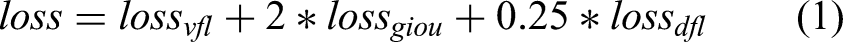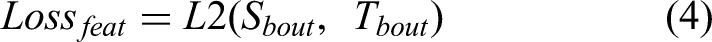Abstract
Background
When applying for teleconsultations, medical laboratory reports are usually photographed with a mobile phone, and the photographic results are uploaded as teleconsultation application materials. It is very meaningful to extract the content of the image medical laboratory report and store the content digitally. There are already applications of OCR technology for medical text file recognition, but no researchers have recognized the format of the medical laboratory report and obtained the report content as a serialized process to digitize the image report. This article proposes a serialization method to digitize the medical laboratory report image.
Materials and Methods
This article first collects 330 image-based medical laboratory reports, annotates the format of the medical laboratory reports, and forms a training dataset for the layout analysis model. Then, using the pre-trained model, the dataset is trained to obtain a layout analysis model that can correctly recognize the format of the medical laboratory report. Then, the layout of the input image-based medical laboratory report is analyzed, and the layout analysis results are used to call the text detection and text recognition models to obtain the digital content of the image report. Finally, adjusting the layout of the digital content and storing the digital content as a docx file.
Results
After training the layout analysis model, integrating layout analysis, text detection, and text recognition, we have obtained a serialization method that digitizes the content of the image medical laboratory report, restores the report format, shields sensitive and irrelevant content, and digitizes the report content of interest.
Conclusions
By digitizing the image medical laboratory report through the serialization method, we can correctly display the content of the medical laboratory report for teleconsultation, while removing irrelevant content in the report, such as user names, examination equipment numbers, etc.
Introduction
Medical laboratories have been widely used, and the importance of these tests is well known. 1 When applicants send the required information of patient during teleconsultation between different hospitals, due to the lack of system-level interconnection, it is not possible to export the original electronic medical laboratory report normally. Therefore, applicants usually use mobile phones to take images of patients’ medical laboratory reports, and stored them in the form of pictures, and these report images are often uploaded as an attachment. 2
There are some limitations to image reports in the form of picture attachments. It is impossible to directly obtain the digital content on the medical laboratory report, nor is it possible to store the image attachments in a structured manner. 3 Medical data involves personal privacy protection and cannot be processed using existing online recognition tools. Prior studies have explored deep learning-based text recognition in medical imaging, 4 but their approaches primarily focused on single-modality document processing and lacked adaptability to multi-format report.5–8 Real-world deployments of similar systems in clinical settings face significant challenges, including integration into existing Electronic Health Record workflows, handling diverse report formats, and ensuring privacy compliance.9,10 There are various formats, and the size of layout elements is not uniform. The medical laboratory report has personalized characteristics, and different hospitals have different types of report templates. 11
Making papery document digitalized mainly relates to OCR technologies (Optical Character Recognition), especially text detection and text recognition. 12 Text detection and text recognition have been applied in healthcare. Text detection and text recognition have received continuous attention with the emergence of many application scenarios, as the emergence of high-end hardware has facilitated the development based on deep learning algorithms.13–16 OCR technologies achieve recognition rate higher than 99% on scanned document.17,18 Usually, text detection and recognition are treated as a whole, and computer vision and learning methods are used to process the problem. 19 In healthcare, deep neural networks are used to improve the accuracy of OCR engines for transcribing scanned medical reports. 20 OCR and NLP technologies are used to structure and digitize medical records. 21
PP-OCRv2 is a lightweight OCR system, which balances the accuracy against the efficiency. Multiple pre-trained models are provided by PP-OCRv2 for use, and lightweight optimization is performed for CPU usage scenarios. However, existing models cannot accurately recognize medical laboratory reports’ layout. 22
This article is based on the PP-OCRv2 framework and establishes own digital recovery application for medical laboratory reports. The independent processes of layout analysis, text detection, text recognition, and layout recovery are established as a sequential series to digitize image-based medical laboratory reports. This solves the problem of the inability to digitize image-based reports, protects patient privacy, and adapts to different report formats.
Methods
The method proposed in this article is shown in the Figure 1. Existing layout analysis models cannot meet the requirements. Due to medical data security and other reasons, there are publicly available medical laboratory reports. We collected 330 medical laboratory reports and trained our own layout analysis model based on a pre-training model. For input image-based medical laboratory reports, after processing by the layout analysis model, a list of results containing different labels in the image is detected. For each label obtained, the text detection and text recognition models are called to obtain the position information and content. Finally, the obtained results are written into a docx file.

Sequential series of reports recovery.
Training layout analysis model
PP-OCRv2 provides multiple pre-trained models. However, existing models cannot meet the requirements for layout analysis. Figure 2 shows the result of using pre-trained model. For image-based medical laboratory report taken with a mobile phone, existing models cannot accurately detect box information for key contents such as “diagnostic opinions” and “impressions”. Due to information security considerations, we do not want to analyze and recognize boxes containing contents such as “patient name” and “examination equipment number”. Therefore, we trained a layout analysis model to meet our requirements.

The result of using the pre-trained model, containing wrongly detected box, “diagnostic opinions” not labelled.
Data annotation
We collected 330 image-based medical laboratory reports taken by mobile phones, and used the annotation tool, LabelMe to annotate and label each image of medical laboratory report, which is a widely used labelling system, and allows users to draw polygons and constructs training sets. 23 The 330 medical laboratory reports were collected from the First Affiliated Hospital of Zhengzhou University in Henan, China, covering five major categories of reports (CT reports, MRI reports, pathology reports, and ultrasound reports) with distinct template formats. This diversity ensures the model's adaptability to institutional variations in report layouts. However, to further validate the robustness of the model, future work will involve collecting data from multiple healthcare institutions with different documentation styles. Then converted these annotations to COCO format. 24 After LabelMe is launched, using “Create Polygons” to circle the area to be recognized and setting the area label. A corresponding annotation JSON format file will be generated once a report is labelled. Figure 3 depicts the interface of LabelMe after launched.
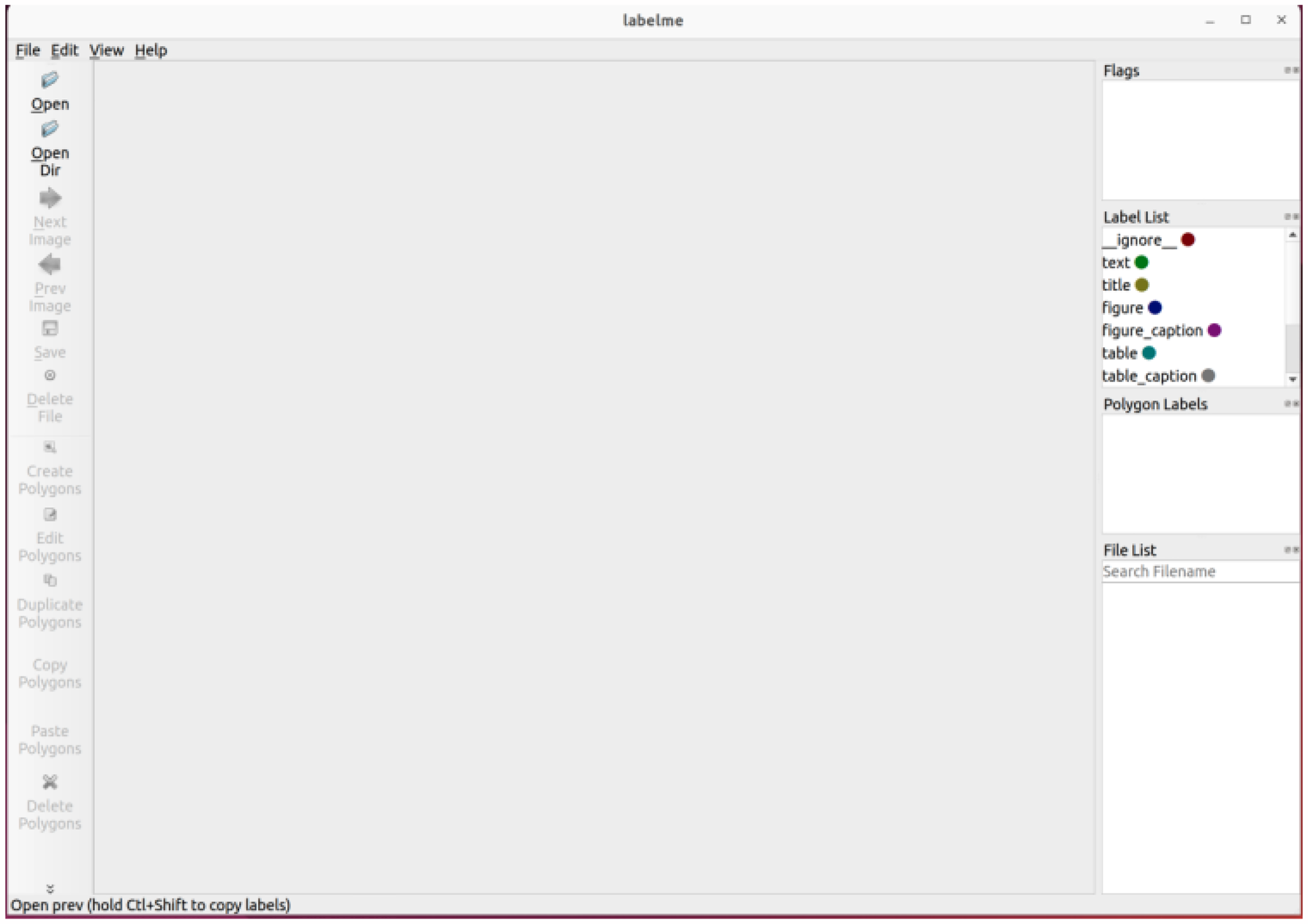
LabelMe interface.
Training layout analysis model
Layout analysis refers to the regional division of documents in the form of images, common images are stored in the form of pictures (jpg, JPG format) or PDF files. The goal is to identify key areas such as text, titles, and figures.
PP-StructureV2 provides a pre-trained layout analysis model based on PP-PicoDet,25,26 which is a lightweight detector that balances accuracy and efficiency well. It uses Enhanced ShuffleNet (ESNet) as the backbone, with SE modules added to each ES Block for better channel weighting. The neck of the network uses CSP-PAN for feature concatenation and fusion, reducing computational costs by using 1*1 convolution.27,28 As shown in Figure 4.
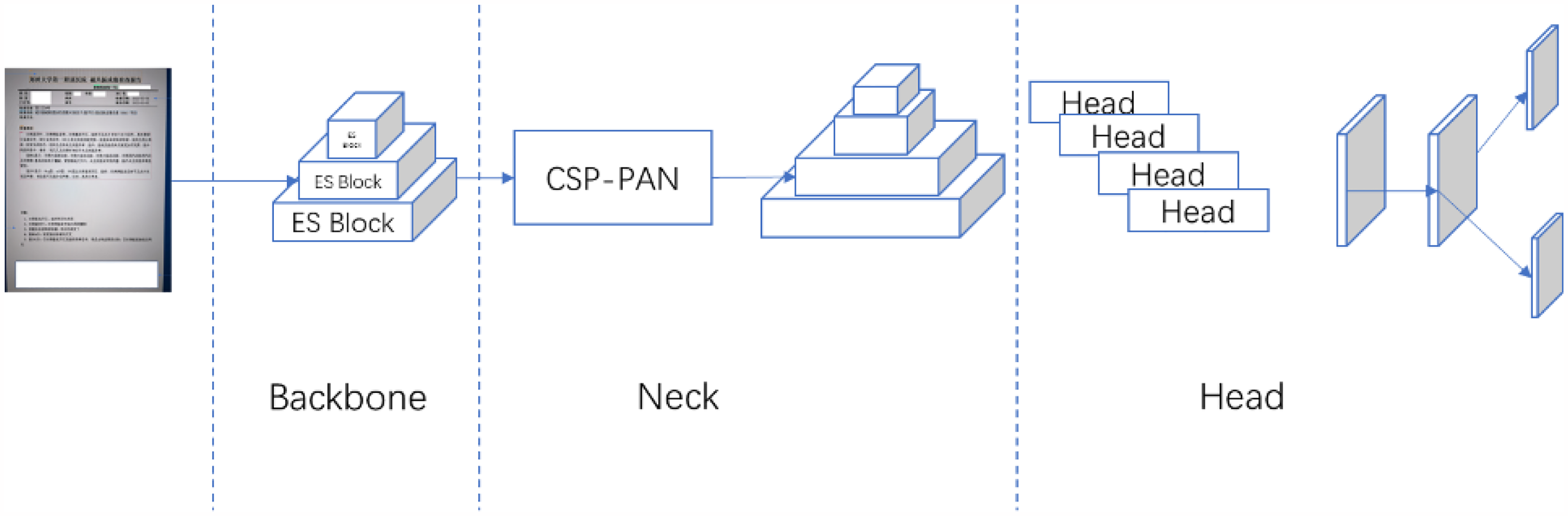
PP-PicoDet architecture. The backbone is ESNet which outputs 3 feature maps to CSP-PAN. CSP-PAN acts as neck that outputs 4 feature maps. PP-PicoDet uses SimOTA dynamic label assignment strategy to optimize our training process.
During training, the dataset is resized to different resolutions, and a random interpolation method is used. SimOTA dynamic label assignment strategy optimizes the training process, and various loss functions (varifocal loss, GIoU loss, and Distribution Focal Loss) are used to improve model performance. In the head of PP-PicoDet, calculating varifocal loss to couple classification prediction and quality prediction. Correspondingly, for regression, using GIoU loss and Distribution Focal Loss. The formula is as follows:
Model export
Compared with the checkpoints model, that saved in training process and only saved the parameters of temporary model, the inference model also needs structural information. Therefore, we export the layout analysis model, including model structure and model parameters in solidified files. This way performs superiorly in prediction deployment, is flexible and convenient, and is suitable for actual system integration.
Sequential series to digitize image-based medical laboratory reports
Text detection and text recognition
The PP-OCRv2 framework supports bilingual recognition (Chinese/English) and common medical abbreviations (e.g., AD, CT, MRI) through its hybrid language model architecture. 22 This capability enables accurate interpretation of both localized terminology and international standardized abbreviations.
In the text detection model, PP-OCRv2 uses CML between three models, allowing the large model to guide the small model. The method of knowledge distillation is commonly used in deployment. By guiding the small model with the large model, the accuracy of the small model can be further improved under the condition of unchanged prediction time, thereby enhancing the actual deployment experience. The structure figure of CML is shown in the Figure 5. The entire training process consists of three loss functions: GT loss, DML loss, and Distill loss. Sub-student models learn from each other with reference to DML method.
29
DML loss function in the CML is as follows:

CML framework, the teacher model guides the student model.
The total loss function in CML is as follows.
In the text recognition model, PP-OCRv2 uses the U-DML knowledge distillation method. Based on the traditional DML strategy, PP-OCRv2 further adds the intermediate output feature map supervision signal as a loss function to the final output layer of the text recognition model. As shown in Figure 6, the teacher model and the student model have the same network structure but different initialization parameters. A supervision mechanism for feature map is introduced, and feature loss is added. The Feature loss uses L2 loss, and the specific calculation method is shown below:
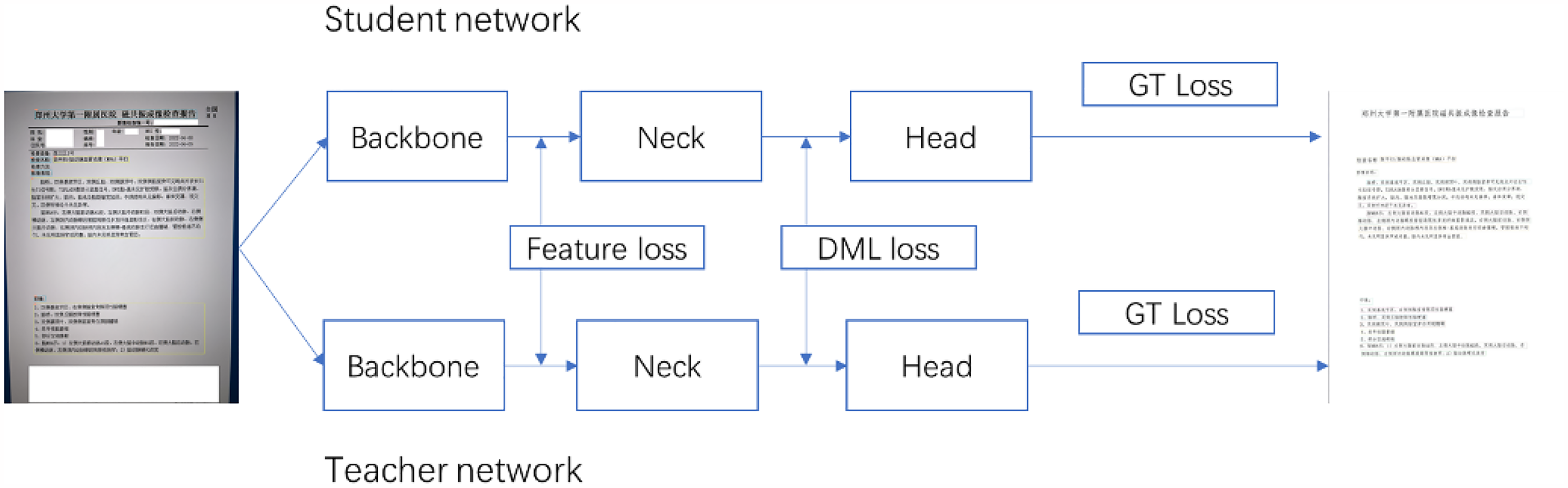
U-DML framework, teacher model is identical with student model in structure, and feature loss is added.
Finally, the total loss is the following:
Converting image to docx format
This section is about converting the analyzed results into the docx file format. By iterating through the result collection returned by the previous steps, different docx file display modules are generated based on the result category, such as center alignment and title generation. During the development process, the document results that do not match the layout of the original image display are refined to ensure the correct layout of the displayed content. Finally, the code is written to combine the results and store them as a docx document.
Results
It took 8 months to build this sequential series method. According to the sequence function, the entire process is divided into several steps: collecting medical laboratory reports, annotating the collected reports, training the layout analysis model, text detection and text recognition, converting the analysis results into docx documents, and so on. Next, we will list the experimental results of each stage.
Dataset and data annotation
We collected 330 photos and annotated them. We only annotated the medical content modules, such as ‘imaging performance’, ‘diagnostic opinions’, ‘impressions’, etc. For some information that is not related to medical treatment, such as ‘patient name’ and ‘examination equipment number’, we do not annotate them due to information security reasons. Since the layout analysis model is trained on a pre-trained model of Chinese CDLA dataset for Chinese paper scenarios, 30 we specified a label file, which consists of 10 categories, including Text, Title, Figure, Figure caption, Table, Table caption, Header, Footer, Reference, Equation. Figure 7 depicts the interface of labelling process.
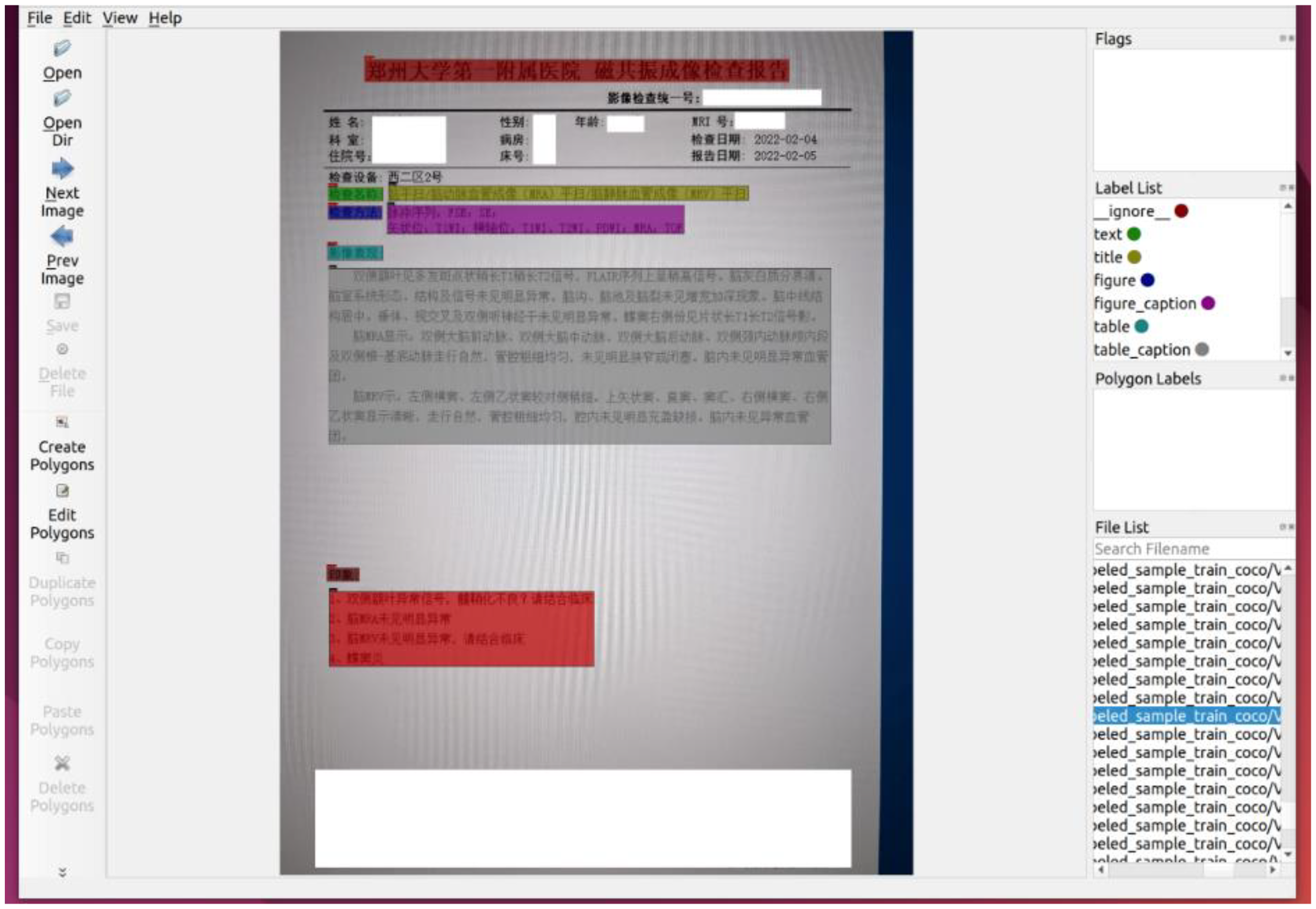
Using labelMe to label medical laboratory reports, we only label the parts needed, and desensitize some sensitive fields.
As shown in figure 8. At a ratio of 9:1:1, 330 reports were divided into training set, validation set and test set, and the training set, validation set were labelled respectively. To facilitate training, the annotation results are converted to the COCO data format. Each annotated image corresponds to a JSON format annotation file. After conversion to the COCO format, all annotation files are merged into an “annotations.json” file.

Depicting the process of data splitting, labelling images, and converting to “annotations.json” file.
Layout analysis model training results
Comparative experiments with Tesseract OCR 5.0 demonstrated superior performance of our method in handling complex medical reports (Table 1). The proposed system achieved 92.4% F1-score versus 78.1% in recognizing medical abbreviations from low-quality mobile images.
Performance comparison.

Export the optimal evaluation model as an inference model. The inference model contains model parameter information as well as model structure information. This facilitates the integration of the layout analysis model and makes model deployment easier. The process is shown in Figure 9.
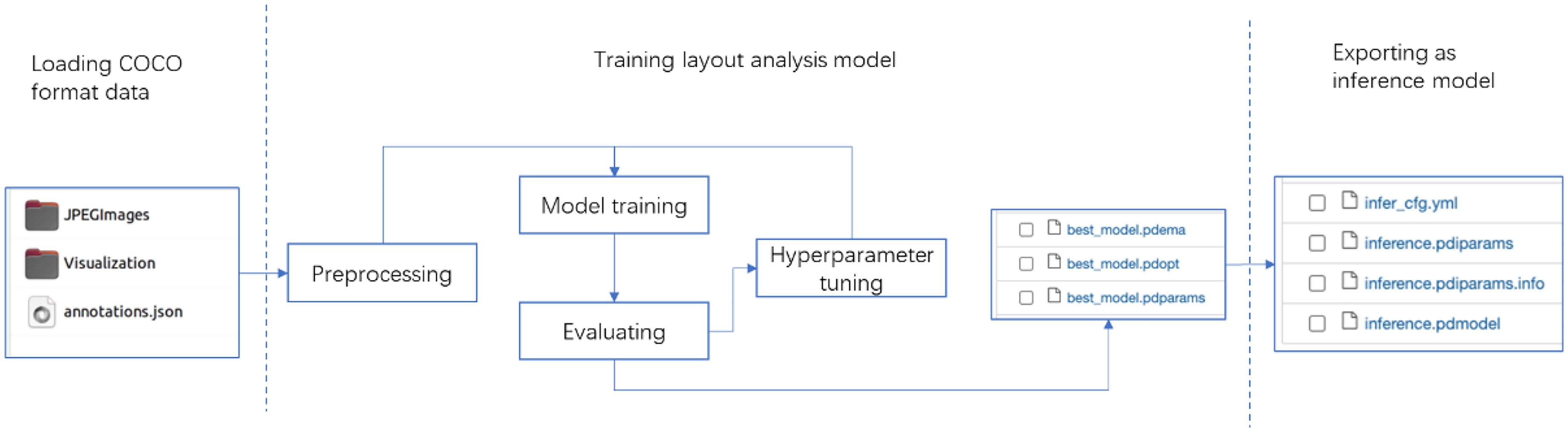
After training the layout analysis model, we export the best evaluation model as inference model.
Sequentially digitizing image-based medical laboratory reports
For an image-based medical laboratory report, the layout analysis, text detection, and text recognition are called in sequence to digitize the content of the report in image form, and finally convert the content of the report from an image to a docx file.
Performing layout analysis on the image report. Calling the layout analysis model, which returns a list of “box coordinates, labels”. For each box coordinate area, text detection model is performed to obtain a list of text area coordinates within the box. Performing text recognition on each text area. Until all the contents of the “box coordinates, labels” list are processed, programmatically adjusting the mismatched areas and converting the result into a docx file. The detailed process is shown in Figure 10.

Workflow of sequentially digitizing image-based medical laboratory report. After performing layout analysis, a list of “box coordinates, labels” returns. Iterating the list, by using text detection and recognition. Finally, converting the previous result into a docx file.
Discussion
This study presents a serialized approach for digitizing image-based medical reports, addressing three critical challenges in telemedicine: 1) Privacy-aware processing through selective annotation and identifier exclusion; 2) Cross-categorical adaptability via a hybrid model architecture; 3) Computational efficiency enabling CPU-based deployment. Compared with existing medical OCR systems that primarily focus on scanned documents, 31 our method demonstrates superior performance in handling mobile-captured images with complex layouts (Table 1).
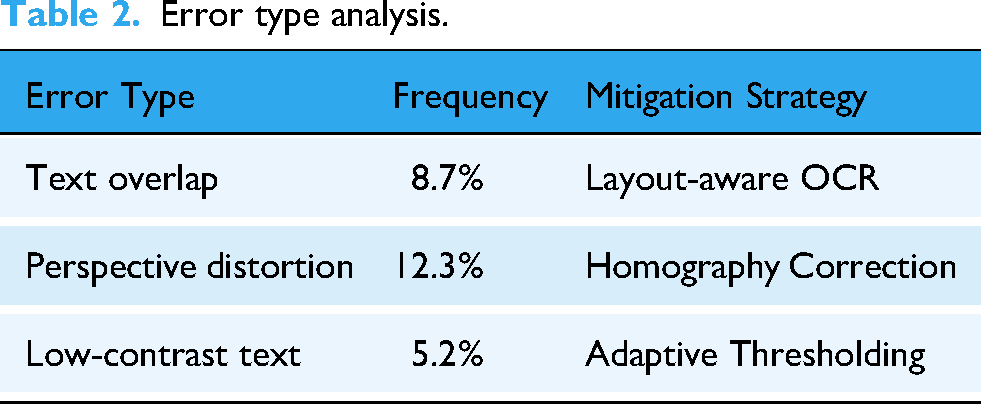
Error type analysis.

Practical deployment considerations
The system achieves 2.3 s average processing time per report on standard CPU hardware (Intel Xeon E5-2680v4), meeting real-time requirements for teleconsultation scenarios. This efficiency enables cloud-based batch processing deployment paradigm: Handling 43 reports/minute on NVIDIA T4 GPU.
Error pattern analysis
Common recognition errors occurred in cases of severe image distortion (12.3% error rate) and overlapping text annotations (8.7% error rate). These issues could be mitigated through image preprocessing techniques like perspective correction. These are shown in Table 2.
These challenges were empirically evaluated, and the following results were obtained:
Homography correction: Reduced perspective distortion errors by 47.5%. Adaptive thresholding: Improved low-contrast text recognition accuracy by 30.1%.
Limitations and future directions
The current model shows decreased performance (72.1% AP) when processing reports from unseen hospital templates. This limitation stems from the single-institution dataset (330 reports from one hospital).
32
Future works should:
Expand dataset diversity: Collect cross-institutional reports through multicenter collaboration System integration: Implement HL7/FHIR interfaces for direct EHR (Electronic Health Record) insertion, as proposed in our conclusion
The proposed serialization method provides a foundation for privacy-preserving telemedicine infrastructure, particularly valuable in developing regions lacking standardized EHR systems.
Conclusion
This study presents a serialization method for digitizing image-based medical reports with three key advantages: 1) Privacy protection through selective annotation; 2) Cross-format adaptability via customized layout analysis; 3) CPU-friendly deployment. Future work will focus on: 1) Expanding multilingual support for global telemedicine applications; 2) Developing HL7/FHIR interfaces for hospital information system integration; 3) Implementing federated learning frameworks to enhance model generalizability across institutions.
Footnotes
Acknowledgements
We would like to thank all the participants, for their valuable suggestions and support during the completion of the research.
Ethical considerations
All data and images included in this study were fully anonymized prior to analysis. No personally identifiable information (including patient names, ID numbers, or addresses) was retained in any materials. Therefore, informed consent from participants was not required for the use of these anonymized data, in accordance with guidelines on the use of de-identified medical records.
Author contributions/CRediT
All authors approved the final manuscript. Author contributions are as follows: Xiaoyang Ren: Conceptualization, Methodology, Software, Resources, Writing. Dongwei Dou: Validation, Data Curation, Supervision, Project administration, Editing. Xianying He: Validation, Editing. Fangfang Cui: Validation, Editing. Jie Zhao: Funding acquisition, Project administration.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Key Science and Technology Program in Henan Province, Key Scientific Research Project of Colleges and Universities in Henan Province, (grant number 201400210400, 23A520018).
Conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.