Abstract
Aims
New-onset atrial fibrillation (NOAF) occurs in approximately 23% of patients with sepsis and is independently associated with increased mortality. Therefore, early prediction of NOAF has significant clinical value. However, current artificial intelligence (AI) models predominantly rely on tabular data. These unimodal AI models face limitations in predicting NOAF as they fail to fully utilize the predictive potential arising from the interplay of multimodal data.
Methods
We reviewed current Machine Learning (ML) and Deep Learning (DL) approaches for atrial fibrillation (AF) prediction. It summarizes the selected features in ML models for predicting AF in ICU patients, and the advantages of time-window selection in DL models using electrocardiogram (ECG) signals. Notably, we compared these models in terms of feature selection, prediction horizons, and performance when applied to tabular data and ECG signal features. To enhance the predictive capability of ML for NOAF in patients with sepsis, we drew inspiration from multimodal models developed for other diseases, such as Alzheimer's disease, and proposed integrating tabular data and ECG signal data within a multimodal framework.
Results
This study systematically analyzed the application of ML and DL in AF prediction. After screening, 12 studies (6 ML, 6 DL) were included. ML models, based on electronic medical records (EMR) or ECG features, achieved prediction windows ranging from minutes to hours with AUCs of 0.74–0.90. DL models processing raw ECG signals extended prediction windows to days, achieving AUCs of 0.74–0.96, with performance improving with larger datasets. A Transformer-based multimodal model (integrating clinical data and ECG) was proposed to enhance AF prediction in sepsis patients, though further validation is needed for cross-modal data fusion feasibility.
Conclusions
Transitioning from unimodal predictive models to multimodal frameworks that combine tabular clinical data and raw ECG signals is feasible within the current deep-learning framework. This approach has the potential to significantly improve the early prediction capabilities of NOAF in sepsis patients.
Introduction
Atrial fibrillation (AF) is the most prevalent cardiac arrhythmia with a lifetime risk ranging from 22% to 26%, and its prevalence increases with age. 1 It is a major risk factor for heart failure, ischemic stroke, and increased mortality, imposing a heavy burden on the healthcare system. 2 For intensive care units (ICU) patients, new-onset atrial fibrillation (NOAF) serves as both an indicator of disease severity and a potential contributor to adverse prognosis, occurring in approximately one in every six patients.
Sepsis is a severe and potentially fatal systemic response to infection that threatens millions of lives worldwide each year. As a common complication of critical illness, AF is the most predominant arrhythmia observed in patients with sepsis. Studies have indicated that the incidence of NOAF in patients ranges from 20% to 30%%.2,3 If not promptly detected and treated, it can lead to severe health consequences including progressive organ dysfunction and even death. Moreover, its occurrence in septic patients is not merely a marker of severity but is independently associated with a significantly increased risk of in-hospital stroke and mortality. 4 Consequently, the prediction and early detection of NOAF are imperative in patients with sepsis. Several models have been established to predict and detect NOAF in ICU patients, and the AF score is currently recognized as the gold standard. However, its clinical application is limited owing to its low accuracy.
ML involves algorithms that utilize statistical and optimization techniques to learn from historical data. The primary objective of ML is to identify or predict significant outcomes from large-scale datasets. When applied to medical datasets, ML often performs better than the traditional clinical scoring systems.5,6 In recent years, ML models have been increasingly employed to predict NOAF, emphasizing the crucial necessity of effective feature engineering and model validation to improve the predictive accuracy and clinical applicability.7,8 These models can be divided into two categories: the early warning model to predict the occurrence of AF during sepsis and the real-time detection model. Researchers in the former often rely on tabular data, such as demographics, physical signs, and test results, to train models for predicting the risk of AF during sepsis. Raw ECG signals that provide critical information are often overlooked. In contrast, the latter studies are mostly based on bedside ECG data, which mainly focus on the automatic diagnosis of AF when it occurs.9,10 Despite their high accuracy, these prediction models may not provide clinicians with sufficient information to prevent the onset of AF, due to their short prediction horizons—often limited to only a few minutes before onset.11,12 Additionally, the integration of ECG data has been demonstrated to significantly enhance predictive capabilities, allowing for a more comprehensive assessment of patients and improved early intervention strategies.
This article presents a systematic review (SR) by searching the PubMed database to identify studies focused on ML models for predicting NOAF, with a particular emphasis on multimodal models. We compared these models in terms of feature selection using tabular data and ECG signals, clinical applications, and predictive performance. Additionally, we referenced the application of multimodal models to other diseases (such as Alzheimer's disease) to evaluate their potential value for NOAF prediction.
Materials and methods
Protocol and registration
This SR was conducted based on the recommendations of the Preferred Reporting and Items for Systematic Reviews and Meta-Analyses (PRISMA) and was recorded in the PROSPERO registry for SR ((ID number: CRD420250654679). The full protocol is available at: https://www.crd.york.ac.uk/prospero/.
Search strategy
We conducted a systematic search of PubMed, Embase, Medline and Scopus databases covering publications available until Feb 16, 2025. The search strategy employed the following terms: (atrial fibrillation OR new-onset atrial fibrillation) AND (artificial intelligence OR machine learning OR Deep Learning) AND (intensive care unit OR ICU OR Sepsis).
The identification and removal of duplicate records were conducted using Zetero prior to the screening process. Data extraction was subsequently performed using Excel software. Two independent reviewers (W.X. and B.J.) carried out the initial title and abstract screening. The full-text review and subsequent screening were conducted by B.J. and Y.F.L., with all exclusion reasons being systematically documented during the screening process. During title/abstract screening and full-text review, any disagreements between the three independent reviewers were resolved through consensus discussions.
Inclusion and exclusion criteria
Studies were eligible for inclusion if they met the following criteria:
Study objective: Focused on predicting clinical outcomes of AF, including both pre-existing and new-onset cases. Study Population: Included adult patients (age > 18 years) admitted to ICU or diagnosed with sepsis. Methodology: Utilized ML or DL techniques for predictive modeling. Model Description: Provided detailed descriptions of the ML/DL models and predictor variables used for risk prediction. Study Design: Included randomized controlled trials (RCTs), observational studies, cohort studies, case-control studies, and review studies. Language: Published as full-text articles in English.
Exclusion criteria encompassed case reports, comments, letters to the editor, editorials, study protocols, and replies. Studies involving pediatric patients and those not published in English were also excluded. The studies were required to provide a description of the ML or DL models and predictor variables used in prediction and/or detection of NOAF in sepsis patients. Data collected from the studies included author information, year of publication, variables used in the model, population studied, prediction model type, and model performance metrics including the area under the receiver operating characteristic curve (AUC), accuracy, sensitivity, and specificity positive predictive values, negative predictive values, area under the precision–recall curve, precision and recall, F1 score of the ML or DL algorithms in predicting and detecting the occurrence of NOAF.
Risk of bias assessment in studies
The risk of bias assessment was systematically conducted by evaluating five critical parameters: an sufficiently sized cohort, appropriate cross-validation methods, an external validation dataset, blinding of participants and personnel, and the handling of incomplete outcome data. For data extraction, a predefined form was employed to capture essential information including study design, sample size, patient demographics, intervention or exposure specifics, feature selection, outcome measures, model performance, comparator details, follow-up duration, funding sources, and potential conflicts of interest.
Results
Study selection and description
A comprehensive literature search was conducted in November 2024, yielding 2497 records related to various aspects of AF, including risk factors, prediction, treatment, prophylaxis, monitoring, detection, and outcomes. Among these, 2302 articles were identified as potentially relevant for examining the application of ML or DL in AF prediction and detection. Two independent expert reviewers performed the screening process, ultimately selecting 66 articles from the initial pool. A secondary screening was then conducted to exclude studies that lacked detailed descriptions of their ML/DL models. Any disparities between the reviewers were resolved through discussion and consensus. Ultimately 12 studies, each aligning with at least one of the designated outcomes, were determined to be eligible for inclusion in this SR. These studies were subsequently categorized into ML-based AF prediction models (n = 6) and DL-based AF prediction models (n = 6). The study selection process is detailed in the PRISMA flowchart presented in Figure 1.

PRISMA diagram of the literature search and exclusion process flowchart. AI: artificial intelligence; ML: machine learning; NOAF: new-onset atrial fibrillation; SR: systematic review; ICU: intensive care unit; DL: deep learning.
The ML models reviewed in this study were trained using datasets derived from several large-scale, publicly or institutionally available sources. These included the Intensive Care (MIMIC-II, 13 III, 14 IV 15 ) databases, the Kensington General Hospital (KGH) database, 16 AmsterdamUMCdb, 11 the AF Prediction Challenge database, 17 the Ghent University Hospital database, 11 and the Belgian hospital ICU database. 11 The diversity of these databases guarantees a broad range of patient characteristics and clinical scenarios, thus facilitating a robust model evaluation. A brief description of databases used in this study presented in Supplementary material.
The current ML-based AF prediction models
The types of features utilized in different supervised ML models and their performances for AF prediction are summarized in Table 1.
Comparison of current ML-based AF prediction models using tabular data.
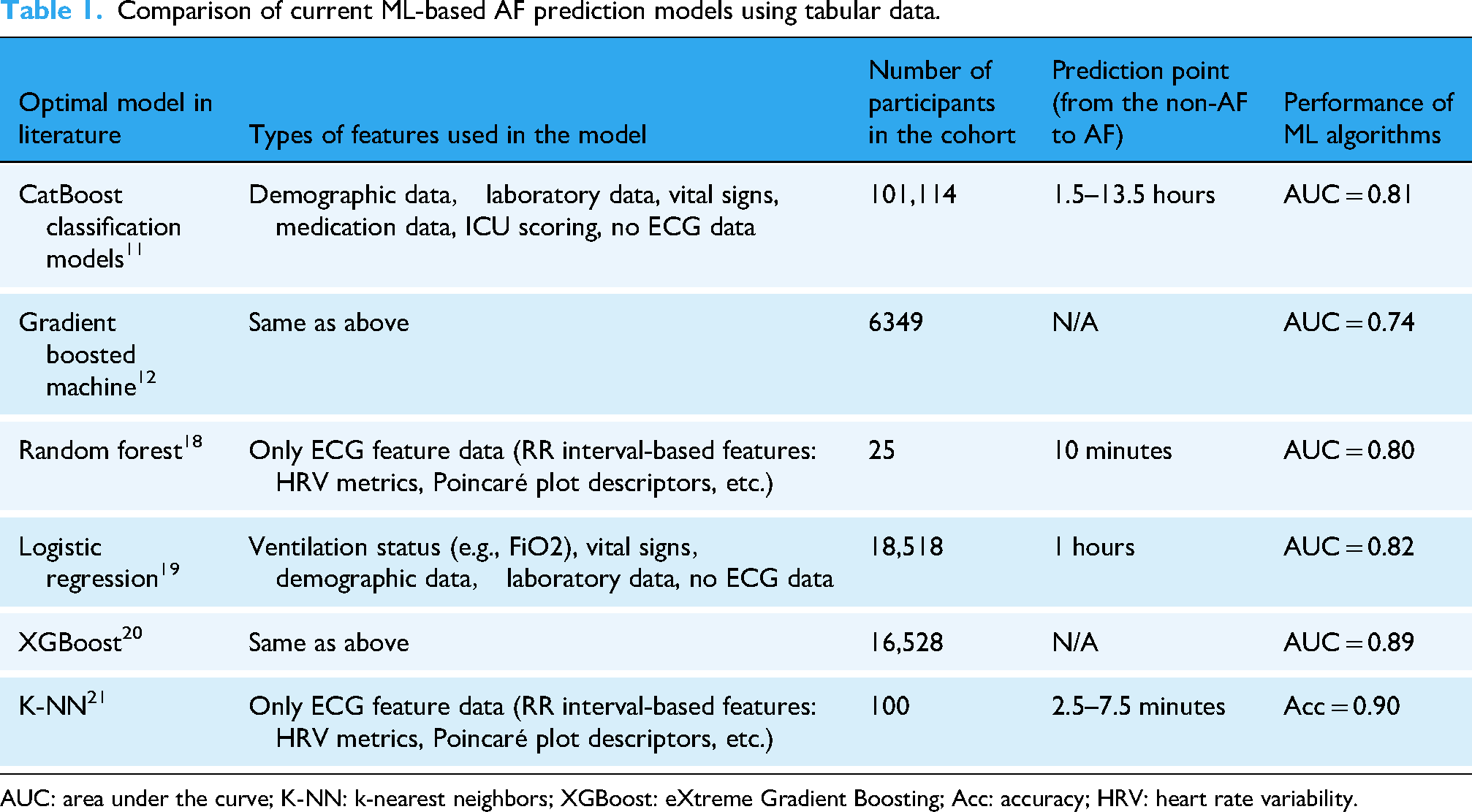
AUC: area under the curve; K-NN: k-nearest neighbors; XGBoost: eXtreme Gradient Boosting; Acc: accuracy; HRV: heart rate variability.
Among the models, CatBoost 11 utilized the most comprehensive feature set, excluding ECG data. Tested on a cohort of 101,114 participants, it achieved an AUC of 0.81 with a prediction window of 1.5–13.5 hours. Similarly, the Gradient Boosted Machine 12 used the same feature set, but was tested on a smaller cohort of 6349 participants, resulting in a lower AUC of 0.74. In contrast, Random Forest 18 focused exclusively on ECG features, tested on only 25 participants, and achieved an AUC of 0.80 with a short prediction window of 10 minutes. Logistic Regression 19 employed a diverse feature set, including ventilation parameters (e.g., FiO2), vital signs, demographic data, and laboratory results, while excluding ECG data. It achieved an AUC of 0.82 in a cohort of 18,518 participants with a 1-hour prediction window. XGBoost 20 used a similar feature set, achieving the highest AUC of 0.89 across 16,528 participants, demonstrating its ability to handle complex, multivariate data. Finally, K-Nearest Neighbors (K-NN) 21 relied solely on ECG features and achieved an accuracy of 0.90 in a cohort of 100 participants with a prediction window of 2.5–7.5 minutes. However, its reliance on accuracy as a metric may not adequately reflect the model performance in imbalanced datasets.
In this study, tabular data are defined as a structured representation of features, typically organized in a table format, where rows correspond to individual samples (participants) and columns represent feature variables. These features are classified as either numerical (e.g., laboratory results and vital signs) or categorical (e.g., demographics and medication usage). This format is versatile, allowing for the integration of various data types, such as demographic information, laboratory results, vital signs, medication records, and ICU scoring systems (e.g., APACHE II and SOFA). However, although tabular data provide a standardized approach for feature utilization, they often lack the temporal resolution needed to fully capture sequential patterns, such as the dynamic variations observed in ECG signals.
Table 1 indicates that the six existing NOAF prediction models can be broadly classified into two categories based on the type of feature data utilized. The first category includes demographic information, laboratory results, vital signs, medication data, ventilation status, and ICU scoring,11,12,19,20 which are typically derived from electronic medical record (EMR) systems and encompass comprehensive patient health information. Moreover, tabular data can be conveniently collected and organized from EMR, facilitating effective data analysis and model development. Other researchers’ AF testing models showed that the AUC of AF detection was 0.79 during a 6-month follow-up. This performance is comparable to that of several recent non-AI-based clinical AF risk scores, which exhibit an AUC range from 0.71 to 0.79.22–24 In terms of time prediction windows, these models generally predict the timing of NOAF occurrence at an hourly resolution. Thus, AI-EHR-based predictive tools may eventually play a role in clinical practice for predicting the occurrence of NOAF. 25
In comparison, the second category focuses exclusively on ECG data, encompassing various HRV-related features, as shown in Figure 2. These features include the standard deviation of the heart rate data (SDNN), the total number of consecutive heart rate differences exceeding 50 ms (NN50), ratio of NN50 to the total number of RR intervals (pNN50), skewness and kurtosis of the heart rate data, root mean square of successive differences (RMSSD), Poincaré plot features, sample entropy (SE), multiscale entropy (ME), approximate entropy (AE), very-low-frequency power (VLF), low-frequency power (LF), high-frequency power (HF), and their respective power spectral densities (PSDs). These features are extracted using methods such as the tunable Q-factor wavelet transform (TQWT), variable frequency complex demodulation (VFCDM), and time-domain and nonlinear feature extraction techniques. Although ECG data contain features directly related to NOAF and may implicitly include signal patterns that precede the onset of AF, the second type of model converts the original ECG signal into tabular data through feature extraction during the training process, which commonly referred to as shallow models “shallow models.” ML-processed ECG features are insufficient to capture subtle patterns preceding AF onset and fail to incorporate the molecular mechanisms leading to AF. Consequently, their prediction horizon remains limited to a minute-level timeframe.
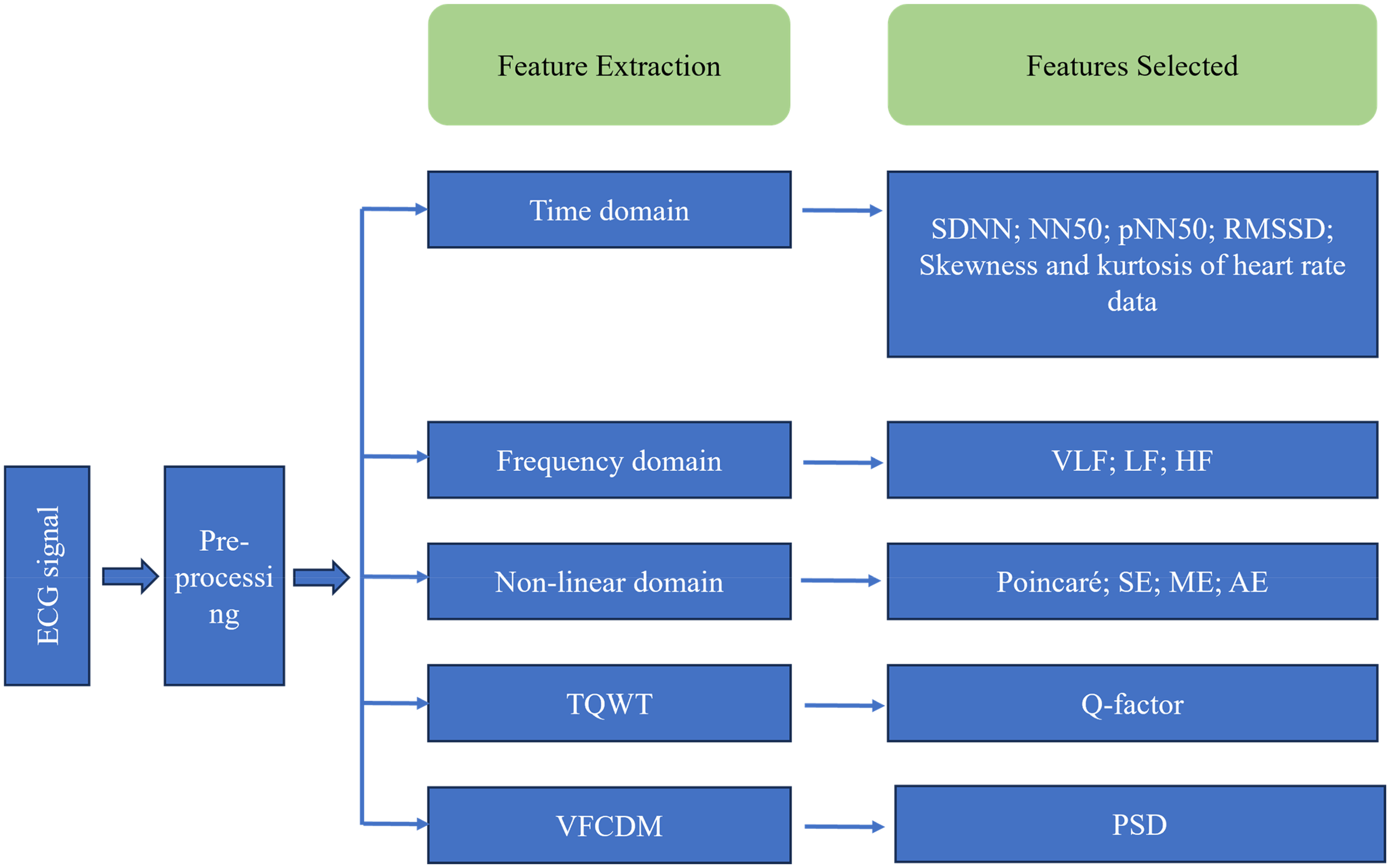
Feature extraction of HRV signals using TQWT, VFCDM, and time domain and nonlinear methods. SE: sample entropy; AE: approximate entropy; ME: multiscale entropy; LF: low-frequency power; HF: high-frequency power; PSDs: power spectral densities; TQWT: the tunable Q-factor wavelet transforms; VFCDM: variable frequency complex demodulation; SDNN: the standard deviation of the heart rate data; NN50: the total number of consecutive heart rate differences exceeding 50 ms; pNN50:ratio of NN50 to the total number of RR intervals; RMSSD: skewness and kurtosis of the heart rate data, root mean square of successive differences.
Indeed, the arrangement and dispersion of myocardial fibers play a crucial role in shaping the heart's anisotropic conductive properties, which in turn affect the occurrence and maintenance of AF. By employing DL algorithms to directly extract features from 12-lead (or multi-lead) ECG signals, these features may contain signal patterns related to parameters such as QRS duration, QT interval, and Tpeak-Tend interval, rather than directly calculating these parameters. This method of data analysis capitalizes on the advantage of DL in automatically performing feature engineering.
In summary, both types of models have limited prediction resolution. Therefore, we introduce a DL model in the next section that can capture the original ECG signal features from several days prior as input, thus extending the time window from an hourly resolution to a weekly scale.
The current DL-based AF prediction models
In recent years, DL has achieved remarkable progress in the medical field, with its applications in disease diagnosis, treatment prediction, and personalized medicine expanding steadily. 26 In the area of medical imaging analysis, breakthroughs in DL algorithms, particularly convolutional neural networks (CNNs), have enabled models to achieve performance in disease detection (e.g., tumor segmentation and lung nodule detection) that rivals or even surpasses that of human experts.27,28 In the field of natural language processing (NLP), DL-based models are widely used to process electronic health records, medical literature, and physician notes, thereby facilitating automated information extraction, diagnostic support, and decision-making. 29 Additionally, recurrent neural networks (RNNs) and their advanced variants (e.g., LSTM and Transformer) have demonstrated exceptional capabilities in analyzing physiological signals such as ECG and electroencephalograms (EEG), effectively identifying complex temporal patterns and supporting early disease warning systems. 30 For example, Thivya Anbalagan's study proposed a DNN-based approach that converts 1D ECG signals into 2D patterns via Chirplet/Stockwell transforms for identifying AF in the presence of noise and other beats. Their ensemble model (ShuffleNet + AlexNet) achieved 93.7% accuracy on the CinC 2017 dataset by leveraging spectral-temporal features, demonstrating the potential of hybrid signal processing and lightweight architectures. 31 The developed model can identify the severity of AF conditions effectively and helps reducing mortality over time due to arrhythmia.
It is worth noting that ECG feature data fundamentally belong to the category of tabular data. Using these features, traditional ML models can effectively predict NOAF. However, these shallow models cannot directly process raw ECG signals containing a wealth of complex patterns. By utilizing DL methods, it is possible to analyze these raw signals and predict AF occurrences several days in advance. Numerous researchers have proposed neural network architectures that are specifically designed to process raw ECG signal data. A summary of these methodologies is presented in Table 2.
Comparison of current DL-based AF prediction models for long term.

Based on the data presented in the table, we analyzed the performance of the DL models in the analysis of ECG signals. Firstly, we observed the performance of models using ResNet and 12-lead ECG in different studies. H. Zhu et al.'s study 32 utilized 49,300 records with a predicted time frame of 30 days, achieving a high AUC value of 0.96. In contrast, P. Melzi et al.'s study, 36 which also employed ResNet with 12-lead ECG, had a reduced number of records at 26,657, with an extended prediction time of 6 months, resulting in a lower AUC value of 0.79. This suggests that an increase in the number of ECG raw signal samples correlates with an enhancement in the predictive performance of the model, as measured by the AUC value.
Further analysis reveals that N. Yuan et al.'s study, 34 with the highest number of records (907, 858), had a prediction time of 31 days and achieved an AUC value of 0.86. This is an improvement over T. Habineza et al.'s study, 37 which had 7566 records, a prediction time of 40 weeks, and an AUC value of 0.85, despite the latter's longer prediction horizon. Additionally, M. Gadaleta et al.'s study, 35 with a relatively smaller dataset of 459,889 records, had a prediction time of only 14 days and still achieved an AUC value of 0.80. These data points support the hypothesis that an increase in the number of ECG raw signal samples is associated with higher AUC values, whereas an extended prediction time does not necessarily ensure an increase in AUC values. This may be related to the complexity of the model, quality and quantity of the training data, and the generalization capabilities of the model. The longer the prediction time, the lower the AUC value, which may indicate that the predictive accuracy of the model decreases as the prediction time increases.
In summary, DL models demonstrate greater predictive accuracy when dealing with a larger number of ECG raw signal samples. However, the relationship between the length of the prediction time and the AUC value is not linear, indicating that when designing and evaluating DL models, it is necessary to consider a comprehensive range of factors, including the amount of data, the prediction time frame, and the complexity of the model's architecture.
Multimodal data integration for predicting NOAF in sepsis patients
The aforementioned ML or DL models were not specifically developed for predicting NOAF in sepsis patients, and these models were all trained using unimodal data. The integration of multimodal data plays a crucial role in the prediction of AF. By combining different types of data sources, such as ECG features, with clinical laboratory data, predictive models can more comprehensively capture the various factors that trigger AF. This fusion of data not only provides a more holistic perspective but also helps to enhance the accuracy and specificity of predictive models.
However, to date, no AI model has been able to effectively integrate ECG data with clinical laboratory data to predict AF. In the following chapters, we will explore in detail the potential application of clinical tabular data and electrophysiological data in AF prediction models for sepsis patients as well as the feasibility of DL model architectures in this field.
In the healthcare field, a review of multimodal ML approaches indicated that traditional ML methods in healthcare have primarily focused on single-modality data, which restricts their ability to integrate diverse information sources in clinical practice to improve decision-making. 38 In the field of disease diagnosis, research progress in multimodal models has been rapid. Although the integration of multimodal data faces challenges in precision oncology, it indicates a direction for future development. For example, by combining medical imaging, clinical records, and genomic data, a more comprehensive understanding of tumor characteristics can be achieved, 39 thereby enhancing predictive capabilities for risk assessment, cancer progression, and treatment response. In the assessment of Alzheimer's disease, multimodal DL models have been developed that can process a variety of clinical information, including demographic data, medical history, neuropsychological tests, neuroimaging, and functional assessments, matching the diagnostic accuracy of practicing neurologists and neuroradiologists. 40 Moreover, AI-based multimodal data differential diagnosis models have shown potential in the diagnosis of dementia etiologies as they can effectively handle and integrate heterogeneous data from different sources, such as medical imaging, clinical records, and biomarkers, to improve the accuracy and efficiency of diagnosis. 41
Inflammatory factors are significantly associated with the occurrence of AF in patients with sepsis. 42
The systemic inflammatory response triggered by sepsis leads to the release of various proinflammatory factors. These factors can trigger a range of reactions, including the activation of the clotting pathway, which may contribute to the development of AF. 43 The presence of inflammatory markers is associated with atrial remodeling, which further increases the risk of AF. 44 Systemic inflammation contributes to widespread endocardial and endothelial dysfunction, promoting atrial structural and electrical remodeling, which may lead to AF. 42 Various inflammatory biomarkers have been identified as potential predictors of AF, including C-reactive protein (CRP),45,46 interleukins (e.g., IL-6), and red blood cell distribution width (RDW). 45 International normalized ratio (INR) has been identified as an independent predictor of NOAF in sepsis patients and serves as a key variable in NOAF prediction models. 47 Existing studies suggest that these biomarkers can serve as important features in multimodal predictive models.
Moreover, integrating ECG signals is essential for accurate prediction of AF in patients with sepsis. ECG data provide real-time insights into electrical and structural changes in the atria, allowing for the detection of early atrial remodeling signs that may precede AF. By combining ECG data with inflammatory markers, prediction models can more effectively capture the interplay between electrical activity and inflammation-driven atrial changes, enhancing the model's predictive power and specificity. This multimodal approach highlights the importance of ECG feature fusion, which may offer critical early warning signs and facilitate timely intervention in managing AF risk among sepsis patients.
Thus, it is evident that integrating tabular data and ECG signals for multimodal NOAF prediction is mechanistically feasible. To enhance the capability of ML in predicting NOAF in sepsis patients, we drew inspiration from multimodal modeling approaches used in other diseases, such as Alzheimer's disease, 41 and proposed a multimodal implementation strategy for AF prediction in sepsis patients. We propose a Transformer-based multimodal model that integrates four key data modalities to predict NOAF in sepsis patients as illustrated in Figure 3.

An overview diagram of a multimodal prediction model for NOAF in sepsis patients. NOAF: new-onset atrial fibrillation; ECG: electrocardiogram signals; RP: recurrence plot; Emb: embedding; MMSE: minimum mean square error; MLP: multilayer perceptron.
Tabular clinical data: It includes demographic information, laboratory results (e.g., inflammatory markers like CRP, IL-6), vital signs, and ICU scores (e.g., SOFA, APACHE II). These features capture systemic inflammation and organ dysfunction, which are mechanistically linked to atrial remodeling and NOAF. ECG Signals: Raw 12-lead ECG data provide direct insights into electrical activity and atrial abnormalities. Temporal patterns in ECG signals (e.g., P-wave morphology, QT intervals) are critical for early AF detection. ECG-Derived Images: Recurrence plots (RPs) transform 1D ECG signals into 2D representations, highlighting nonlinear dynamics (e.g., chaos, periodicity) that may precede AF onset. Textual Data: Clinical notes or reports (e.g., radiology findings) contextualize the patient's condition, such as lung injury severity, which may indirectly influence cardiac load.
In the terms of feature embedding and cross-modal fusion: Tabular data: Categorical variables (e.g., gender) are one-hot encoded, while numerical features (e.g., INR levels) are projected into embeddings via a linear layer. ECG signals: Processed using DDxNet (a pre-trained CNN) to extract high-level embeddings. ECG images: Converted into embeddings via ResNet, capturing spatial patterns in RPs. Textual data: Processed using a clinical BERT model to generate context-aware embeddings. The transformer's self-attention mechanism dynamically weights interactions between modalities.
RP is a powerful method for analyzing the periodicity, chaotic behavior, and non-stationary properties of time series, and is commonly utilized for visualizing dynamic systems.48–51 Zhu et al. 32 successfully employed the RP technique to predict the occurrence of AF up to 30 days in advance. By converting a 1D time series into a 2D recurrence matrix through recursive relationships, the RP method enhances the model's ability to identify complex patterns and improves the recognition accuracy.
The medical community has increasingly made available various multimodal datasets that integrate medical images, textual reports, and clinical annotations to advance applications in diagnosis, image segmentation, and report generation. MedTrinity-25M 52 is a large-scale dataset released by leading institutions, containing over 25 million medical images with detailed, multi-granularity annotations to support multimodal model training and evaluation. MedPix 2.0 is an improved version of the original MedPix dataset, 53 providing both medical images and corresponding clinical reports and diagnostic descriptions, serving as a valuable resource for training large-scale multimodal medical models. M3D datasets 54 focus on 3D medical imaging and associated textual information, making them ideal for tasks such as image-text retrieval, report generation, and visual question answering.
In addition to these, the MIMIC-IV Clinical Database captures comprehensive clinical data-HER-including demographics, diagnoses, treatments, and laboratory tests. Complementing this, the PhysioNet Waveform Database55,56 contains high-resolution physiological signals such as ECG waveforms. By leveraging shared patient IDs and hospitalization details, researchers can match patients’ ECG signals with their corresponding clinical records. This integrated dataset is particularly valuable for training predictive models for conditions like AF, as well as other diseases that are associated with physiological waveform signals.
Transformers require input tokens in the form of embeddings. Therefore, medical data must be adapted into a unified embedding format that the model can process. We categorized multimodal medical data into four types: imaging data compressed into embeddings via ResNet; fixed-length ECG signals (e.g., for 24-hour-ahead prediction) converted into embeddings using DDxNet 57 ; categorical data encoded with one-hot embeddings; and numerical data projected into the transformer's input space through a single linear layer to preserve the original data structure. ResNet and DDxNet are pre-trained classification models specifically designed for ECG signals from patients without AF and those experiencing their first AF episode, respectively, with the embeddings extracted from the final layer of each network.
Discussion
This study demonstrates that machine learning (ML) and deep learning (DL) exhibit outstanding performance in cardiovascular disease risk prediction. The reviewed ML/DL models achieved AUC values of 0.74–0.96 in predicting new-onset atrial fibrillation (NOAF), comparable to the predictive perforsepsis patientsmance (AUC 0.71–0.96) for sudden cardiac death (SCD) reported by Barker's team. 58 The AI-based electrocardiogram analysis conducted by Attia et al. 59 further substantiated the pivotal value of ECG in arrhythmia prediction. By innovatively integrating clinical tabular data with ECG signals, the developed multimodal prediction framework aligns with the trend of multi-source data fusion in medical AI. This approach echoes both the multi-data integration strategy for Alzheimer's disease diagnosis proposed by Qiu et al. 40 and the multidimensional SCD prediction concept advanced by Barker et al. 58 The findings also reveal a positive correlation between DL model performance and ECG sample size. Notably, Zhu et al. 32 achieved an AUC of 0.96 using 49,300 ECG records, a finding corroborated by Goldstein et al.'s 60 analysis of extensive dialysis data (n = 22,000,000). Hannun et al. 61 further demonstrated that with over 90,000 ECG samples, DL models attained diagnostic accuracy comparable to that of cardiologists, unequivocally proving the determinant role of big data in AI model performance.
This study focuses on predicting sepsis-associated new-onset atrial fibrillation (NOAF), involving systemic inflammation (e.g., CRP, IL-6)-induced atrial electrical remodeling—a pathophysiological mechanism distinct from the myocardial electrical instability and structural abnormalities underlying SCD prediction mechanisms examined by Barker et al. 58 In terms of the research population, while Attia et al.'s 59 atrial fibrillation model targeted general cases, this study emphasizes sepsis-specific pathophysiological characteristics, particularly infection-mediated myocardial injury. Methodologically, we innovatively employed recurrence plot (RP) transformation of ECG signals to capture nonlinear dynamic features, an approach novel to NOAF prediction conceptually aligned with Mathunjwa et al.'s 49 RP application in arrhythmia classification. In contrast, Tse et al.'s 62 SCD prediction studies predominantly rely on conventional ECG parameters (e.g., QT interval), whereas our study additionally incorporates clinical text data (e.g., radiology reports). Architecturally, our Transformer-based multimodal data processing contrasts with the traditional random forest or support vector machine approaches predominantly used for SCD prediction, as reviewed by Barker et al. 58
Study limitations: While this systematic review highlights the potential of multimodal ML/DL models for predicting NOAF in sepsis patients, several limitations should be acknowledged. First, the included studies predominantly utilized retrospective data from heterogeneous sources, which may introduce biases in model generalizability. Second, the current evidence is limited by the small number of studies (n = 12) and the lack of external validation for most models, particularly those integrating multimodal data. Third, the proposed transformer-based multimodal framework, though theoretically promising, requires empirical validation in prospective clinical cohorts to assess its feasibility and performance. Additionally, the interpretability of DL models remains a challenge, potentially limiting their clinical adoption.
Future directions: While a few sepsis-specific clinical risk scores exist, 47 the application of advanced ML/DL models tailored to the unique pathophysiology of sepsis remains an under-explored area with significant potential. To enhance the reproducibility and generalizability of multimodal models, we recommend adopting the PROBAST-AI framework to standardize reporting protocols, followed by multicenter validation studies leveraging established databases such as PhysioNet (Research Resource for Complex Physiologic Signals database) and MIMIC-IV (Medical Information Mart for Intensive Care IV database). Such standardization is essential for facilitating clinical translation. Furthermore, integrating wearable technologies with temporal modeling approaches (e.g., LSTM networks) could enable continuous ECG monitoring and extended risk prediction in septic patients, potentially overcoming current limitations in temporal prediction and enabling personalized therapeutic interventions. To improve clinical utility, interpretability tools such as SHAP method 63 should be incorporated to elucidate model decision-making processes. Enhanced AI transparency is critical for fostering clinician trust and guiding treatment strategies.
Conclusion
This systematic review comprehensively evaluates machine learning (ML) and deep learning (DL) models for predicting new-onset atrial fibrillation (NOAF) in sepsis patients, highlighting the potential of multimodal data integration. Traditional ML models (AUC 0.74–0.90) using tabular data (e.g., demographics, lab results) or ECG-derived features offer short-term prediction (minutes to hours), while DL models (AUC 0.74–0.96) processing raw ECG signals extend horizons to days, with performance scaling with dataset size. The proposed transformer-based multimodal framework, integrating clinical data and ECG signals, offers a promising avenue to enhance predictive performance by capturing systemic inflammation and atrial remodeling dynamics. Despite limitations like retrospective data heterogeneity, the proposed framework offers valuable insights for addressing common challenges in cardiovascular AI research particularly regarding data harmonization and clinical implementation.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076251384237 - Supplemental material for Prediction of new-onset atrial fibrillation in sepsis patients by machine learning: A systematic review
Supplemental material, sj-docx-1-dhj-10.1177_20552076251384237 for Prediction of new-onset atrial fibrillation in sepsis patients by machine learning: A systematic review by Shuxuan Ye, Wan Xu, Zhiyu Jiang, Yujie Zhan, Yuqiang Shen, Lan Su, Keda Yang and Bin Ju in DIGITAL HEALTH
Supplemental Material
sj-docx-2-dhj-10.1177_20552076251384237 - Supplemental material for Prediction of new-onset atrial fibrillation in sepsis patients by machine learning: A systematic review
Supplemental material, sj-docx-2-dhj-10.1177_20552076251384237 for Prediction of new-onset atrial fibrillation in sepsis patients by machine learning: A systematic review by Shuxuan Ye, Wan Xu, Zhiyu Jiang, Yujie Zhan, Yuqiang Shen, Lan Su, Keda Yang and Bin Ju in DIGITAL HEALTH
Footnotes
Ethical approval
Not applicable.
Contributorship
Keda Yang and Bin Ju conceived of the study idea and designed the search strategy. Shuxuan Ye and Wan Xu screened abstracts and full texts. Bin Ju and Lan Su performed the analysis and drew figures. Lan Su revised the manuscript. Yujie Zhan and Yuqiang Shen drafted the introduction, materials and methods sections. All authors cowrote and revised the manuscript for intellectual content. All authors provided their final approval for manuscript submission. All authors agree to be accountable for all aspects of the work.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Medical and Health Research Project of Zhejiang Province (2023KY1039). Higher Education Research Planning Project of the China Association of Higher Education 2024 (24YJ0403).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
