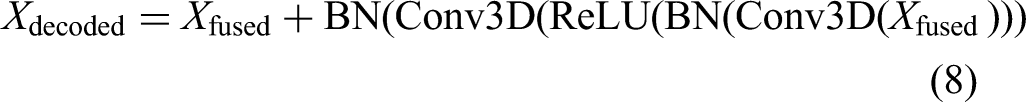Abstract
Objective
Accurate magnetic resonance imaging-transrectal ultrasound (MRI-TRUS) registration is essential for improving prostate cancer detection and biopsy guidance. This study presents a weakly supervised learning framework to enhance image alignment while minimizing reliance on extensive labeled data.
Methods
We developed a two-stage weakly supervised framework integrating an attention-enhanced U-Net for prostate segmentation and a residual-enhanced registration network (RERN) for MRI-TRUS alignment. The segmentation model was trained and evaluated on public MRI datasets (MSD Prostate, Promise12, µ-RegPro) and TRUS datasets (µ-RegPro), with additional testing on a clinical cohort of 32 MRI-TRUS pairs. Performance was assessed using dice similarity coefficient (DSC), accuracy (Acc), precision (Pre), recall (Rec), and the 95th percentile Hausdorff distance (HD95). The registration model was trained and tested using MRI-TRUS pairs from µ-RegPro and the clinical cohort, with performance evaluated based on HD95 and target registration error (TRE). Additionally, clinical validation included PI-RADS-like grading, Likert confidence scoring, and receiver operating characteristic (ROC) analysis to assess diagnostic impact.
Results
The segmentation model demonstrated high accuracy (DSC: MRI 0.9154, TRUS 0.9384) and strong generalizability to clinical data (MRI 0.9010, TRUS 0.9173). The registration model achieved robust performance, with HD95 of 10.18 mm on public datasets and 11.18 mm on clinical data, and TRE below 8.64 mm. Clinical validation confirmed that registered MRI images preserved diagnostic integrity, as no significant differences were observed in radiologists’ diagnostic performance (AUC: junior 0.706, senior 0.781, p > 0.05). Moreover, registered images enhanced diagnostic confidence among senior radiologists (Likert score: 3.032 vs. 3.548, p = 0.015), highlighting their potential to support expert-level decision-making.
Conclusions
The proposed weakly supervised framework achieves high-precision MRI-TRUS registration with minimal annotation, ensuring strong generalizability and clinical applicability.
Keywords
Introduction
Prostate cancer is one of the most common malignancies among men worldwide and a leading cause of cancer-related mortality in males.1–3 According to global cancer statistics, the incidence of prostate cancer has been continuously rising in recent years, particularly in aging societies.4,5 Although the disease progresses relatively slowly, early identification and accurate assessment of its malignancy are crucial for optimizing treatment strategies and improving patient survival. 6 The early detection and precise localization of clinically significant prostate cancer (csPCa) are essential for improving patient prognosis, underscoring the clinical significance of exploring more precise and reliable diagnostic techniques.
Magnetic resonance imaging (MRI) and transrectal ultrasound (TRUS) are the primary imaging modalities for prostate cancer diagnosis.7,8 MRI offers excellent soft tissue contrast and can clearly delineate prostate anatomy and tumor boundaries, making it widely used for preoperative evaluation.9,10 However, MRI has certain limitations, including long acquisition times, high costs, and the inability to provide real-time guidance during biopsy procedures.11,12 In contrast, TRUS is widely utilized for guiding prostate cancer biopsy due to its simplicity, lower cost, and real-time imaging capabilities. However, TRUS has relatively poor soft tissue resolution, making it challenging to distinguish between normal and malignant tissues. 13 Additionally, TRUS images are often affected by anatomical overlaps and artifacts, which can limit the precision of needle placement and consequently impact biopsy success rates and diagnostic accuracy.14,15 Therefore, improving imaging guidance quality can not only reduce false-negative and false-positive rates but also minimize patient discomfort and complications. High-quality image registration techniques, particularly MRI-TRUS fusion-based registration, have the potential to significantly enhance biopsy guidance accuracy, 16 thereby facilitating early prostate cancer detection and personalized treatment. Accurate MRI-TRUS registration plays a particularly important role in patients with a history of prior negative biopsies, where multiparametric MRI (mpMRI) has been shown to exhibit reduced diagnostic sensitivity and increased variability in lesion interpretation. A recent study by Barone et al. 17 demonstrated that the detection rate of clinically significant prostate cancer (csPCa) in previously biopsied patients was significantly lower than in biopsy-naïve individuals, emphasizing the necessity of precise image fusion to improve lesion targeting in repeat biopsy scenarios.
In recent years, artificial intelligence (AI), particularly deep learning, has demonstrated immense potential in medical image analysis. In the context of MRI-TRUS image registration, deep learning-based methods can automatically extract high-dimensional features and optimize the accuracy of cross-modal image matching. 18 Currently, mainstream AI-based registration approaches can be categorized into three paradigms: supervised learning, unsupervised learning, and weakly supervised learning.19–21 Supervised learning methods rely on large amounts of labeled data, which yield high registration accuracy but come at a significant annotation cost. 22 Unsupervised learning methods do not require registration labels but may suffer from insufficient accuracy due to the absence of explicit supervision signals. 23 Weakly supervised learning, by integrating the strengths of both paradigms, leverages partial annotations to guide the learning process, thereby reducing the need for extensive manual labeling while maintaining high registration accuracy. 24 However, existing weakly supervised MRI-TRUS registration methods still face challenges such as poor model adaptability and limited generalization capabilities, necessitating further optimization.
In this study, we propose a weakly supervised MRI-TRUS image registration framework to address the challenges associated with MRI-TRUS alignment and improve registration accuracy. Our framework incorporates an improved U-Net-based segmentation model combined with a weakly supervised registration model, where segmentation labels guide the registration process to reduce dependence on large-scale annotated datasets while enhancing registration precision. We utilize publicly available datasets (MSD Prostate, 25 Promise12, 26 µ-RegPro 27 ) alongside clinical datasets for model training and evaluation, ensuring robustness across different data environments. Additionally, we introduce a novel evaluation method based on objective registration metrics such as dice similarity coefficient (DSC), 95th percentile Hausdorff distance (HD95), and target registration error (TRE) to assess the readability of registered MRI images and their potential clinical value.
Methods
Study datasets
The dataset used in this study consists of the MSD Prostate dataset (48 prostate MRI cases), the Promise12 dataset (50 prostate MRI cases), the µ-RegPro dataset (73 paired MRI-TRUS cases), and an additional 32 paired MRI-TRUS cases from Zhangzhou Affiliated Hospital of Fujian Medical University, which serve as the clinical cohort. All MRI images were acquired using T2-weighted imaging (T2WI) sequences and were utilized for image analysis and registration tasks. The clinical data were collected between April 2023 and July 2023, with a mean patient age of 75.13 ± 5.69 years (mean ± SD). This study was conducted as a retrospective analysis and was approved by the Institutional Review Board (IRB) of Zhangzhou Affiliated Hospital of Fujian Medical University (IRB: 2024LWB332). Given that the data were derived from previously recorded clinical imaging and did not involve any direct patient intervention, the IRB granted a waiver of informed consent. For cases from Zhangzhou Affiliated Hospital of Fujian Medical University, lesion assessment was performed according to the PI-RADS v2.1 criteria to ensure consistency and standardization in image interpretation.
Inclusion criteria: (a) underwent prostate MRI for prostate-related conditions; (b) aged ≥50 years; (c) had a documented PI-RADS v2.1 score in their medical records; (d) had MRI images meeting quality requirements, including artifact-free T2WI sequences and other essential sequences, along with complete TRUS images that clearly delineated the prostate contour and structural features. Exclusion criteria: (a) presence of severe psychiatric or cognitive disorders; (b) prior treatment for prostate-related conditions that could affect imaging or clinical assessment, such as radical surgery, radiotherapy, chemotherapy, or hormone therapy; (c) recent participation in other clinical studies that might interfere with the study results; (d) presence of severe comorbidities or poor imaging quality (e.g. severe artifacts) that rendered the images unsuitable for analysis.
Study design
This study adopts a weakly supervised learning strategy to address the MRI-TRUS image registration task and proposes a two-stage approach, consisting of an initial segmentation phase followed by a registration phase, to achieve high-precision non-rigid registration. In the segmentation phase, a segmentation model was trained and evaluated to extract the prostate regions from both MRI and TRUS images, providing structured anatomical information for subsequent registration tasks. The MRI images were obtained from the MSD Prostate, Promise12, and µ-RegPro datasets, comprising a total of 171 cases (11,624 images), with approximately 80% used for training and 20% for testing. The TRUS images were sourced from the µ-RegPro dataset, consisting of 73 cases (6424 images), and were split into training and validation sets in a roughly 9:1 ratio. In the registration phase, both MRI and TRUS images, along with their segmentation labels, were used as weak supervision signals to guide the registration model in learning non-rigid deformation fields of the prostate. Specifically, the µ-RegPro dataset, which contains 73 paired MRI-TRUS cases, was divided into training and test sets (approximately 9:1 split). The model leveraged the structural information provided by segmentation labels to align MRI images to TRUS images, thereby enhancing spatial consistency in cross-modal registration. To further evaluate the generalizability of the proposed model, an external clinical test set of 32 paired MRI (512 images) and TRUS (2816 images) cases was included for model adaptation and robustness assessment. After excluding one outlier case, a total of 31 cases were analyzed in the clinical validation study, which aimed to assess the usability of registered images in real-world diagnostic scenarios. The overall workflow of this study is illustrated in Figure 1.

The flowchart of this study included three elements: (a) segmentation, (b) registration, and (c) clinical assessment, integrating public datasets, weakly supervised MRI-TRUS alignment, and expert evaluation via PI-RADS-like grading.
Data acquisition and preprocessing
For the clinical dataset, prostate MRI and TRUS images were acquired following standardized imaging protocols using multiple imaging systems. MRI scans were obtained using Siemens Healthineers MAGNETOM Sola and Siemens Healthineers MAGNETOM Altea scanners, with magnetic field strengths of 1.5 T and 3.0 T, respectively. TRUS images were captured using the Esaote MyLabClass C ultrasound system.
Given the inherent characteristics of MRI and TRUS imaging, certain slices from the upper and lower regions of MRI scans and frames from the initial and final portions of TRUS sequences often failed to adequately depict prostatic structures. These regions contained redundant information that contributed little to model training. To optimize computational efficiency and maximize the relevance of input data, we preprocessed the images by removing the top and bottom 10% of MRI slices and discarding the first and last 25% and 12.5% of TRUS frames, respectively. This refinement process not only improved the quality of input data but also enhanced the overall accuracy of model predictions.
The preprocessing pipeline involved format conversion, image standardization, and data augmentation. MRI and TRUS images, originally stored in DICOM format, were processed using pydicom to extract image metadata and subsequently converted into NIfTI format using nibabel for efficient storage and handling. To ensure spatial consistency across different imaging modalities, we standardized MRI images to 160 × 160 pixels and TRUS images to 118 × 88 pixels through cropping and resampling. Additionally, multiple data augmentation techniques—including random rotation, translation, and scaling—were implemented to enhance model generalizability. These augmentation strategies increased data diversity and improved the robustness of both segmentation and registration models.
For the clinical validation, further preprocessing techniques, including histogram equalization, gamma correction, and Laplacian filtering, were applied to adjust image contrast and align feature distributions with those of the public dataset. After segmentation, the original images were used as inputs to the registration network, while segmentation masks were leveraged as auxiliary supervision signals during training, facilitating a segmentation-guided weakly supervised registration framework.
Development of the segmentation model
This study proposes a semantic segmentation network based on an attention mechanism, incorporating spatial attention (SA) in the encoder and channel attention (CA) in the decoder to optimize feature extraction and fusion (Figure 2(a)). This design allows the model to accurately capture target region information while enhancing semantic representation across different channels, ultimately improving segmentation accuracy and generalization ability.
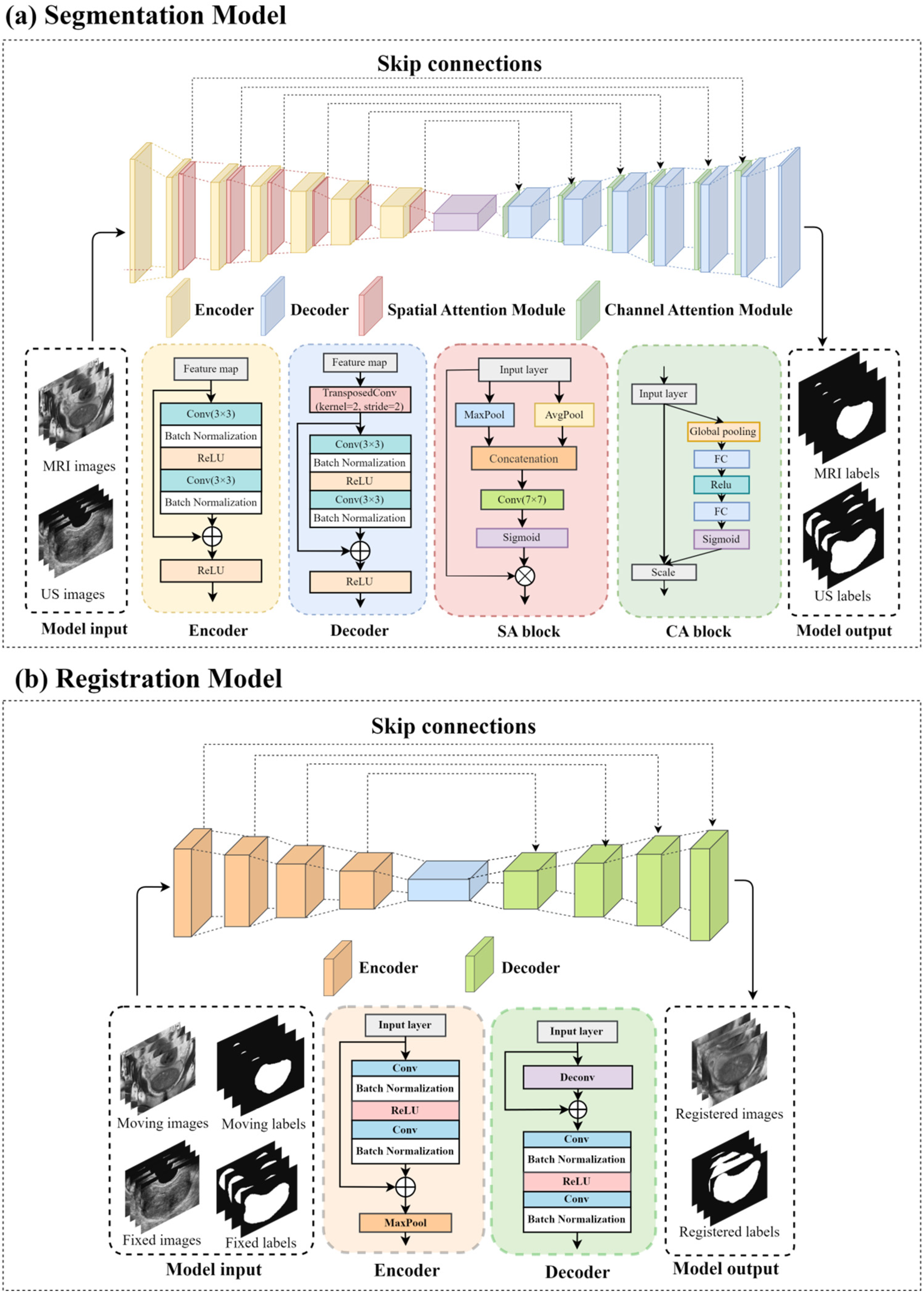
Schematic illustration of the network architecture: (a) MRI/TRUS image segmentation network, and (b) MRI-TRUS image registration network.
In the encoder, the output of each stage undergoes processing by a SA module to enhance spatial feature extraction. Given an input feature map
In the decoder, each stage input is processed by a CA module to optimize cross-channel information interaction. Given an input feature map
Additionally, the model adopts a progressive spatial restoration decoder structure with skip connections to facilitate feature fusion between shallow and deep layers. Specifically, at each decoding stage, the decoder output is element-wise added to the corresponding encoder feature:
Development of the registration model
To enhance registration accuracy and robustness, this study proposes a residual-enhanced registration network (RERN), which is based on a U-Net architecture. RERN employs residual learning and skip connections to facilitate feature extraction and fusion, optimizing the learning of deformation fields for more precise medical image alignment (Figure 2(b)).
Similar to conventional U-Net architectures, the encoder in RERN utilizes residual learning for feature extraction. Each encoding layer consists of convolutional operations (Conv), batch normalization (BN), and ReLU activation, followed by max pooling to reduce computational complexity and increase the receptive field of the model. Given an input feature map
Initially, a double convolution block is employed for feature extraction, followed by skip connections for residual fusion, ensuring stable gradient propagation. Max pooling reduces the spatial resolution, thereby enhancing the robustness of feature extraction.
The decoder is responsible for restoring spatial resolution and generating a dense displacement field (DDF). In the decoding phase, 3D deconvolution (Deconv3D) is first applied to upsample the feature maps and recovers spatial resolution. Subsequently, skip connections integrate the upsampled features with the corresponding encoder outputs, mitigating gradient vanishing and promoting complementary fusion between shallow and deep features. The skip connections employ element-wise addition, ensuring effective feature combination across different hierarchical levels. Given a decoder input
In this framework, deconvolution serves to restore spatial resolution, while element-wise addition fuses decoder and encoder features. Finally, a residual convolution block refines the deep feature representation, enhancing the expressive power of the registration network.
Clinical evaluation of segmentation model
During the model training phase, separate segmentation models were trained for MRI and TRUS images to ensure that each model independently learned and adapted to its respective imaging modality. Upon completing training, segmentation performance was first evaluated on public datasets by computing five key metrics: DSC, accuracy, precision, recall, and the HD95. The accuracy of the segmentation models was assessed by comparing the predicted segmentation results with the ground truth segmentation labels available in the public datasets.
To further evaluate the generalization ability of the segmentation models, we selected 32 cases from the clinical dataset at Zhangzhou Affiliated Hospital of Fujian Medical University, including T2-weighted MRI scans with complete sequences and TRUS images with high-quality acquisition. These cases were used to independently test the MRI and TRUS segmentation models. A senior radiologist manually delineated the prostate region using 3D Slicer software (version 5.6.2) to generate segmentation labels, which served as the GT for evaluation. Subsequently, the 32 MRI and TRUS images were processed through their respective pre-trained segmentation models, generating automatic segmentation results. The model predictions were then quantitatively compared against the GT labels using the five evaluation metrics, allowing us to assess segmentation performance on clinical data.
Evaluation of the registration model
Following model training, the registration performance was evaluated using three key metrics: DSC, HD95, and TRE. Description of the TRE can be found in Appendix S2. The model's accuracy was first assessed on public datasets and subsequently validated on a clinical dataset consisting of 32 MRI-TRUS cases from Zhangzhou Affiliated Hospital of Fujian Medical University. One case exhibiting abnormal prostatic hyperplasia was excluded, leaving 31 cases for final evaluation. The model's generalization ability was examined by comparing performance across different datasets and assessing its consistency with existing registration methods. This comprehensive evaluation provided insights into the model's effectiveness and cross-dataset adaptability.
In addition to evaluating registration accuracy, we further examined the clinical applicability of MRI-TRUS registered images in prostate disease diagnosis. Based on 31 patients, the initial PI-RADS scores assigned during diagnosis were used as reference (PI-RADS 4/5 classified as positive, 2/3 classified as negative). Two radiologists (junior and senior) independently re-evaluated the registered MRI images under blinded conditions. Since this assessment was based solely on T2-weighted sequences, without incorporating diffusion-weighted imaging (DWI) or dynamic contrast-enhanced (DCE) sequences, a modified PI-RADS-like scoring system was employed, following standard PI-RADS evaluation principles. Description of the PI-RADS-like Scoring System Based on T2-weighted MRI can be found in Appendix S1.
To further investigate the impact of image registration on diagnostic confidence, both radiologists rated their diagnostic certainty using a 7-point Likert scale (1 = highly uncertain, 7 = highly certain), Details related to the confidence score can be found in Appendix S3. Finally, a receiver operating characteristic (ROC) analysis was conducted to compare diagnostic accuracy between pre- and post-registration MRI images. Additionally, the inter-reader agreement for PI-RADS-like scoring and Likert ratings was analyzed to evaluate the effect of image fusion on diagnostic confidence.
Statistical analysis
Statistical analyses were conducted by two experienced biostatisticians, one with 10 years and the other with 7 years of expertise. A p-value < 0.05 was considered statistically significant, and 95% confidence intervals (CIs) were used to represent the reliability of the results. All statistical analyses were performed using SPSS software (version 26.0, IBM Corporation, Armonk, USA) and RStudio software (version 2024.09.0 + 375, RStudio, Inc., Boston, USA).
For non-paired sample data, the Mann–Whitney U test (two-tailed) was applied, while paired data were analyzed using the Wilcoxon signed-rank test (two-tailed). To evaluate differences in AUC values between junior and senior radiologists under different conditions for PI-RADS scoring, the Pairwise DeLong test (two-tailed) was used. Kappa statistics were employed to assess the inter-rater agreement between the two groups.
Results
Patient characteristics
Table 1 presents the baseline characteristics and PI-RADS classification of the 32 prostate cancer patients included in the clinical dataset. All patients were male, with a mean age of 75.13 years (range: 58–85 years). PI-RADS grading was used to assess the risk of prostate cancer, where 16 patients had PI-RADS scores of 2 or 3, indicating a lower risk of malignancy, while the other 16 patients had PI-RADS scores of 4 or 5, suggesting a higher risk of prostate cancer.
Patient characteristics of the clinical cohort.
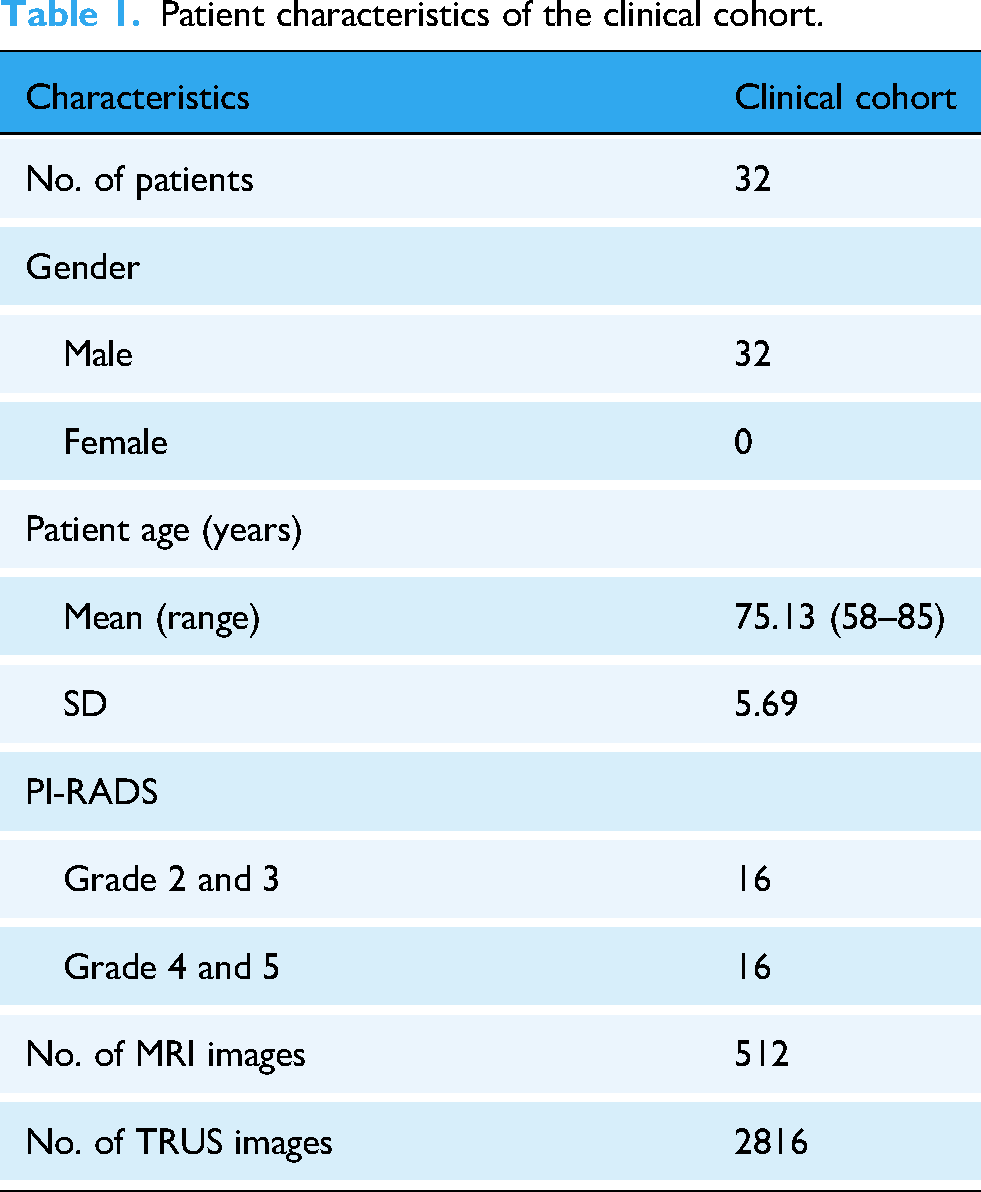
Each patient's dataset contained 16 MRI slices and 88 TRUS images, resulting in a total of 512 MRI slices and 2816 TRUS images in the clinical cohort. These data provide a critical clinical reference for subsequent registration and segmentation model evaluation.
Performance of segmentation models
Table 2 presents the performance evaluation results of the segmentation model across different datasets, demonstrating consistent performance in various test scenarios. On the public MRI dataset, the model achieved a DSC of 0.9154 and an Acc of 0.9914, indicating high segmentation precision for MRI-based prostate structures. While the performance slightly declined in the clinical dataset, with DSC = 0.9010 and Acc = 0.9753, the model still maintained strong segmentation accuracy in real-world clinical applications.
Performance metrics of the segmentation model on different datasets.
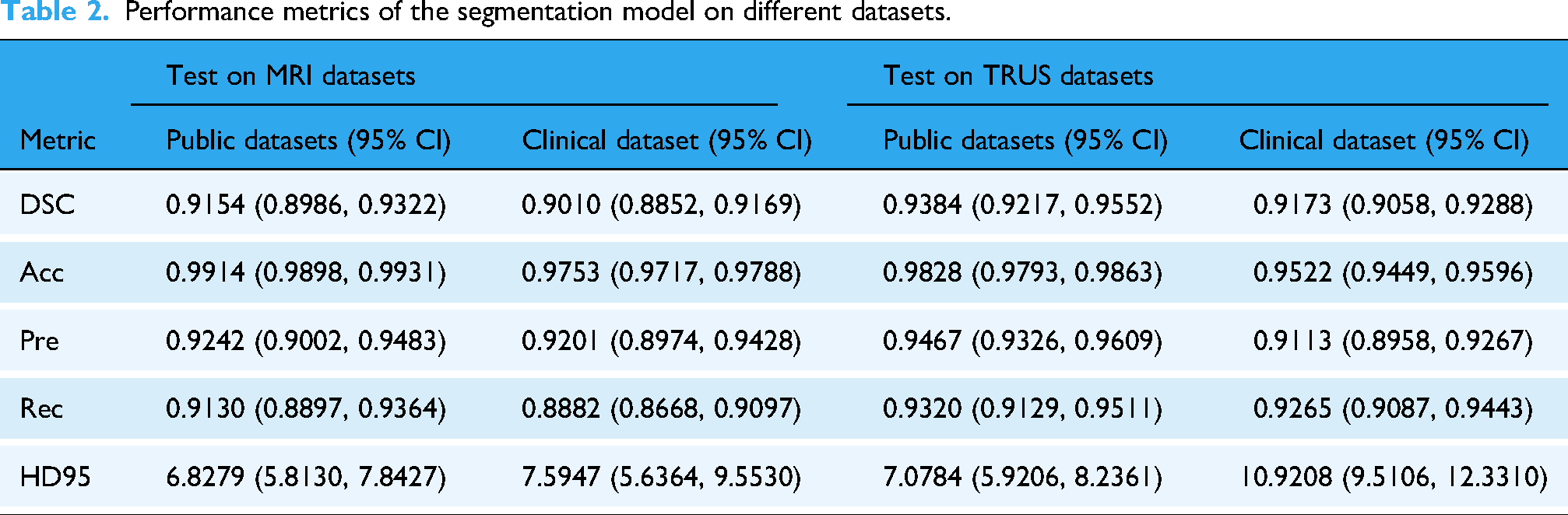
For TRUS images, the segmentation task was more challenging, yet the model still exhibited robust performance. In the public TRUS dataset, the model achieved DSC = 0.9384 and Acc = 0.9828, reflecting its effective segmentation capability for TRUS images. Similarly, in the clinical dataset, despite a slight drop in performance, the model still reached DSC = 0.9173 and Acc = 0.9522, demonstrating strong adaptability to different data domains.
Overall, the segmentation performance on MRI images was slightly superior to that on TRUS images, which may be attributed to better anatomical boundary clarity and higher soft tissue contrast in MRI scans. Regarding boundary segmentation accuracy, as measured by HD95, no statistically significant difference was observed between the public MRI dataset and the clinical MRI dataset (public MRI vs. clinical MRI, p = ns). However, in the TRUS images, the public dataset significantly outperformed the clinical dataset (public TRUS vs. clinical TRUS, p = 0.007). Similarly, while no significant difference in HD95 was found between public MRI and public TRUS datasets (public MRI vs. public TRUS, p = ns), the segmentation performance on clinical MRI images was significantly better than that on clinical TRUS images (clinical MRI vs. clinical TRUS, p < 0.001).
Notably, the segmentation model demonstrated strong and reliable performance across different modalities and datasets, making its segmentation outputs suitable as weak supervision labels for the registration model. Although a performance drop was observed in the clinical dataset, particularly for boundary segmentation, the model exhibited high generalizability, providing structural guidance for registration tasks in real-world clinical applications. Figure 3 visualizes the prostate segmentation results on MRI and TRUS images, alongside expert annotations for comparison.
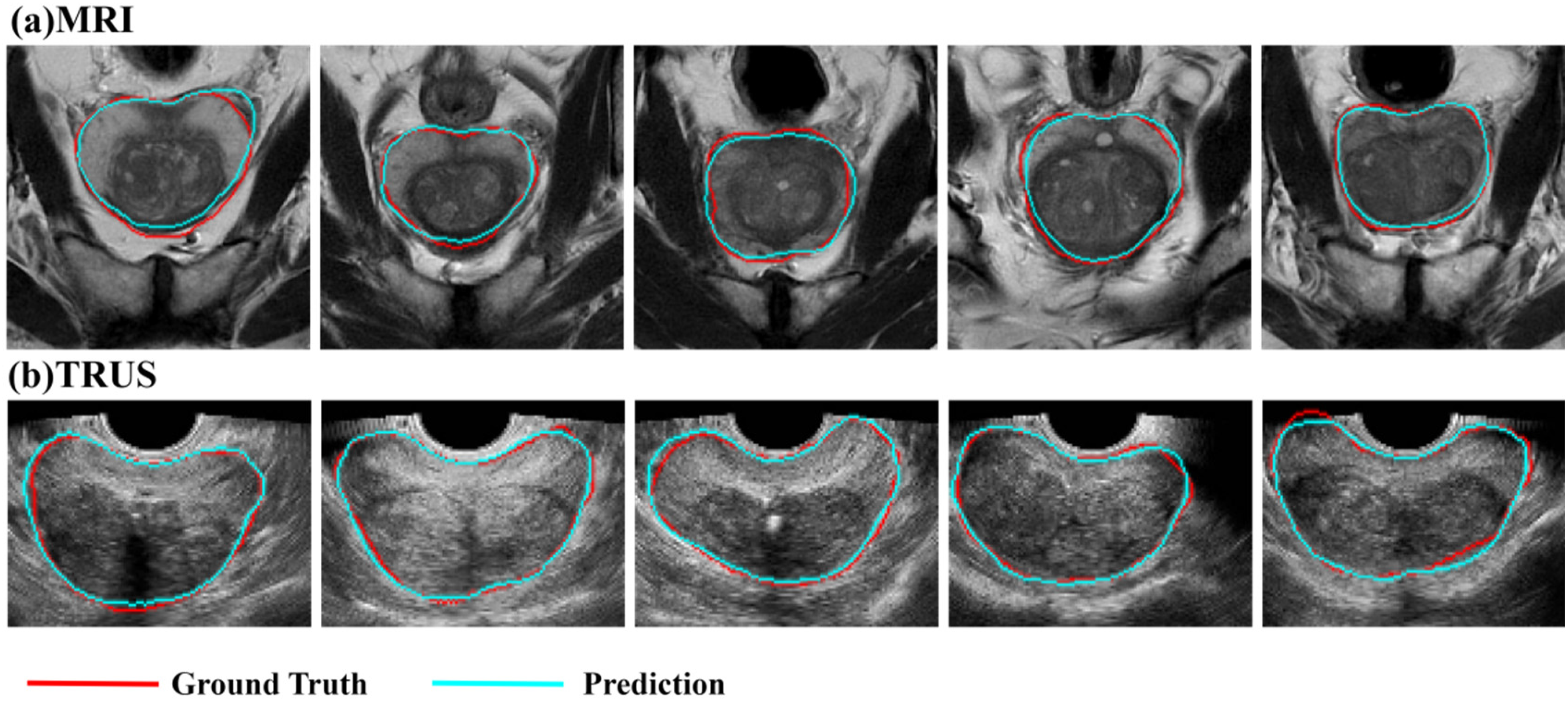
Segmentation results of (a) MRI and (b) TRUS images. Red indicates the GT annotated by expert radiologists, while blue represents the outputs of the segmentation model.
Performance of registration model
Table 3 presents the performance evaluation results of the registration model across public and clinical datasets, focusing on comparing contour alignment and key anatomical points between original MRI images and registered MRI images. The evaluation is based on three key metrics: DSC, HD95, and TRE.
Performance metrics of the registration model on different datasets.

In this study, TRE is used to assess the multimodal registration accuracy between prostate MRI and TRUS images. Specifically, TRE measures the registration error between selected MRI and TRUS coordinates, mathematically defined as:
On the public dataset, the model achieved DSC = 0.7481, HD95 = 10.1839 mm, and TRE = 7.8562 mm, demonstrating high registration accuracy, particularly in DSC and accuracy metrics, confirming the model's ability to achieve precise segmentation and registration of the prostate region. Comparatively, on the clinical dataset, the model achieved DSC = 0.7295, HD95 = 11.1766 mm, and TRE = 8.6307 mm. No statistically significant differences were observed between the public and clinical datasets for DSC, HD95, or TRE, suggesting robust model performance and high registration precision in real-world clinical settings. These findings further validate the generalizability and adaptability of the model in complex clinical scenarios, indicating its capability to maintain high segmentation and registration accuracy for practical clinical applications. The output results of the registration model are shown in Figure 4.

Registration results: (a) Registered MRI images, (b) Original TRUS images (fixed images), and (c) Original MRI images (moving images).
Clinical assessment of the registration model
We conducted ROC analysis to assess the diagnostic accuracy of physicians when interpreting original and registered MRI images, as shown in Figure 5. The AUC for the junior doctor increased from 0.590 to 0.706 (p = 0.2239), while the senior doctor's AUC showed a slight increase from 0.775 to 0.781 (p = 0.941). Neither change was statistically significant, indicating that image registration did not affect overall diagnostic performance nor introduce interpretation bias due to deformation-based alignment or physician reading habits. These findings further validate the stability of the registration process, demonstrating its feasibility in providing reliable imaging support for clinical decision-making without compromising diagnostic accuracy.

Comparison of physicians’ diagnostic performance based on PI-RADS-like grading. (a) ROC curve comparison for the junior doctor before and after MRI registration. (b) ROC curve comparison for the senior doctor before and after MRI registration. (c) Kappa coefficient comparison between junior and senior doctors. (d) Confidence score comparison before and after MRI registration. * p < 0.05.
To further evaluate consistency in PI-RADS scoring, we categorized scores 2 and 3 as low risk (0) and scores 4 and 5 as high risk (1) for simplified analysis. The results showed that the junior doctor's Kappa coefficient was 0.330, indicating low scoring consistency and significant inter-reader variability. In contrast, the senior doctor's Kappa coefficient was 0.784, demonstrating high scoring consistency, suggesting that experienced physicians maintained more stable assessments under the simplified grading system.
We further analyzed physicians’ diagnostic confidence. The junior doctor's mean confidence score decreased after registration (3.613 vs. 3.266, p = 0.022), while the senior doctor's score significantly increased (3.032 vs. 3.548, p = 0.015), with scores becoming more concentrated around 4. These results suggest that, although image registration did not significantly enhance the junior doctor's confidence and may have even introduced some uncertainty, it improved diagnostic confidence and reading stability for the senior doctor.
Discussion
This study proposes a weakly supervised registration model that integrates an attention-enhanced U-Net architecture with label-driven learning to achieve high-precision prostate MRI-TRUS image registration. Experimental results demonstrate that the proposed method exhibits stable registration performance on both public and clinical datasets, with HD95 < 11.18 mm and TRE < 8.64 mm, ensuring spatial consistency and anatomical alignment between MRI and TRUS images. Further analysis revealed no significant difference in ROC curves for junior and senior physicians when evaluating PI-RADS-like grading before and after MRI registration, indicating that registered MRI images maintain the same clinical diagnostic efficacy as the original MRI scans. Additionally, registered images significantly enhanced the diagnostic confidence of senior physicians, improving the consistency of imaging assessments. This study employs weakly supervised learning to achieve high-accuracy MRI-TRUS image registration while reducing dependence on manual annotations. The proposed approach enhances registration stability and improves biopsy guidance accuracy, thereby reducing annotation costs and increasing model generalizability. Ultimately, this method provides technical support for early prostate cancer diagnosis and personalized treatment while contributing to advancements in automated medical image processing.
The results of this study are consistent with previous research 28 on weakly supervised deep learning for MRI-TRUS prostate image registration. Prior studies have demonstrated that deep learning models, particularly U-Net-based segmentation and registration methods, perform well in multimodal medical image registration. 29 This study adopts a weakly supervised strategy to achieve high-accuracy MRI-TRUS image registration while reducing dependence on manual annotations, with registration errors controlled at HD95 < 11.18 mm and TRE < 8.64 mm. However, compared to partially supervised learning methods or traditional landmark-based approaches, 30 our registration errors are slightly higher. This may be due to the weakly supervised approach reducing reliance on precise annotated data, allowing the model to remain stable even with limited annotation availability, thereby enhancing its applicability in real-world clinical settings. 31 In contrast, traditional methods often rely on manually defined anatomical landmarks or statistical deformation models, which in some cases may yield lower registration errors. 32 Nonetheless, it is important to note that traditional optimization-based methods typically involve high computational complexity, making them less feasible for real-time applications. In contrast, our method significantly improves computational efficiency while maintaining high registration accuracy. By employing segmentation-guided weakly supervised registration, our approach leverages segmentation labels as indirect supervision signals to guide non-rigid MRI-TRUS image registration. Compared to direct deformation field prediction, this strategy reduces the need for manual annotations while improving anatomical consistency and computational efficiency, making it more suitable for automated medical image analysis and clinical applications.
Notably, although MRI registration did not lead to significant differences in PI-RADS-like grading performance for both junior and senior physicians (AUC: junior p = 0.224; senior p = 0.941), there was a marked difference in scoring consistency (Kappa: junior 0.330 vs. senior 0.784). Retrospective analysis revealed that junior physicians exhibited lower consistency due to limited experience and insufficient MRI interpretation skills, leading to a decrease in diagnostic confidence after registration. In contrast, senior physicians demonstrated significantly improved diagnostic confidence (original MRI vs. registered MRI = 3.032 vs. 3.548, p = 0.015). This phenomenon may be attributed to several mechanisms. First, MRI registration optimized spatial alignment and anatomical consistency, enabling physicians to visually localize lesion characteristics (e.g. T2 signal abnormalities) more intuitively within standard anatomical structures, thus reducing spatial uncertainty caused by patient positioning variability. 33 Second, the registration process corrected anatomical landmark misalignment due to positional deformation (e.g. blurred prostate boundaries), thereby enhancing visual continuity of key regions (e.g. central gland, capsule). 18 This may have shortened senior physicians’ interpretation time and reduced cognitive load. 34 Lastly, senior physicians’ clinical experience may have made them more adept at leveraging the spatial consistency of registered images to infer biological characteristics of lesions (e.g. multifocality, invasiveness), ultimately improving diagnostic decision-making certainty. In addition to the quantitative evaluation, our findings revealed a noteworthy divergence in diagnostic confidence between junior and senior physicians following the use of registered images. While senior physicians reported significantly improved confidence, junior physicians exhibited a notable decrease. This discrepancy likely arises from differences in clinical experience and familiarity with interpreting spatially fused multimodal images. To address this issue, future efforts may focus on enhancing the interpretability and usability of registered outputs through visual aids such as semi-transparent MRI overlays on TRUS images, contour-based highlighting of anatomical structures, and synchronized navigation tools. Additionally, user interface optimizations—including standardized alignment indicators, confidence heatmaps, and simplified reporting modules—may reduce cognitive burden for less experienced readers. The development of targeted training protocols for image fusion interpretation may further promote consistent diagnostic performance across physician groups. In addition, Barone et al. 17 further highlighted the diagnostic challenges associated with patients who had prior negative biopsies, where mpMRI demonstrated limited sensitivity and inconsistent lesion detectability. These findings underscore the clinical necessity of improving MRI-TRUS registration accuracy, particularly in anatomically complex or previously biopsied regions. By enhancing spatial correspondence between modalities, our proposed approach has the potential to reduce diagnostic uncertainty and improve lesion targeting in these high-risk populations.
This study provides a novel perspective on deep learning-driven medical image registration, particularly in the application of weakly supervised learning methods, expanding and optimizing existing theoretical models of medical image registration. Traditional medical image registration approaches primarily rely on rigid or non-rigid transformations based on optimization techniques, often requiring handcrafted feature extraction or similarity metric computation. 34 In recent years, fully supervised deep learning methods have become mainstream; however, their heavy dependence on large-scale pixel-wise annotated datasets significantly limits their scalability in clinical applications. 35 By introducing a weakly supervised learning strategy, this study reduces dependence on precise annotations while ensuring high-accuracy MRI-TRUS registration, thereby broadening the applicability of deep learning in medical image registration tasks. The results demonstrate that weakly supervised deep learning models can achieve accurate MRI-TRUS alignment under limited supervision, while maintaining spatial consistency. Although registration errors in weakly supervised methods may be slightly higher compared to fully supervised approaches, their advantages in data efficiency, generalizability, and computational complexity control make them more suitable for medical scenarios with limited annotation availability. Furthermore, this study validates the feasibility of self-supervised and weakly supervised learning in medical image analysis and paves the way for future research directions, such as integrating self-supervised learning with contrastive learning-based registration frameworks to further reduce reliance on manual annotations while enhancing registration accuracy. Additionally, the findings of this study can contribute to the further optimization of learning-based medical image registration models, providing theoretical support for MRI-TRUS prostate image registration and other multimodal medical imaging tasks. This work advances the application of deep learning methods in medical image registration, promoting the development of automated and efficient medical imaging solutions.
Although this study validates the effectiveness of weakly supervised learning in MRI-TRUS image registration, achieving high registration accuracy while reducing dependence on manual annotations, certain limitations remain. First, the clinical dataset used in this study was derived from a single-center cohort with a relatively limited sample size, which may impact the model's generalizability. Its robustness in cross-device scenarios (e.g. different TRUS probe frequencies or MRI scanners with varying field strengths) and heterogeneous imaging protocols still requires further validation. 30 Second, while the weakly supervised strategy reduces annotation dependence, it relies on segmentation labels as indirect supervision, which may lead to suboptimal local deformation fields in certain regions (e.g. the prostatic apex or bladder neck). 31 This could partially explain why the registration errors (HD95: 11.18 mm; TRE: 8.64 mm) were slightly higher than those of fully supervised methods. 33 To further illustrate the limitations of our model in anatomically challenging regions, Supplementary Figure S1 presents representative cases of suboptimal registration in the prostatic apex and bladder neck. These examples demonstrate localized deformation mismatches, potentially caused by weak supervision signals and complex boundary structures. Such cases provide valuable insights for future optimization strategies, including refinement of the loss function and incorporation of region-specific adaptive mechanisms. Finally, although the clinical relevance of the registered images was verified using PI-RADS scoring and diagnostic confidence metrics, the biological consistency of the registration results—such as the spatial correlation between registered MRI features (e.g. ADC values) and histopathological findings—has not yet been rigorously quantified.
Future research should prioritize validating the proposed model across multiple centers and diverse imaging environments, as variations in scanner types, imaging protocols, and patient populations may introduce clinically significant variability. To enhance robustness and generalizability, future work should also explore domain adaptation and calibration strategies tailored to cross-institutional settings. Moreover, incorporating anatomical topology constraints into the loss function and developing lightweight architectures could improve spatial coherence in anatomically complex regions and facilitate real-time deployment in resource-constrained clinical workflows. Beyond technical improvements, the integration of accurate MRI-TRUS registration into routine diagnostic pathways holds substantial translational potential. By aligning imaging predictions more closely with underlying pathology, this approach may help bridge the diagnostic gap between preoperative imaging and final histological staging. As highlighted by Rapisarda et al., 36 discrepancies in lesion laterality and Gleason grading between mpMRI-guided biopsies and prostatectomy specimens can affect treatment planning and risk stratification. Enhancing anatomical alignment and lesion localization could therefore reduce sampling errors and improve biopsy targeting accuracy. To preliminarily evaluate this potential, we conducted an additional analysis comparing lesion laterality identified on registered MRI versus original TRUS with subsequent biopsy results. The results showed improved concordance following registration, particularly for left-sided and bilateral lesions. Furthermore, consistency between PI-RADS scores and corresponding Gleason grades supported the alignment between imaging suspicion levels and pathological aggressiveness. These findings, detailed in Supplementary Tables S1–S3, provide early evidence that our registration framework may contribute to more precise lesion localization and clinically meaningful decision-making.
Conclusion
In this study, we proposed a weakly supervised MRI-TRUS image registration framework that integrates an attention-enhanced U-Net segmentation model with a label-guided non-rigid registration network. This approach reduces reliance on pixel-level annotations while achieving high-precision alignment across multimodal imaging. Our method achieved robust performance on both public and clinical datasets, with segmentation Dice score reaching 0.903, and registration errors controlled within HD95 of 11.18 mm and TRE of 8.64 mm. Clinically, the registered images significantly enhanced senior physicians’ diagnostic confidence (confidence score: 3.032 vs. 3.548, p = 0.015) and improved lesion localization accuracy. Additional analysis showed that concordance between registered MRI and biopsy-confirmed tumor laterality increased for left-sided (from 61.5% to 69.2%) and bilateral cases (from 72.7% to 81.8%), while PI-RADS scores correlated well with pathological Gleason grades. These findings suggest that the proposed framework improves anatomical alignment, supports more accurate biopsy targeting, and holds promise for integration into prostate cancer diagnostic workflows. Future studies will focus on multicenter validation, histopathological correlation, and deployment in real-time clinical environments.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076251375870 - Supplemental material for Weakly supervised deep learning for multimodal MRI-TRUS registration: Toward assisting prostate biopsy guidance
Supplemental material, sj-docx-1-dhj-10.1177_20552076251375870 for Weakly supervised deep learning for multimodal MRI-TRUS registration: Toward assisting prostate biopsy guidance by Jiabin Yu, Binggang Xiao, Jiayi Wang, Hongmei Mi, Enyu Wang, Ningjie Huang, You Zhou, Dongping Zhang, Furong Luo, Li Yang, Hanzong Lin, Ruigang Huang, Yingying Ding, Xia Li, Chenjie Xu, Guorong Lyu and Ming Chen in DIGITAL HEALTH
Footnotes
Acknowledgments
The authors would like to express their gratitude to the ultrasound and radiology teams at Zhangzhou Affiliated Hospital of Fujian Medical University, as well as the artificial intelligence team from the Department of Artificial Intelligence at China Jiliang University, for their dedicated efforts.
ORCID iDs
Ethical approval
Ethical approval was obtained from the Independent Ethics Committee of Zhangzhou Affiliated Hospital of Fujian Medical University (Approval Number: IRB- 2024LWB332), with informed consent waived for all participants due to the retrospective nature of the study, in accordance with the ethical principles outlined in the Declaration of Helsinki. In addition, this study involved the use of two publicly available prostate MRI datasets: Promise12 and MSD Prostate. Both datasets are openly accessible for academic research and were utilized in compliance with their respective licensing terms and usage guidelines.
Contributorship
Jiabin Yu did conceptualization, methodology, and formal analysis. Binggang Xiao did methodology, supervision, and project administration. Jiayi Wang did software, validation, and investigation. Ningjie Huang did resources and data curation. Enyu Wang did software, resources, and data curation. You Zhou and Furong Luo did resources and data curation. Dongping Zhang did resources and funding acquisition. Hongmei Mi, Li Yang, Yingying Ding, and Chenjie Xu did conceptualization and investigation. Hanzong Lin, Ruigang Huang, and Xia Li did resources and data curation. Guorong Lyu did supervision and project administration. Ming Chen did methodology, supervision, and project administration.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Foundation of Fujian Province, Key R&D Projects in Hangzhou, Zhejiang Provincial Natural Science Foundation of China, Key R&D Projects in Ningbo, Key Research and Development Program of Zhejiang Province (grant numbers 2023J011826, 2024SZD1A03, LY23F010005, 2024Z114).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
Details of the challenge open data set involved in this study can be found in the link below: MR to Ultrasound Registration for Prostate Challenge (zenodo.org), Medical Segmentation Decathlon (medicaldecathlon.com), Home-PROMISE12-Grand Challenge (grand-challenge.org); for clinical data, data sharing is available upon reasonable request to the corresponding author.
Guarantor
As the corresponding author, Dr Ming Chen assumes full responsibility for the integrity of the entire study, including its design, data collection, analysis, and interpretation. He guarantees that all aspects of the work are accurate and reported with transparency.
Institutional review board statement
This study was conducted according to the guidelines of the Declaration of Helsinki and approved by the Institutional Review Board of Zhangzhou Affiliated Hospital of Fujian Medical University (protocol code 2024LWB332; approval date: 12 Sep 2024).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.