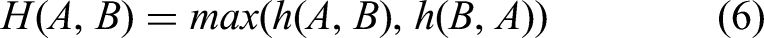Abstract
Background
Cervical spine disorders are becoming increasingly common, particularly among sedentary populations. The accurate segmentation of cervical vertebrae is critical for diagnostic and research applications. Traditional segmentation methods are limited in terms of precision and applicability across imaging modalities. The aim of this study is to develop and evaluate a fully automatic segmentation method and a user-friendly tool for detecting cervical vertebral body using a combined neural network model based on the YOLOv11 and U-Net3 + models.
Method
A dataset of X-ray and magnetic resonance imaging (MRI) images was collected, enhanced, and annotated to include 2136 X-ray images and 2184 MRI images. The proposed YOLO-UNet ensemble model was trained and compared with four other groups of image extraction models, including YOLOv11, DeepLabV3+, U-Net3 + for direct image segmentation, and the YOLO-DeepLab network. The evaluation metrics included the Dice coefficient, Hausdorff distance, intersection over union, positive predictive value, and sensitivity.
Results
The YOLO-UNet model combined the advantages of the YOLO and U-Net models and demonstrated excellent vertebral body segmentation capabilities on both X-ray and MRI datasets, which were closer to the ground truth images. Compared with other models, it achieved greater accuracy and a more accurate depiction of the vertebral body shape, demonstrated better versatility, and exhibited superior performance across all evaluation indicators.
Conclusion
The YOLO-UNet network model provided a robust and versatile solution for cervical vertebral body segmentation, demonstrating excellent accuracy and adaptability across imaging modalities on both X-ray and MRI datasets. The accompanying user-friendly tool enhanced usability, making it accessible to both clinical and research users. In this study, the challenge of large-scale medical annotation tasks was addressed, thereby reducing project costs and supporting advancements in medical information technology and clinical research.
Highlights
Cervical spondylosis is quite common, and an accurate, fully automated vertebral imaging segmentation method is needed to solve the current dilemma of extensive manual labeling.
YOLO-UNet network combined the advantages of the YOLOv11 and U-Net3 + models for automatic cervical vertebral body segmentation was proposed, which showed excellent performance and versatility for both X-ray and MRI images.
A user-friendly tool was designed for the automatic segmentation and manual optimization of cervical vertebral body, providing accessibility to both experts and non-experts.
A reliable automatic segmentation approach and the accompanying software were provided for studying cervical spondylosis and conducting clinical research.
Introduction
Headache and neck pain associated with cervical spondylosis have become increasingly prevalent in contemporary society. Clinical diagnosis and treatment frequently incorporate imaging examinations such as X-ray and magnetic resonance imaging (MRI), which are widely employed across healthcare settings, from primary clinics to tertiary referral centers. 1 X-ray imaging, as a cost-effective fundamental modality, clearly delineates the bony structures of the cervical spine, including vertebral alignment and the presence of fractures. MRI provides detailed visualization of both bony and soft tissues of the spine, such as intervertebral discs and the spinal cord, at a relatively high cost but without the radiation associated with X-rays. Through the evaluation of vertebral structures via lateral X-ray and MRI images, clinicians can assess the curvature of the cervical spine, intervertebral spaces, and related pathophysiological states, thereby determining the presence of degenerative changes or structural abnormalities and informing clinical decision-making. 2
Research indicates that the application of fully automated image segmentation technologies to medical imaging analysis can not only enhance diagnostic efficiency and reduce labor costs but also provide a more robust foundation for clinical decision-making.3,4 In many AI-driven medical scenarios, datasets are typically acquired through manual annotation and segmentation performed by expert clinicians during their limited spare time. However, as medical imaging technologies have advanced and become more refined, the size and complexity of datasets have increased exponentially, leading to greater time consumption by manual segmentation. Concurrently, the demand for large-scale models and diverse market applications continues to grow with evolving socioeconomic dynamics, and collaborating medical experts face intensified clinical workloads. Under these circumstances, relying solely on manual segmentation imposes excessive labor and time costs.5,6 Therefore, there is an urgent need for fully automatic segmentation approaches tailored to specific medical tasks, as well as for efficient and intuitive interactive tools accessible to both experts and non-experts, including information technology professionals and medical staff.
Image segmentation, a classic task within the field of computer vision, involves the extraction of target objects from images. Early methods can be broadly categorized into threshold-based segmentation, region-based segmentation, and edge-detection approaches, which are based on certain image features of the objects to be segmented and thus often exhibit limited generalizability7–9. At the turn of the century, new advancements emerged with the introduction of graph theory-based methods and clustering-based methods.10,11 In recent years, deep learning methodologies have enabled the automatic learning of features directly from images to improve segmentation performance. In 2015, Long et al. introduced a fully convolutional network to achieve end-to-end image prediction. Subsequently, Ronneberger et al. proposed U-Net, a model based on an encoder‒decoder structure, to iteratively extract and reconstruct image features through multiple layers of convolution and pooling.12,13 In 2020, Huang et al. enhanced the U-Net model by introducing full-scale skip connections, resulting in the U-Net3 + model, which demonstrates outstanding performance in medical image segmentation. 14 In 2018, Chen et al. proposed the DeepLabv3 + model based on the DeepLab model by Liang et al., which refined segmentation performance through the adoption of encoder‒decoder architectures, deep separable convolution, and an improved Xception module.15,16
Image segmentation techniques have been extensively applied in the medical field, encompassing bone structure extraction, tissue and organ segmentation, and lesion extraction. 17 For example, M. Emin et al. used convolutional neural network (CNN) technique to diagnose patients with COVID-19 from lung CT images. 18 Medical image segmentation tasks related to the spine have also been the subject of considerable research19–21. For example, Wongthawat et al. used to detect cervical spine fractures from X-ray images; M. Fatih et al. and Yohannes et al. used CNN models to classify intervertebral disc and cervical pain disease types; Wang et al. presented a lumbar vertebrae extraction method based on the U-Net model, whereas Yaseen et al. proposed a YOLOv5-based approach for identifying vertebral bodies in axial MR images. Benjdira et al. introduced a Deeplabv3 + method to extract images of spinal cords from ultrasound images.22–27 However, these studies have primarily focused on the lumbar and thoracic spine, with relatively limited attention given to the cervical vertebral bodies.
In this study, cervical vertebral body are manually labeled on X-ray and MRI images, and vertebral body extraction methods based on the YOLO, U-Net, and Deeplab models are replicated. The YOLOv11, developed by the Ultralytics team, demonstrates superior performance in terms of speed, accuracy, and adaptability. The model employs an enhanced architecture that optimizes the feature extraction network, thereby enabling the detection of objects of various scales and in complex backgrounds. U-Net3 + exhibited significant advantages in processing complex medical image data. Its full-scale skip connections have the capacity to transmit multi-scale feature information, thereby facilitating the capture of fine structures and edge details. Furthermore, it facilitates multi-task learning by integrating image segmentation with other related tasks, thereby enhancing its practical application. The integration of these two models has the potential to mitigate each other's deficiencies.14,28 YOLOv11 can provide object detection information for U-Net3+, thereby facilitating the delineation of segmentation areas and the mitigation of misdetections. We propose a combined network model based on the integration of the YOLOv11 and U-Net3 + models, which uses the YOLOv11 model for vertebral body recognition to localize vertebral structures and the U-Net3 + model for vertebra body extraction to isolate the vertebral body from the background. The resulting approach enables the accurate extraction of vertebral body from both X-ray and MRI images while ensuring high accuracy in vertebral body recognition and improved delineation of vertebral boundaries. This method provides valuable informational support for clinical diagnosis, particularly in assessing the relative positional relationships among the cervical vertebrae. Figure 1 illustrates the overall workflow.
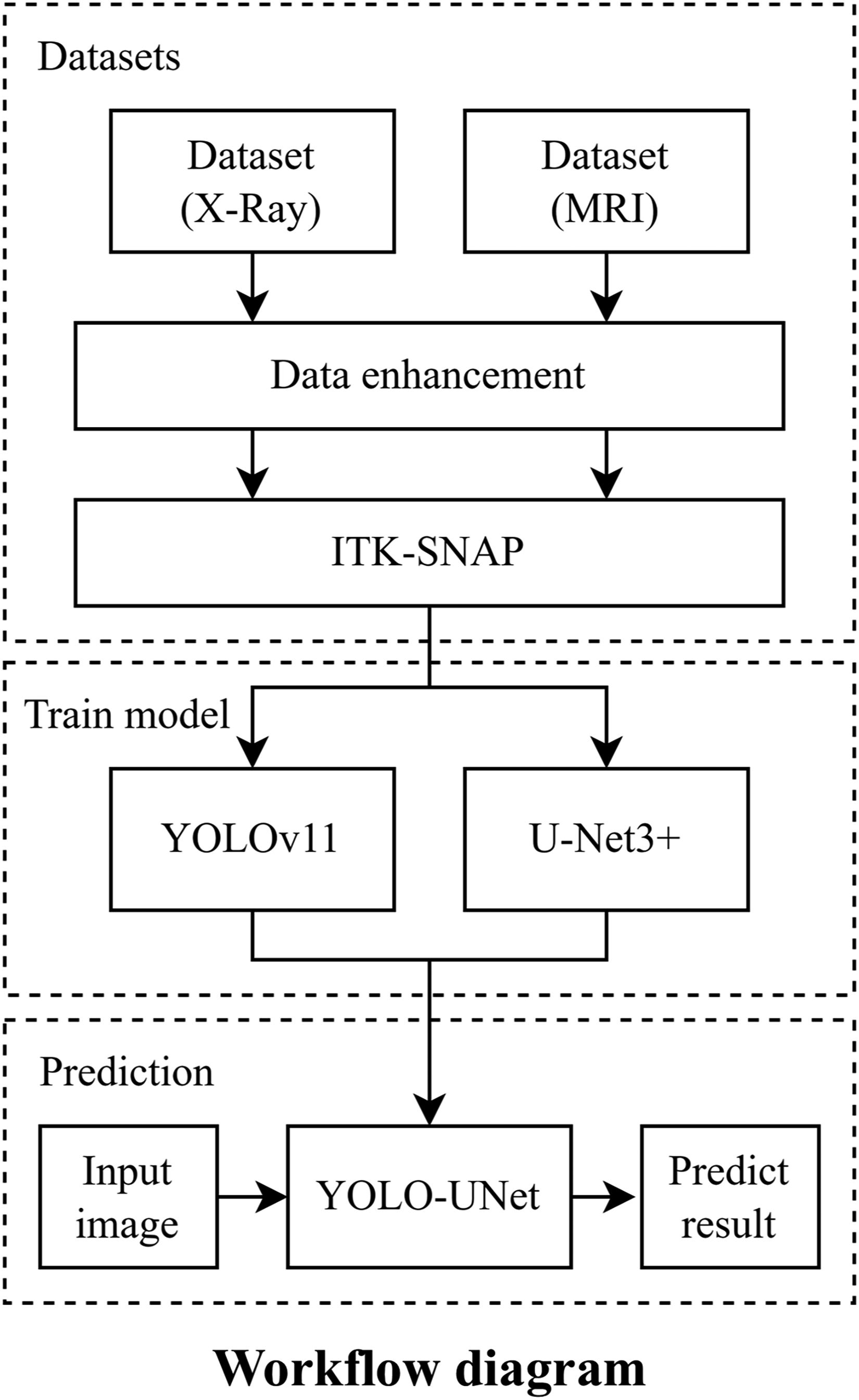
Study workflow diagram.
Materials and methods
Data sources and image preprocessing
The dataset utilized in this study was derived from cervicogenic headache research conducted by Xuanwu Hospital of Capital Medical University and was approved by the Ethics Review Board of Xuanwu Hospital Capital Medical University (KS2023212), waived the informed consent. The imaging data were obtained from the authorized electronic health records and digital imaging database. The dataset comprised cervical spine X-ray and T2-weighted MRI lateral images of 100 patients, collected between January 2022 and December 2023. The dataset included 89 X-ray cases (one image per case, totaling 89 images) and 16 MRI cases, from which 91 images were ultimately selected on the basis of the relative clarity and completeness of the vertebral structures.
Annotations for the original dataset were performed by medical experts at Xuanwu Hospital using ITK-SNAP 4.0.1. 29 The annotation process focused on lateral images of X-ray and MRI middle sequences, marking the anatomical structures of all vertebral body from C2 to C7. The atlas vertebra (C1) was excluded from the labeling task because of absent vertebral body structure. Other anatomical features, such as spinous processes, intervertebral discs, and surrounding musculature, were also omitted.
Because the deep learning algorithm model only accepts images of a specified size for processing, all images and their corresponding ground truth (GT) images were resized to a uniform pixel dimension. Additionally, the GT images were binarized to facilitate model training. Given the inherent ethical, security, privacy, and regulatory challenges associated with acquiring large-scale medical imaging datasets for deep learning applications, image enhancement techniques were utilized as a key strategy to augment data quantity. Various measures were taken to ensure the reliability of augmented images and avoid overfitting. In terms of image enhancement, a combination of diverse transformations such as rotation (± 10 °), translation (± 5 pixels), scaling (90% - 110%), and flipping are employed, while maintaining key features based on the characteristics of medical images to ensure the rationality of enhancement. For data partitioning and validation, k-fold cross-validation is used, where the data is randomly divided into k subsets. One subset is used for validation while the remaining k-1 subsets are used for training in turns, and this process is repeated multiple times to evaluate model performance. In holdout validation, an independent test set is designated for the final evaluation. Through these methods, limited data is fully utilized to monitor the model's performance under different partitions, adjust hyperparameters, and assess the model's generalization capability. This approach helps prevent overfitting to noise, ensures the reliability of augmented images, and enhances the model's reliability and adaptability. The specific effects of these augmentation techniques on the dataset are shown in Figure 2.

Schematic diagram of image augmentation techniques on dataset. Image augmentation techniques were used to expand the dataset, including random cropping, flipping, rotation, and salt-and-pepper noise addition.
Image segmentation method based on the combined network of the YOLOv11 and UNet3 + models
Vertebral body recognition module
The vertebra body recognition module was used to identify and localize the positions of the C2–C7 vertebra body within the images. This module was implemented using the YOLOv11 network, which comprises a backbone network, neck network, detection head, and other supporting components. The structure of the YOLOv11 network is illustrated in Figure 3(a). The backbone network is composed of the CBS, C3K2, SPPF and C2PSA components. CBS consists of a series of convolutional layers, batch normalization, and SiLU activation layers arranged in sequence to downsample the input data. C3K2 is an enhancement of the C2F module that integrates the C3 module into the bottleneck architecture when the C3 K parameter is set to True. The C3 module features three convolutional layers and serves as an advanced feature extraction mechanism. The SPPF component improves the spatial pyramid pooling module by replacing the original parallel max pooling module with a serial and parallel calculation method, enhancing the detection of objects of different sizes. C2PSA comprises two convolutional layers and multiple bottleneck blocks combined with a pointwise spatial attention block, which enhances feature extraction through a multihead attention mechanism and a feedforward neural network.

Architecture of YOLO-Unet vertebral segmentation network. (a) YOLOv11 network structure for vertebral detection. (b) U-Net3 + network structure for vertebral extraction.
The neck network employs the path aggregation network feature pyramid network (PAN-FPN). The PAN upsamples the input data twice to improve the extraction of semantic information, and the FPN structure fuses feature maps of different scales through bottom-up and top-down paths. This process facilitates cross-scale information transmission and enhances the model's ability to detect objects of different sizes.
The detection head, also known as the head network, uses a convolutional phase decoupling structure for detection and classification, the CIoU loss function to predict the location of the target and the BCE loss function to predict the category of the target. Formula 1 shows the calculation of CioU as follows:
The intersection area is the area of intersection between the prediction frame and the real frame, the union area is the area of union between the prediction frame and the real frame, ρ(b, bgt) and ρ(b, bgt) are the Euclidean distances between the center points of the two frames, c is the diagonal length of both the prediction frame and the real frame, a is a balancing parameter, and v is an index that measures the consistency of the shape of the prediction frame with that of the real frame. The BCE loss function is calculated using Formula 2 as follows:
Vertebral body extraction module
The vertebra body extraction module based on the U-Net3 + model is used to separate the cervical vertebra body from the background, distinguishing the foreground from the background on the basis of the predicted vertebrae position. Figure 3(b) shows the network structure of the U-Net3 + model, which consists of two main parts: an encoder and a decoder. Compared with earlier U-Net models, the U-Net3 + model incorporates full-scale skip connections and a depth supervision mechanism. Full-scale skip connections enhance segmentation accuracy and boundary clarity by enabling each decoder layer to fuse feature maps of the same size and smaller from the encoder, as well as feature maps of larger sizes from the decoder. The depth supervision mechanism enhances training stability and model performance by introducing side outputs at each decoder stage, which are supervised using the true classification results.
The loss function used in the U-Net3 + model in the experiment is BCEDiceLoss, which is a combination of BCE loss and Dice coefficient loss. α is a hyperparameter that adjusts the weight between the two losses. BCE is the binary cross-entropy loss. Dice is the reciprocal of the Dice coefficient; thus, the smaller the Dice loss is, the more accurate the prediction result. Formulas 3 and 4 show the calculation process as follows:
X represents the GT segmentation image, and Y represents the predicted segmentation image. The notation ∣X∩Y∣ represents the number of pixels in the same part of the two images, and ∣X∣ and ∣Y∣ indicate the total number of pixels in images X and Y, respectively.
Metrics
In this study, several metrics commonly used in the field of image segmentation were introduced to evaluate the performance of the YOLO-U-Net combined neural network model for segmentation. These metrics include the Dice coefficient, Hausdorff distance, intersection over union (IoU), positive predictive value (PPV), and sensitivity.30,31
The Dice coefficient is a widely used similarity metric for assessing the overlap between two samples. Its value ranges from 0 to 1, with higher values indicating greater similarity. IoU quantifies the degree of overlap between the predicted region and the GT region, which is calculated as the ratio of the intersection to the union of the two regions, as shown in Formula 5. ∣A∩B∣ represents the number of pixels marked as true in both the predicted image and the GT image, and ∣A∪B∣ represents the number of pixels marked as true in either the predicted image or the GT image.
The Hausdorff distance is a measure of similarity between two datasets of points, defined as the maximum of the two unidirectional distances. Two datasets are considered close in distance if every point in one set is close to some point in the other set. This is calculated as shown in Formulas 6 and 7, where h (A, B) is the unidirectional distance from set A to set B, defined as d (a, b), which is the distance metric between points a and b.
In addition, PPV and sensitivity represent the accuracy and recall rates of the model predictions, respectively.
Results
Experimental data
Following image enhancement, the original dataset was expanded to include 2136 X-ray images and 2184 MRI images. These images were randomly divided into training, validation, and test datasets on a patient-by-patient basis in a 6:2:2 ratio. Table 1 presents the structure of the experimental dataset.
Dataset structure and features.

MRI: magnetic resonance imaging.
Model training
During the training of the vertebral body detection model, the hyperparameter single-cls was set to True, and the early termination tolerance was configured at 20 rounds. Accuracy was used as the evaluation metric, with training terminating if the accuracy did not improve after 20 consecutive rounds. For X-ray images, early termination was triggered after 65 rounds, resulting in a maximum accuracy of 99.35%. For MRI images, early termination was triggered after 101 rounds, achieving a maximum accuracy of 98.54%. The training process of the vertebral detection model is illustrated in Figure 4(a).

Training process of YOLO-Unet vertebral segmentation network. (a) The vertebral extraction model training process. (b) The vertebral extraction model training process.
For the vertebral body extraction model, the IoU was used as the evaluation metric for early termination. For X-ray images, early termination occurred after 89 rounds, with the IoU reaching a maximum of 94.38%. For MRI images, early termination occurred after 64 rounds, with the IoU reaching a maximum of 94.47%. The training process of the vertebral extraction model is shown in Figure 4(b).
Comparison with other extraction methods
To evaluate the performance of the YOLO-UNet ensemble model, four additional image extraction models were trained and compared using the same dataset. The direct image segmentation models included YOLOv11, DeepLabV3+, and U-Net3+, as well as the YOLO-DeepLab network image segmentation model, which is composed of the YOLOv11 and DeepLabV3 + models.
As shown in Table 2, the two combined models (YOLO-UNet and YOLO-DeepLab) demonstrated higher accuracy than the three standalone models for both the X-ray and MRI datasets. The combined models also produced more precise depictions of vertebral body shapes, albeit with a slight increase in the time required for image prediction. Among the two combined models, the YOLO-Unet model outperformed the YOLO-DeepLab model across all evaluation metrics. In addition, predictions made using the test dataset of X-ray images consistently achieved better prediction results than those that used MRI images across all models.
Performance of different segmentation networks on X-ray and MRI datasets.

MRI: magnetic resonance imaging; IoU: intersection over union; PPV: positive predictive value.
A parallel comparison of the segmentation results for X-ray and MRI images is shown in Figure 5. The results highlight the superior performance of the combined models in accurately predicting vertebral body locations and edges. Additionally, the YOLO-UNet model avoided producing segmentation results unrelated to vertebrae, which were observed in the results of the DeepLabV3 + and U-Net3 + models. It also addressed the limitation of low segmentation accuracy inherent to the YOLOv11 model. Among all the models tested, the YOLO-U-Net ensemble achieved the closest resemblance to the GT images and demonstrated superior generalizability across both the X-ray and MRI datasets.

Comparative experimental results of the proposed YOLO-UNet model with other segmentation models on X-ray and MRI datasets. Shown from left to right are the input original image (X-ray/MRI), the ground truth (GT), and the corresponding predictions of YOLO, U-Net, DeepLab, YOLO-DeepLab, and YOLO-UNet (ours). MRI: magnetic resonance imaging.
Discussion
In this study, the YOLO-UNet combined network based on the YOLOv11 and U-Net3 + models was proposed for extracting cervical vertebra body from cervical lateral X-ray and MRI images. Although several neural network models in the field of deep learning can be used for medical image segmentation when trained on specialized datasets, the YOLO-UNet network presented here demonstrates superior segmentation performance by combining the strengths of both the YOLO and U-Net models.
The YOLO-UNet model combines the accuracy of the YOLO model in predicting the position of vertebra body with the accuracy of the UNet model in predicting the shape and edges of the target. Compared with the YOLO-UNet model, the YOLO model itself has difficulty precisely depicting the edge shape of the vertebra body when predicting the vertebra body and is not suitable for subsequent practical applications, such as calculating the degree of cervical lordosis using the geometric relationship between the vertebrae. The U-Net model is prone to missing, over-detecting, and mis-detecting vertebral body because of the lack of a prediction process for the position of the vertebral body, especially for MRI images with clearly distinguishable cervical vertebrae and thoracic vertebrae. Compared with the YOLO-DeepLab model, the YOLO-UNet model can more accurately delineate vertebral boundaries, which combines the advantages of both the YOLO and U-Net models, achieving excellent and robust segmentation performance across both X-ray and MRI datasets and delivering results closely aligned with GT images.
With the advancement of information technology and the popularization of mobile devices, the population affected by cervical spine-related diseases is becoming younger, with more widespread and occupation-related occurrences. This trend, particularly among computer workers, adolescents, and students with sedentary lifestyles, has resulted in a growing economic burden and societal labor loss. To address the need for large-scale research on the cervical spine, we have designed an interactive tool based on the proposed YOLO-U-Net model. This tool enables users to apply the pretrained model for automatic segmentation of cervical vertebral body in lateral cervical spine images through desktop software. The software is compatible with the Windows environment and supports X-ray and/or MRI images in DICOM formats. When the software is used, images can be imported; it automatically reads the image information, performs vertebral body segmentation, displays the results on the interactive interface, and provides tools for the manual refinement of the segmentation predictions. If users are satisfied with the result, the data can be exported in two formats: a labeled image showing segmented vertebral body alone and an annotated image merged with the original input. This functionality meets the diverse needs of both expert and non-expert users, including computer professionals and healthcare staff. The software package and detailed user manual instructions are provided in the Supplementary Material, and the user interface is illustrated in Figure 6. This tool can also serve as a precursor for large-scale AI-driven medical tasks, such as clinical decision-making and cervical spine curvature calculation for the diagnosis of spinal-related diseases.
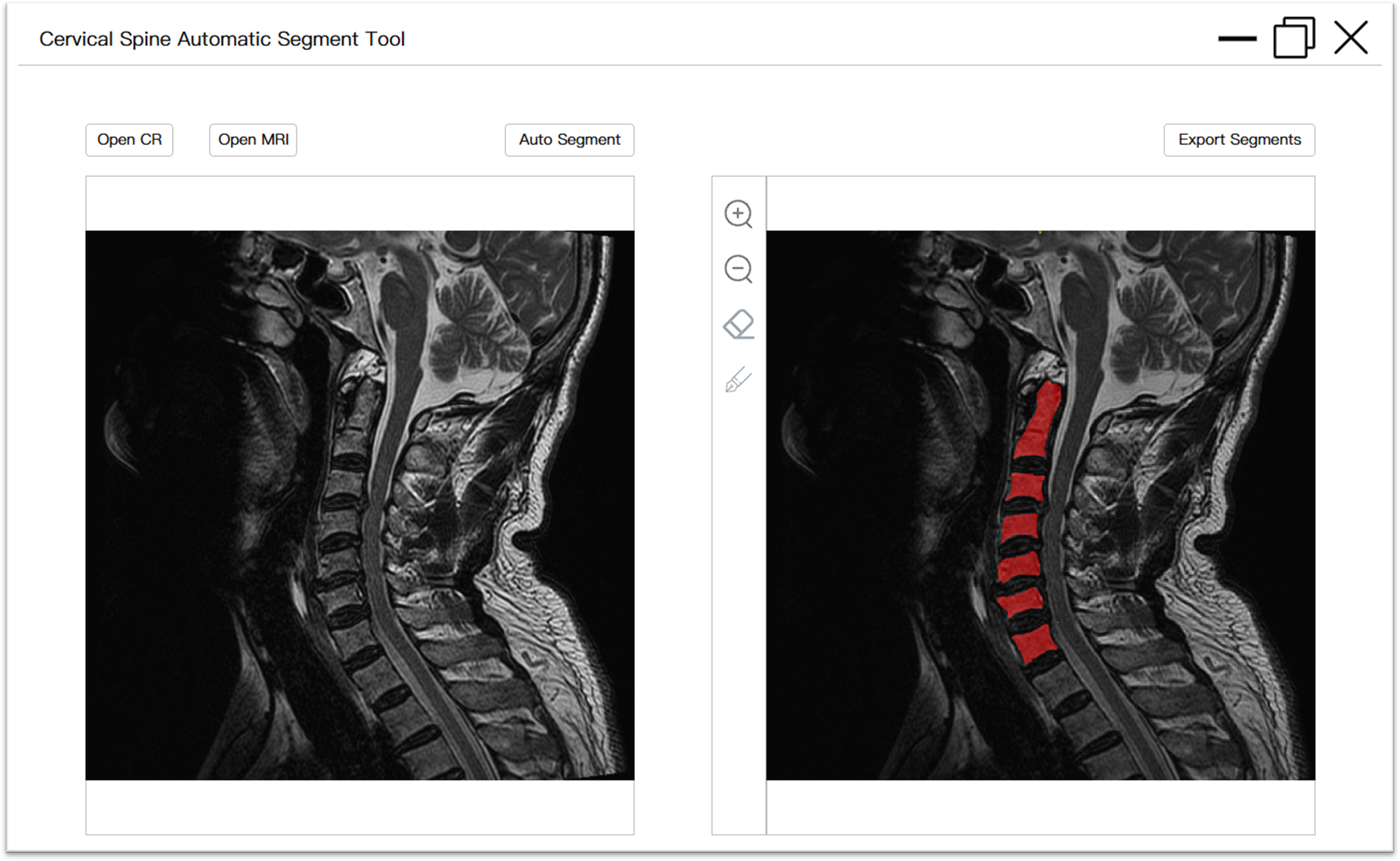
User interface of interactive tool for cervical vertebrae automatic segmentation. The image on the left depicted the original image that was imported, while the image on the right illustrated a preview of the segmentation result with annotations (which can be modified). Furthermore, there were interactive buttons, including input, auto-segment, zoom, manual refinement, and export.
This study had certain limitations. In some cases, the YOLO-U-Net model exhibited errors in vertebral localization, including missed or multiple identifications. Further image analysis revealed that in some cases, the C7 vertebral body was occluded by the clavicle or other tissues in X-ray images, or the first thoracic vertebra (T1) was misidentified as a cervical vertebra in MRI images. These issues may be due to the binary labeling approach used during image preprocessing, where all vertebral bodies are uniformly labeled “vertebrae” limiting the model's ability to account for intervertebral relationships; the limited number of patient images used for model training is also a likely influencing factor. In future research, it will be necessary to improve the accuracy of predicting the positions of vertebrae, such as by refining the cervical vertebrae into a multiclassification task for each vertebra (C2–C7) to improve the model's ability to recognize the relationships between vertebrae. On the other hand, based on verifying the feasibility of the method presented in this paper, it can be improved by cervical spine image of more patients and incorporating a wider variety of medical imaging data to enhance its versatility in different environments and meet the needs of more application scenarios.
Conclusion
In this paper, a novel fully automatic segmentation method for cervical spine images that combines the YOLOv11 and U-Net3 + models for vertebral body segmentation is presented. The experimental results demonstrate that this approach outperforms traditional medical image segmentation methods in extracting cervical vertebrae, exhibiting higher accuracy and greater versatility across both X-ray and MRI images. Additionally, an executable interactive program has been developed to facilitate its application. This tool is designed for use by both professionals and nonprofessionals, enabling the automated extraction of vertebrae from lateral cervical spine images. By providing a user-friendly and efficient image processing solution, more research related to cervical vertebral body can be conducted, such as cervical curvature. And the challenge of large-scale medical annotation tasks was addressed, more patient image data can be collected, and the approach can be improved with a larger sample set to support future research topics in clinical decision-making and diagnostic workflows.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076251347695 - Supplemental material for Cervical vertebral body segmentation in X-ray and magnetic resonance imaging based on YOLO-UNet: Automatic segmentation approach and available tool
Supplemental material, sj-docx-1-dhj-10.1177_20552076251347695 for Cervical vertebral body segmentation in X-ray and magnetic resonance imaging based on YOLO-UNet: Automatic segmentation approach and available tool by Hongyan Wang, Jie Lu, Song Yang, Yin Xiao, Liangliang He, Zhi Dou, Wenxing Zhao and Liqiang Yang in DIGITAL HEALTH
Footnotes
Acknowledgements
Not applicable.
Ethical considerations
This study was approved by the Ethics review board of Xuanwu Hospital Capital Medical University (KS2023212).
Informed consent statement
Waived informed consent for approval.
Author contributions
HW contributed to conceptualization, methodology, formal analysis, writing—original draft. JL contributed to formal analysis and writing—review and editing. SY contributed to software, investigation, and visualization. YX contributed to software and methodology. LH contributed to supervision and validation. ZD contributed to data curation and validation. WZ contributed to supervision and methodology. LY contributed to project administration, funding acquisition, and resources. HW, JL, and SY contributed equally to this work as co-first author.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study is financially supported in part by Capital's Funds for Health Improvement and Research (CFH2024-2-20111), National Health Commission Capacity Building and Continuing Education Center (PMT1003-3) and Beijing Municipal Administration of Hospital Incubating Program (PX2024035).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.