Abstract
Background and Objective
The rapid development of computer technology has led to a revolutionary transformation in artificial intelligence (AI)-assisted healthcare. The integration of whole-slide imaging technology with AI algorithms has facilitated the development of digital pathology for lung cancer (LC). However, there is a lack of comprehensive scientometric analysis in this field.
Methods
A bibliometric analysis was conducted on 197 publications related to digital pathology in LC from 502 institutions across 39 countries, published in 97 academic journals in the Web of Science Core Collection between 2004 and 2023.
Results
Our analysis has identified the United States and China as the primary research nations in the field of digital pathology in LC. However, it is important to note that the current research primarily consists of independent studies among countries, emphasizing the necessity of strengthening academic collaboration and data sharing between nations. The current focus and challenge of research related to digital pathology in LC lie in enhancing the accuracy of classification and prediction through improved deep learning algorithms. The integration of multi-omics studies presents a promising future research direction. Additionally, researchers are increasingly exploring the application of digital pathology in immunotherapy for LC patients.
Conclusions
In conclusion, this study provides a comprehensive knowledge framework for digital pathology in LC, highlighting research trends, hotspots, and gaps in this field. It also provides a theoretical basis for the application of AI in clinical decision-making for LC patients.
Introduction
Lung cancer (LC) is responsible for the highest number of cancer-related fatalities worldwide, with more than 1.8 million people succumbing to the disease each year.1–3 Timely and accurate diagnosis is essential for increasing the likelihood of a positive prognosis for LC patients. However, the current standard method for histopathological evaluation of LC based on tissue slices requires pathologists to identify microscopic histopathological features in highly complex tissue images. This time-consuming and subjective process can produce inter- and intra-observer variation.4,5 The emergence of digital pathology provides a new solution to this problem that shows promise in simplifying the diagnostic process and reducing the risk of error and variation. Digital pathology is based on deep learning or machine learning methods to extract features from pathological images of diseases, characterizing various attributes of the disease for precision treatment. Now, digital pathology has been widely used in cancer diagnosis,6–11 pathology grading,6–12 prognosis prediction,13–18 and efficacy evaluation. 19
In recent years, there have been significant and beneficial explorations in the field of digital pathology in LC. With the continuous emergence of new discoveries, the understanding of literature and attention to the forefront of research related to digital pathology in LC have also been progressively improving. 20 However, there is still a lack of global and comprehensive reports summarizing the current research status of this filed. Bibliometric analysis provides a broad and objective overview, enabling researchers to quickly map out the research landscape, identify the current state and key trends, patterns of multidisciplinary integration and collaboration, as well as knowledge gaps in the field of digital pathology in LC. It has now been applied in the fields of comparison between segmental resection and lobectomy, 21 triptolide treatment of non-small cell LC 22 and anti-PD-1/PD-L1 treatment of LC 23 and has promoted the sorting of knowledge systems in these fields to a certain extent.
Here, we collected studies related to digital pathology in LC from the Web of Science Core Collection (WoSCC) database and conducted bibliometric analysis to visualize current research findings, hotspots, frontiers and gaps, which will provide valuable information for researchers and institutions to identify meaningful research directions and provide references for the application of artificial intelligence (AI) in clinical decision-making in LC.
Methods
Data collection
A literature search was performed on the WoSCC database from 1 January 2004 to 31 December 2023. The search formula was as follows: (cancer OR carcinoma OR adenocarcinoma OR tumour OR tumor OR malignanc* OR neoplas*) AND (lung OR pulmonary OR respiratory OR respiration OR aspiration OR bronchi* OR alveol* OR pneumocytes OR “air way” OR NSCLC) AND (whole-slide OR “whole slide” OR “WSI” OR “pathology image” OR “pathology imaging” OR “digital pathology” OR “histopathology image” OR pathomics OR “digital pathology” OR WSI OR “artificial intelligence” OR “pattern recognition” OR “convolutional neural networks” OR “artificial neural networks”). The document was limited to articles and reviews and was downloaded in plain text format. Three authors independently conducted the search and screening. In case of any discrepancies among them, the fourth author made the final decision.
Inclusion and exclusion criteria
The inclusion criteria were as follows: (1) studies related to LC; (2) research related to digital pathology; and (3) literatures were written in English.
The following records were excluded: (1) literature unrelated to pathology; (2) literature unrelated to LC; (3) literature mentioning LC and pathology, but not as the main subject; (4) meeting abstracts, proceeding papers, book chapters, etc.; and (5) literature not written in English.
Research methods
VOSviewer (version 1.6.18), CiteSpace (version 6.1.3), and the Bibliometrix package (version 4.2.1) in R were used to perform the bibliometric analysis. VOSviewer is mainly used for co-occurrence analysis of countries/regions, institutions, journals, authors and references. 24 CiteSpace is used to conduct double map superposition for journals, cluster analysis, timeline distribution and emergence analysis for references and keywords, and then analyze the structure, rules, and distribution of scientific knowledge. 25 The Bibliometrix package provides a web interface for comprehensive scientific mapping analysis for the distribution of documents over the years and countries/regions. 26
Results
A total of 197 articles were finally included in this study (Figure 1). Full records including main text, references, publication year, country/region, institution, source publication, research direction, funding institution, author, and other relevant information of each literature are exported in the form of plain text for further analysis.
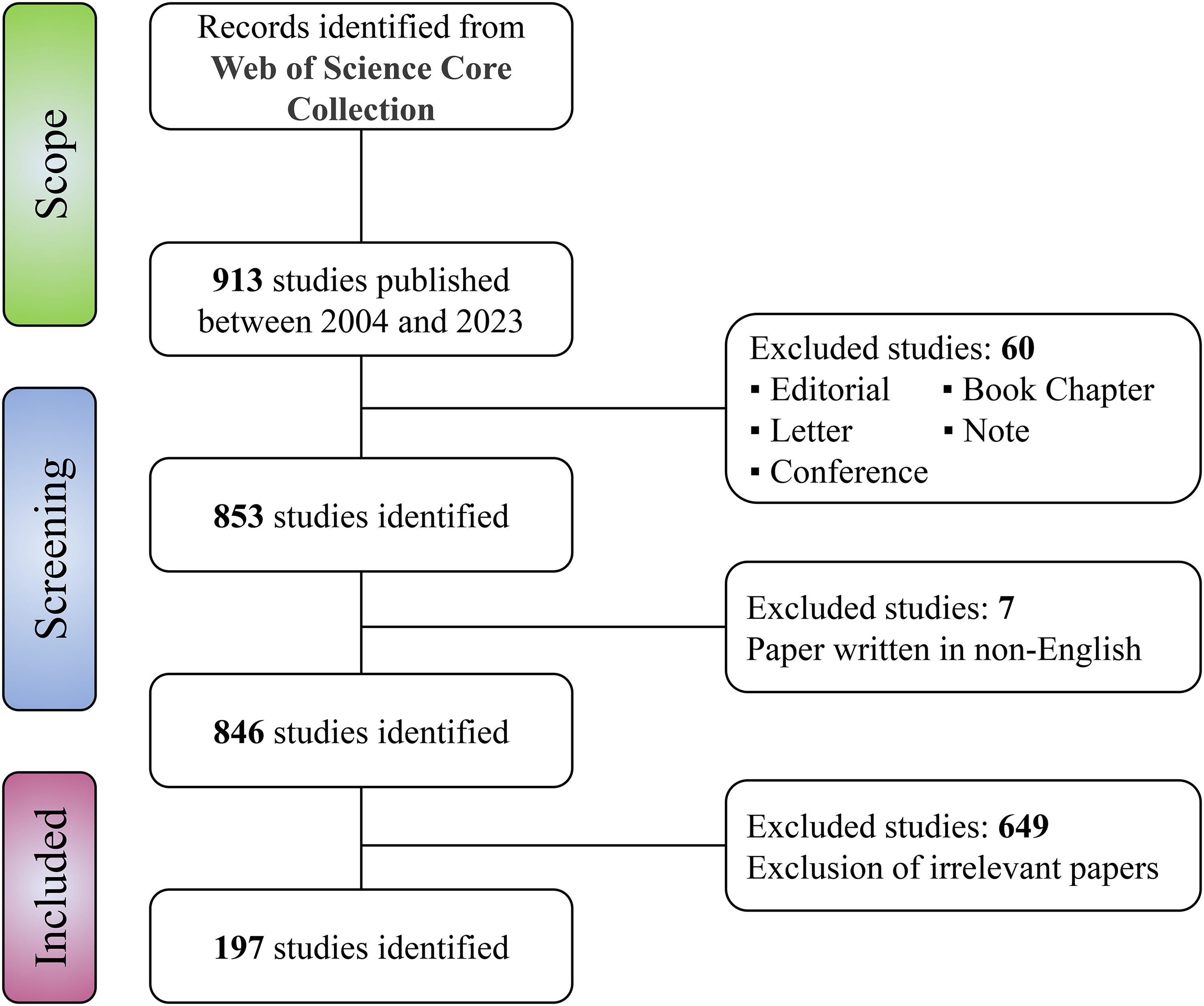
Flow chart of publication screening.
Global publishing trends
It shows the annual number of publications and citations per year in the field of digital pathology in LC (Supplemental Figure S1). The first article was published in 2004; however, until before 2016, only a very small number of studies were published in this field. From 2016, the number of publications increased significantly, initiating a prosperous period in the field of LC pathology: from five publications in 2016 to 45 in 2021. The largest increase occurred in 2021, with a growth rate of 136.8%, indicating the emergence of a substantial number of studies based on several years of research accumulation. It is noteworthy that while the number of publications has been increasing, citation patterns may not always follow a linear trajectory. In 2018, a peak in annual citation was recorded. This may be attributed to breakthrough research published in 2018, which played a significant role in advancing the development of the field and was widely cited. In summary, these findings indicate that research of digital pathology in LC has become a focus of attention for pathologists and has entered a period of rapid development.
International cooperation between influential countries/regions
The spatial distribution of research of digital pathology in LC reflects the influence and cooperation of various countries. A total of 39 countries have contributed to publications in this field. Figure 2(a) presents the top five countries with the highest publication output and corresponding citation counts. The United States leads with 69 publications and 4090 citations, followed by China with 69 publications and 976 citations. Japan, England, and India follow with 19, 16, and 13 publications respectively, with England having the third-highest citation count of 563, while Japan and India have received 333 and 74 citations respectively. These findings provide a comprehensive understanding of the global research landscape of digital pathology in LC, highlighting the leading countries and their impact in the field.

Publication status of researches related to digital pathology in lung cancer. (a) The annual number of scientific productions on digital pathology for lung cancer from 2004 to 2023. (b) Clustering of collaboration among countries. Larger nodes indicate more publications.
Country network visualizes collaborations and research trends at a global level. By analyzing this network, readers can discern international collaborations, predominant countries in the research field, and global research patterns. The analysis of co-authorship included all 39 countries that contributed to the publication. However, three countries were excluded due to their lack of connection with other countries. Ultimately, a network of 36 countries was used to analyze the collaboration among nations (Figure 2(b)). The United States had the highest total link strength with 51 times, followed by England with 35 times, indicating their dominance in this research field. Nevertheless, it is noteworthy that research related to digital pathology in LC is still predominantly conducted independently by each country, and international collaboration remains limited. The dominance of a few countries and the lack of broad-based international cooperation can lead to the creation of knowledge silos, where similar approaches and perspectives circulate in a closed loop, potentially stifling innovation and the diversity of research methods. Cooperation at the international level can promote innovation and development in the field, accelerating the application of artificial intelligence technologies in clinical decision-making for lung cancer. Therefore, strengthening cooperation between countries is very necessary.
International cooperation between institutions
Construction of a collaboration network in the field of digital pathology in LC facilitates the comprehension of international cooperation among prominent institutions and authors. In this study, 502 institutions that have contributed to research related to digital pathology in LC were identified, and a collaboration network mapping analysis was conducted using VOSviewer (Supplemental Figure S2). Specifically, by setting a minimum publication threshold of two, we included 123 eligible institutions. The top three most productive institutions are the University of Texas MD Anderson Cancer Center, South China University of Technology, and the University of Texas Southwestern Medical Center at Dallas. These institutions form close collaborative groups and hold a leading position in the field of digital pathology in LC. Likewise, institutions from China have also formed their own collaborative groups. From a temporal perspective, research groups from the United States are more mature in this field, with greater research impact, while collaborative groups from China and other Asian countries are relatively younger and more active.
Recognition of high-impact journal
As of now, articles related to digital pathology in LC are distributed among 97 different journals. We extracted core journals through source clustering analysis based on Bradford's Law. Bradford's Law indicates that if scientific journals are ranked in descending order of the number of papers published on a specific subject, the core part with the highest article rate and the subsequent part with the same number of articles as the core part can be distinguished, and the number of journals in each part is in a ratio of n:n1:n2:n3 (n > 1). Table 1 lists the basic information and recent impact factor (IF) of these core journals. Approximately 35.03% of publications in the field of digital pathology in LC are published in these journals, which are mainly classified into the fields of oncology, pathology, computer and information systems science, and engineering biology. All journals are classified as Q1 or Q2 in the Journal Citation Reports, with Q1 journals being the majority. Additionally, we found that these eight core journals are all high-impact journals (defined as IF greater than 3), indicating that research of digital pathology in LC can be recognized by outstanding journals.
The core journals publishing researches related to digital pathology in lung cancer.

IF: impact factor; JCR: Journal Citation Reports.
Identification of most active authors
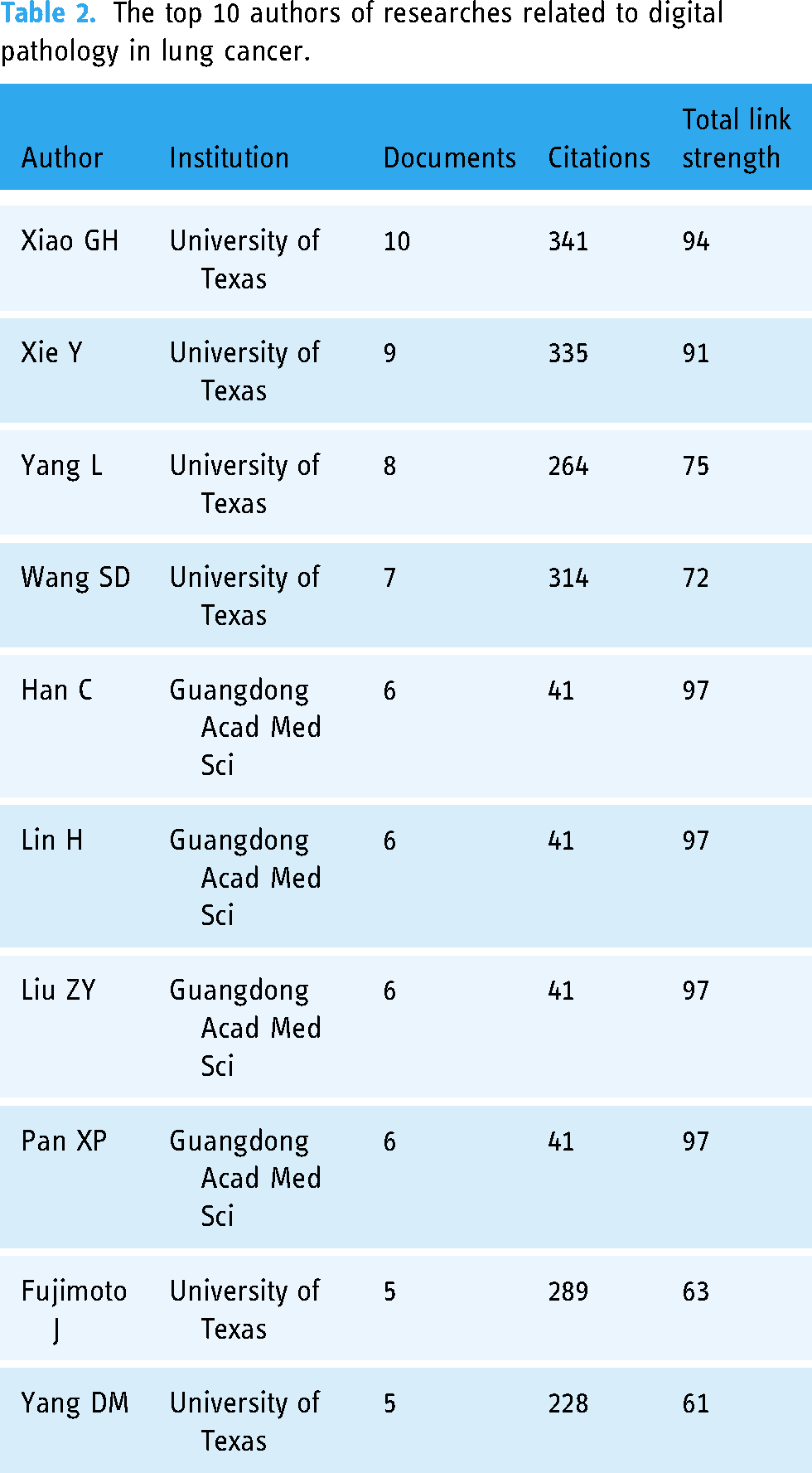
A total of 1330 authors were involved in the 197 studies. As shown in Table 2, the top 10 active authors contributed a total of 68 publications, accounting for 34.52% of the total number of studies. It is worth noting that six of these authors are from the University of Texas, an active institution in the field of digital pathology in LC. Additionally, the other four authors are also from Guangdong Acad Med Sci, another active institution in the field. The co-authorship analysis map shows that the research of digital pathology in LC is still dominated by single research groups (Supplemental Figure S3). The co-author clusters formed are mostly from the same institution or research team, and a few studies involving multiple institutions are often led by a single institution, with other institutions serving as affiliates. Currently, collaborative research involving multiple centers or research groups in this field is extremely scarce.
The top 10 authors of researches related to digital pathology in lung cancer.
Identification of articles with most local citations
Local citations are not global citations across different fields, but citations within a specific subset of the literature, such as within the field of digital pathology in LC. Through the analysis of local citations, we can identify the most influential studies on the direction and development of digital pathology in LC. The local citations score (LCS) was used to represent the number of citations from local sources, reflecting the internal citation volume of the field of digital pathology in LC. The LCS measurement provides insights into the active development of this field within the research community. Figure 3(a) displays the top ten articles with the highest LCS, a metric indicating the number of citations from local sources. The study conducted by Coudray N et al. 27 in 2018 obtained the highest LCS score of 73, signifying a notable level of citation within the geographic region of the study. The authors developed a multi-task classification Convolutional Neural Networks (CNNs) model to predict the ten most common genetic mutations in H&E-stained whole-slide images, including frequently mutated genes in LC tissue such as STK11, EGFR, FAT1, SETBP1, KRAS, and TP53. This study was the first to apply AI techniques to predict gene mutations from histopathological tissue sections.
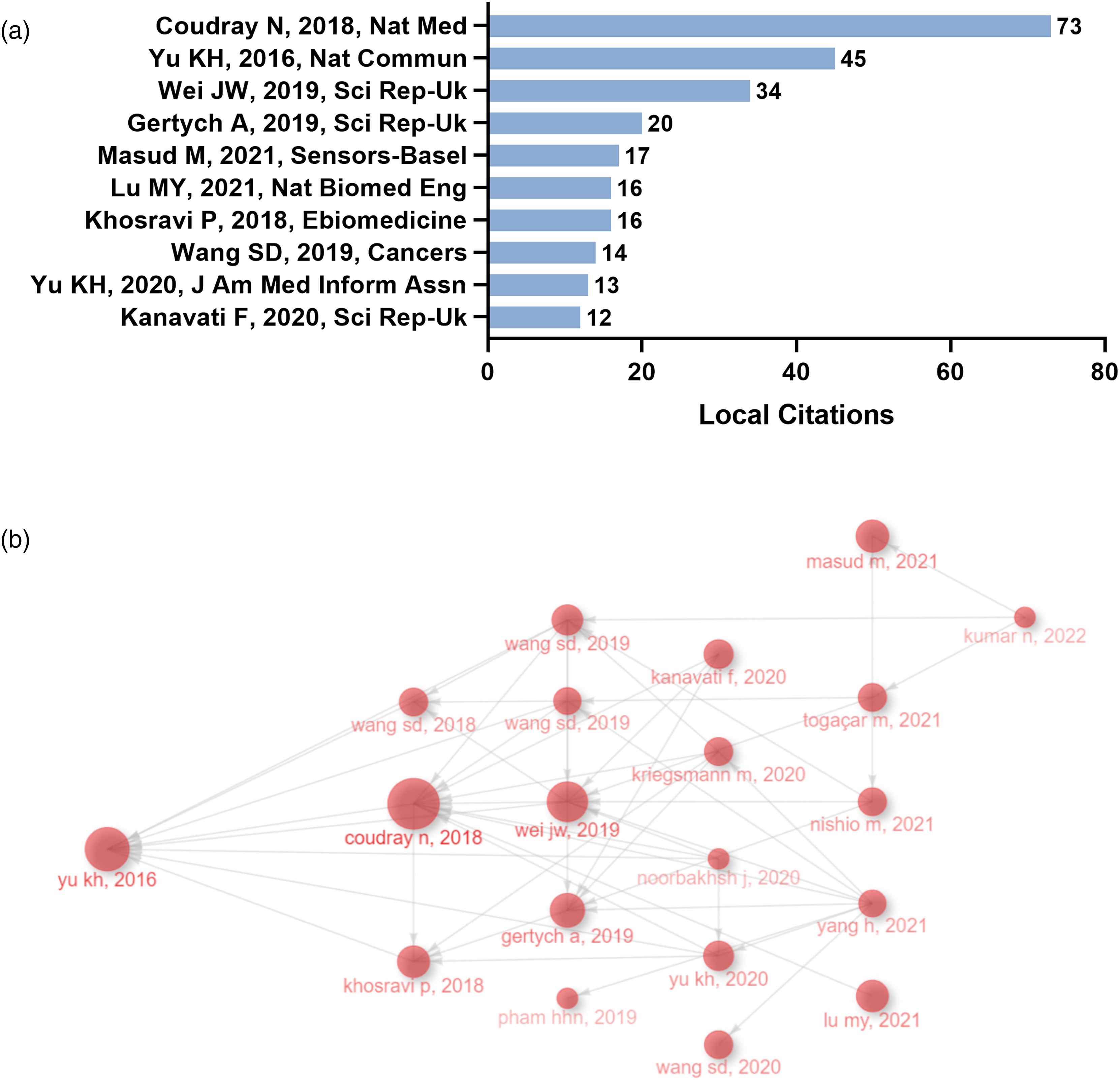
Publications with the highest number of local citations and academic inheritance. (a) The top 10 publications with the highest local citations. (b) Historiograph Map illustrating the academic inheritance relationships among publications in the field of digital pathology in lung cancer. Each node represents a literature, and the size of the node represents the local citation score of the literature.
To investigate the academic inheritance relationships in the field of digital pathology in LC, we plotted a historiograph map arranged based on the publish year (Figure 3(b)). Each node represents a literature, and the size of the node represents the local citation score of the literature. The connecting lines between two nodes represent the inheritance relationship between two studies. The study by Yu KH et al. 28 in 2016 was the earliest and most pioneering study in the field of digital pathology in LC, which promoted and opened up the prosperity of the field. This study proposed the use of quantitative tissue morphology to analyze the local anatomical structure and overall characteristics of cancer cells, and combined these automatically extracted features with machine learning classifiers to develop a prognostic model for early stage non-small cell LC. Furthermore, Wei JW et al., 29 Khosravi P et al., 30 and Gertych A et al. 31 have addressed the issues of multi-label classification in digital pathology in LC, the identification of cancer subtypes and transferability in the presence of extensive tumor heterogeneity using CNN, and the identification of four different growth patterns (acinar, micropapillary, solid, and lepidic) of lung adenocarcinoma in surgical resection specimens using CNN. In summary, these innovative studies have made outstanding contributions to the advancement of computational pathology in the diagnosis and prediction of LC.
Research hotspots and emerging trends
Temporal evolution of research hotspots according to co-cited references
Co-cited references not only provide core literature that plays a crucial role in the evolution of a field but also reveal the changing trends of research hotspots. 32 Figure 4(a) presents a density visualization Map that highlights the 62 most frequently cited references, with citation counts of 10 or higher, where references with darker colors and larger fonts are considered more important. Many co-cited references are from journals in pathology and computer science. The study published by Coudray N et al. 27 in 2018 has the highest number of citations, being cited 73 times, followed closely by the study of Yu KH et al. 28 published in 2016, which has been cited 45 times. Notably, these two articles also have the highest LCS among all analyzed publications. Additionally, a timeline map was constructed to show the significant clusters of co-cited references (Figure 4(b)). In terms of temporal distribution, most clusters are concentrated between 2015 and 2022. Among these clusters, cluster 7 (differential diagnosis), cluster 8 (laser capture microdissection), cluster 13 (automated morphological classification), and cluster 16 (self-organizing neural network) had earlier starting times and represented early research directions in digital pathology in LC. Clusters 0 (deep learning), 1 (tumor mutation burden), 5 (predicting non-small cell lung cancer prognosis), 6 (mutation prediction), 9 (convolutional neural network), 10 (automated tumor analysis), and 11 (supervised graph) were more mature topics. Clusters 2 (survival prediction), 3 (colon cancer), and 4 (non-small cell lung cancer) had relatively later starting times and represented emerging topics in the field of digital pathology in LC.
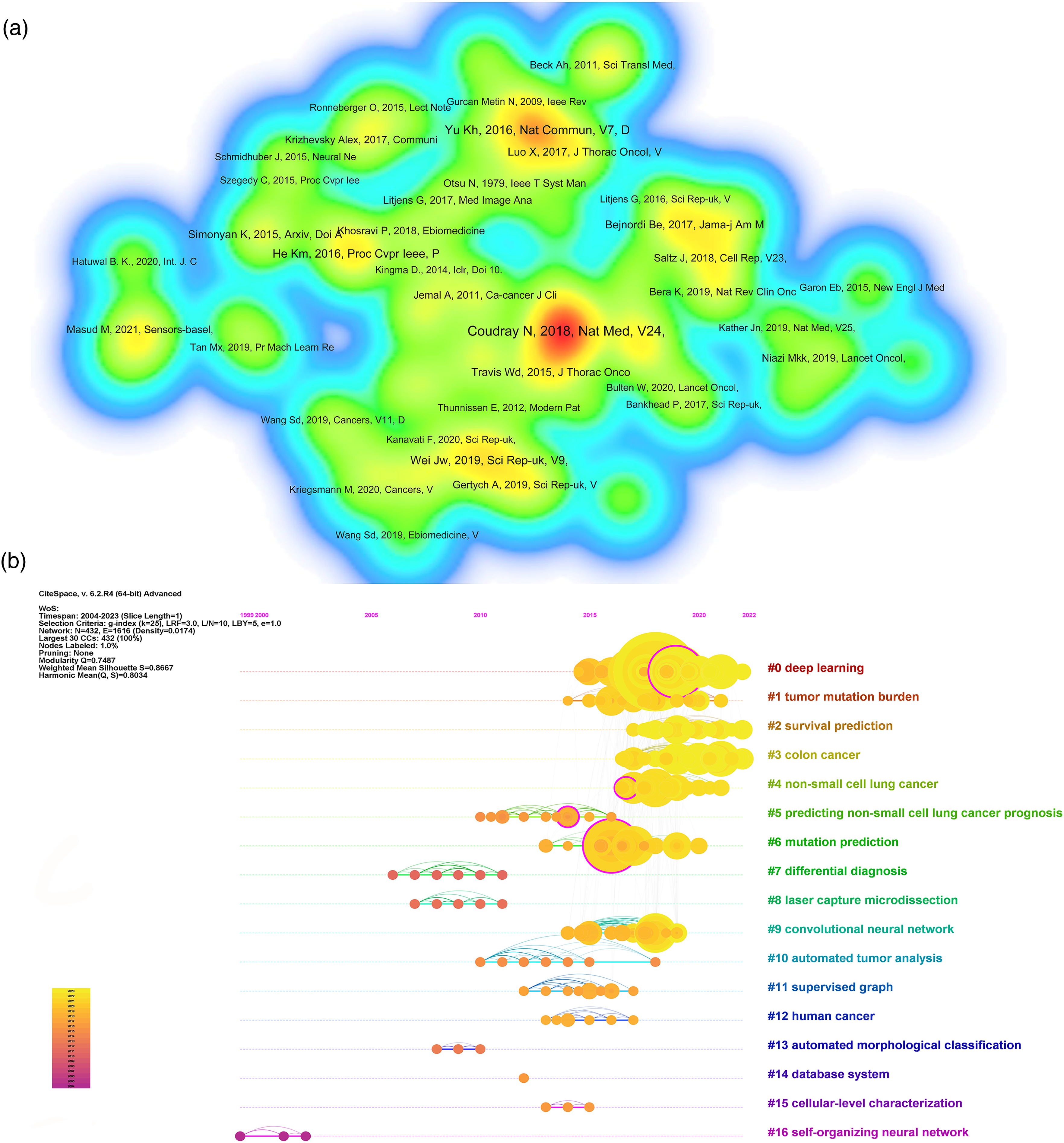
Co-cited references analysis. (a) Density visualization of co-cited references. Larger nodes and darker colors indicate higher citation counts for the corresponding publication. (b) Timeline distribution of the 14 clusters extracted from the co-cited references analysis in the field of digital pathology in lung cancer.
Evolution of research hotspots in digital pathology in LC
Keywords are closely related to each other and can form different clusters based on their affinity. We included 86 keywords with a minimum frequency of three to draw a strategic map of research hotspots (Figure 5(a)). The size of the circle reflects the frequency of the keyword, and the distance between two circles reflects the strength of their association. Keywords that are closer together are grouped into the same cluster, which can roughly reflect the main research content of the field of digital pathology in LC. The results showed that the blue cluster 1 mainly focuses on traditional pathological diagnosis methods of LC and terms related to tumor treatment, such as “diagnosis,” “image analysis,” and “therapy.” The red cluster 2 is mainly about the development and prognosis of LC, such as “prognosis” and “recurrence.” The remaining clusters 4–6 are closely related to machine learning and artificial intelligence, with the green cluster 4 being the center, which focuses on various functions of machine learning, such as “deep learning,” “neural networks,” “feature extraction,” and “classification.” The cyan cluster 5 is related to image analysis, such as “whole slide image” and “computer-aided diagnosis.” The purple cluster 6 focuses on machine learning algorithms, especially “convolutional neural networks,” “image analysis,” and “biomarkers.”
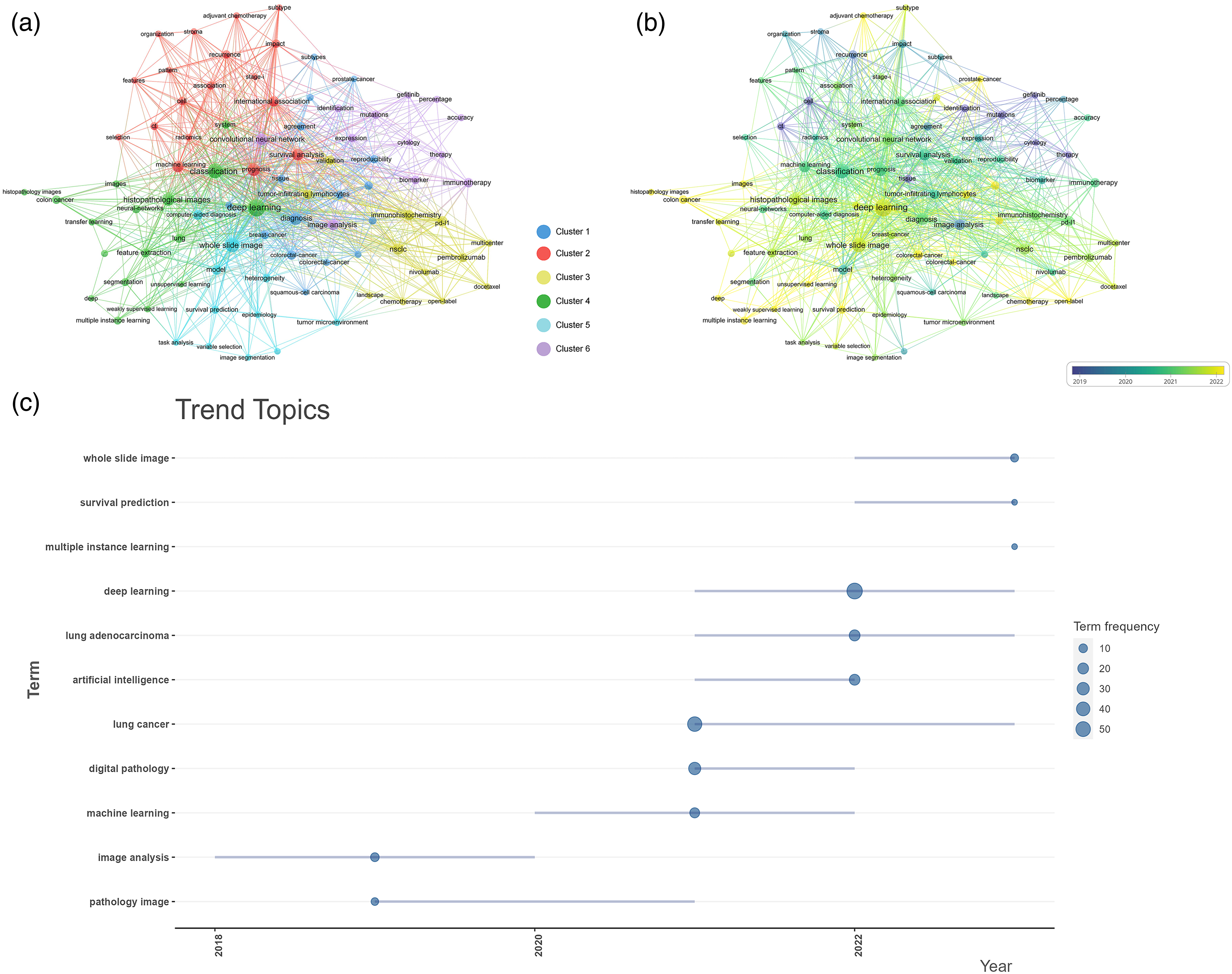
Co-occurrence network and evolution of research hotspots in researches related to digital pathology in lung cancer. (a) Network visualization of keywords. The size of the node reflects the frequency of the keyword, and the distance between two nodes reflects the strength of their association. Keywords that are closer together are grouped into the same cluster, which can roughly reflect the main research content of the field of digital pathology in lung cancer. (b) Overlay visualization of keywords. The latest keywords were highlighted in a dotted box. (c) Trend topics graph of keywords plus.
We further analyzed the changes in hot topics and constructed an overlaid time-evolving visualization map (Figure 5(b)). The early appearing keywords mainly focused on cluster 6, such as “identification,” “gefitinib,” and “therapy,” which were the main topics in the early stage. The keywords “deep learning,” “weakly supervised learning,” “transfer learning,” “PD-L1,” “survival analysis,” and “pembrolizumab” are recent hot topics, and they all cluster in clusters 3 and 4. These results indicate that research in digital pathology in LC has shifted from using image analysis for low level tasks of single tumor feature recognition to using deep learning algorithms to perform high-level tasks, including tumor type recognition, survival analysis, and immunotherapy efficacy evaluation.
In addition, we analyzed the evolution trend of research topics in digital pathology in LC and constructed a strategic coordinate map based on annual changes to capture the trend of topics in this field (Figure 5(c)). The algorithm for tumor identification based on cell structure and immunohistochemistry staining, represented by “pathology image,” is an early topic. Digital pathology based on “whole slide imaging” and “survival analysis” is currently the most mature topic. “Deep learning” is an emerging field, with CNNs being the most popular techniques in this field. In recent years, most of the research on LC digital pathology has been based on CNN algorithms. “Multiple instance learning” is a brief emerging topic, with the aim of improving the transfer learning capabilities of machine learning models.
Discussion
Since the official introduction of the concept of “artificial intelligence” in 1956, 33 it has gradually been applied in various fields. With the widespread application of AI in pathological research, the concept of “digital pathology” has emerged. As LC is the leading cause of cancer deaths worldwide, 1 its combination with digital pathology has received attention from researchers. In 2004, the first research related to digital pathology in LC was published, which used a self-organizing feature mapping neural network architecture to identify non-small cell carcinoma. 34 However, this field was quiet in the following five years, which may be attributed to the researchers still adapting to this new research field, as well as the time required to collect a large number of pathological slides. Due to advances in whole-slide imaging (WSI) and AI, this quiet period has been broken since 2010, and from 2010 to 2018, digital pathology in LC entered a stable development period. In 2018, Coudray et al. 27 developed a deep learning framework, DeepPATH, for analyzing histopathological images and validated it using WSIs of LC. They discovered that compared to pathologists, this model could accurately diagnose LC and differentiate between histological types of LC with an area under the curve at least 0.97. Additionally, the model was capable of predicting common genetic mutations in lung cancer WSIs. This research was widely cited and laid the groundwork for subsequent studies in digital pathology for LC. From 2018 to 2023, improved algorithms, hardware, and rapidly increasing available data volumes 35 have led to a rapid development of digital pathology in LC, indicating that this filed has achieved a certain level of maturity in the theoretical aspect. In addition, changes in health policy, funding allocations and the establishment of cancer research consortia may have an impact on publication trends. The emphasis on precision medicine and personalized treatment strategies may also contribute to the surge in interest.
Research on digital pathology in LC varies across different countries. United States is the most influential and productive country in this field, with citation numbers far exceeding those of any other country. LC is one of the most common diseases in the United States, providing American researchers with a large sample size for their studies. The United States has also collaborated extensively with many countries, and its researchers and institutions have been studying this field for the longest time, which is one of the reasons why the United States has published many high-quality articles. China, as the largest developing country, has published the second largest number of articles on LC pathology after the United States. This is closely related to China's recent increase in investment in scientific research in fields such as public health and the application of artificial intelligence in the medical field. Additionally, it has been observed that research in digital pathology for LC is predominantly conducted by single institutions, lacking substantive international collaboration. This can lead to knowledge silos, impede the development of artificial intelligence technology, weaken the generalization ability of AI models, and hinder the practical application of digital pathology in clinical decision-making for LC patients. Key strategies to address this issue include fostering the establishment of global research consortia, promoting open science and data sharing, building global digital sharing platforms, strategically funding collaborative projects, and increasing the visibility of diverse research.
The development and application of WSI technology is the cornerstone of the advancement of digital pathology. WSI technology involves scanning and converting entire histological and pathological glass slides into high-resolution digital images. This allows for the viewing, analyzing, and managing of these images on a computer. WSI technology has revolutionized the field of pathology by enabling more efficient workflows, facilitating remote consultations, and enhancing educational and research opportunities.36,37 Early WSI analysis primarily relied on feature engineering, where researchers extracted key features from pathological images to construct models for disease identification. 38 Manual feature extraction is a time-consuming and labor-intensive process, susceptible to individual differences and fatigue, leading to variability in judgments among pathologists, which affects the consistency and repeatability of feature extraction. Due to these limitations, coupled with the immense potential of CNNs in image recognition, researchers have shifted focus to exploring CNN-based deep learning models for WSI analysis. These models can autonomously and rapidly identify and extract features from large datasets, significantly enhancing processing speeds and efficiency, and are capable of handling a far greater number and complexity of images than manual methods. 39 This advancement substantially improves the level of intelligence in digital pathology for clinical decision-making in lung cancer and fosters its application in diversified clinical tasks. 40 The development of more sophisticated and precise deep learning algorithms has thus become a central focus in scientific research. Key contributions include Yu KH et al., 28 who designed an automated workflow for LC prognosis prediction, has attracted attention in the field of digital pathology in LC and promoted its prosperity. Other significant studies by Coudray N et al., 27 Simonyan K et al., 41 and Wei JW et al., 29 who employed deep convolutional neural networks for LC classification, prognosis prediction and mutation prediction. Moreover, He KM et al. 42 constructed a residual learning framework, and Campanella G et al. 43 used a deep learning system with multi-instance learning to evaluate and classify entire slide images of cancer pathology, which may have great potential in the diagnosis, treatment, and prognosis of cancer. These advances hold immense potential for improving cancer diagnosis, treatment, and prognosis. However, despite these advances, deep learning algorithms face limitations, notably their “black box” nature, which obscures the understanding and interpretation of internal decision-making processes. 40 Future research must thus focus on demystifying these processes and integrating the outcomes of deep learning models with the biological characteristics of lung cancer to enhance their clinical utility.
Immunotherapy has become a key treatment method for lung cancer. 44 PD-1 and PD-L1 are key negative regulatory factors in immunotherapy, and identifying their expression in tumor tissues can be used to evaluate the efficacy of immunotherapy in patients. 45 Currently, immunohistochemical staining is the main detection method for both factors, and its interpretation mainly relies on pathologists’ evaluation of immune cell staining in tumor areas, which is inherently subjective. 46 To address this, many researchers have proposed WSIs-based immune histochemical feature prediction systems for use in immunotherapy-related fields, such as evaluating PD-1/PD-L1 expression47,48 and identifying parameters related to CD8 + cancer-infiltrating lymphocytes to assess the efficacy of immunotherapy for patients with LC. 49 However, due to the complexity of the tumor microenvironment, current research may not fully capture the full spectrum of immune responses or the heterogeneity within LC. Additionally, although WSIs provide extensive data, existing computational models may not fully utilize this data due to limitations in image analysis algorithms or computational power. Future research should focus on a more comprehensive study of the interactions among various immune cells within the tumor microenvironment, develop more advanced image analysis algorithms and computational models to more effectively utilize the data from WSIs. Additionally, multimodal models should be constructed, integrating information from digital pathology, immunology, computational biology, and clinical medicine to develop more precise and personalized immunotherapy strategies.
The integration of multiple omics studies such as pathomics, radiomics, transcriptomics, genomics, and proteomics represents a promising research frontier. 50 This area is closely related to technological advancements as well as the increasing global burden of cancer and the demand for multidisciplinary collaboration for precise patient diagnosis and treatment. The work of Yu KH et al.,28,51,52 which combines digital pathology in LC with transcriptomics and proteomics, and explores their correlations with patient outcomes, has gained international recognition. Their publications are highly cited locally, reflecting the global research community's value placed on multidisciplinary integration and collaboration in digital pathology for LC. Despite significant potential in cancer treatment, multimodal models face challenges including the complexity of integrating diverse omics data, high computational demands for processing large datasets, and the difficulty of interdisciplinary collaboration. 53 Future research should focus on developing advanced data integration techniques, creating efficient computational solutions, enhancing interdisciplinary collaboration through AI, and improving real-time data processing capabilities. These advancements are crucial for enabling precise diagnostics and personalized treatment, ultimately improving outcomes for cancer patients.
This investigation represents the inaugural bibliometric examination of the scientific corpus pertaining to digital pathology in LC, marking a significant stride in the scholarly understanding of this field. Through comprehensive analysis, this research outlines the future research trajectories and trends in the field of digital pathology in LC. These include strengthening international cooperation, improving existing deep learning algorithms to enhance the accuracy and generalization capabilities of AI models, integrating multi-omics data to advance the development of personalized medicine, and enhancing the application of digital pathology in the immunotherapy of LC patients. This study contributes to the advancement of digital pathology in the diagnosis of LC by pathologists, the prediction of lung cancer prognosis, and the assessment of treatment efficacy. It also provides a theoretical basis for clinical decision-making for LC patients. It not only charts a course for future academic efforts in the field of LC digital pathology but also aids in optimizing the strategic allocation of resources to foster areas of high potential and emerging importance. However, this study has several limitations. Firstly, the data utilized in this study comprised only articles and reviews from the WoSCC database, which may lead to some omissions. Secondly, the bibliometric analysis software employed might have overlooked certain information, as it cannot analyze the full texts of publications. Thirdly, there are inconsistencies in various aspects, such as differences in the abbreviation of author names and the use of different names by the same institution at different times. Additionally, the relatively small number of documents included in this study may not comprehensively represent all research trends in the field, potentially increasing the randomness of the results and decreasing the stability and reproducibility of the findings. Moreover, the limited number of documents may restrict our in-depth analysis of specific subtopics or newer technologies. Therefore, the conclusions of this study should be interpreted with caution and regarded as preliminary observations within the context of the current dataset.
Conclusion
The application of digital pathology in LC holds great promise for enhancing our understanding of the molecular mechanisms underlying the disease and improving prognostication. This article provides a summary and analysis of the global research trends in digital pathology in LC over the past 20 years. Overall, the trend of publications has been increasing year by year, and interest in this field is also growing. However, it is important for academic collaboration between countries to be strengthened to advance research in this field. The improvement of AI algorithms for enhancing the accuracy of pathology classification, prognosis prediction, and treatment efficacy assessment in LC remains a focal point for researchers. Immunotherapy-related studies and multi-omics research may be the future hot research directions in digital pathology in LC. By integrating the vast amounts of data generated by various omics technologies with histopathological data, digital pathology can enable a more comprehensive analysis of LC, facilitating the identification of novel biomarkers and potential therapeutic targets.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076241277735 - Supplemental material for Global bibliometric mapping of the research trends in artificial intelligence-based digital pathology for lung cancer over the past two decades
Supplemental material, sj-docx-1-dhj-10.1177_20552076241277735 for Global bibliometric mapping of the research trends in artificial intelligence-based digital pathology for lung cancer over the past two decades by Dan-dan Xiong, Rong-quan He, Zhi-guang Huang, Kun-jun Wu, Ying-yu Mo, Yue Liang, Da-ping Yang, Ying-hui Wu, Zhong-qing Tang, Zu-tuan Liao and Gang Chen in DIGITAL HEALTH
Footnotes
Abbreviations
Acknowledgements
We thank the Guangxi Zhuang Autonomous Region Clinical Medicine ResearchCenter for Molecular Pathology and Intelligent Pathology Precision Diagnosi for technical support.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author (CG) on reasonable request.
Contributorship
Conceptualization: DX and GC; Methodology: DX, RH, and ZH; Software: KW and YM; Validation: YL, DY, and YW, ZT and ZL; Formal analysis: RH and ZH; Investigation: DX, RH and ZH; Resources: KW and YM; Data curation: KW and YM; Writing—original draft preparation: DX; Writing—review & editing: GC; Visualization: KW and YM; Supervision: GC; Project administration: GC; Funding acquisition: GC. All authors reviewed and edited the manuscript and approved the final version of the manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
The Ethics Committee of the First Affiliated Hospital of Guangxi Medical University approved this study.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Guangxi Zhuang Autonomous Region Medical Health Appropriate Technology Development and Application Promotion Project (S2020031), the Guangxi Higher Education Undergraduate Teaching Reform Project (2022JGA146), the Guangxi Educational Science Planning Key Project (2022ZJY2791), the Guangxi Medical High-level Key Talents Training “139” Program (2020), and the Guangxi Medical University Key Textbook Construction Project (GXMUZDJC2223).
Guarantor
CG.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
