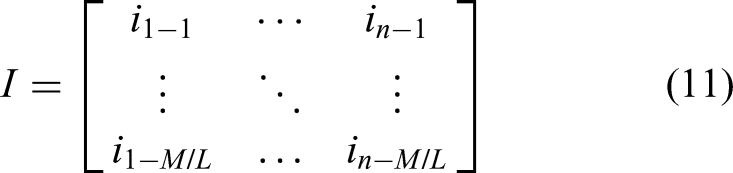Abstract
Objective
In the field of digital health, emotion recognition is vital for human–computer interaction and clinical neuropsychology. However, existing deep-learning models often lack interpretability, limiting their ability to provide insights into brain activity.
Methods
This study proposes an innovative brain-inspired neuronal competition model for emotion recognition from electroencephalography (EEG). First, a discrete wavelet transform is employed to extract five brain rhythms (delta, theta, alpha, beta, and gamma) as feature inputs, maximizing the preservation of signal information. Then, the proposed model simulates excitatory and inhibitory competitive dynamics between neurons, where a rhythm attention encoder dynamically focuses on key brain rhythms and uses a leaky integrate-and-fire model to simulate the neuronal decision-making process. Finally, emotion recognition is implemented based on a winner-take-all principle.
Results
Ten-fold cross-validation experiments on three widely used datasets, DEAP, SEED, and DREAMER, achieve average accuracies of 86.64–92.70% on DEAP, 81.89% on SEED, and 80.38–82.70% on DREAMER, respectively. These results indicate that different brain rhythms exhibit distinct characteristics in various emotion recognition tasks.
Conclusions
This study presents a biologically plausible paradigm for EEG emotion recognition, laying the foundation for understanding the neurophysiological basis of brain activity and developing reliable tools for healthcare applications, such as mental health monitoring and depression detection.
Keywords
Introduction
Emotion is a complex phenomenon that influences fundamental mental functions. 1 Disruptions in the affective processes are recognized as a core feature of numerous neuropsychological and psychiatric conditions, impacting an individual's mental health and overall well-being. 2 In the field of digital health, an insightful understanding of the cognitive mechanisms that underpin emotional experience is meaningful for advancing clinical practice. Hence, developing objective and precise tools to recognize emotional states from neurophysiological data is a vital pursuit. These advancements present the promise of illuminating the intricate interplay between cognition and emotion, facilitating the clinical identification of diverse psychological distress, such as anxiety 3 and depression. 4
While traditional emotion assessment often relies on overt behavioral cues such as facial expressions and vocal prosody, these methods are susceptible to voluntary control or interpretive bias. 5 To gain a more objective window into emotional states, cognitive neuropsychology has increasingly utilized intrinsic physiological signals, with electroencephalography (EEG) being a particularly effective modality. The high-temporal resolution of EEG is especially valuable, as it enables the capture of fast-paced neural dynamics that characterize the interplay between emotion and cognition. 6 Nevertheless, EEG presents a significant challenge, as emotions are complex neural phenomena arising from the synergistic activity of distributed brain networks, 7 making their direct interpretation from signal signatures inherently difficult. This complexity necessitates the development and application of advanced computational approaches. In related disciplines, such as medical imaging, several methods are tackling similar challenges by designing sophisticated models that integrate diverse information. For instance, Khan et al. 8 demonstrated that fusing features from multiple deep-learning architectures and utilizing spatial attention mechanisms to indicate clinically relevant regions significantly improves diagnostic accuracy for diseases such as tuberculosis.
Furthermore, for such systems to be viable in daily healthcare applications, they must strike a balance between performance and practicality. That means computational efficiency achieved through lightweight models and clinical trust fostered by explainable artificial intelligence techniques are paramount for real-world deployment.
9
Thus, a critical aim in affective neuroscience of digital health is to develop accurate and explainable systems capable of recognizing these intricate neural patterns, which provide insights into emotional states and pave the way for daily applications in healthcare. Based on this, a brain-inspired neuronal competition model is designed in this study by considering the intrinsic operational principles of the brain. Particularly, it models a fundamental principle of competitive interaction among neuronal populations (processing information through excitatory and inhibitory dynamics in response to stimuli). Such a brain-inspired approach aims to bridge the current gap between high-performance emotion recognition and its biological plausibility, providing a deeper understanding of neural dynamics underlying emotional states and fostering the development of more trustworthy solutions for enhancing affective neuroscience and clinical neuropsychology. In short, the main contributions of this study are listed below:
A model grounded in the biological principle of neuronal competition is designed, which simulates decision-making mechanisms inspired by cortical neurons and offers a plausible mechanism for how brain rhythms are utilized to achieve EEG emotion recognition. The proposed model reduces information loss and computational overhead associated with complex hand-crafted feature selection by using brain rhythms as dynamic feature inputs. Therefore, it helps preserve the intrinsic characteristics of EEG signals, which are essential for recognizing various emotional states. The weight encoding module and the stimulation control module are implemented, where the former directs the attentional focus toward brain rhythms of greater significance for emotion discrimination, and the latter enables the model to shift its focus from a global to a local temporal perspective, allowing for the more granular analysis of brain dynamics across multiple timescales.
The rest of the paper is organized as follows: the second section presents the related work. The third section describes the experimental datasets and details the proposed brain-inspired neuronal competition model, including the brain rhythm separator, rhythm attention encoder, pulse elicitor, and neuronal decision unit. The fourth section shows the experimental results. The penultimate section provides an insightful analysis of emotional activities and a comparative study with recent work, facilitating a thorough discussion of the proposed method. Finally, the last section provides the conclusion of this study.
Related work
The existing techniques suggest that reliable EEG emotion recognition typically relies on the pivotal steps of extracting valuable features that characterize emotional states. Generally, EEG features are conceptualized as neurophysiological signatures of emotions within brain activities, as they reveal specific patterns indicative of emotional variations. EEG feature extraction methods are categorized into time domain, frequency domain, and time–frequency domain. Time-domain features typically focus on the statistical properties and morphological variations of signals. For instance, statistical metrics such as mean, variance, skewness, kurtosis, and event-related potential amplitude and latency characterize overall signal energy fluctuations and transient neural dynamics. Frequency-domain features represent the signal energy distribution across various frequency bands. Through the calculation of parameters such as power spectral density, relative power, and rhythm-specific energy within defined frequency bands,10–12 the distinct contributions of different neural oscillations related to emotional processes are investigated, facilitating the selection of appropriate features for classification tasks accordingly. Nonetheless, the inherent non-stationarity of EEG signals often limits the efficacy of purely time-domain or frequency-domain methods in fully showing their dynamic characteristics. Consequently, time–frequency domain approaches have been used to simultaneously resolve dynamic information across both the temporal and spectral dimensions. Typical methods include short-time Fourier transform, Wigner–Ville distribution (WVD), and discrete wavelet transform (DWT), among others, which decompose EEG signals into multiple frequency sub-bands using different time windows. 13 For example, Yeh et al. 14 employed WVD to extract emotion-related features by quantifying energy per second within specific frequency bands. Beyond the transform-based approaches, adaptive signal processing techniques, such as empirical mode decomposition and its variants, 15 which adaptively decompose EEG signals into a series of intrinsic mode functions, can be adopted to characterize complex dynamic properties.
To study emotional and cognitive functions based on EEG, brain rhythms have shown particular advantages, which are defined as neural oscillatory activities within specific frequency bands, that are, 0.5–4, 4–8, 8–13, 13–30, and 30–70 Hz, denoted as delta, theta, alpha, beta, and gamma, respectively. Indeed, substantial evidence corroborates the correlation between brain rhythm activities and variations in emotional states. For instance, Iyer et al. 16 found a positive correlation between pleasant and delta rhythm energy near the midline of the prefrontal cortex. Li et al. 17 claimed that emotional stimuli significantly amplified high-frequency gamma rhythm activities during arousal assessment. Zhang et al. 18 investigated that noise exposure, which often induces negative emotions, is usually associated with increased delta power and decreased theta or alpha power. These findings insightfully indicate the significant potential of brain rhythms as valuable biomarkers of emotion.
In addition to conventional feature extraction techniques, recent advances in deep learning have spurred the widespread adoption of novel feature extraction approaches. Architectures such as convolutional neural networks (CNNs) 19 and long short-term memory (LSTM) 20 automatically learn and extract complex representations directly from EEG signals. Whether EEG features are meticulously hand-engineered through conventional signal processing techniques or automatically extracted using deep-learning networks, they constitute emotion recognition at the neurophysiological level. Subsequently, the primary objective of the classification task is to categorize these representations accurately and map them to human-interpretable emotional categories. Therefore, once methods for extracting features from EEG signals have been elucidated, whether through traditional approaches or by adopting deep learning, the study's focus logically shifts to developing robust classification models. Indeed, selecting an appropriate model is vital, as it directly determines the accuracy of emotion recognition and the overall practical value of the system. 21
Particularly, deep-learning models usually fulfill a dual role in emotion recognition, as they can function as potent feature extraction, converting complex data into more discriminative classifiable representations, or they can be architected as end-to-end learning systems that directly predict emotional states from input signals, tightly integrating feature extraction and classification within the network. For instance, Thiruselvam and Reddy 6 presented an approach employing reduced correlation in frontal EEG signals for pre-screening emotion-eliciting segments, which were then classified using a compact CNN model. This method yielded an average accuracy of 80.87% in subject-specific classification tasks, providing an improvement over the 70.50% accuracy achieved with unscreened EEG segments, and demonstrating the importance of strategic data selection in enhancing method performance. Cai et al. 22 proposed an EEG input representation that integrates spatial- and frequency-domain features, which was subsequently processed by an EEG swin transformer model. In subject-independent experimental paradigms, this model acquired accuracies of 80.07% and 66.72% for three-class and four-class tasks, respectively, showing the suitability of the transformer for processing emotional EEG signals to explore brain activities. Cruz-Vazquez et al. 23 studied the synergy between feature transformation techniques and deep-learning models. They utilized quantum rotations to transform EEG features before inputting them into the models, successfully classifying three emotions (sadness, happiness, and neutral) with an accuracy and F1-score of approximately 95%, which reveals good performance in integrating advanced feature extraction with deep-learning models.
Alongside deep learning, traditional machine-learning methods remain relevant in EEG emotion recognition due to comparatively lower computational demands and better interpretability. For example, Yao et al. 24 developed a technique that involves extracting network theory-based features, such as average node degree (AND) and node degree entropy (NDE). Then, a support vector machine (SVM) was employed for classification. They yielded accuracies of 91.39% (AND) and 85.39% (NDE), respectively, demonstrating the effectiveness of integrating network theory-based feature extraction with traditional classifiers for modeling the topological properties of functional brain networks during emotional activities. Similarly, Colafiglio et al. 25 utilized MiniRocket as a feature extraction method for EEG emotion recognition. While MiniRocket incorporates concepts from convolutional operations, it is not a fully-fledged deep-learning model designed to generate features. Specifically, MiniRocket-extracted features were applied to an SVM, achieving 75–80% accuracy in a four-class task, even with emotional EEG signals from only four channels. Except for these individual classifier approaches, ensemble learning methods, which amalgamate multiple predictions, have also been adopted in previous works.26–28 Usually, compared with a single classifier model, combining various models beneficially enhances overall performance and cross-subject robustness, establishing a promising approach to analyzing complex and highly emotional EEG signals. Recently, Adhikari et al. 29 demonstrated the effectiveness of an approach that uses variational mode decomposition to extract frequency-domain features, which were subsequently classified using a random forest classifier. It shows the continued relevance and viability of traditional feature engineering pipelines.
Methodology
Overall workflow
Figure 1 illustrates the overall workflow of the proposed brain-inspired neuronal competition model for EEG emotion recognition. It receives data input from EEG signals and provides an output with the emotion recognition result. The key processing architecture that links the input and output is constructed from four main operational modules: (i) brain rhythm separator, (ii) rhythm attention encoder, (iii) pulse elicitor, and (iv) neural decision unit.

Proposed method's modular design follows a sequential path, as shown by the arrows, with the output of each stage feeding into the next.
EEG acquisition
The SEED dataset 30 contains emotional data from 15 subjects (seven males and eight females, with an average age of 23.27 ± 2.37 years), comprising 15 trials per subject. Each trial contains a 5-s cue, a 4-min film presentation, a 45-s self-assessment period, and a 15-s rest. The stimuli are film clips associated with one of the three pre-defined emotional labels (positive, negative, or neutral), resulting in a three-class classification task. A 62-channel Neuroscan system was employed for the acquisition of EEG data. These raw signals were originally sampled at 200 Hz and then band-pass filtered within the 0–75 Hz. To maintain methodological consistency across the different datasets, note that the method validation in this study only performs the last 30-s segments of EEG corresponding to each stimulus, as this period avoids noise or irrelevant information that may be caused by fluctuations, focusing more on analyzing emotional response sensitively.31,32
The DREAMER dataset 33 contains recordings from 23 individuals (14 males and 9 females) aged 22 to 33. In this dataset, each subject watched 18 audio-visual film excerpts of varying lengths (65–393 s), intended to induce a range of emotions. After watching each clip, subjects provided ratings for their emotional states on the dimensions of arousal, valence, and dominance. It permits binary classification tasks on these three emotional dimensions, as well as a four-class task derived from the combined arousal–valence dimensions. For data collection, a 16-channel Emotiv wireless system recorded EEG signals at a sampling rate of 128 Hz, with M1 and M2 electrodes serving as ground and reference, respectively. Additionally, 14 scalp EEG channels were available for analyzing emotional responses. Again, the last 30 s of each trial were segmented for method validation in this study.
The DEAP dataset 34 includes EEG recordings from 32 subjects (17 male and 15 female), aged between 19 and 37 years, who were presented with 40 1-min music videos designed to elicit a range of emotional responses. After viewing each video, subjects rated their emotional states on various dimensions, including arousal, valence, and dominance, using a 1–9 scale. This protocol facilitates binary classification tasks for each dimension. As for data acquisition, EEG signals from 32 scalp channels were recorded using a 10–20 system at a sampling rate of 128 Hz.
Note that all datasets used in this study, including DEAP, SEED, and DREAMER, have undergone a standard pre-processing stage that removes explicit artifacts and noise, a crucial step for ensuring the reliability of emotional EEG signals.35,36 Therefore, the datasets provided pre-processed versions that can be employed directly. By utilizing these artifact-free signals, it is beneficial to evaluate the performance of the proposed method, since meticulous pre-processing and innovative model design are complementary, both of which are essential for advancing the field of EEG emotion recognition.
Brain rhythm separator
Considering the relationship between emotion and brain rhythm activity, the proposed model adopts five specific neural oscillations: delta (0.5–4 Hz), theta (4–8 Hz), alpha (8–13 Hz), beta (13–30 Hz), and gamma (30–70 Hz). Thus, the function of the initial module is to extract these five brain rhythms. To this end, DWT is employed, as it has demonstrated effectiveness in the time–frequency decomposition of non-stationary signals such as EEG and its inherent multi-resolution analysis capabilities. The fundamental concept of DWT is the iterative separation of a signal into its low-frequency and high-frequency components across various scales. The decomposition is technically realized through a multi-level filter bank containing complementary low-pass and high-pass filters, as described mathematically below:
Daubechies-4 (db4) is selected as the mother wavelet function for the DWT in this study. Specifically, the low-pass and high-pass filter coefficients with db4 are shown below:
As outlined in Figure 2, a four-level DWT decomposition is performed in this study, where the decomposition operates recursively. The input EEG signals are processed through the high-pass filter during the first level to generate the initial high-frequency detail coefficients (cD1) and low-frequency approximation coefficients (cA1). This procedure is then repeated with the approximation coefficients from each level serving as the input for the next, continuing until the fourth level. The iterative application results in a final set of wavelet coefficients (cA4, cD4, cD3, cD2, and cD1). Following the decomposition, these specific coefficients are utilized to map the five brain rhythms, that are, delta (cA4), theta (cD4), alpha (cD3), beta (cD2), and gamma (cD1), correspondingly. Additionally, to provide a better intuitive understanding of the brain rhythms utilized as features in this study, an example of each rhythm is illustrated in Figure 3.

Four-level DWT is applied to extract corresponding coefficients for five brain rhythms. DWT: discrete wavelet transform.

Variations of extracted brain rhythms and original emotional EEG signals. EEG: electroencephalography.
Furthermore, to provide a clear neurophysiological insight into how the proposed method analyzes emotional states, the relationship between raw EEG signals and their corresponding brain rhythms under different emotions is depicted in Figure 4, where the EEG signals and their brain rhythms exhibit distinct patterns when a subject expresses a low-arousal (LA) state compared to a high-arousal (HA) state. During HA, the amplitude of the high-frequency beta and gamma rhythms appears more pronounced, while an LA state is often associated with a more dominant alpha rhythm. This visual evidence not only demonstrates the effectiveness of the DWT-based approach in extracting vital brain rhythms but also provides an interpretable link between these rhythms and emotional states.

Visualization of the EEG signals and their corresponding brain rhythms in low-arousal (LA) versus high-arousal (HA), where the data is from subject S1 in the DEAP dataset. It demonstrates that different levels of emotional arousal are reflected in the characteristics of specific brain rhythms, providing key evidence of the proposed method in this study. EEG: electroencephalography.
Rhythm attention encoder
Since not all features support emotion recognition, the five brain rhythms extracted are processed through a weight encoding mechanism, which aims to shift the model's analytical focus from a holistic examination of the entire rhythms to a more localized inspection of key temporal windows. Meanwhile, it provides the data appropriately for the subsequent processing stage and mitigates the risk of overfitting. Figure 5 presents the rhythm attention encoder, which takes a brain rhythm of length M as input, supplied by the previously described feature extraction. A detailed explanation is as follows:

Rhythm attention encoder used in the proposed model.
Let R denotes the brain rhythm, W means the weight matrix, and N refers to the target emotional classes to be recognized (e.g. N = 2 for the binary classification). The R and W are provided below:
By using this module, the R is transformed into an encoded matrix, denoted C, as expressed below:
Next, the W is updated dynamically during the training phase, contingent upon classification performance. The specific update strategy for W is:
Pulse elicitor
Figure 6 illustrates the pulse elicitor, which functions as an intermediary that processes the encoded brain rhythms supplied by the rhythm attention encoder to generate the final classification output. Here, two principal operations are consolidated: stimulation control and pulse counter.
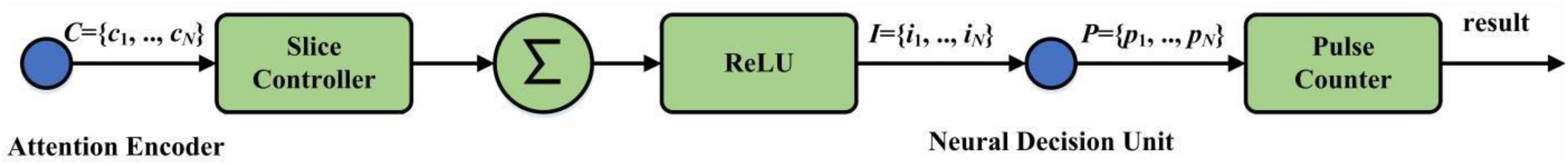
Architecture of the pulse elicitor.
The stimulation control function includes a slice controller, an accumulator (∑), and a rectified linear unit (ReLU) activation function. Overall, it processes the encoded brain rhythm inputs. The whole process starts with the slice controller partitioning the incoming rhythms into segments of a specified length. Each of these segments is then passed through the accumulator and the ReLU activation function to obtain a stimulus value, which is subsequently dispatched by the stimulation outputter.
cn is partitioned into M/L sub-sequences (sk), where k is an integer ranging from 1 to M/L. The operational pipeline for each cn is as follows: first, the slice controller sequentially outputs each sk; next, each sk is fed into an accumulator, wherein its constituent elements are summed to determine an initial stimulus magnitude. The output from the accumulator is then passed through a ReLU aimed at rectifying the stimulus by setting any negative magnitudes to 0, yielding the non-negative electrical stimulus, as expressed below:
Pulse accumulation is a task executed by a pulse counter. Specifically, it tallies the neuronal pulses generated by each class-specific neuron operating within the subsequent neuronal competitive module. Such a tallying process culminates in a set of accumulated pulse counts, denoted as P = {p1, p2, …, pN}, where pN refers to the total pulse count associated with the n-th emotional class over a defined integration period (n = 1, 2, …, N). The final classification is then achieved based on these pulse counts, according to the winner-take-all principle implemented by the argmax function:
Neuronal decision unit
The innovative brain-inspired mechanism within the proposed model is the neuronal competitive module, designed to simulate a key operational principle in cortical information processing. When a stimulus is encountered, specialized groups of neurons in the cerebral cortex often enter a competitive dynamic, a process orchestrated primarily by inhibitory synaptic connections. The resolution of these interactions, which manifests as neuronal firing patterns, is foundational to how the brain makes decisions and categorizes inputs. A classic illustration can be found in the visual system. When the retina registers an image, populations of neurons tuned to distinct visual features, such as vertical or horizontal orientations, can become simultaneously active. Following this initial co-activation, a mechanism known as reciprocal inhibition comes into play, where the currently active neurons diminish the activity of competing neurons through the release of inhibitory neurochemicals. Consequently, the group of neurons that exhibits the most vigorous response to the incoming stimulus prevails. In detail, the designed neuronal competitive module contains three main stages. First, the continuous adjustment of the membrane potential for each neuron representing a specific class is driven by the incoming electrical stimuli it receives. Then, reciprocal inhibitory interactions occur among these different neuronal units. Lastly, a neuronal unit fires an output pulse if its membrane potential reaches a specified threshold of activation.
Given the set of stimuli, each component Ij, which originates from the j-th class and is associated with neuron Nj, exerts an influence on that specific neuron. For instance, when a particular neuron (N1) is driven by its corresponding input stimulus I1, its membrane potential will shift from a resting state to a more depolarized state. This change refers to an elevation in the membrane potential, which is governed by the leaky integrate-and-fire (LIF) neuron model, as described below:
In order to elucidate the inhibition mechanism, consider the specific interaction where an active N1 inhibits N2, at the synaptic junction, N1 could release inhibitory neurotransmitters, which subsequently bind to receptors on the membrane of N2. This binding action typically causes an influx of negative ions (Cl) or an efflux of positive ions (K+). Such ion movement leads to a hyperpolarization or a shunting effect that decreases the membrane potential of N2. Thus, the total inhibitory synaptic current Isyn,j(t) received by neuron j from all other competing neurons k is expressed as below:
At the end of each dt, the membrane potentials for all neurons are updated according to the LIF model. If any Vj exceeds a pre-defined firing threshold Vthresh, then Nj will generate an output pulse. Upon firing, its pulse count
Finally, the accumulated pulse count from each Nj is transmitted to the spike statistics function within the pulse elicitor.
Results
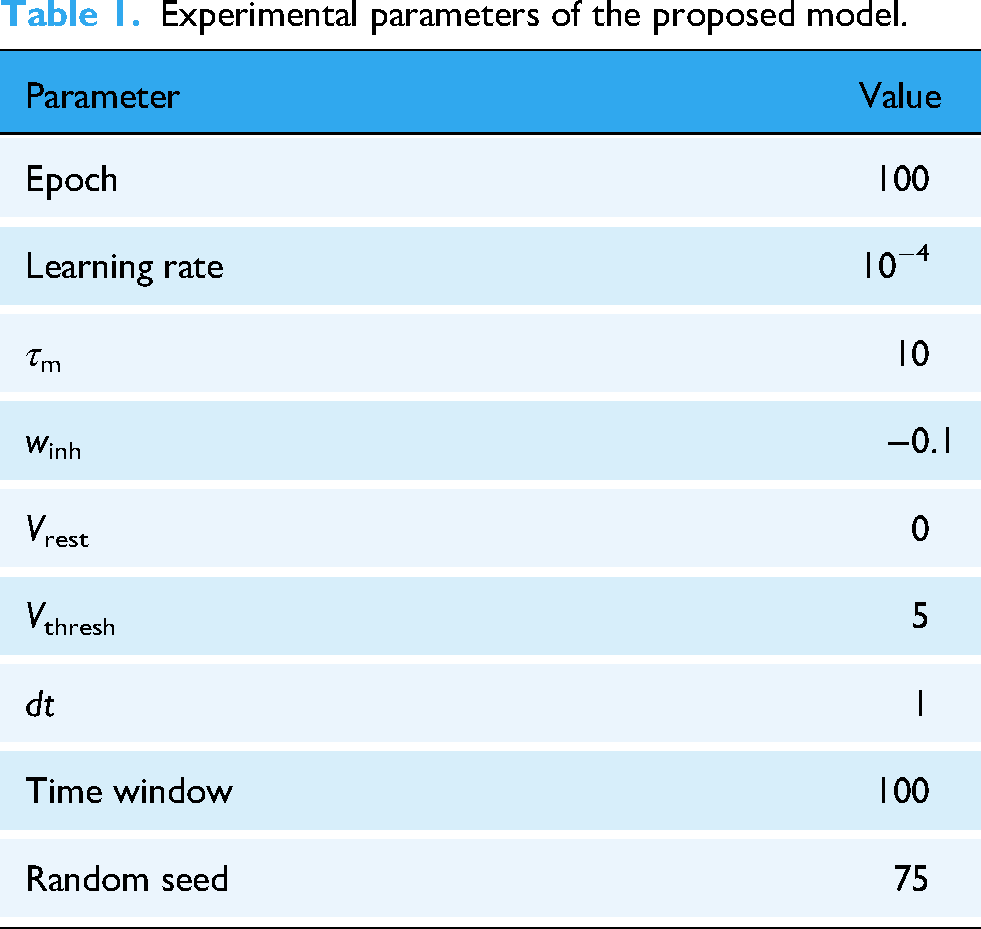
In the experiment, EEG signals were downsampled to 128 Hz for each dataset, and 30-s epochs were segmented to serve as input for the proposed model. Additionally, the evaluation was conducted subject-dependent, meaning that separate training and testing sets were created for each subject from the individual data. Meanwhile, to ensure that the evaluation results were not skewed by the randomness of a single data split, a 10-fold cross-validation was employed. The experimental configurations are listed in Table 1. Furthermore, Tables 2 to 5 present the evaluation metrics (accuracy, F1-score, precision, and recall) from the three public EEG datasets (DEAP, SEED, and DREAMER) using the proposed model.
Experimental parameters of the proposed model.
Classification accuracy (%) using different brain rhythms through the proposed model.

F1-score (%) using different brain rhythms through the proposed model.

Precision (%) using different brain rhythms through the proposed model.

Recall (%) using different brain rhythms through the proposed model.

Regarding the DEAP dataset, which uses music videos as emotional stimuli, a distinct pattern emerges where the gamma rhythm demonstrates better performance, as it achieves the highest accuracy and F1-score for both arousal (accuracy: 91.63%, F1-score: 93.87%) and valence (accuracy: 92.70%, F1-score: 92.69%), demonstrating a remarkable performance that is also presented by its precision and recall. It suggests that for emotions elicited by dynamic music video stimuli, high-frequency neural activity associated with complex cognitive processing and sensory integration in the gamma provides the most discriminative features. The alpha and beta rhythms also yield impressive results, consistently surpassing 85% in accuracy and F1-score for valence classification. Conversely, and in line with findings from the other datasets, the delta rhythm performs the poorest, reinforcing its limited utility for emotion recognition under such experimental conditions.
Concerning the SEED dataset, where emotional stimuli are labeled as positive, negative, or neutral, the alpha and theta rhythms are the most effective. From Tables 2 and 3, the alpha rhythm achieves an average accuracy of 81.89% and an F1-score of 90.85%. Then, although the theta rhythm shows a slightly lower accuracy of 80.72%, its F1-score is the highest among all brain rhythms, reaching 92.03%. These results indicate that for emotional states elicited by film clips, neural oscillatory patterns associated with relaxation and internal attention in the alpha rhythm, 37 as well as emotion processing in the theta rhythm, 38 become distinguishing features for recognizing emotions. The performances of the beta rhythm (accuracy: 78.01%, F1-score: 88.11%) and the gamma rhythm (accuracy: 74.76%, F1-score: 84.21%), while satisfying, are inferior to those of the alpha and theta rhythms in this context.
Since the DREAMER dataset also utilizes film clips as stimuli, its results closely resemble those observed with SEED. The alpha and theta rhythms perform well in the binary classification tasks for arousal, valence, and dominance. For example, the alpha rhythm achieves an accuracy of 82.70% and an F1-score of 81.52% for valence. The theta rhythm also performs well in terms of the arousal and dominance dimensions, with approximately 80% accuracy. Then, the beta rhythm exhibits impressive performance in the arousal and valence dimensions, also providing around 80% accuracy, as it is typically associated with active cognition and alertness. 39 Lastly, consistent with previous findings, the delta rhythm performs poorly across all metrics, as it is more effective in infants under music-directed rather than audio-visual stimuli in adults. 40 Regarding the four-class task on the DREAMER dataset, the alpha rhythm (accuracy: 75.76%) and theta rhythm (accuracy: 75.23%) achieve relatively good performances, with the beta rhythm following closely behind. The gamma rhythm is further diminished in this scenario. The delta rhythm performs worse, with an accuracy of 16.96% and an F1-score of 6.50%, which are drastically below the chance level (i.e. 25% for a four-class task), indicating that delta-based features possess almost no discriminative information and may have even introduced noise into the data.
To further illustrate the classification performance, Figure 7 presents the confusion matrices from 10-fold cross-validation on the DREAMER dataset. Specifically, Figure 7(a) displays the confusion matrix for the binary classification of arousal (high-arousal or low-arousal, denoted as HA or LA), and Figure 7(b) illustrates the results of the four-class classification of arousal valence. For the binary arousal task, the proposed model demonstrates a high true positive rate for both HA and LA states, with only a small number of samples being misclassified. It indicates that the proposed model can effectively discriminate between different arousal levels. In the four-class arousal–valence scenario, most samples are correctly assigned to their corresponding categories. However, some degree of confusion is observed between adjacent classes (e.g. HA/LA vs. LA/LV), indicating that while the model retains good discriminative capability in the multi-class setting, emotional states with similar physiological patterns are inherently more challenging to distinguish.

Confusion matrices from 10-fold cross-validation on the DREAMER dataset: (a) binary arousal classification and (b) arousal–valence four-class classification.
Discussion
Emotional activities analysis
Synthesizing the experimental results from three public datasets enables a comprehensive analysis of neural responses and brain activity across diverse emotional states. Meanwhile, it allows an exploration of potentially generalizable neurophysiological signatures associated with EEG emotion recognition. Therefore, a rhythmic analysis of emotional activities is conducted.
A key observation from the results of the proposed model is that the modality of the emotion-eliciting stimulus influences the dominant brain rhythm activity associated with emotional states. When emotions are elicited by audio-visual stimuli, alpha and theta rhythms are most informative for classification. Film clips, representing a more complex and ecologically rich form of stimulation, typically engage multi-faceted cognitive-affective processes, including scene comprehension, narrative immersion, empathic resonance with storylines, and associated memory retrieval and affective experiences. 41 The prominence of alpha rhythm activity is commonly associated with attentional engagement (e.g. focusing on-screen content) and states of cognitive immersion or relaxation (e.g. absorption in the storyline). Similarly, theta rhythm activity, which plays a significant role in episodic memory encoding, the integration of emotional information, and introspection, also exhibits heightened importance. Consequently, alpha and theta rhythms emerge as vital neural biomarkers for differentiating emotional states under these stimulus conditions. Such characteristics can also be interpreted as a group-level brain activity driven by the stimulus modality.
While providing valuable information, gamma rhythm activity is not consistently the predominant or most effective one for classification across all emotional dimensions on these film-based datasets. Nonetheless, it is generally associated with advanced cognitive and emotional processing. 42 This activity is consistent at the group level, indicating that gamma rhythm acts as a neural substrate for decoding certain elicited emotional states, particularly those involving high-level cognitive engagement.
Beta rhythm activity provides useful discriminative information across both stimulus sources, often ranking second to the best for those scenarios. Considering that beta rhythm is typically linked to cognitive engagement, including concentration, alertness, and mental effort (often considered part of motor preparation but also general cognitive readiness), this finding implies that beta rhythm may reflect a neurocognitive component related to active engagement or cognitive load during emotional experiences, irrespective of stimulus modality in emotional processing.
In contrast, the delta rhythm offers limited utility across various datasets. It is consistent with the established neurophysiological role of delta rhythm, primarily associated with deep sleep and certain atypical brain functional states. Therefore, delta rhythm may not contain discriminative information for the emotional states explored in this study. In the four-class task on the DREAMER dataset, the classification accuracy based on delta rhythm is far below the chance level, suggesting that including delta-based features may decrease performance in certain specific emotional scenarios. Therefore, further investigation to determine its underlying causes is meaningful.
Another commonality observed is the impact of classification task complexity on EEG emotion recognition, where the overall accuracy decreases across all datasets as the complexity of the classification task increases. This trend highlights a well-established principle in affective neuroscience: the finer the granularity of emotional state differentiation and the more dimensions combined, the more challenging it becomes to reliably recognize these states from EEG signals. Nevertheless, the classification derived from the dominant brain rhythms (e.g. alpha and theta rhythms in the SEED and DREAMER datasets and the gamma rhythm in DEAP for music stimuli) exhibits a relatively robust performance with increasing task difficulty. Such results reveal that the specific rhythmic activities are valuable neural correlates representing core emotional characteristics, which can be achieved using the proposed brain-inspired neuronal competition model.
Comparative study
This study aims to propose a biologically plausible and interpretable EEG emotion recognition model by simulating neuronal competition. To comprehensively evaluate its advantages and limitations, a comparative study is conducted with recent works, as listed in Table 6, with the best results of each case highlighted in bold and underlined. These compared methods contain techniques such as spiking neural networks (SNNs), transfer learning, domain adaptation, graph neural networks, attention mechanisms, and multi-task learning.
Comparative study with recent works based on classification accuracy (%).

SSTNAS: spiking spatio-temporal neural architecture search; SNN: spiking neural network; STGATE: spatial-temporal graph attention network with a transformer encoder; DACB: multi-channel automatic classification model; LGEP: local and global feature extraction for emotion prediction; FBSTCN: filter-bank spatio-temporal convolutional network; MTLFN: multi-task learning fuse network.
First, regarding studies employing SNNs, the proposed model operates similarly on SNN-like principles but focuses on simulating the biological mechanism of neuronal competition within cortical information processing. spiking spatio-temporal neural architecture search 43 automatically explores optimal SNN architectures for EEG emotion recognition using a genetic training-free neural architecture search algorithm. It incorporates spiking convolutional neural networks and spiking long short-term memory for feature extraction. However, its overemphasis on architectural search leads to limited biological interpretability. On DEAP, it achieves only 63.52% (arousal) and 60.94% (valence), which is approximately 28–32% lower than the proposed model's results (91.63% and 92.70%). On DREAMER, its arousal (76.33%) and valence (64.01%) are also lower than those of the proposed model (80.57% and 82.70%). The fractal SNN 44 captures multi-scale spatio-temporal spectral information from EEG signals through a novel fractal rule and an inverted drop-off. While fractal SNN aims for automation and broad feature exploitation, the proposed model extracts specific brain rhythms through DWT as dynamic feature inputs, processing their dynamic characteristics through a weight encoding module and pulse accumulation mechanisms. Thus, the fractal SNN aims for automation and breadth, whereas this study seeks to delve deep into specific biological principles for profundity and interpretability.
Second, compared to graph neural network-based approaches, the proposed model offers a distinct biological simulation. Spatial-temporal graph attention network (STGATE) 45 uses a transformer learning block to capture time–frequency features and a spatial-temporal graph attention mechanism that adaptively learns intrinsic connections between EEG channels using a dynamic adjacency matrix, achieving 90.37% accuracy on the SEED dataset and 77.44% for valence and 75.26% for arousal on the DREAMER. Moreover, a local and global feature extraction for emotion prediction (LGEP) 47 model focuses on mitigating heterogeneity in temporal scales among cortical regions. It utilizes modified deep-wise separable convolution for multi-scale temporal features, coordinate attention for local spatial feature optimization, and a retention transformer combined with a graph convolutional network for global temporal and spatial features, respectively. Its performances reported 80.79% accuracy for arousal and 82.78% for valence on the DREAMER. While STGATE and LGEP excel at modeling channel correlations and temporal dynamics through complex graph structures and attention mechanisms, they remain largely black-box models. Conversely, this study directly models competitive interactions among neurons, providing a biologically plausible explanation for how brain rhythms contribute to decision-making, which is a level of interpretability not typically found in these data-driven graph models.
Third, those models incorporating advanced attention mechanisms or feature fusion strategies usually offer high performance but differ in their underlying principles. For instance, a DACB model 46 is a multi-channel automatic classification model that integrates dual attention mechanisms, CNNs, and bidirectional LSTM networks. It extracts features in both temporal and spatial dimensions, utilizing a squeeze-and-excitation attention mechanism for channel importance and a dot product attention mechanism for spatio-temporal feature importance. On the DREAMER dataset, accuracies of 85.40%, 84.26%, and 85.02% were reported for arousal, valence, and dominance, respectively (all results from 10-fold cross-validation). Then, FBSTCNet 48 is a spatio-temporal convolutional network that integrates power and connectivity features through a filter bank and spatio-temporal filters, employing a cropped decoding strategy. It achieved 74.35% accuracy for three-class emotion decoding on the SEED dataset. Li et al. 49 presented a multi-task learning fuse network (MTLFN), which focuses on deep latent feature fusion of EEG signals and multi-task learning. It employs a variational autoencoder for unsupervised spatio-temporal latent features and a graph convolutional network-gated recurrent unit network for supervised spatio-spectral features, fusing these to form a comprehensive representation. MTLFN trains on three related tasks using focal loss and triplet-center loss to address class imbalance and enhance feature separability. While it reached 83.33% accuracy for arousal and 80.43% for valence on the DREAMER, its performance on DEAP is substantially lower: 73.28% for arousal and 71.33% for valence, 18.35–21.37% below the proposed model's DEAP results (91.63% and 92.70%). This discrepancy highlights MTLFN's difficulty in capturing the nuanced neural patterns underlying DEAP's audio-visual emotional stimuli, as its latent feature fusion prioritizes statistical patterns over biologically meaningful brain rhythms. While such models demonstrate high accuracy through complex architectures, they often operate as black-boxes, lacking explicit links to the underlying neurophysiological processes. By simulating the dynamics of neuronal competition and decision-making through brain rhythms, the proposed model presents a biologically plausible and inherently interpretable framework, providing a clearer understanding of how different brain rhythms contribute to EEG emotion recognition.
In short, while acknowledging the impressive performance of other complex models in the field, the proposed model demonstrates distinct advantages by addressing the critical trade-offs between performance, interpretability, and computational efficiency. It achieves high interpretability by directly simulating the neuronal competition for emotion recognition, that is, a brain-inspired design rooted in the neuronal competition that introduces a novel biological perspective to emotion recognition, and computational efficiency through brain rhythms extracted from EEG signals as dynamic feature inputs, maximizing the preservation of their inherent temporal information and reducing overhead from handcrafted feature extraction. Meanwhile, it offers a dynamic attentional focus, in which the weight-encoding module guides attention to brain rhythms that are more significant for emotional discrimination, facilitating a more granular analysis of neural dynamics across multiple timescales.
Conclusion
This study establishes a neurobiologically grounded model for EEG emotion recognition by computationally formalizing the dynamics of neuronal competition. By integrating brain rhythm extraction through DWT with brain-inspired modules, the proposed method achieves satisfying emotion recognition results while preserving interpretable rhythmic characteristics. Specifically, it improves conventional feature extraction and black-box deep learning by simulating how excitatory–inhibitory interactions facilitate emotional categorization through winner-take-all dynamics. Experimental validation across the DEAP, SEED, and DREAMER datasets demonstrates the neurophysiological plausibility of this approach, in which alpha and theta rhythms emerge as dominant biomarkers for audio-visual emotional stimuli, reflecting their roles in attentional engagement and memory–emotion integration. In contrast, delta rhythm is found to be neurologically incongruent with conscious affective processing.
Regarding the experimental results, on the SEED dataset, the proposed model performs an average emotion classification accuracy of 81.89%. On the DREAMER dataset, it acquires accuracies of 80.57% for arousal, 82.70% for valence, and 80.38% for dominance. Notably, the evaluation on the DEAP dataset achieves remarkable performance, with accuracies of 91.63% for arousal, 92.70% for valence, and 86.64% for dominance, demonstrating improvements over these results. For example, 14.52% higher than LGEP 47 and 15.29% higher than MTLFN. 49 It also aligns with neurophysiological relationships, linking alpha rhythms (dominant in low-arousal states) and theta rhythms (critical for valence judgment) to classification performance. Collectively, these quantified results show that the proposed method performs impressively compared to other data-driven models across three benchmark databases, effectively validating the generalizability of this study.
The innovation of the proposed method lies not in outperforming complex data-driven models but in providing a way to explore the neurocomputational basis of emotion. Its modular design offers inherent adaptability, and future work will explore domain-generalization techniques to address cross-subject variability and implement them on neuromorphic hardware for real-time deployment. It is also noted that, despite the promising results, this study has certain limitations, including the inherent uncertainties in pre-processed EEG recordings and the simplified nature of the approach compared to actual biological processes. Consequently, integrating high-level neuroscience encoding into classification models will yield profound insights and reliable applications in EEG emotion recognition for digital health.
Footnotes
Acknowledgments
The authors would like to appreciate the special contributions from Anhui Provincial Key Laboratory of Multimodal Cognitive Computation, ZUMRI-LYG Joint Laboratory, and Guangdong Provincial Key Laboratory of Intellectual Property and Big Data.
Author contributions
JL, GF, WL, XH, and SZ designed the research. JL, GF, and RC performed the research. LW, KL, and XZ analyzed the data. JY, XZ, and RC interpreted the results. JL, GF, and XH revised the paper. KL, JY, SZ, and XZ conceptualized the method. JL, XH, and RC administrated the project and made some contributions to the figures. JL, GF, WL, XH, SZ, and RC investigated the dataset and supported funding acquisition. LW, KL, JY, and XZ provided computing resource and supervised the research. JL, GF, WL, and XH wrote the manuscript. All authors contributed to editorial changes in the manuscript. All authors read and approved the final manuscript. All authors have participated sufficiently in the work and agreed to be accountable for all aspects of the work.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported in part by the Guangzhou Science and Technology Plan Project under Grants 2024B03J1361 and 2023B03J1327, in part by the Open Research Fund of Anhui Provincial Key Laboratory of Multimodal Cognitive Computation under Grant MMC202305, in part by the Sichuan Science and Technology Program under Grant 2025ZNSFSC0780, in part by the Foundation of the 2023 Higher Education Science Research Plan of the China Association of Higher Education under Grant 23XXK0402, in part by the Sichuan Applied Psychology Research Center of Chengdu Medical College under Grant CSXL-25102, in part by the Neijiang Philosophy and Social Science Planning Project under Grant NJ2025ZD007, in part by the Guangdong Province Ordinary Colleges and Universities Young Innovative Talents Project under Grant 2023KQNCX036, in part by the Scientific Research Capacity Improvement Project of the Doctoral Program Construction Unit of Guangdong Polytechnic Normal University under Grant 22GPNUZDJS17, in part by the Graduate Education Demonstration Base Project of Guangdong Polytechnic Normal University under Grant 2023YJSY04002, in part by the Open Research Fund of State Key Laboratory of Digital Medical Engineering under Grant 2025-M10, in part by the Open Project Program of State Key Laboratory of Computer-Aided Design and Computer Graphics (CAD&CG) under Grant A2533, and in part by the Research Fund of Guangdong Polytechnic Normal University under Grants 2023SDKYA004 and 2022SDKYA015.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of data and materials
The EEG data analyzed during the current study are from three public datasets: DEAP (https://eecs.qmul.ac.uk/mmv/datasets/deap), SEED (https://bcmi.sjtu.edu.cn/#seed/index.html), and DREAMER (![]() ). Other code will be made available from the corresponding authors on reasonable request.
). Other code will be made available from the corresponding authors on reasonable request.