalogram (EEG) signals.
Methods
The KAN-SleepNet consists of two primary components: (1) a ConvKAN block that integrates convolutional neural networks with KAN to effectively extract discriminative features from EEG signals, and (2) a Bidirectional Long Short-Term Memory layer to capture temporal dependencies across sleep stages. The model was trained and evaluated using two publicly available datasets: the SleepEDF-78 dataset, comprising 153 recordings from 78 subjects, and the ISRUC-S1 dataset, consisting of 100 recordings from 100 subjects. SleepEDF-78 was annotated according to the Rechtschaffen and Kales criteria, whereas ISRUC-S1 followed the American Academy of Sleep Medicine guidelines. Performance was assessed using accuracy, F1-score, and Cohen's Kappa (κ), and results were compared against baseline models, including SleepEEGNet, DeepSleepNet, TinySleepNet, AttnSleep, GraphSleepNet, and MSTGCN.
Results
Experimental results demonstrate that KAN-SleepNet outperforms existing baseline models across both datasets (all p < 0.05 except vs. AttnSleep, p = 0.051). The KAN-SleepNet achieved an accuracy of 85.1%, F1-score of 80.0%, and Kappa of 0.792 on SleepEDF-78, and an accuracy of 82.8%, F1-score of 80.5%, and Kappa of 0.778 on ISRUC-S1. The model also exhibited strong performance in the challenging N1 stage, with F1-scores of 53.2% and 57.4% on SleepEDF-78 and ISRUC-S1, respectively.
Conclusion
The KAN-SleepNet demonstrates superior performance in automated sleep staging, highlighting its potential as an efficient and supportive tool for clinical sleep analysis.
Introduction
Sleep plays a vital role in maintaining physical and mental health. Quality sleep is essential for physical recovery and cognitive function, while poor sleep is associated with an increased risk of cardiovascular diseases, cognitive decline, metabolic abnormalities, and immunological dysfunctions.1–4 Globally, sleep disorders are highly prevalent. It is estimated that approximately 10% of adults worldwide suffer from insomnia, while obstructive sleep apnea (OSA) affects nearly one billion people.5,6 The prevalence of OSA is approximately 5.7–6.0% in adult males and 2.4–4.0% in adult females.7,8 This highlights the substantial public health burden and emphasizes the importance of accurate sleep analysis for diagnosis and treatment.
Sleep staging, which classifies sleep into distinct stages, is fundamental to evaluating sleep quality and diagnosing disorders. Polysomnography (PSG) is the gold standard for assessing sleep, capturing multiple physiological signals, including electroencephalogram (EEG), electrooculogram (EOG), electromyogram (EMG), and electrocardiogram (ECG). 9 Scoring rules from the Rechtschaffen and Kales (R&K) and the American Academy of Sleep Medicine (AASM) standardize manual annotation.10,11 However, full PSG with manual annotation is not only expensive and labor-intensive but also time-consuming, often taking several hours to score a single night's data, which poses a significant impediment to large-scale applications. 12 These limitations underscore the need for simplified approaches, particularly single-channel EEG, that reduce cost and patient burden while retaining clinical relevance.
Automated sleep staging has been widely studied to overcome the inefficiencies of manual scoring. Deep learning models, such as Convolutional Neural Networks (CNNs) and Long Short-Term Memory (LSTM) networks, extract features from EEG signals effectively.13–19 More recently, Transformer-based models have gained attention for modeling long-range dependencies.20–24 Yet, these approaches often require large training datasets and high computational resources. Moreover, many prior works focus heavily on algorithmic design while underemphasizing the translational and clinical aspects necessary for adoption.
Recent developments in single-channel and wearable EEG systems, including in-ear sensors and dry electrode technologies, have shown promising performance compared to PSG.25,26 These technologies open the way for home-based testing and longitudinal monitoring, addressing clinical accessibility barriers. However, real-world adoption requires robust validation and clinician acceptance. 27
Accurate N1 stage detection remains particularly challenging due to its subtle EEG features and overlap with wakefulness, leading to lower stage-specific accuracy for N1 compared to other stages.28,29 Clinically, N1 is crucial in conditions such as insomnia and periodic limb movement disorder (PLMD), where subtle sleep–wake transitions and fragmented sleep structure are diagnostically significant.30,31 Improved staging of N1 extends the impact of automated sleep analysis beyond OSA, toward insomnia, PLMD, and parasomnias. 31
To address these challenges, we propose Kolmogorov–Arnold Networks (KAN)-SleepNet, a hybrid deep learning model designed for automated sleep staging from single-channel EEG. The KAN-SleepNet integrates KAN and Bidirectional LSTM (BiLSTM) layers to leverage their respective strengths. 32 The KAN extracts features flexibly through adaptive B-spline activations, while BiLSTM captures temporal dependencies across stages. By combining these elements, our approach seeks to overcome limitations of baseline models, especially in early N1 stage classification, and to advance clinically applicable, resource-efficient automated sleep staging.
Materials and methods
Datasets and preprocessing
We evaluated the performance of the proposed model using two publicly available datasets: SleepEDF-7833,34 and ISRUC-S1. 35 These datasets were acquired in different experimental settings and annotated according to distinct sleep scoring guidelines, thereby providing a robust basis to assess the model's generalizability. Both datasets are de-identified and openly accessible, and their use does not require ethical approval under national regulations. All analyses in this study were conducted in compliance with the terms and conditions established by the dataset providers. The key EEG acquisition parameters for the Sleep-EDF-78 and ISRUC-S1 datasets are summarized in Supplementary Table S1.
The SleepEDF-78 dataset consists of 153 PSG recordings from 78 healthy individuals (37 males and 41 females), aged between 25 and 101 years (Table 1). For this study, only the Fpz-Cz EEG channel was used, recorded at a sampling rate of 100 Hz. Preprocessing was conducted following the procedure described by Supratak et al. to ensure fair comparison. 36 Each recording contained extended periods of wakefulness (stage W) at the beginning and end, during which the subject was not asleep. To focus on sleep periods, we included only 30 min of wakefulness immediately before sleep onset and after sleep termination. The PSG signals were segmented into nonoverlapping 30-s epochs and manually scored by expert sleep technologists according to the R&K criteria. In accordance with previous studies,37,38 stages S3 and S4 were merged into a single N3 stage, while MOVEMENT and UNKNOWN epochs were excluded (Figure 1). Consequently, each epoch was categorized into one of five stages: W, N1, N2, N3, and REM. Consistent with previous work,13,36,39 no denoising or filtering procedures were applied to the raw EEG signals.

Terminology used by R&K and AASM for sleep stage classification. In the R&K criteria, the sleep stage is classified into W, S1, S2, S3, S4, and REM. In the ASSM criteria, S3 and S4 are merged into a single stage N3. AASM: American Academy of Sleep Medicine; R&K: Rechtschaffen and Kales.
Clinical characteristics of SleepEDF-78 and ISRUC-S1 datasets.
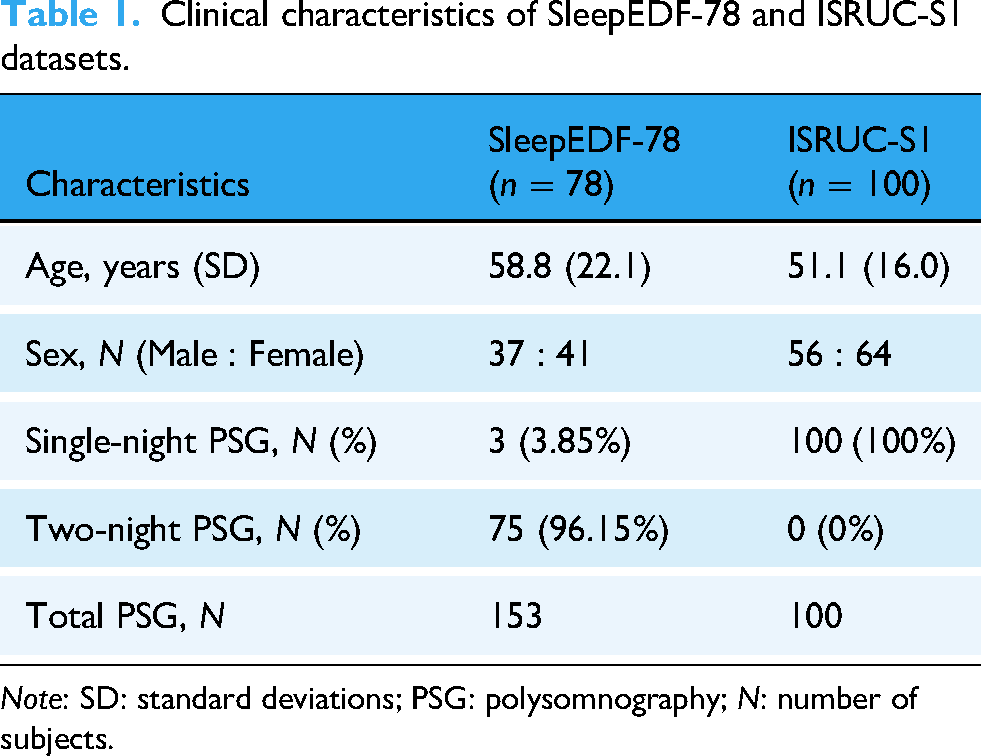
Note: SD: standard deviations; PSG: polysomnography; N: number of subjects.
The ISRUC-S1 dataset comprises 100 PSG recordings from 100 individuals (55 males and 45 females), aged 20–85 years (Table 1). Each PSG recording was independently annotated by sleep experts following the AASM standards, with every 30-s epoch classified into one of five sleep stages: W, N1, N2, N3, and REM. The EEG F3-A2 channel was chosen for model evaluation, and the EEG signals were resampled at 200 Hz. Preprocessing steps have already been performed on the EEG signals in the ISRUC-S1 dataset, including the removal of unwanted noise and Direct Current offset to improve signal quality and signal-to-noise ratio. Specifically, a notch filter was applied to suppress 50 Hz powerline interference, and a bandpass filter (0.3–35 Hz) was used to retain physiologically relevant frequencies. Epochs labeled as MOVEMENT or UNKNOWN were excluded from the analysis, as they are not considered part of the five canonical sleep stages under the AASM criteria.
The distribution of samples across sleep stages for both the Sleep-EDF-78 and ISRUC-S1 datasets is summarized in Table 2, with more detailed information provided in Supplementary Table S2 and Supplementary Figure S1.
Sample distribution of SleepEDF-78 and ISRUC-S1 datasets for sleep staging.
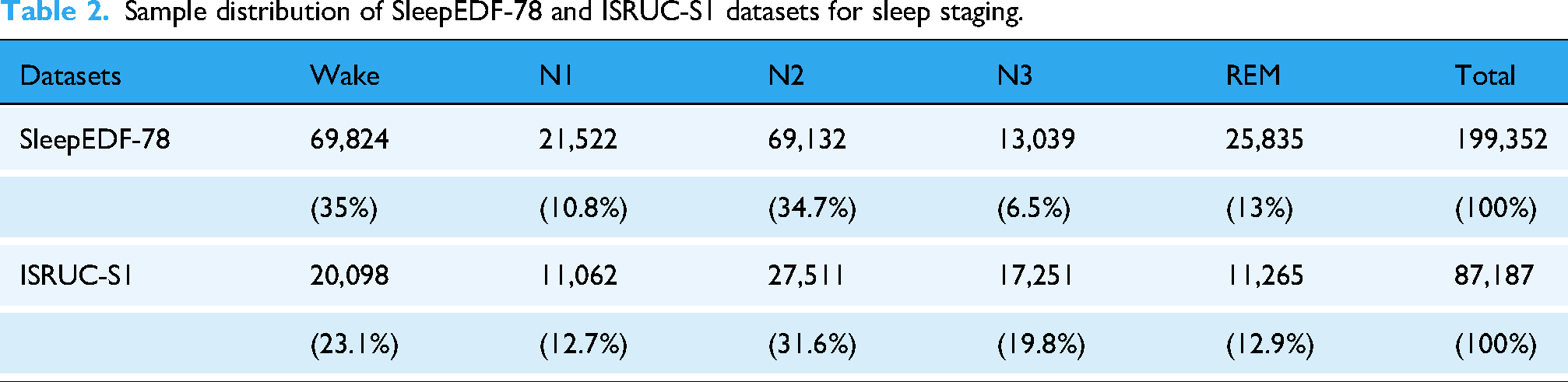
Overview of the proposed model
The proposed model, KAN-SleepNet, combines a KAN layer with a BiLSTM layer for automated sleep stage classification using EEG signals, as illustrated in Figure 2. The model consists of multiple stages designed to extract features from EEG data, enhancing its ability to differentiate between various sleep stages effectively. Implementation code is available at https://github.com/xzhenliang/KAN-SleepNet.
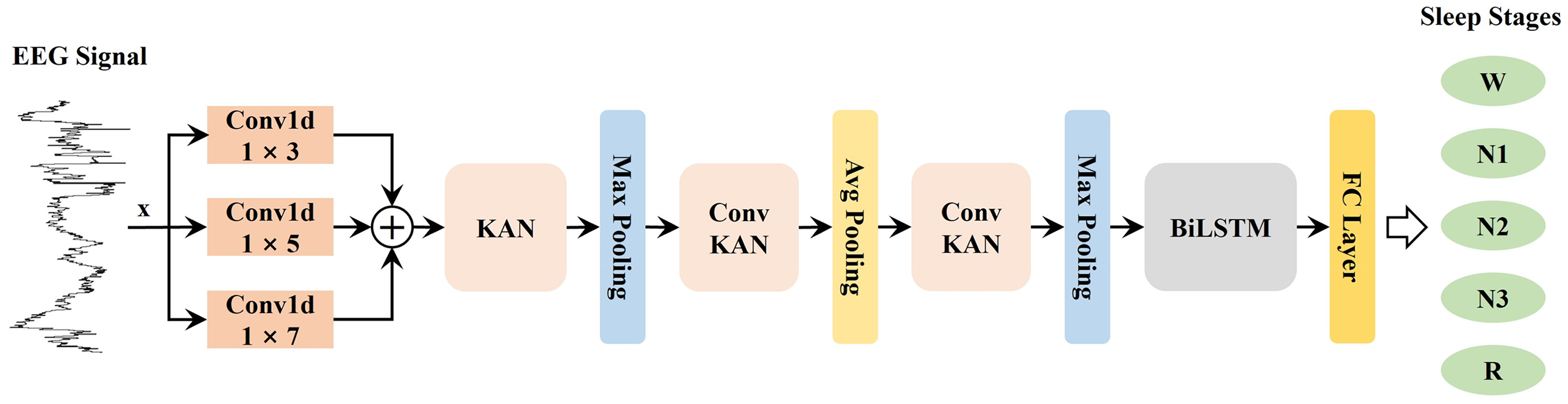
The overall architecture of the proposed model. BiLSTM: Bidirectional Long Short-Term Memory; EEG: electroencephalogram; FC: Fully Connected; KAN: Kolmogorov–Arnold Networks.
Firstly, EEG signals are fed into three parallel 1-dimensional convolutional layers with kernels of sizes 1 × 3, 1 × 5, and 1 × 7. These layers perform convolution operations with different receptive fields, enabling the model to extract multiscale spatial features, thereby capturing both short- and long-term dependencies within the signal. The outputs of these convolutional layers are then summed element-wise, consolidating information across the different filter sizes.
Secondly, the combined features are passed through a KAN layer, which applies learnable B-spline functions along with the edges. This allows the model to adjust activations locally based on the characteristics of the input signal, making it more adaptable to subtle variations in the data. After the KAN layer, max pooling is applied to reduce the dimensionality and retain the most salient features.
Thirdly, the downsampled features are processed through a ConvKAN block (Figure 3), which integrates convolutional operations with KAN. This is followed by average pooling to further distill the feature representations. Subsequently, an additional ConvKAN block and max pooling are applied. This sequential combination of convolution, KAN layers, and pooling operations progressively refines the extracted features, highlighting critical patterns essential for distinguishing between sleep stages.

Schematic representation of the Kolmogorov–Arnold Networks (a) and the ConvKAN Block (b). BN: Batch Normalization; KAN: Kolmogorov–Arnold Networks.
Finally, the refined features are input into a BiLSTM layer (Figure 4), which captures temporal dependencies across consecutive sleep stages. The BiLSTM layer's output, containing both forward and backward context, is finally passed through a fully connected layer to predict the five sleep stage classes (W, N1, N2, N3, and R).

Schematic representation of the Bidirectional Long Short-Term Memory network. LSTM: Long Short-Term Memory.
Kolmogorov–Arnold Networks layer
Inspired by the Kolmogorov–Arnold representation theorem, 40 KAN replace traditional fixed activation functions and linear weights with learnable spline-based activation functions applied along with the network edges. This innovative design enables KAN to effectively capture complex, high-dimensional relationships, enhancing both efficiency and accuracy in function approximation.
The theoretical foundation of KAN lies in the Kolmogorov–Arnold theorem, which states that any continuous multivariate function 
Unlike Multi-Layer Perceptrons, which rely on fixed nonlinear activations at each node, KAN incorporates learnable activation functions along with the edges. Each edge's activation is a trainable spline function, allowing finer control over individual input transformations. This spline-based activation function can be expressed as shown in equation (2):

Here,
Each KAN layer updates node representations by first passing messages and then aggregating and updating nodes. Specifically, the information from node i to node j is represented as shown in equation (4):
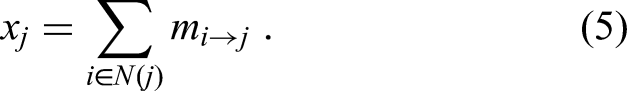
This structure allows KAN to stack multiple layers to increase expressive power and adapt to complex data patterns.
ConvKAN block
Inspired by the Squeeze-and-Excitation Networks (SENet), 41 we propose the ConvKAN block (Figure 3), a novel module designed to enhance feature extraction by combining convolutional operations with the KAN for adaptive feature learning. This block captures both spatial features and complex, nonlinear relationships in the EEG signals, making it highly effective for sleep stage classification.
The ConvKAN block starts with two sequential one-dimensional convolutional layers (Conv1d), each accompanied by Batch Normalization (BN) and a Rectified Linear Unit (ReLU) activation function. These convolutional layers employ one-dimensional kernels to learn local features along with the time dimension of the EEG signal. The BN helps stabilize and accelerate training by normalizing the activations, while ReLU introduces nonlinearity, allowing the model to learn complex patterns.
The output of the second convolutional layer undergoes one-dimensional average pooling (AvgPool1d) to reduce the feature dimensionality and focus on the most representative features. Following the pooling layer, a KAN layer applies learnable B-spline functions, allowing the network to flexibly model localized patterns and variations in the EEG data. This adaptiveness is crucial for capturing subtle, stage-specific features that may vary across sleep stages.
The processed features are then passed through a Sigmoid activation function to produce a gated output. Inspired by SENet, this Sigmoid output is applied element-wise to the original features from the convolutional layers, selectively emphasizing important information while suppressing less relevant details. The gated features are further refined using a ReLU activation function, preparing them for subsequent processing in the network.
By combining convolutional operations, KAN-based adaptiveness, and a gating mechanism, the ConvKAN block efficiently extracts discriminative and stage-relevant features from EEG signals. This block is instrumental in enabling the KAN-SleepNet model to achieve high classification performance across different sleep stages.
Bidirectional LSTM layer
Long Short-Term Memory networks, introduced by Hochreiter and Schmidhuber, 42 have significantly advanced the handling of sequential data by addressing the challenges of vanishing and exploding gradients typical of traditional recurrent neural networks. While LSTMs are effective at capturing long-term dependencies through their memory cells and gating mechanisms, BiLSTMs offer an additional enhancement for sequence modeling tasks.
Bidirectional LSTMs extend the capabilities of standard LSTMs by processing the input sequence in both forward and backward directions. This dual-processing approach allows the network to capture information from both past and future contexts relative to each time step, providing a more comprehensive understanding of the sequence. In a BiLSTM network, two separate LSTM layers operate in parallel: one processes the sequence from the beginning to the end, while the other processes it from the end to the beginning. The outputs of these two layers are then combined, allowing the model to effectively leverage information from both directions, as illustrated in Figure 4.
This bidirectional processing is particularly valuable for tasks where the context from both preceding and succeeding elements in a sequence can enhance the model's performance. Applications such as natural language processing, 43 speech recognition, 44 and time-series forecasting 45 benefit greatly from this approach, as it provides a richer representation of the input data and improves the model's ability to capture dependencies that span across different segments of the sequence. Hence, in this study, our model incorporates Bidirectional LSTMs for sleep staging using EEG signals.
Loss function
To address the issue of class imbalance within the dataset, a weighted cross-entropy loss function was applied. The weight coefficients were determined based on the sample distribution of each class and the relative difficulty in classification. The weighted cross-entropy loss function is defined as equation (6):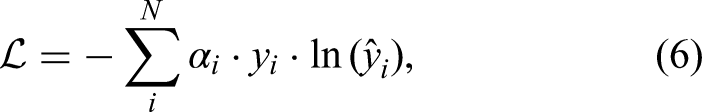
Evaluation metrics
To assess model performance for sleep staging, we employed three primary metrics: accuracy (ACC), macro-averaged F1-score (MF1), and Cohen's Kappa (κ). Additionally, we calculated the per-class F1-score (F1) to evaluate the classification of each sleep stage individually. These metrics were computed by treating each sleep stage as a positive class while treating all others as negative classes in a binary classification manner.
Accuracy (ACC) measures the proportion of correctly predicted outcomes in relation to the total sample size, offering an overall assessment of model performance across all classes. The accuracy is defined as follows (equation (7)):
Precision reflects the proportion of true positive classifications among all predictions made as positive, indicating the model's accuracy in classifying each stage without excessive false alarms. The precision is defined as follows (equation (8)):
Recall is defined as the ratio of correctly identified positive instances to the total actual positives. This metric evaluates the model's ability to identify all relevant instances for each sleep stage. It is calculated as follows (equation (9)):
The F1-score, which represents the harmonic mean of precision and recall, is calculated as follows (equation (10)):
This metric provides a balanced evaluation of precision and recall, particularly useful in the presence of class imbalance, as often observed in sleep staging.
Cohen's Kappa (κ) measures the agreement between predicted and true labels beyond what is expected by chance. It provides a robust metric for assessing classification performance. The Kappa coefficient is defined as follows (equation (11)):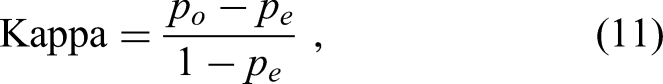


Experiment details
Our KAN-SleepNet model was developed using PyTorch v1.11.0, and all experiments were conducted on an NVIDIA A100 80GB PCIe GPU. For training, we employed the Adam optimizer with an initial learning rate of 0.001. The learning rate was dynamically adjusted using an exponential decay schedule with
In order to maintain the independence between the training and test sets, we adopted a subject-wise data splitting strategy. The dataset was partitioned into training (85%) and test (15%) sets at the subject level. Furthermore, 15% of the training set was allocated as a validation set. Crucially, for subjects with two-night PSG recordings, all data from the same individual were exclusively assigned to either the training, validation, or test set, thereby preventing any data leakage across splits. To avoid unnecessary training, early stopping was employed, halting training when the validation loss showed no improvement for 20 consecutive epochs. The model checkpoint with the highest validation accuracy was selected as the final model. The batch size is set to 16, and the maximum number of epochs is 150. The model's performance was evaluated on the test set. A paired t-test was applied to assess the statistical significance of the performance differences between KAN-SleepNet and the baseline models across all metrics, and results with p < 0.05 were considered statistically significant.
To address class imbalance, a common issue in sleep staging, we employ a weighted cross-entropy loss function during training. This adjustment helps mitigate bias towards more frequent classes and enhances the model's sensitivity to less-represented stages, particularly the N1 stage.
Results
Overall performance
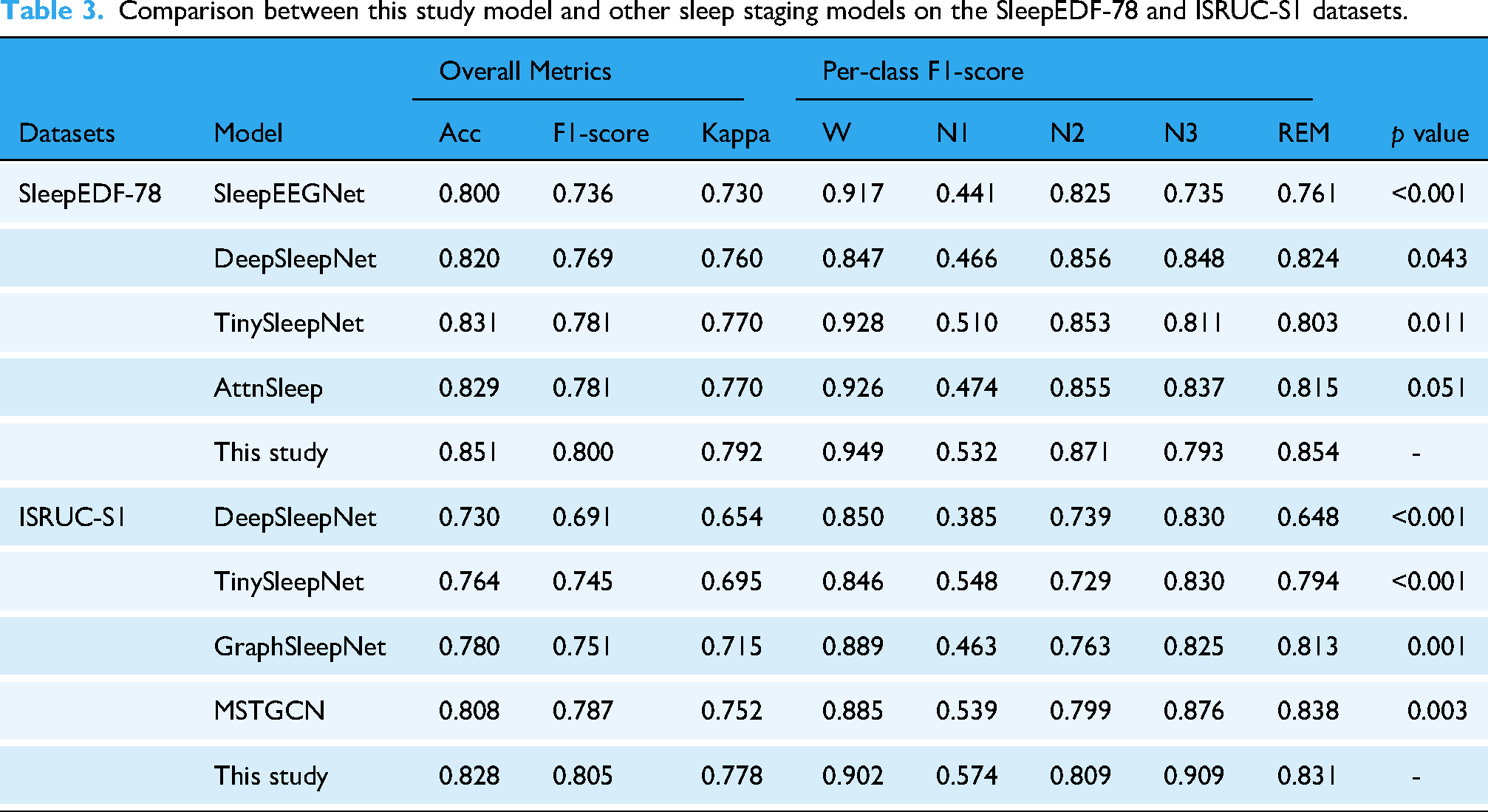
The proposed KAN-SleepNet model was evaluated on the SleepEDF-78 and ISRUC-S1 datasets, achieving promising performance in automated sleep staging. On the SleepEDF-78 dataset, KAN-SleepNet achieved an accuracy of 85.1% and an F1-score of 80.0%, while on the ISRUC-S1 dataset, it attained an accuracy of 82.8% and an F1-score of 80.5% (Table 3). KAN-SleepNet was compared with baseline models, including SleepEEGNet, 46 DeepSleepNet, 39 TinySleepNet, 36 AttnSleep, 20 GraphSleepNet, 47 and MSTGCN 48 (Table 3 and Figure 5). KAN-SleepNet exhibited superior performance, particularly in detecting the challenging N1 stage (Figure 6). Moreover, it achieved balanced classification across all sleep stages, representing a significant improvement over these methods. The overall and per-class performance metrics with 95% confidence intervals for the KAN-SleepNet are provided in Supplementary Table S3.

Performance comparison of various sleep staging models on the SleepEDF-78 and ISRUC-S1 datasets. The models are evaluated using overall Accuracy, Macro F1-score, and Cohen's Kappa coefficient.

Comparison of per-class F1-scores achieved by different models across sleep stages on the SleepEDF-78 and ISRUC-S1 datasets.
Comparison between this study model and other sleep staging models on the SleepEDF-78 and ISRUC-S1 datasets.
Per-class performance
The confusion matrices, presented in Figures 7 and 8, provide comprehensive results of the model's performance across all sleep stages. For the wake (W) stage, KAN-SleepNet achieved per-class accuracies of 94.9% and 90.2% on the SleepEDF-78 dataset and ISRUC-S1 dataset, respectively (Table 3). The N1 stage, challenging due to class imbalance and subtle EEG features, achieved per-class accuracies of 53.2% on the SleepEDF-78 dataset and 57.4% on the ISRUC-S1 dataset (Table 3).

Confusion matrix (left panel) and normalized confusion matrix (right panel) on the SleepEDF-78 dataset.

Confusion matrix (left panel) and normalized confusion matrix (right panel) on the ISRUC-S1 dataset.
Ablation study
An ablation study was conducted on the SleepEDF-78 dataset to further evaluate the contributions of individual components (Table 4). The results showed a significant performance drop when critical components were removed. Specifically, removing the ConvKAN block reduced the model accuracy to 67.7%, indicating the importance of the KAN layer in extracting discriminative features from EEG data. Similarly, excluding the BiLSTM layer resulted in a reduction in accuracy to 78.6%, highlighting the crucial role of temporal modeling in sleep stage classification.
Ablation study conducted on the SleepEDF-78 dataset.

In addition, experiments were conducted to evaluate the impact of varying the number of ConvKAN blocks on model performance. It was observed that using four ConvKAN blocks yielded the best performance, achieving an accuracy of 85.1% and an F1-score of 80.0%.
Discussion
In this study, we proposed KAN-SleepNet, a hybrid deep learning model for automated sleep staging from single-channel EEG. Across two publicly available datasets (SleepEDF-78 and ISRUC-S1), the model consistently outperformed several representative baselines in terms of accuracy, macro F1-score, and Cohen's κ. In particular, KAN-SleepNet achieved balanced classification across all five sleep stages, with noticeable improvements in the challenging N1 stage compared with existing methods. These results demonstrate the robustness of the model across datasets that differ in population characteristics, scoring criteria, and recording settings.
The improved performance of KAN-SleepNet can be attributed to the synergistic design of its core components. A critical innovation of this research is the incorporation of ConvKAN blocks into the model architecture. Unlike traditional CNNs, which rely on predefined filter structures, KAN layers are capable of approximating complex mathematical functions. This allows the model to effectively capture subtle and nonlinear patterns in EEG signals, which is particularly critical for sleep staging due to the highly dynamic nature of EEG data and significant intersubject variability. 49
The temporal modeling capabilities of the BiLSTM block further enhance the model's performance by capturing the sequential dependencies inherent in sleep stage transitions. Sleep staging is a time-dependent process, where transitions between stages often follow a predictable pattern. 50 For example, transitions from REM sleep to N1 or from N2 to N3 are influenced by physiological processes. By integrating BiLSTM, the model can account for these temporal dependencies, as evidenced by the significant performance drop when the BiLSTM block is excluded.
Another important contribution is the model's ability to handle class imbalance, a prevalent issue in sleep staging. The N1 stage presents unique challenges due to subtle EEG features, with traditional models often underperforming.20,36,39,46,51 KANSleepNet achieved F1-scores of 53.2% (SleepEDF-78) and 57.4% (ISRUC-S1) for N1 classification. These results demonstrate significant improvement over baseline approaches. This improvement can be attributed to the use of a weighted cross-entropy loss function, which compensates for the imbalance by assigning higher weights to underrepresented classes. These findings demonstrate that addressing class imbalance is critical for achieving balanced performance across all sleep stages. While this may hold potential relevance in clinical contexts, further validation is required to confirm its utility for early detection of sleep disorders.
Furthermore, this study explored the impact of varying the number of ConvKAN blocks on model performance. The results indicate that four ConvKAN blocks yield the best trade-off between model complexity and performance. Adding more ConvKAN blocks did not lead to significant improvements, suggesting that overparameterization may introduce redundancy and increase the risk of overfitting. This observation aligns with existing findings in deep learning, 52 where excessively deep architectures may not always translate to better performance, particularly when dealing with limited datasets.
The superior performance of KAN-SleepNet across multiple datasets also highlights its generalizability. Many models perform well on a single dataset but fail to replicate their performance on other datasets due to differences in data distribution, signal acquisition protocols, and demographic factors. By demonstrating consistent results on both SleepEDF-78 and ISRUC-S1 datasets, KAN-SleepNet shows promise for real-world applications.
It is worth noting that the proportion of N3 sleep differs between SleepEDF-78 (6.5%) and ISRUC-S1 (19.8%). This discrepancy is likely attributable to age-related variations in deep sleep. Prior studies have shown that N3 sleep decreases significantly with age.53,54 SleepEDF-78 includes a larger proportion of elderly participants, who typically exhibit reduced deep sleep, whereas ISRUC-S1 involves subjects across a broader age range, including more younger adults. The difference highlights the heterogeneity across datasets, potentially affecting model performance and generalizability.
This study has several limitations. Firstly, the model was trained and evaluated on single-channel EEG signals. While this setting simplifies the architecture and reduces hardware complexity, it excludes modalities such as EOG and EMG, which are well known to provide critical information for REM specificity. The absence of these channels may reduce the accuracy of REM detection compared with full PSG. Secondly, the two datasets employed in this work follow different scoring standards. While this heterogeneity offers an opportunity to evaluate generalizability across distinct labeling criteria, it may also introduce inconsistencies that limit the direct comparability of results. Thirdly, the datasets used in this study are relatively limited in size, which may constrain the model's ability to capture broader interindividual variability and reduce its generalizability to larger or more diverse populations.
Future work should focus on validating KAN-SleepNet on larger and more diverse datasets, incorporating multichannel EEG recordings, and exploring transfer learning strategies to further enhance model generalizability. Beyond algorithmic improvements, several practical directions merit attention. Prospective validation in clinical cohorts, such as patients with insomnia, obstructive sleep apnea, and pediatric populations, will be crucial for establishing clinical robustness and reliability. Moreover, integrating automated staging systems into commercial PSG analysis platforms could facilitate clinical adoption and streamline routine workflow. Finally, applying KAN-SleepNet to longitudinal, home-based sleep monitoring may open new avenues for personalized and precision sleep medicine.
Conclusion
In summary, this study proposed KAN-SleepNet, a hybrid deep learning model that integrates ConvKAN blocks and BiLSTM for automated sleep staging using single-channel EEG. The model achieved consistent improvements over representative baselines across two public datasets, particularly in the challenging N1 and REM stages, which are clinically important for detecting sleep fragmentation and REM-related disorders. These gains suggest a potential to reduce manual scoring time and support more efficient workflows, although such benefits remain to be demonstrated in practice.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076251398440 - Supplemental material for KAN-SleepNet: A deep learning model combining Kolmogorov–Arnold Networks and bidirectional LSTM for automated sleep staging using EEG signals
Supplemental material, sj-docx-1-dhj-10.1177_20552076251398440 for KAN-SleepNet: A deep learning model combining Kolmogorov–Arnold Networks and bidirectional LSTM for automated sleep staging using EEG signals by Zhenliang Xiong, Yuxuan Gou, Yinglin Zhou, Yuwan Yang, Yunsong Peng, Yan Gong, Rui Xu, Rongpin Wang and Xianchun Zeng in DIGITAL HEALTH
Footnotes
Ethical considerations
This study utilized publicly available datasets, and no ethical approval was required.
Contributorship
ZX contributed to writing—original draft, writing—review & editing, conceptualization, methodology, investigation, and visualization. YG contributed to validation, data curation, conceptualization, and writing—review & editing. YZ contributed to validation, data curation, and writing—review & editing. YY contributed to validation, data curation, and writing—review & editing. YP contributed to validation and writing—review & editing. YG contributed to validation, data curation, and writing—review & editing. RX contributed to validation, funding acquisition, and writing—review & editing. RW contributed to validation, resources, and writing—review & editing. XZ contributed to supervision, resources, project administration, funding acquisition, and writing—review & editing.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Guizhou Provincial High-level Innovative Talent Training Program (QianKeHe Platform Talent-GCC [2023] 083), the National Natural Science Foundation of China (Grants No. 82460344 and 82060314), and the Natural Scientific Project of Guizhou Province (QianKeHe Foundation - ZK [2022] General 263).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The data used in this study are publicly available and can be accessed from the SleepEDF-78 dataset (https://www.physionet.org/content/sleep-edfx/1.0.0/) and the ISRUC-S1 dataset (![]() ). These datasets are freely available for research purposes in accordance with the respective usage terms.
). These datasets are freely available for research purposes in accordance with the respective usage terms.
Guarantor
All authors take full responsibility for the content of this article, including the accuracy and appropriateness of the reference list.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
