Abstract
Purpose
Prostate cancer (PCa) is the second most common cancer in males worldwide, requiring improvements in diagnostic imaging to identify and treat it at an early stage. Bi-parametric magnetic resonance imaging (bpMRI) is recognized as an essential diagnostic technique for PCa, providing shorter acquisition times and cost-effectiveness. Nevertheless, accurate diagnosis using bpMRI images is difficult due to the inconspicuous and diverse characteristics of malignant tumors and the intricate structure of the prostate gland. An automated system is required to assist the medical professionals in accurate and early diagnosis with less effort.
Method
This study recognizes the impact of zonal features on the advancement of the disease. The aim is to improve the diagnostic performance through a novel automated approach of a two-step mechanism using bpMRI images. First, pretraining a convolutional neural network (CNN)-based attention-guided U-Net model for segmenting the region of interest which is carried out in the prostate zone. Secondly, pretraining the same type of Attention U-Net is performed for lesion segmentation.
Results
The performance of the pretrained models and training an attention-guided U-Net from the scratch for segmenting tumors on the prostate region is analyzed. The proposed attention-guided U-Net model achieved an area under the curve (AUC) of 0.85 and a dice similarity coefficient value of 0.82, outperforming some other pretrained deep learning models.
Conclusion
Our approach greatly enhances the identification and categorization of clinically significant PCa by including zonal data. Our approach exhibits exceptional performance in the accurate segmentation of bpMRI images compared to current techniques, as evidenced by thorough validation of a diverse dataset. This research not only enhances the field of medical imaging for oncology but also underscores the potential of deep learning models to progress PCa diagnosis and personalized patient care.
Introduction
Prostate cancer (PCa) remains the second most common cancer among men worldwide, presenting a significant public health challenge. The American Cancer Society estimated over 248,530 new cases and 34,700 deaths from PCa in the United States in 2023 alone, highlighting the crucial need for early detection and accurate diagnosis. Early identification of clinically significant PCa (csPCa) is essential for effective treatment planning and patient management, potentially saving lives and reducing the burden of this disease. The diagnostic process for PCa has evolved over the years, with magnetic resonance imaging (MRI) playing a pivotal role in the detection, staging, and monitoring of this malignancy. To ensure a standardized global approach to prostate multiparametric MRI (mpMRI) interpretation, the latest iteration of the Prostate Imaging Reporting and Data System (PI-RADS 2.1) amalgamates existing evidence to assign scores to objective findings in each sequence. 1 Nevertheless, the interpretation of bi-parametric MRI (bpMRI) remains time-consuming, contingent on expertise, 2 and often characterized by significant interobserver variability,3,4 especially in nonspecialized centers. 5 Moreover, like any human-based decision-making process, MRI interpretation is susceptible to errors, a vulnerability that may be exacerbated by cognitive impairments such as mental stress. 6 Computer-aided diagnosis (CAD) systems represent a particularly promising avenue of research in medical imaging. CAD has demonstrated successful applications across various medical contexts, offering potential benefits including expedited diagnosis, reduced diagnostic errors, and enhanced quantitative analysis.7–10 Various early diagnostic systems that assist radiologists in accurate lesion segmentation have been proposed, marking the inception of this field.11,12 While these early methods laid the groundwork, they were constrained by several limitations, including insufficient evaluation, absence of expert comparison, and inadequate dataset sizes. The landscape shifted with the emergence of deep learning, particularly deep convolutional neural networks (DCNNs), 13 which swiftly supplanted traditional classification methods across various image analysis domains, including medicine. For prostate MRIs, a pivotal moment occurred during the PROSTATEx Challenge of 2016.14–16 This challenge revolved around csPCa classification based on tentative lesion locations within mpMRI scans. Researchers employing deep learning approaches first extracted a region of interest (ROI) surrounding the lesion position within the mpMRI, and a CNN architecture inspired by VGG was trained on smaller patches to classify the MRIs. 17
From a radiological perspective, the prostate is typically divided into two distinguishable zones on MRI scans: the central gland (CG) and the peripheral zone (PZ). Our research aims to incorporate anatomical knowledge of these zones into CAD systems. The initial step in this process is the automated segmentation of these zones as this is essential for CAD methods that specifically target the PZ. 11 Among the various MRI modalities, we opted to use bpMRI because of its reduced acquisition time, cost-effectiveness, and the potential to decrease false positive rates compared to mpMRI. bpMRI streamlines the imaging procedure by prioritizing two essential sequences: T2-weighted (T2w) images and diffusion-weighted images (DWIs), ensuring adequate diagnostic precision for detecting PCa without the need for contrast agents. Although there are benefits, the process of detecting PCa in bpMRI pictures is still difficult because of the inconspicuous and diverse appearance of malignant tissues, the intricate structure of the prostate gland, and the existence of other noncancerous illnesses that can imitate cancer.
Accurate segmentation of PCa is essential for evaluating tumor size, location, and invasion extent, which are crucial criteria in making treatment decisions and assessing prognosis. This research follows a two-stage implementation exploring multiple datasets for a rigorous analysis. Data preprocessing including converting all datasets into a uniform NifTI format, standardization, resizing, and intensity normalization is carried out. To address overfitting of the models, data augmentation is performed to increase the number of images. The prostate zonal segmentation model is developed to automate the PCa detection system. The second stage of our methodology involves training an attention-guided U-Net for lesion segmentation. Our approach of training multiple pretrained models with different architectures (VGG19, ResNet201, and SEResNet152) and then training an attention-guided U-Net from scratch allowed for a comprehensive comparison. The performance of our models compared to these baselines architectures is presented. Several evaluation metrics including the dice similarity coefficient, IoU, sensitivity, specificity, and AUROC, are utilized for a comprehensive assessment of the model performance. Our results demonstrate significant improvements in both prostate zonal segmentation and lesion detection, showcasing the efficacy of incorporating anatomical knowledge into deep learning models.
Related work
Over the years, several research has been done for the automated diagnosis of PCa including both segmentation and classification. Xu et al., 18 Yoo et al., 19 and Cao et al. 20 employed similar methodologies for lesion segmentation and classification on individual slices of mpMRIs. Yoo et al. 19 utilized a ResNet20 architecture to classify each slice, subsequently aggregating the probabilities of each slice to produce a final score. Xu et al. 18 also used a ResNet-based architecture for segmenting individual slices. Similarly, Cao et al. 20 applied a slice-wise segmentation CNN for csPCa prediction and Gleason grade group mapping. Litjens et al. 21 introduced a method that combines anatomical, intensity, and texture data to distinguish between the CG and PZ, achieving mean dice coefficients of 0.89 ± 0.03 and 0.75 ± 0.07, respectively. Saha et al. 22 compared the diagnostic performance of artificial intelligence (AI) models with that of radiologists, csPCa with an AUROC of 0.88 ± 0.01 and a sensitivity of 76.38 ± 0.74%.
Jia et al. 23 proposed a segmentation method utilizing a two-step approach, incorporating deep neural networks and ensemble learning, achieving a dice ratio of 0.910 ± 0.036. Le et al. 24 presented a multimodal CNN approach, achieving a sensitivity of 89.85% and specificity of 95.83% for cancer detection and 100% sensitivity and 76.92% specificity for discriminating between indolent and clinically relevant malignancies. Aldoj et al. 25 developed a Dense-2 U-net model, integrating DenseNet and U-net architectures, to accurately segment the prostate and its zonal structures from MRI scans, achieving average dice score of 92.1%. Fütterer JJ et al. 26 evaluated the diagnostic accuracy of mpMRI for detecting cs-PCa, reporting a range of diagnostic accuracy from 44% to 87%. Mehrtash et al. 27 created a 3D CNN model to categorize the clinical importance of PCa observations based on mpMRI data, achieving an area under the curve (AUC) of 0.80. Ghorashi et al. 15 proposed a method utilizing transrectal ultrasonography images and a Hidden Markov Model classifier for the identification of PCa, through statistical supervised learning and texture analysis. Niaf et al. 13 evaluated the probability of PCa in the PZ using mpMRI. This imaging technique included T2w, diffusion-weighted, and dynamic contrast-enhanced MRI at 1.5T. Xin Yang et al. 28 introduced a new method for detecting PCa using mpMRI data, which includes apparent diffusion coefficient (ADC) and T2w images, utilizing cotrained CNNs.
Pellicer-Valero et al. 29 utilized deep learning techniques to automatically detect, segment, and estimate the Gleason grade of PCa in mpMRI images, achieving AUC, sensitivity, and specificity of 0.96, 1.00, and 0.79, respectively. De Vente et al. 30 utilized a neural network to both detect and classify PCa based on bpMRI data, achieving voxel-wise weighted kappa of 0.446 ± 0.082 and a dice similarity coefficient of 0.370 ± 0.046 for accurately segmenting clinically significant cancer. Valerio et al. 31 conducted a systematic review comparing the detection rate of clinically relevant PCa using software-based MRI-ultrasound (MRI-US) fusion targeted biopsy with traditional biopsy. Chan et al. 32 developed a statistical classifier utilizing multiple channels from T2w MRI, T2-mapping, and line scan diffusion imaging to detect PCa. Seah et al. 33 introduced a DCNN method for lesion identification on mpMRI of the prostate, achieving an AUC of 0.84 by incorporating innovative techniques such as “auto-windowing” and extensive data augmentation. Cao et al. 34 presented FocalNet, a CNN model capable of detecting PCa lesions and predicting their Gleason score using mpMRI, demonstrating exceptional sensitivity in lesion detection and accuracy in Gleason score classification. Toth et al. 35 introduced the Multi-Feature Landmark-Free Active Appearance Model, tailored for prostate MRI segmentation. This approach utilizes a level set representation and integrates multiple image-derived attributes to improve segmentation accuracy. They achieved an average dice similarity coefficient of 88% and an average surface error of 1.5 mm. Woznicki et al. 36 showcased the potential of integrating radiomics, PI-RADS, and clinical characteristics in multiparametric MRI for improving PCa identification. Utilizing machine learning models, they achieved AUC values of 0.889 for distinguishing malignant from benign lesions and 0.844 for identifying clinically important cancer. Gupta et al. 37 detailed the evolution of the PI-RADS from v1 to v2.1. Yoo et al. 19 developed a computerized pipeline using a DCNN for identifying clinically relevant PCa, achieving AUC values of 0.87 and 0.84 at the slice and patient levels, respectively. Xu et al. 18 demonstrated the effectiveness of ResNets in accurately detecting and segmenting potentially cancerous spots in mpMRI images where the ResNet model achieved a binary accuracy of 93% and an average Jaccard score of 71% in lesion detection. Liu et al. 38 introduced XmasNet, a deep-learning method tailored for the classification of PCa lesions using 3D mpMRI data. Trained end-to-end with data augmentation techniques, XmasNet significantly outperformed conventional machine learning models, achieving an impressive AUC score of 0.84 in the PROSTATEx Challenge. Yuan et al. 39 presented Z-SSMNet, a method for detecting csPCa in bpMRI and demonstrated excellent performance showcasing its potential in diagnosing csPCa using bpMRI. Mahapatra et al. 40 segmented the prostate from MRI images using a combination of random forests and graph cuts. Karimi et al. 41 proposed a CNN approach for prostate segmentation in MRI scans, integrating statistical shape models. Their model achieved a dice score of 0.88. Aldoj et al. 42 introduced a semiautomatic method for classifying PCa, utilizing a multichannel 3D CNN on mpMRI data. Their findings demonstrated optimal performance with an average AUC of 0.897. Arif et al. 43 developed an automated technique for detecting csPCa in low-risk individuals using a 3D CNN on multi-bpMRI data. Their model showed promising sensitivity (82%–92%) and specificity (43%–76%), with AUC ranging from 0.65 to 0.89.
Some commonly used architectures in the field include U-Net, DeepSegNet, and versions that integrate recurrent neural networks and long short-term memory networks. 44 In this work, 45 the researchers evaluated the effectiveness of a deep learning system based on U-Net with the clinical assessment. A different research 46 employed a VGG-16 CNN to extract features from mpMRI, with a specific emphasis on ADC, high b-value (hbv), and T2W images. The collected features were then analyzed using an ordinal class classifier with J48 as the underlying classifier to account for the ordinal nature of PCa grading. The objective of this approach was to categorize PCa into five grade groups, resulting in a modest quadratic weighted kappa score of 0.4727.
Despite the advances brought by these approaches, a notable limitation is the necessity for the localization of ROIs, which restricts their practical utility in clinical environments. Additionally, it has been observed that two-dimensional (2D) slice-wise CNNs often underperform in lesion detection tasks compared to true three-dimensional (3D) CNNs in terms of time and resource complexity. The heightened time and memory complexity in 3D CNNs arises from their requirement to handle complete volumetric data instead of separate 2D slices. This task entails managing larger 3D feature maps, which greatly increases the amount of memory needed, especially GPU RAM. This is because the network needs to store and manipulate data in three dimensions (height, width, and depth) at the same time. Furthermore, the computations involved in 3D CNNs are more demanding, resulting in increased processing durations for both training and inference. The convolutional and pooling processes, which need to be performed along all three dimensions, lead to an increased computational burden, causing slower forward passes and backpropagation during training. As a result, 3D CNNs necessitate more powerful computational resources, such as high-performance GPUs or clusters, to effectively handle the increased memory requirements and longer processing durations, in contrast to the more efficient but less contextually aware 2D CNNs. Occasionally, researchers choose for 2D models over 3D models due to these constraints. Our approach involves training several pretrained models with diverse architectures, all utilizing ImageNet as the backbone. We selected VGG19, ResNet201, and SEResNet152 as baseline models for comparison. Additionally, we trained an attention-guided U-Net from scratch to evaluate and contrast the performance between pretrained models and a model trained from scratch. Although several previous studies may present comparable findings, this research distinguishes itself from the existing literature through several significant variations. The present work utilizes a two-stage process in which the initial step concentrates on the segmentation of prostate zones, followed by the segmentation and categorization of lesions. In contrast to numerous previous research, which frequently consider segmentation and classification as a unified task or fail to highlight the significance of zonal awareness in the segmentation process. The integration of attention mechanisms into the U-Net architecture represents a notable progress. This enables the model to concentrate on pertinent regions of the images, hence improving the accuracy of segmentation. The study utilizes several datasets for both training and validation, guaranteeing that the model can be applied to different MRI modalities with a high level of accuracy and reliability. The disparities emphasize the impact of the present text on the domain of PCa identification and emphasize its capacity to enhance diagnostic precision and individualized patient treatment.
Methodology
Our proposed methodology consists of two stages to achieve precise segmentation of PCa. First and foremost, it is crucial to define the return on investment, specifically in relation to the prostate in our situation. Not doing this hinders the second stage model's capacity to properly learn significant information about lesion segmentation. The study utilizes a two-stage strategy that greatly improves the precision of PCa detection. The first step involves segmenting the prostate region, which is crucial for the following detection of lesions. This approach allows the model to focus on the relevant anatomical features, which reduces the occurrence of incorrect positive results and improves the accuracy of lesion categorization. The initial segmentation phase is crucial because it defines the ROI, enabling the succeeding model to extract and acquire significant information for accurate lesion segmentation. Figure 1 depicts the entire methodology of the research.

The proposed methodology of this study.
To begin with, data preprocessing including converting all datasets into a uniform NifTI format, standardization, resizing, and intensity normalization is carried out. The input shapes were adjusted to fit our specific requirements, which in this case is 192 × 192 × 17. To address overfitting of the models, data augmentation is performed to increase the number of images. This facilitates the development of a generalized model capable of extracting ROIs from various MRI modalities, including T2w, ADC, sagittal (sag), coronal (cor), and hbv imaging, particularly for bpMRI dataset. Subsequently, for the training of the second-stage model tasked with lesion segmentation, we utilize the output of the first model. This involves feeding the input MRIs into the initial model, which generates a mask for each modality. These predicted masks are then utilized to extract the prostate region from the original MRI data. The resulting dataset, comprising the newly delineated prostate areas, serves as the training data for a second attention-guided U-Net model tailored specifically for lesion segmentation.
To achieve automatic prostate segmentation, we employ an attention-guided U-Net architecture trained from scratch across diverse datasets, as reported in Figure 2. The attention U-Net enhances segmentation by incorporating attention gates (AGs) within U-Net's skip connections. These AGs apply an attention mechanism at each layer, calculated as:

The structural details of the attention mechanism.
Here, 
Datasets
PROSTATEx dataset: The PROSTATEx dataset, as annotated by Cuocolo et al. 47 provided zonal masks essential for prostate-related research, encompassing lesion detection and zonal segmentation. With 204 labeled examples, this dataset served as a foundational component for training the first-stage model.
Prostate158 dataset: Prostate158 dataset, 48 comprising T2WI and DWI bpMRI images, along with corresponding masks, contributed 158 examples for training the initial model.
MSD prostate dataset: The MSD prostate dataset, sourced from Radboud University Medical Center, 49 encompassed multiparametric MRIs consisting of T2WI, DWI, and DCE series, focusing on the prostate's PZ and transition zone. Utilizing the 32 cases available in the public dataset, we integrated this resource into the training data for the first-stage model.
Multisite prostate segmentation dataset: The multisite prostate MRI segmentation dataset 50 was specifically curated to facilitate generalization in prostate segmentation, multisite learning, and lifelong learning. Collected from six different data sources, this dataset predominantly comprises T2w MRIs featuring six distinct image variations, namely RUNMC, I2CVB, BIDMC, BMC, HK, and UCL.
Initially, our model was trained with all variations; however, we observed that certain variations hindered model performance. Consequently, we refined our approach by retraining the model exclusively with RUNMC and I2CVB variations to optimize its performance. In total 49 images were used for training the initial model.
The PI-CAI challenge, a multinational comparative study, examining independently developed AI models using a large multicenter cohort of 10,207 patient exams. Preliminary findings suggest that even with training on just 1500 cases, AI models can achieve diagnostic accuracy on par with radiologists reported in literature. The PI-CAI study protocol, established in collaboration with 16 experts across prostate radiology, urology, and AI, involved the retrospective analysis of prostate MRI exams from four European tertiary care centers. This encompassed imaging data from 9129 patients suspected of harboring PCa, acquired using diverse MRI scanners. The challenge invites global researchers to design models for detecting csPCa in bpMRI using 1500 training cases. Within the public training and development dataset, comprising 1500 cases, 328 cases originate from the PROSTATEx Challenge. Notably, 1075 cases exhibit benign tissue or indolent PCa, labeled as all zero, while 220 malignant cases are manually labeled by trained investigators or radiology residents, under expert supervision. An additional 205 malignant cases are labeled by an AI model. Each patient is provided with bpMRI scans, including Axial T2WI, Axial hbv DWI, and axial ADC, acquired using Siemens Healthineers or Philips Medical Systems-based scanners with surface coils. All annotations are harmonized to match the dimensions and spatial resolution of corresponding T2WI images.
Data preprocessing
For training the Stage 1 model several preprocessing steps are carried, a total of 443 samples taken from the merged PROSTATEx, prostate158, MSD-prostate and multisite prostate data are used to train the prostate segmentation model. Preprocessing steps involve converting all datasets into a uniform NifTI format using the SimpleITK package, and standardizing the images to have the same shape across height, width, and depth dimensions across their corresponding original T2W scans. The input shape is set to (192, 192, 17), where the depth size was selected after calculating the minimum depth size out of all the examples. The spatial resolution of the images was set to 0.5 × 0.5 × 3 mm. Furthermore, images underwent min–max normalization to the (0, 1) range.
For training the Stage 2 model the same preprocessing steps are performed as that for the first stage model, the spatial dimension was set to (0.5 × 0.5 × 3) mm as mentioned in the preprocessing step for the dataset.
Data augmentation
Data augmentation is applied to increase the training examples and make the model generalize to unseen out of place samples during inference. Augmentation is applied in a randomized way, a 20° rotation is applied, a horizontal flip and some random noise is applied to the input image but not to the mask, as shown in Figure 3.

(First row) original image, (second row) horizontal flip, and (third row) rotation 20°.
Model architecture and training
When training the prostate segmentation model, we have used a train-test split ratio of 5:1 to make the model as generalized as possible without overfitting. The model is a U-Net architecture with multiple encoder blocks, a bottleneck block and multiple decoder blocks. Skip-connections connect the learning from the encoder blocks to the decoder blocks, before the residual learning is passed to the decoder blocks, we apply an attention block to focus on important parts in the residual learning. The incorporation of the attention mechanism greatly improves the model's performance by enabling it to concentrate on the most pertinent regions of the input images, hence enhancing feature selection and overall accuracy. Within the framework of the attention-guided U-Net architecture, AGs are incorporated into the skip connections, allowing the model to dynamically assess the significance of various characteristics throughout the training phase. The selective focus employed by the model allows it to prioritize crucial anatomical features and lesions while reducing the impact of extraneous background information. This is especially advantageous in intricate medical imaging tasks such as PCa identification. Consequently, the model attains greater accuracy in segmenting and classifying, as demonstrated by its superior performance metrics in comparison to the baseline models. Adam is used as the optimizer and the initial learning rate was set to 0.01 gradually decreasing on plateau. The loss function used is the dice coefficient loss for segmentation of the prostate zone. The maximum epoch is set to 20 and the best results for test data was achieved during epoch 15.
When training the prostate lesion segmentation model, we used a five-fold cross validation split provided by the dataset organizer. It is the same U-Net architecture as the one used for prostate segmentation with multiple encoder blocks, a bottleneck block and multiple decoder blocks. Skip-connections connect the learning from the encoder blocks to the decoder blocks, before the residual learning is passed to the decoder blocks, we apply an attention block to focus on important parts in the residual learning. Adam is used as the optimizer where the initial learning rate was set to 0.01 gradually decreasing on plateau. The loss function used is the cross-entropy loss and dice coefficient loss for the segmentation of the lesions. The maximum epoch is set to 200.
Result analysis
Dice similarity coefficient (DSC) and IoU51,52 are utilized for evaluating the outcomes of the prostate segmentation model. DSC typically ranges from 0 to 1, with 1 indicating perfection, these metrics compare ground truth data with predicted data. It measures the overlap between two segmentations, computed as twice the volume of their intersection divided by the sum of their volumes, usually aggregated across patients using the mean. The IoU assesses the overlap between two bounding boxes, calculated as the volume of their intersection divided by the volume of their union.
The metrics utilized for evaluating the outcomes of the lesion segmentation model are DSC, sensitivity, specificity, and AUROC. These measures can be adjusted based on the confidence threshold associated with each prediction, affecting the sensitivity-specificity balance. The ROC curve summarizes sensitivity and specificity across various threshold values, representing the AUC of sensitivity plotted against 1-specificity. The assessment of patient-level diagnosis performance employs the AUROC metric, while the evaluation of lesion-level detection performance relies on the average precision (AP) metric. The composite score utilized to rank the Stage 2 model is the mean of both task-specific metrics.
Regarding the prostate zonal segmentation model, developed specifically to automate the PCa detection system, the results for the combined dataset are presented in Table 1 including dice similarity score, IoU. Figure 4 shows the plot of dice similarity score over training phase of the model and it is noticed that the scores ranged from 0.70 to 0.75. More information during the training process can be found in the Supplemental Materials.

Dice-coefficient score during the training process.
Results for prostate segmentation model.

The performance of patient-based diagnosis and lesion-level detection on the testing set is detailed in Table 2, respectively. For each combination of the 3D CNN models, we observed improvements in performance compared to the other baseline models. Table 2 shows the performance of the attention U-Net model and the other baselines on the test data in terms of the AP and AUROC metric. Figure 5 illustrates that the best-performing model on the test set is the U-Net architecture enhanced with AGs within the skip connections, regularized with batch normalization and dropout. As a baseline, we also evaluated a pretrained 3D ResNet201 model with an ImageNet backbone, which achieved the second-best performance. Additionally, a pretrained 3D VGG19 with an ImageNet backbone performed slightly below the ResNet201, while the pretrained 3D SEResNet152 model with an ImageNet backbone had the lowest performance among the models tested.

(1) Precision and recall. (2) AUC curve for attention U-Net and baselines.
Results for tumor segmentation model.

DSC: dice similarity coefficient; AP: average precision.
For a more rigorous analysis, additional experiments have been conducted by showing the specific DSC values for the central zone (CZ) and PZ to understand the model's performance on different prostate zones. For a more comprehensive assessment of segmentation quality, additional metrics have also been considered including Hausdorff Distance (HD95) and relative volume distance (RVD). Table 3 showcases the results.
Results for different prostate zones.

DSC: dice similarity coefficient; RVD: relative volume distance.
Table 3 presents the segmentation performance of the model in different prostate zones: the CZ and the PZ. The DSC shows better performance in the CZ (0.75) compared to the PZ (0.70), indicating a higher overlap between predicted and ground truth segmentations in the CZ. HD95 is lower for the CZ (4.5 mm) than for the PZ (5.2 mm), suggesting that the model achieves closer boundary alignment in the CZ. Additionally, the RVD is smaller in the CZ (0.08) than in the PZ (0.12), indicating that the predicted volumes are more accurate in the CZ. Overall, the model performs better in the CZ compared to the PZ, which is typically more challenging to segment accurately.
Moreover, a statistical analysis has been conducted showing
Statistical analysis across different methods with our method.

Qualitative results
Figure 6 shows the output of the model evaluated on the output generated by the first model. The qualitative analysis of Figures 6 and 7 offers additional insights into the model's performance. The visual outputs of the prostate segmentation and lesion detection models reveal that our attention-guided U-Net can accurately delineate the prostate zones and identify clinically significant lesions. These qualitative results complement the quantitative metrics, providing a holistic view of the model's capabilities.

(First row) Original image (second row) ground truth mask (third row) predicted mask.

(First row) Original image, (second row) ground truth, and (third row) prediction with the mask to extract the prostate ROI.
Comparison with existing literatures
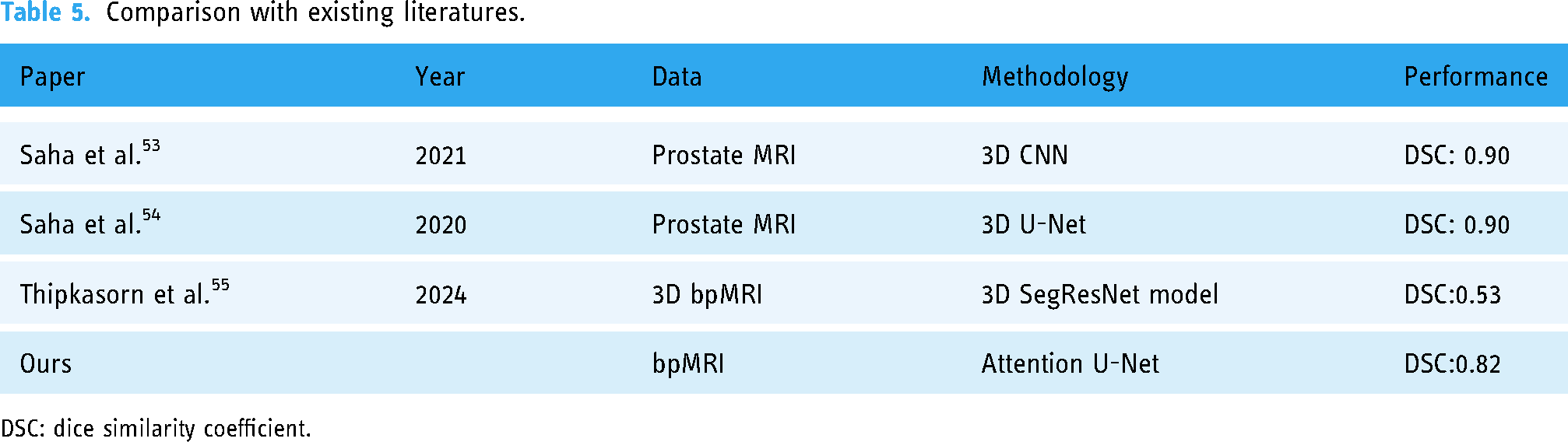
Table 5 summarizes the performance comparison of some existing literatures conducted in the similar task using MRI.
Comparison with existing literatures.
DSC: dice similarity coefficient.
Our 2D attention U-Net has several distinct advantages as compared to previous 3D models. Although Saha et al. produced a DSC of 0.90 using 3D CNNs and U-Nets, our 2D attention U-Net, with a DSC of 0.82, shows comparable performance despite the difference in dimensions. An important advantage of our 2D model is its decreased computational complexity and memory demands, enabling more efficient training and inference without significantly compromising accuracy. This is especially crucial in clinical environments where prompt decision-making is vital. Furthermore, our research utilizes a merged dataset approach, where data from many sources are combined. This strategy improves the model's capacity to generalize across diverse patient populations and imaging situations. This strategy reduces the risk of overfitting that may arise when working with smaller and less varied datasets, therefore enhancing the resilience and dependability of the model in real-world medical applications. Our performance meets the clinical requirements necessary for decision-making. Despite being slightly lower than many 3D models, the DSC of 0.82 is still within a useful range for clinical applications. This is especially true when considering the tradeoff between accuracy and computing economy. The balance between performance and resource management is essential for the successful integration of the model into real-world processes, as it is necessary for timely and accurate diagnosis.
Ablation study
In this study, we included an ablation analysis to compare the performance of different model configurations, emphasizing the impact of our proposed two-stage attention-guided U-Net. The
As reported in Table 6, the results demonstrate that the two-stage approach, particularly with the inclusion of attention mechanisms, significantly enhances segmentation accuracy. The two-stage attention-guided U-Net achieved a DSC of 0.82, an AUROC of 0.85, and an AP of 0.80, outperforming other models. In comparison, the one-stage U-Net and one-stage attention U-Net models achieved DSCs of 0.63 and 0.64, respectively, indicating that a single-step process is less effective for precise segmentation. Additionally, the two-stage U-Net without attention yielded a DSC of 0.70, reinforcing the conclusion that the combination of a two-stage architecture and attention mechanisms is crucial for improved segmentation and lesion detection in PCa. This ablation study underscores the advantage of incorporating zonal awareness and attention mechanisms in deep learning models for medical imaging tasks.
Ablation study on the two-stage and attention component of proposed model.

DSC: dice similarity coefficient; AP: average precision.
Discussion
This study aimed to enhance the accuracy of PCa detection in bpMRI by integrating zonal awareness into an attention-guided U-Net architecture. Our findings suggest that our model performs well in segmenting the prostate zones, a critical step in improving the accuracy of subsequent lesion detection models. The qualitative results, illustrated in Figure 7, provide visual confirmation of the model's effectiveness in various bpMRI images. Our model consistently outperformed the pretrained baseline architectures, with the attention-guided U-Net achieving the best test set results. Specifically, the U-Net architecture with AGs and regularization techniques such as Batch Normalization and dropout demonstrated superior performance in both patient-based diagnosis and lesion-level detection.
The baseline models provided a benchmark, and the enhancements achieved by our attention-guided U-Net highlight the importance of integrating domain-specific anatomical knowledge. The pre-trained models, while effective to some extent, could not match the performance of a model specifically tailored and trained for the task at hand. The attention mechanisms incorporated in our U-Net architecture proved particularly beneficial in focusing on relevant areas of the images, thereby improving the segmentation accuracy. The high scores of our model across a wide range of evaluation metrics indicate its robustness and reliability in clinical applications. The improvements observed in the attention-guided U-Net model over the baseline models underscore the value of custom architectures tailored to specific medical imaging tasks.
Validation of the proposed method was conducted using several datasets specifically designed for cancer segmentation in bpMRI. In order to ensure the robustness and applicability of our findings, we utilized a wide array of datasets that encompassed differences in patient characteristics, types of tumors, and imaging settings. To conduct the training and testing stages, we adopted a split ratio of 70:30. Consequently, 70% of the data was allocated for training the model, enabling it to acquire knowledge from a wide range of situations, while the remaining 30% was set aside for testing, so ensuring a dependable assessment of the model's performance on unfamiliar data. This methodology not only improves the model's capacity to generalize to many situations but also guarantees that it is evaluated against a representative sample of the varied medical conditions found in clinical practice. Through meticulous validation of the model in this way, our objective is to enhance its validity and dependability, so leading to improved results in the diagnosis and planning of therapy for PCa. Histology is key to the justification for developing and validating the suggested instrument in the proposed study. Radiology, especially bpMRI pictures, provides noninvasive insights into prostate architecture and probable pathology, but histology provides the ultimate diagnosis by studying the lesions’ cellular and tissue-level features. Histological data is used in the automated approach to ground imaging findings in biology. Histology validates prostate lesion presence, nature, and grade, which affects imaging-based diagnostic tool accuracy and reliability. Training and validating the attention-guided U-Net model against histology outcomes, the most accurate diagnosis, is possible. This improves diagnostic performance by detecting anomalies with high sensitivity and matching tissue pathology to expectations. Histological data can also reveal prostate lesion heterogeneity, helping the model learn more about cancer types and situations. These findings can improve segmentation, resulting in more accurate and therapeutically relevant predictions. Radiology is a noninvasive way to discover potential abnormalities, but histology is the gold standard for diagnosis. Integrating histological insights into your model ensures accuracy and reliability, boosting clinical decision-making.
Limitations and future work
While our model demonstrates significant improvements, it is important to acknowledge certain limitations. The requirement for manual localization of ROIs in some of the baseline approaches remains a constraint. Additionally, the model's performance, although robust, may vary with different MRI scanners and imaging protocols. Future work should focus on further refining the model to enhance its generalizability across diverse datasets and imaging conditions. Incorporating additional data augmentation techniques and exploring advanced neural network architectures could further improve performance.
Conclusion
This study demonstrates a substantial advancement in the identification of PCa using bpMRI, by incorporating zonal awareness into the attention-guided U-Net architecture. Our research not only tackles the complex task of precisely diagnosing and categorizing PCa but also presents a new method that utilizes a detailed understanding of the prostate's zonal structure to improve the accuracy of diagnosis. The incorporation of zonal awareness into deep learning models for medical imaging, namely in the segmentation of PCa, has the potential to significantly enhance the precision and dependability of diagnostic procedures. Our model's efficacy in detecting csPCa is superior because of its incorporation of anatomical knowledge of the prostate gland. This superiority has been proven by comprehensive testing across multiple bpMRI datasets. This progress highlights the possibility of combining deep learning technologies with domain-specific knowledge to expand the limits of personalized medicine and patient-specific diagnostics.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076251314546 - Supplemental material for Enhancing prostate cancer segmentation in bpMRI: Integrating zonal awareness into attention-guided U-Net
Supplemental material, sj-docx-1-dhj-10.1177_20552076251314546 for Enhancing prostate cancer segmentation in bpMRI: Integrating zonal awareness into attention-guided U-Net by Chao Wei, Zheng Liu, Yibo Zhang and Lianhui Fan in DIGITAL HEALTH
Footnotes
Acknowledgments
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author contributions
Chao Wei (first author): conceptualization, methodology, data curation, software, formal analysis, and writing—original draft and editing. Lianhui Fan (corresponding author): project administration, conceptualization, supervision, and writing—review and editing. Zheng Liu: validation, review, and editing. Lianhui Fan: supervision, validation, and review and editing. Yibo Zhang: writing—original draft and editing. Zheng Liu: visualization and writing—original draft and editing. Lianhui Fan (corresponding author): validation and writing—review and editing. Yibo Zhang: conceptualization and validation.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
