Abstract
Stroke is a leading cause of mortality and disability worldwide, requiring early detection and timely intervention to improve patient outcomes. However, in resource-limited locations, the lack of specialists often leads to delayed and inaccurate diagnoses. To address this, we propose an AI-driven stroke identification and treatment system that integrates expert knowledge with machine learning, enabling healthcare providers to make informed decisions without direct specialist input. The data for this study were obtained from Debre Berhan Referral Hospital through expert interviews, prescriptions, and from a public dataset in the Kaggle platform. Feature selection was performed using decision trees, Chi-Square tests, Elastic Net coefficients, and correlation analysis. Additionally, we applied to Shapley Additive Explanations to demonstrate the feasibility of feature selection in AI model development. Machine learning models, including Decision Tree, Random Forest, and Support Vector Machine, were evaluated, and Random Forest classifier achieved the highest accuracy of 99.4% using k-fold cross-validation technique. Expert knowledge was encoded in Prolog, while machine learning models were implemented in Python to develop a hybrid expert system. Medical professionals evaluated the system, confirming its effectiveness as a decision-support tool for stroke diagnosis and treatment. This approach demonstrates the potential of AI-driven expert systems to enhance stroke management, particularly in regions with limited access to specialized care.
Introduction
Stroke is a noncommunicable illness that plagues all world populations, regardless of their development. 1 It most often harms the brain through blood vessels that burst, so medical attention must be immediately sought to stop the damage from reaching a chronic state or death. Stroke is a leading cause of disease burden in the world: 12.2 million new cases and 101 million current cases per year, according to the 2019 Global Burden of Diseases, Injuries, and Risk Factors Study (GBD). 2 It causes 143 million disability-adjusted life years (DALYs) to be lost and 6.5 million deaths—rendering stroke second in death worldwide, after ischemic heart disease. The rising prevalence of stroke is due to population ageing and factors with controllable risk factors such as high blood pressure, diabetes, poor diet, obesity, smoking, air pollution, alcohol, high cholesterol, and lack of exercise. Strokes disproportionately kill men in low-income countries, accounting for 86.0% of deaths and 89.0% of DALYs. 3
In reports issued by the American Heart Association and the National Institutes of Health, 4 the burden of heart disease and stroke is made even more prominent globally. These combined kills more people every year than cancer and lower respiratory conditions combined. Cardiovascular disease caused 33% of global mortality in 2019, attributed to ischemic heart disease and stroke for 85% of deaths. Although the stroke mortality rate is down from 354.5 stroke deaths per 100,000 in 1990 to 239.9 strokes per 100,000 in 2019, 5 it is still high. Stroke occurs in the United States alone approximately every 40s, and more than 795,000 people are diagnosed each year. 6
More than 90% of strokes are caused by risk factors that can be modified. Feigin et al. 7 suggest that good prevention would make a world of difference. These include a healthy lifestyle, body mass index (BMI) control, and heart and kidney health. Diagnosis and treatment must be performed early if we want to minimize deaths and disability. However, the traditional diagnostic approach depends so much on clinical expertise that it can be subject to bias and variability. The application of machine learning in healthcare is already powerful, and we have found new ways to identify and anticipate strokes through the application of expertise. 8 Algorithms for predictions support clinical decisions, aid diagnosis, and increase patient care by delivering knowledge that had previously been unavailable. In the same vein, knowledge-based systems (KBS) make use of AI to solve tough medical issues. By aggregating different knowledge sources, such systems support decision-making and learning, which can help to drive more effective operational efficiency, cost reduction, and quality of care.9,10
These Clinical Decision Support Systems (CDS) offer the modern-day answer to medical diagnosis and care. With the help of institutions such as the Office of the National Coordinator for Health Information Technology, 11 CDS systems improve healthcare quality, safety, and efficiency by combining AI and automation. Knowledge-based systems and machine learning as artificial intelligence subdomains can make good decisions by mimicking the actions of human professionals working in fields through advanced algorithms. 12
Several aspects of stroke research have been looked at, including socioeconomic characteristics, prevalence, incidence, and expert-systematic prevention and treatment.13–15 However, there are still major holes to fill in the puzzle of linking computer technology and artificial intelligence to solve the process of learning and to explain clinical surveillance of automated machines. This research aims to propose a system of knowledge-based stroke diagnosis and treatment using domain knowledge and machine learning models. The goal here is to build a comprehensive system to improve diagnostic quality and help clinicians make better choices. Combining computers with AI creates objective, effective, and cost-effective options for reducing morbidity and mortality. The main contribution of this work includes the following:
Integration of Expert Knowledge and Machine Learning: The research filled the gap by developing a hybrid system with the integration of domain knowledge and machine learning. Improved Model Performance: Feature selections and k-fold optimizer were applied to improve machine learning performances. Use of Diverse Datasets: Data from both expert interviews, prescription archives, and publicly available datasets were combined to enhance the system's applicability and robustness. Multimodal Data Processing: The research applied word embedding techniques to integrate numerical and textual data, enabling efficient and precise classification tasks.
This work is organized as follows: the literature section reviews the related literature; the methodology section presents the proposed approach; the results section reports the findings; the discussions section provides a discussion of the results; the limitations of the study section outlines the limitations of the study; and the final section concludes the paper and suggests future research directions.
Literature review
In Kleindorfer et al., 16 medical knowledge (clinical guidelines, expert opinions, and evidence) was established to get a powerful knowledge base. This synthesis ensures that the system reflects evidence and domain knowledge. Siala et al. 17 point to the power of hybrid systems integrating knowledge logic and machine learning algorithms. These systems have shown great promise for diagnosing stroke more accurately and planning for treatment, combining the best of both worlds. Zhang et al. 18 conducted a study on the automatic diagnosis and prevention of ischemic strokes. The researchers collected 5668 brain MRI scans randomly selected between 2017 and 2019, along with clinical imaging reports from 300 cases. Professional neurologists meticulously labeled all lesion areas. To automatically detect lesions in MRI images, three object detection networks were designed and implemented: Faster R-CNN, You Look Once V3 (YOLOv3), and SSD. The study found that lesion detection accuracy (mAP) was comparable for Visual Geometry Group (VGG-16), Residual CNNs (ResNet-101), and YOLOv3, each achieving 74.9%. However, SSD outperformed the other models, achieving an accuracy of 89.77%. While experts were utilized solely for labeling MRI scans, their in-depth knowledge of ischemic stroke could have been crucial for enhancing diagnostic and preventive capabilities.
Tazin et al. 19 evaluated the efficiency of various machine learning algorithms in accurately predicting strokes based on physiological variables. The study employed a stroke prediction dataset from Kaggle, consisting of 5110 patient records with 12 attributes. The findings revealed that the random forest classifier outperformed other methods, achieving a classification accuracy of 96% using cross-validation metrics. However, this study focused exclusively on evaluating machine learning algorithms, without integrating domain expert knowledge. Such expertise could enhance stroke prediction accuracy and improve diagnostic and treatment planning. Thammaboosadee and Kansadub 20 proposed a stroke risk prediction model utilizing three datasets: demographic data, medical screening data, and their combined application. The model employed three classification algorithms—Naive Bayes, Decision Tree, and Artificial Neural Network (ANN)—and was evaluated using metrics such as accuracy, Area Under Curve (AUC), False Positive Rate (FPR), False Negative Rate (FNR), and 10-fold cross-validation. The ANN model with integrated data demonstrated the best performance, achieving an accuracy of 84%, an FPR of 12%, an FNR of 25%, and an AUC of 90%. While this study successfully incorporated multiple datasets for stroke risk prediction, it did not account for domain expert insights, which are critical for enhancing the interpretability and relevance of such models.
Tavares et al. 21 used machine learning for stroke prediction models. The authors used a Kaggle dataset with 5110 records and 10 features. The dataset was divided into training data for model development and testing data for performance evaluation. The model with the highest performance was the random forest algorithm (96%), with an accuracy of 92%, F-score of 92%, and AUC of 96%. While this work proved the utility of machine learning for stroke prediction, gaining knowledge about the subject matter by having it shared by domain experts could be used to enhance health systems’ efficiency and effectiveness.
Among the studies reviewed, there have been some impressive advances in stroke diagnosis and prediction via machine learning. But one weakness of these models that we see common to them is that they do not sufficiently integrate domain expert knowledge which would make the models more comprehensible, precise, and robust. Furthermore, there are no robust data preprocessing tools like parameter tuning, feature selection, and embedding which limits the ability of these models to apply broadly to many datasets.
In this work, to bridge these gaps, we propose a system for stroke diagnosis and treatment that will combine machine learning prediction models with subject matter knowledge. It also tries to use sophisticated data preprocessing to enhance the model's performance. This methodology is designed to empower clinicians to make more informed and consistent clinical decisions for the benefit of patients and healthcare.
Methodology
This study employs the Design Science Research Methodology (DSRM), 22 an iterative approach focused on creating and evaluating artifacts such as models, frameworks, and systems to address real-world problems. The primary aim is to develop innovative solutions that advance both theoretical understanding and practical applications. In this research, health workers from Debre Birhan Referral Hospital (DBRH) were actively involved in data preparation and system performance evaluations, and the knowledge base was evaluated through experiments and parameter adjustments to ensure its robustness and applicability.
Design science research process model
This study adopts the DSRM Process Model proposed by van der Merwe et al. 23 This model comprises the following initiative stages: problem identification, setting objectives, artifact design and development, demonstrating, evaluating, and reflection. This iterative process not only ensures that the artifact meets practical requirements but also contributes to advancing existing knowledge in the domain of healthcare technologies.
At the design and development stage, an artifact—framework, prototype, or tangible solution—is created based on existing knowledge and theoretical concepts to address the identified problem and fulfill the defined requirements. For this study, primary data was collected from DBRH, where stroke patients receive medical services. These primary data sources were integral to the implementation of the knowledge base, as they involved direct input from domain experts to support the development of predictive models and the selection of the optimal model for the KBS. Besides, secondary data was retrieved from the Kaggle healthcare dataset 24 and relevant documents, including clinical guidelines and medical reports. The study employed multiple data collection methods, such as interviews, questionnaires, observations, document analysis, and knowledge discovery techniques. These methods facilitated the acquisition of domain knowledge and machine learning data, supported by literature reviews and curated datasets.
System framework and data preprocessing
Figure 1 shows the proposed system architecture for stroke prediction and treatment integrates machine learning-based predictive modeling with expert domain knowledge to deliver a comprehensive, user-friendly solution. The system involves three main tasks: dataset knowledge extraction, domain expert knowledge extraction, and a knowledge-based system integrating from machine learning and expert knowledge.

Framework for proposed system.
The system begins with data acquisition from a healthcare database of DBRH (located at latitude 9.67858° and longitude 39.5351°), which includes patient medical records, clinical data, and other relevant features. This raw data undergoes a rigorous preprocessing stage to ensure quality, consistency, and readiness for model training. These are detecting outliers, removing noisy values, handling missing values, normalization of the attribute value, and balancing sample size for each corresponding class label. In this study, instances that have extremely high or extremely low values detected as having outliers were rejected from the dataset. The logical step for detecting outliers is using the Inter Quantile Range (IQR), which is mathematically represented in Equation (1) and Algorithm 1:
where Q1 and Q3 are the first quartile (
The preprocessed dataset was divided into training and testing subsets in various ratios, with the optimal split determined to be 80% for training and 20% for testing. The training dataset is utilized to develop a predictive algorithm, which leverages a pretrained model as a foundational framework for enhanced performance and efficiency. The validation dataset aids in fine-tuning the algorithm, while the testing dataset evaluates the generalizability and robustness of the resulting predictive model. The final stroke prediction model is then capable of accurately assessing patient stroke risks.
Simultaneously, the system incorporates qualitative knowledge derived from interviews and document analysis, such as clinical guidelines and expert opinions. This information is represented and modeled into structured domain rules, ensuring alignment with established medical principles. The domain rules are fused with the machine learning model to create a hybrid system that combines data-driven predictions with rule-based decision-making. The outcome is a user-friendly treatment system that not only predicts stroke risk but also provides actionable, evidence-based recommendations for healthcare practitioners. This fusion of predictive analytics and expert knowledge ensures the system's utility in real-world clinical settings by enhancing its interpretability, reliability, and usability.
Inter quantile range (IQR) outliers detecting algorithm.

where
In the noise-removing stage, the QcleanNOISE algorithm was implemented to identify noisy data, as illustrated for the ith instance in Equation (2):
Encoding and embedding to convert categorical data into numerical representations. One-Hot Encoding technique is one of the common, which is used to represent the categorical variable in binary vectors. However, this technique is not effective for complex and larger datasets. Besides, it causes a “curse of dimensionality,” making the dataset more sparse and harder to work with, potentially slowing down learning algorithms and requiring more memory. To fill these gaps, we applied the word embedding technique, which is used to represent complex data types, such as images, text, or audio, in a way that machine learning algorithms can easily process.
26
This embedding improves the performance of machine learning algorithms on text data by capturing the meaning and context of words in a way that is more accurate than traditional bag-of-words representations, better handling vocabulary words, provides more interpretable results than traditional bag-of-words representations by showing the relationships between words in a visual space, and efficient computation. The technique converts words as vectors in a low-dimensional space (Word2Vec). Word2Vec works by representing the data precisely in the embedding space with a larger dataset by the Continuous Bag-of-Words Model (CBOW). The CBOW model takes n words before and after the target word (

Synthetic Minority Over-sampling Technique (SMOTE) is used to balance a significant difference in the dataset class distribution. 27 The technique emphasizes the importance of domain problem understanding in healthcare system planning and strategies, particularly in the construction of stroke treatment and diagnosis system, as it aids in understanding the required dataset. The SMOTE is shown in Algorithm 2.
SMOTE algorithm.

In this research, domain experts were involved and observing the existing working processes, leading to easily extracting knowledge from experts and written documents. The public datasets found at Kaggle were used as reference to check and understand experimental data structures. In this work, we identified stroke treatment and diagnosis determinant attributes as illustrated in Table 1.
Selected attributes from the business domain.

During feature selection for model developments, we made a correlation between the target variable and other features. Besides, we computed the multicorrelation between the variables to know the highly determinant variable and reduce redundancy. Although the tendency of correlations varies, most of the features have strong correlations, as shown in Figure 2. It can be observed from the heatmap that age, hypertension, and heart disease are more influencing features to predict stroke than BMI or average glucose. However, the overall correlations are not that strong, and complex models will require machine learning-to capture nonlinear patterns for stroke prediction.

Correlations of variables.
For feature selection, we utilized multiple techniques to identify high-relevance features with minimal bias, as shown Figure 3. Specifically, Figure 3(a) integrates the Chi-Square test, Elastic Net coefficients, and correlation coefficients in a collaborative manner. These methods consistently identified age, glucose level, and BMI as the most significant predictors of stroke disease. Figure 3(b) presents the feature importance derived from the Random Forest model, showing that the importance ranking of glucose level and age has shifted. However, the top three most influential features remain largely consistent across both figures. The chart illustrates the relative importance of features, with age emerging as the most influential factor, followed by glucose level and BMI. This ranking is based on Shapley Additive Explanations (SHAP), 28 as depicted in Figure 4, and is further supported by the correlation matrix presented in Figure 6.
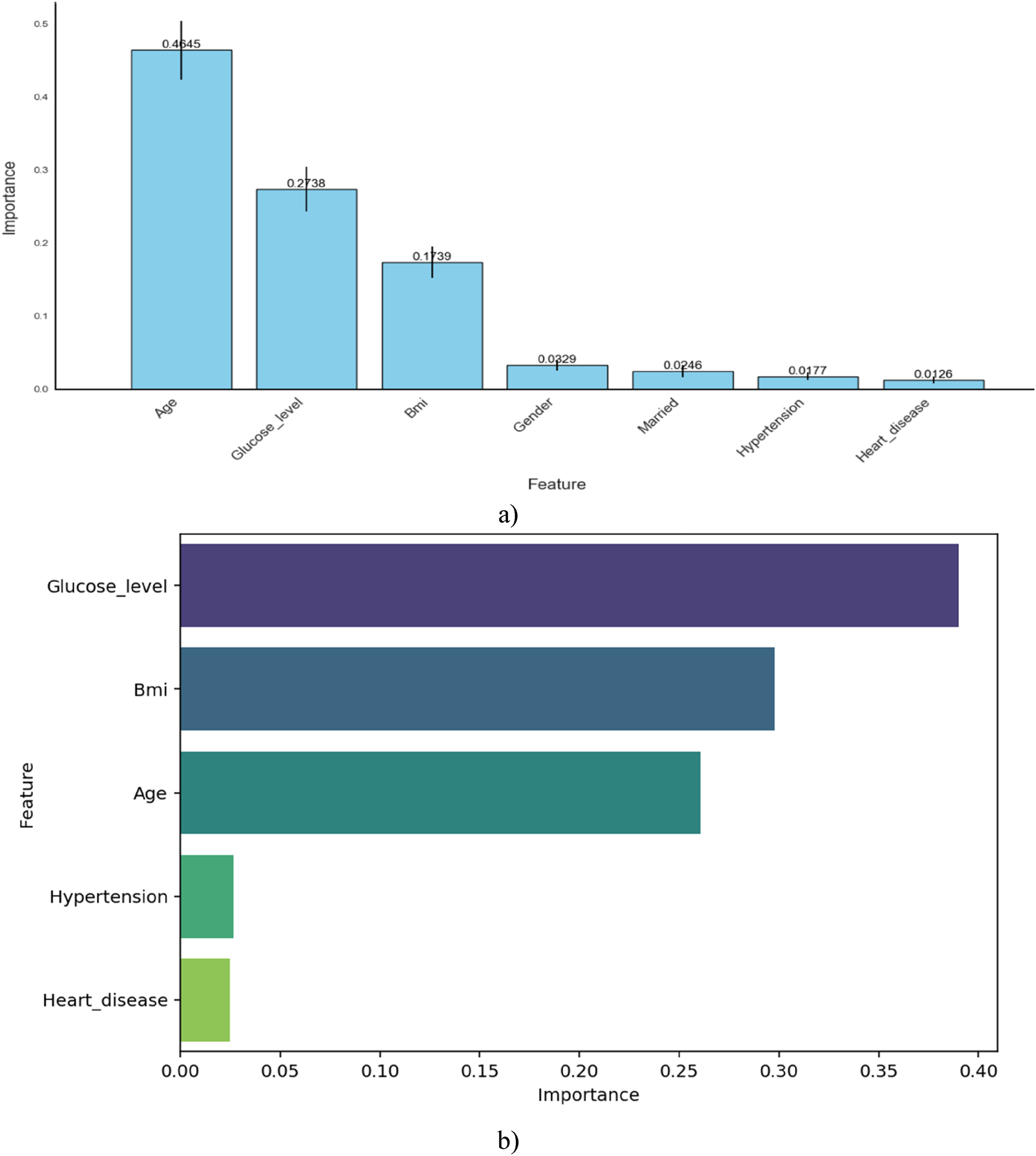
Feature importance: (a). (decision tree, Chi-Square, Elastic Net coefficients, and correlation Coefficient), (b). Random Forest-based feature importance.

SHAP dependance plots: (a) Effect of age on stroke prediction, (b) effect of glucose level on stroke prediction, (c). SHAP for glucos level. SHAP: Shapley Additive Explanations.
Figure 4 presents two SHAP dependance plots that illustrate how age and glucose level influence stroke risk predictions. To improve the interpretability of the machine learning models, SHAP—a technique for evaluating feature importance—was employed.29–31 In Figure 4(a), age has minimal impact on stroke risk before 40, but the SHAP values rise steadily after age 60, indicating a significant increase in stroke risk with ageing. Additionally, the color gradient shows that higher glucose levels (reddish dots) often appear at older ages and align with higher SHAP values, suggesting a combined effect of age and glucose on stroke risk. Figure 4(b) focuses on glucose levels, revealing that glucose levels below 120 mg/dL generally have little to no impact on stroke prediction. However, as glucose levels increase—especially beyond 150 mg/dL—the SHAP values tend to rise, indicating a greater contribution to stroke risk. Notably, many of these higher glucose cases are associated with older individuals (yellow dots), reinforcing the interaction between high glucose and age in increasing stroke risk.
Figure 4(c) shows a SHAP dependance plot for the feature Glucose_level, with data points colored by BMI, in the context of stroke disease prediction. This visualization illustrates how Glucose_level influences the model's output (i.e., the likelihood of stroke), as indicated by the SHAP values. Each point represents an individual prediction, where the color gradient reflects the associated BMI—blue for lower BMI and red for higher BMI. From the plot, it is evident that higher glucose levels (especially above ∼150 mg/dL) contribute more positively to stroke risk prediction (i.e., higher SHAP values), implying a stronger association with stroke likelihood. Additionally, the color variation suggests that BMI may interact with Glucose_level, further modulating the model's prediction. Overall, these plots highlight that both age and glucose level are important risk factors, with their combined effect playing a critical role in stroke prediction.
Domain expert knowledge extraction
Expert interviews, reviewing clinical guidelines and research, medical records analysis, collaboration with stroke centers, and observation was made for knowledge-acquisition methods, and then domain experts were involved to extract knowledge that was used for identifying stroke disease and preparing a rule. This extracted knowledge from text data is modeled by decision tree modeling for analyzing patient data, provides accurate recommendations, and guides treatment decisions. The rule-based systems technique was used for the representation of expert knowledge. These techniques were used by “IF-THEN” rules to describe conditions and corresponding actions, while decision trees represent knowledge in a tree-like structure, as shown in Figure 5. These techniques help capture symptoms, risk factors, and medical guidelines, determining stroke likelihood and suggesting appropriate treatment options. The rule was written using SWI-Prolog and Sublime Text as tools for creating a knowledge base sourced from experts, supplemented by a graphical user interface (GUI). Python was utilized for implementing classifier algorithms for machine learning predictions. Flask, a Python-based micro framework, is employed for back-end development, facilitating the creation of applications and websites with HTML integration. Access to the knowledge base is provided through SWI-Prolog and Sublime Text editor interfaces. To map knowledge into a knowledge-based system, we employed machine learning methods and expertise from domain specialists for stroke diagnosis and treatment. It amalgamates insights acquired from both experts and machine learning techniques, leveraging this combined knowledge to construct the system. A framework is devised to seamlessly integrate the extracted knowledge into the knowledge-based system. This framework reads predictive models from machine learning and expert knowledge, subsequently generating Prolog rules for system implementation.

Decision tree for stroke treatment acquired from domain expert.
Model selection experiments
A total of three experiments were conducted using classification machine learning techniques to derive accurate and reliable stroke diagnosis and treatment, as shown in Table 2. For this research, we collected 5110 records from fieldwork, and 4610 from publicly available sources Kaggle from Tazin et al. 19 The methodology involved preprocessing with 9720 records that split into training (7776) and testing (1944) sets with three conventional machine learning algorithms. Experiments are conducted with three scenarios, one with all attributes and K-fold, the second using selected attributes with K-fold, and the third is with selected attributes without applying K-fold.
Experiments on different features and optimizers.

The researcher compiled the extracted knowledge from domain experts into a set of rules using production rule methodology, utilizing Prolog programming language with SWI-Prolog 7.6.4 open-source software. These rules were then converted into a Python-compatible format for integration with rules derived from the random forest classifier model. The random forest algorithm, alongside decision tree rule induction from domain experts, generated multiple rules crucial for developing a knowledge-based system. For implementation, Python programming language with Anaconda distribution's Jupiter Notebook was employed to construct the rule base module. Additionally, HTML, created using the Sublime Text editor, was utilized for developing the GUI. The connection between the knowledge base constructed in Python and the HTML GUI was established by incorporating the PySWIP file into the Python library, acting as an interface between Python and Prolog.
Results
Model performance evaluation
Three machine learning models—Decision Tree, Random Forest, and Support Vector Machine (SVM)—were evaluated using selected feature configurations and K-fold cross-validation techniques (Table 3). The models were assessed using five key metrics: precision, recall, F1-score, accuracy, and macro-average F1-score.
Performance metrics for decision tree, random forest, and SVM models.
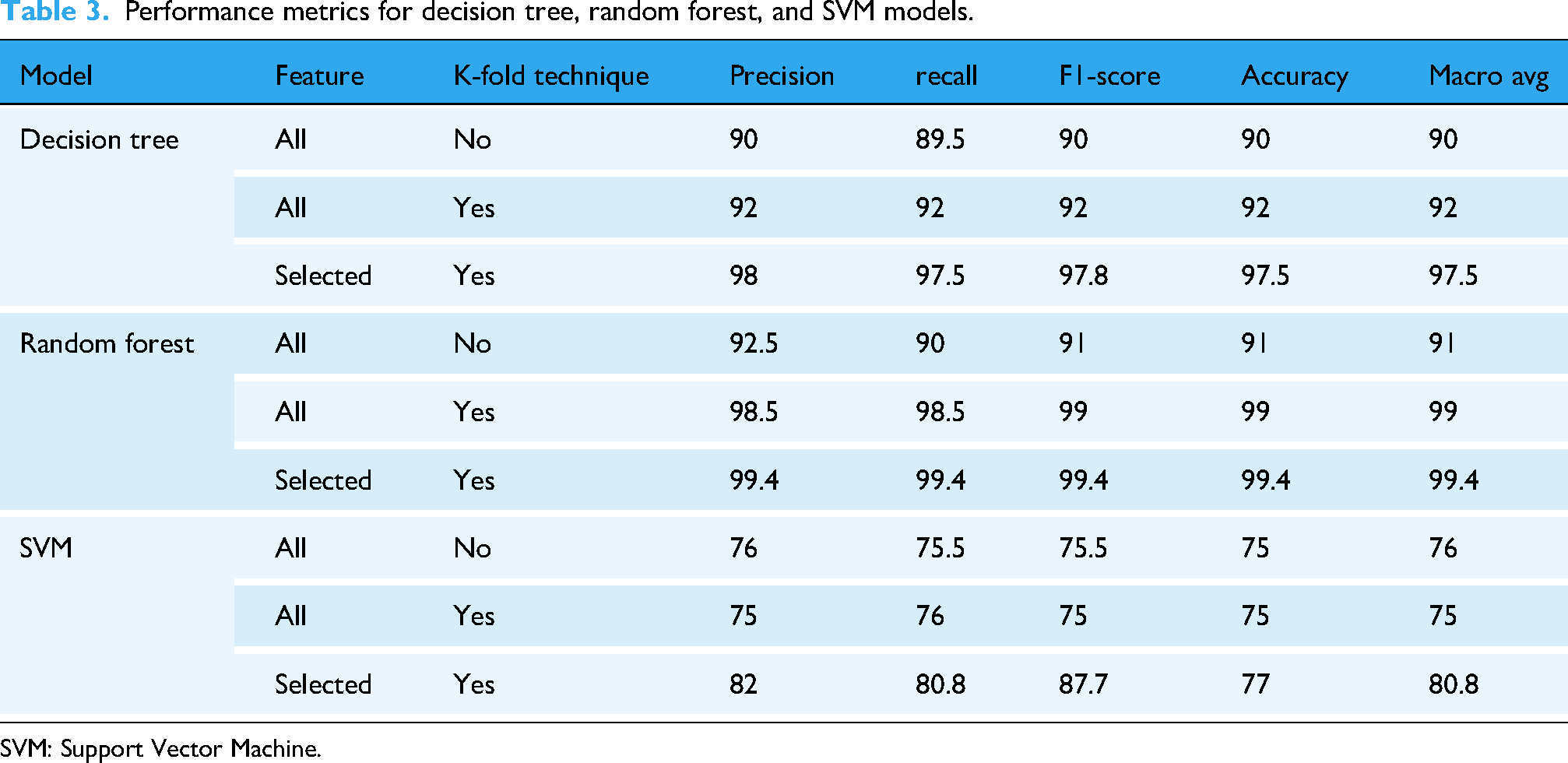
SVM: Support Vector Machine.
The Random Forest model demonstrated superior performance across all metrics. With selected features and K-fold validation, it achieved 99.4% for precision, recall, F1-score, accuracy, and macro-average score. The Decision Tree model also showed strong results with an accuracy of 97.5% and an F1-score of 97.8% under the same conditions. Conversely, the SVM model achieved lower performance, with a maximum of 77% accuracy, despite showing improvement with feature selection and validation.
Figure 6(a) illustrates the confusion matrix for the Decision Tree model after feature selection and applying the K-fold optimization technique. The results highlight the importance of optimizing both data and algorithms for accurate predictions. Out of 1944 testing samples, only 45 items were misclassified, demonstrating the model's reasonable performance. In comparison, Figure 6(b) shows the performance of the Random Forest model, which significantly outperforms the Decision Tree. The Random Forest model misclassified only 16 items from the entire testing dataset, underscoring its superior accuracy and robustness in diagnosing stroke disease. This result reinforces Random Forest as a reliable machine learning technique for medical applications. Conversely, Figure 6(c) presents the performance of the SVM. The confusion matrix clearly indicates that SVM is less stable and performs poorly compared to the Decision Tree and Random Forest models.

Model performances. (a) Confusion matrix of Decision Tree Model, (b) Confusion matrix of Random Forest Model, (c) Confusion matrix of SVM Model. SVM: Support Vector Machine.
Figure 7 shows the association the effect of age and glucose level separately as well as when the combine. Figure 7(a) demonstrates a clear positive association between age and stroke probability, with risk increasing as age advances. While both linear and quadratic models show this trend, the quadratic model more accurately captures the nonlinear escalation in stroke probability, particularly after age 50. Figure 7(b) shows the risk of increasing glucose levels rising in both models. Stroke probability remains low at normal glucose levels but increases more rapidly beyond 150 mg/dL, indicating that elevated glucose levels may contribute to an accelerated stroke risk. Figure 7(c) is a 3D surface plot illustrates the combined effect of age and glucose level on the predicted probability of stroke. A clear nonlinear interaction is observed between the two predictors. Stroke probability remains low across most of the surface for younger individuals with normal glucose levels, as indicated by the flat, dark purple area. However, stroke probability increases substantially when both age and glucose levels are high. Notably, the model captures the gradual increase in risk with age and a sharper rise with increasing glucose, especially past 150 mg/dL.

Associations between predictor and dependent variables. (a). Age and stroke association. (b). Glucose level and stroke associations. (c). predicted stroke probability by age and glucose level.
Figure 8 presents precision-recall and ROC curves. The Random Forest achieved an AUC of 1.00, indicating perfect classification capability, while the Decision Tree followed closely with 0.99. SVM, however, recorded a lower AUC of 0.80, reflecting its reduced ability to distinguish stroke cases.

Performance evaluation of different models. (a). Precision-recall graph, (b). True Positive–False positive graph.
Comparison with previous works
Table 4 shows a comparison performance highlighting several of the algorithms applied in previous literature works and their respective accuracies. The proposed system, based on Random Forest with optimization, outperforms earlier works, including: Cheon et al.: 32 DNN—84.03, Zhang et al.: 18 SSD—89.77%, Tazin et al.: 19 RF—96%, Almadani and Alshammari: 33 C4.5–95.25%, and Chun et al.: 34 Ensemble—80%. However, the proposed system achieved 99.4% accuracy, demonstrating a performance improvement of over 3% compared to the best results in prior studies.
Performance comparison with previous studies.

Usability testing of the knowledge-based system
The proposed knowledge-based system was evaluated for usability, functionality, and acceptability through user acceptance testing conducted with five health professionals (two neurologists and three general practitioners) at DBRH. The evaluation was carried out using a structured questionnaire based on the ResQue (Recommender Systems Quality of User Experience) model, which assessed various aspects such as interface usability, knowledge adequacy, decision accuracy, and overall system functionality. Before the evaluation, participants received training on the system's operation and were then presented with a set of test cases. The evaluation results, summarized in Table 5, indicated strong system performance.
Expert evaluations for treatment system.

The prototype system achieved an accuracy of 95%, while domain experts rated its performance at 88%. These results underscore the system's technical precision and its perceived effectiveness in practical applications.
Discussions
For evaluating the stroke disease, three machine learning models were compared in different scenarios, and the result revealed that Random Forest consistently outperforms Decision Tree and SVM models. This great performance might be because it works as an ensemble, which lowers overfitting and boosts generalization, especially when used with feature selection and K-fold validation. While Decision Tree has lower performance than Random Forest, it shows more powerful performances compared to the SVM model. In contrast, the SVM model has poorer performance than the two models due to the non-linearity of the data. Even after applying K-fold optimizer feature selection, SVM could not match the robustness of the ensemble models.
The influence of glucose level and age on the risk of stroke becomes much larger when both are elevated. The combined effect of older age and high glucose is not merely additive but amplifies the risk substantially, suggesting a strong interaction between the two predictors. These findings emphasize the importance of closely monitoring glucose levels in older adults to mitigate the heightened risk of stroke.
The confusion matrix, ROC graph, precision-recall curves, and true positive-false positive curve further validate the superiority of Random Forest. The result also shows the model minimizes false negatives and false positives, which are critical for medical applications.36,31 The AUC score of 1.00 confirms that the Random Forest model can reliably identify strokes with high confidence, making it highly suitable for medical diagnostic applications. Moreover, compared to prior research, the proposed system's accuracy of 99.4% represents a significant improvement, showing that integrating optimized preprocessing and validation with ensemble models can dramatically enhance diagnostic accuracy. This establishes the model not only as statistically effective but also clinically viable.
Moreover, the usability evaluation conducted with healthcare professionals confirms that the system is not only technically accurate but also it can be practically implemented. The personal rating across selected evaluation criteria demonstrates the potential performance of the proposed technique in hospital settings. The positive reception of the knowledge-based system highlights the importance of clinician-in-the-loop design, ensuring the system addresses real-world workflows and clinical decision needs. The feedback collected during iterative development phases played a vital role in improving the future system's utility and effectiveness.
In conclusion, the proposed Random Forest-based stroke diagnostic system, supported by intelligent feature selection and robust validation, offers a reliable, accurate, and user-friendly tool for enhancing stroke diagnosis and treatment planning in clinical environments.
Limitations of the study
In this study, despite the promising results being obtained, there are several limitations that should be acknowledged. First, the current system primarily utilizes structured clinical data and does not integrate radiological imaging or detailed neurological examination, which are key components of clinical stroke diagnosis. This limits the diagnostic capabilities in differentiate stroke types or the detection of transient ischemic attacks.
A second basic limitation is the lack of focus on etiological classification in ischemic stroke. Determining the underlying cause—whether it is cardioembolic, atherothrombotic, or cryptogenic—is a crucial challenge in clinical practice, since it significantly influences therapeutic approach. The current version of the system does not yet accommodate this level of diagnostic refinement. Moreover, the current data set used, which was relatively small and from a single clinical site, also limits the generalizability of the results. Thus, the model's performance in other health care settings with different populations or diagnostic workflow traits may vary.
Lastly, the usability testing was conducted in a small group of five clinicians in one institution. Although the feedback was very positive, more widespread evaluation across institutions and user groups would provide a clearer image of the system's true effectiveness and real-world acceptance. Based on the limitations presented, most of the issues will be addressed in our going on work.
Conclusion
Stroke remains a leading cause of death and disability worldwide, necessitating early detection and intervention to improve patient outcomes. This study addresses this critical need by developing a hybrid knowledge-based system that integrates expert knowledge with machine learning predictions. Using data from Debre Berhan Referral Hospital and one publicly available from Kaggle, the proposed system applies Design Science Research Methodology to model the expert knowledge into production rules implemented by decision tree models. A random forest classifier with feature selection resulted in the best accuracy of 99.4% among the classifiers tested. The KBS was developed using Python and Prolog, embedding the power of machine learning with domain expertise for reliable diagnosis and to guide treatment in stroke cases. This new approach offers a useful tool for health care providers in enhancing diagnosis and clinical decision-making independently. By leveraging diverse datasets, multimodal data processing, and model optimization, the system demonstrates robust performances. In future studies, we would like to apply deep learning to large datasets incorporating radiological imaging.
Footnotes
Author contributions
Conceptualization: T.K.Y. and A.B.A; methodology: T.K.Y. and A.B.A.; software: A.B.A.; validation: A.B.A.; formal analysis: T.K.Y. and A.B.A; investigation: T.K.Y. and A.B.A; resources: T.K.Y. and A.B.A; data curation: T.K.Y. and A.B.A; writing—original draft preparation: T.K.Y. and A.B.A; writing—review and editing: A.B.A.; visualization: A.B.A. All authors have read and agreed to the published version of the manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.

