Abstract
Objectives
The study aimed to propose a multimodal model that incorporates both macroscopic and microscopic images and analyze its influence on clinicians’ decision-making with different levels of experience.
Methods
First, we constructed a multimodal dataset for five skin disorders. Next, we trained unimodal models on three different types of images and selected the best-performing models as the base learners. Then, we used a soft voting strategy to create the multimodal model. Finally, 12 clinicians were divided into three groups, with each group including one director dermatologist, one dermatologist-in-charge, one resident dermatologist, and one general practitioner. They were asked to diagnose the skin disorders in four unaided situations (macroscopic images only, dermatopathological images only, macroscopic and dermatopathological images, all images and metadata), and three aided situations (macroscopic images with model 1 aid, dermatopathological images with model 2&3 aid, all images with multimodal model 4 aid). The clinicians’ diagnosis accuracy and time for each diagnosis were recorded.
Results
Among the trained models, the vision transformer (ViT) achieved the best performance, with accuracies of 0.8636, 0.9545, 0.9673, and AUCs of 0.9823, 0.9952, 0.9989 on the training set, respectively. However, on the external validation set, they only achieved accuracies of 0.70, 0.90, and 0.94, respectively. The multimodal model performed well compared to the unimodal models, achieving an accuracy of 0.98 on the external validation set. The results of logit regression analysis indicate that all models are helpful to clinicians in making diagnostic decisions [Odds Ratios (OR) > 1], while metadata does not provide assistance to clinicians (OR < 1). Linear analysis results indicate that metadata significantly increases clinicians’ diagnosis time (P < 0.05), while model assistance does not (P > 0.05).
Conclusions
The results of this study suggest that the multimodal model effectively improves clinicians’ diagnostic performance without significantly increasing the diagnostic time. However, further large-scale prospective studies are necessary.
Introduction
Skin diseases, as common and prevalent diseases, have a negative impact on people's health and quality of life. 1 There are thousands of varieties of skin disorders, 2 and there are significant differences in prognosis. 3 Accurate and timely diagnosis is crucial to the diagnosis and treatment of skin diseases. Skin disease diagnosis depends not only on physical examination results but also on skin tissue biopsy results (considered the gold standard for diagnosing skin diseases),4–7 but dermatologists themselves require at least 2–3 years of specialist training to differentiate a skin pathology. Therefore, diagnosing skin diseases by integrating and processing data from multiple modalities is challenging for both general practitioners and dermatologists. Furthermore, in China, the dermatologist–patient ratio can reach 1:60,000; the majority of experienced dermatologists work in large cities, and general practitioners have limited knowledge of dermatological specialties. 8 Due to the insufficient number of dermatologists and the wide variation in levels of experience, many cases have been missed or misdiagnosed, resulting in the clinical diagnosis of skin diseases being far less accurate than necessary.9,10 Hence, developing computer-aided diagnosis (CAD) systems to assist dermatologists in improving efficiency and expertise is of great significance to meet the needs of medical care.
Most CAD systems currently in use are built using machine learning or deep learning (DL) methods. Among machine learning (ML) methods, support vector machines (SVM) are widely used in constructing classifiers. Celebi et al. 11 proposed an SVM-based method for detecting pigmented skin lesions from dermoscopy images. They applied this method to a dataset containing 564 images and obtained a specificity of 92.34% and a sensitivity of 93.33%. Maqsood et al. 12 proposed a new framework for detecting skin diseases, which uses neural networks to extract image features at multiple stages and finally feeds them into SVM for classification. Results on multiple datasets demonstrated that this framework had achieved state-of-the-art (SOTA) performance. Compared to ML, DL relies on deep neural networks to automatically extract features and is receiving increasing attention. The majority of current DL methods are based on convolutional neural networks (CNNs).13,14 For example, Binol H et al. 15 proposed a deep CNN called Ros-NET for the automatic identification of rosacea lesions. Abayomi-Alli et al. 16 designed an improved data augmentation model for the effective detection of melanoma. Nawaz et al. 17 presented an improved DL-based approach, specifically the DenseNet77-based UNET model, for efficient melanoma segmentation. Their results on the public datasets ISIC-2017 and ISIC-2018 demonstrate that the method is robust in skin lesion segmentation and can accurately recognize moles of varying colors and sizes. In addition, some studies have also confirmed the excellent capabilities of CNNs in other skin diseases, such as atopic dermatitis, 18 skin cancer, 19 onychomycosis, 20 and psoriasis.6,21 While these methods have excellent performance, the limited receptive fields restrict their ability to extract global features from images, a limitation that ViT, proposed in 2020, 22 compensates for. Sarker et al. 23 presented a transformer-based model for classifying skin lesion and achieved excellent accuracy on the HAM10000 dataset. SeATrans, proposed by Wu J et al. 24 also outperformed a wide range of SOTA segmentation-assisted diagnosis methods in several tasks. However, the majority of current CAD systems were based on unimodal data. While a unimodal design approach has the advantages of objectivity and reproducibility, multimodal data cannot be fully utilized in clinical diagnosis scenarios using this method.
Clinicians use multimodal data to make a diagnosis, underscoring the need for multimodal methods. Multimodal fusion CAD solutions can help models learn complex and comprehensive clinical feature representations and fully utilize clinical data from real diagnostic scenarios, thus assisting clinicians in making accurate diagnoses. In 2018, Yap et al. 25 effectively improved the detection accuracy of five skin tumors by encoding dermatoscopic and macroscopic images separately using ResNet50 26 and then fusing them with patient metadata. In 2020, Bi et al. 27 proposed a hyper-connected network, HcCNN, for classifying benign and malignant skin lesions. They utilized multi-scale attention blocks to prioritize the semantically more important regions of the two modalities and achieved an average accuracy of 74.9%. In 2022, Tang et al. 28 proposed a multimodal algorithm (FusionM4Net) for multi-label skin lesion classification, dividing feature extraction and classification decision into two stages, and ultimately achieving an average accuracy of 78.5% on a seven-point checklist dataset. Recently, Tian et al. 29 created a multi-view non-tumorous facial pigmentation dataset. They then used multi-view CNN to diagnose these indistinguishable diseases and obtained a great performance. 29 These studies have effectively advanced the progress of multimodal models in the field of dermatology, but they solely focused on enhancing model performance and neglected to carry out additional prospective research.
A few models have been tested in a prospective real-world setting.10,30–32 For example, Tschandl et al.
33
found that good quality artificial intelligence (AI)-based support of clinical decision-making, through the interaction of online test raters with different forms of AI-based decision support, improves diagnostic accuracy more than either AI or physicians alone. They also observed that the least experienced clinicians benefit the most from AI-based support.
33
In 2022, Han et al. reported the first randomized, prospective clinical trial in dermatology that evaluated the performance of physicians collaborating with AI.
34
They further verified that AI could enhance the accuracy of nonexpert physicians in real-world settings.34,35 Although these studies showed AI's potential for improving the performance of nonspecialists in diagnosing skin diseases, they only used dermoscopic or macroscopic image datasets to train the unimodal CAD systems. However, these CAD systems may not accurately simulate the behavior of dermatologists making diagnoses based on multimodal data. As a result, the results of prospective studies based on unimodal models may not be representative of the performance of multimodal models when applied in real-world settings. To the best of our knowledge, there are no reported prospective studies in the field of dermatology that investigate whether the decisions made by multimodal CAD systems can truly influence clinician's decision-making. In this study, to validate the superiority of the mtultimodal model in clinical settings, we constructed three unimodal models and a transformer-based multimodal model using soft voting strategy. Subsequently, we conducted clinician testing to measure the impact of various assistance methods on the diagnostic accuracy and time required for clinicians with different levels of experience. The main contributions of our study can be summarized as follows:
We collected and created a new multimodal dataset, called HuaqiaoDerm-SD5, which includes 1302 macroscopic lesion images, 3056 dermatopathological images, and patient metadata for five skin disorders: lichen planus (LP), eczema (Ecz), psoriasis (Pso), seborrhoeic keratosis (SK), and nevus (Nv). We employed a soft voting method to construct a multimodal model that integrates macroscopic images (skin lesion images) and microscopic images (dermatopathology images) based on the clinical diagnostic behaviors of dermatologists. The proposed multimodal model successfully mitigates the problem of over-training in unimodal models and exhibits excellent performance on test data. Logit and linear regression models were used to analyze the impact of different auxiliary methods on clinicians’ diagnostic accuracy and time, and quantitatively validated the greater superiority of the multimodal model. This fills a research gap in prospective studies of multimodal models in dermatology.
Materials and methods
This section includes: dataset construction, model construction, clinician study, and statistical analyses.
Figure 1 shows the overall flowchart of this study.

Research process flowchart. LM: low magnification; HM: high magnification.
Dataset construction
Data collection
The study was authorized by the Ethics Committees of Jinan University's First Affiliated Hospital and carried out in accordance with the Helsinki Declaration. Individual consent was waived for this retrospective analysis. (Approval number: KY-2023-130; Approval Date: 2023-03-23).
In order to validate the theoretical analysis suggesting that the multimodal diagnostic model is superior to the unimodal diagnostic model, we selected five common but diagnostically challenging skin disorders: LP, Ecz, Pso, SK, and Nv. This selection aims to provide empirical evidence for the results of the theoretical analysis. This study retrospectively collected data from 1561 patients who underwent skin tissue biopsy examinations at the First Affiliated Hospital of Jinan University from 2006 to 2022, which confirmed the presence of LP, Ecz, Pso, SK, and Nv, five skin disorders. Two professional dermatologists from the First Affiliated Hospital of Jinan University removed the following two types of images: 1. skin lesions may be partly or entirely obscured or covered; 2. excessive exudate results in loss of surface appearance and original texture. Finally, we obtained 1311 images of macroscopic skin lesions, 3056 histopathological images (containing 1534 high-magnification dermpathology images and 1522 low-magnification dermpathology images), and metadata (including age, gender, and biopsy site) from 1543 patients for the experiment. The discrepancy between the number of images and patients is because some patients had images that did not meet the inclusion criteria and were excluded, and there were also a few patients with missing images of one modality. Table 1 displays the distribution of disease categories and the demographic information of the patients, respectively. We also present some examples of our dataset in Figure 2.

Example images from our private multimodal dataset, HuaqiaoDerm-SD5. Five categories of skin disorders are abbreviated as: LP: lichen planus; Ecz: eczema; Pso: psoriasis; SK: seborrhoeic keratosis; and Nv: nevus; LM: low magnification; HM: high magnification.
Demographics and other details of multimodal skin disease dataset (HuaqiaoDerm-SD5).

Abbreviations: LP: lichen planus; Ecz: eczema; Pso: psoriasis; SK: seborrhoeic keratosis; Nv: nevus; No.: number.
DL models require sufficient data for training. However, to fairly compare the diagnostic capabilities of models and clinicians, it is necessary to ensure that the images used for model test and clinician test are the same. Therefore, we first randomly selected 50 patients (each containing 50 macroscopic skin lesion images, 50 low-magnification dermatopathological images, and 50 high-magnification dermatopathological images) from which the modality intact is used for external validation of the model and for the clinicians’ study. The remaining patients are divided into training and validation sets in a ratio of 9:1. Specifically, there are 1185 macroscopic skin lesion images, 1375 low-magnification dermpathology images, and 1386 high-magnification dermpathology images for training, along with 126 macroscopic skin lesion images, 147 low-magnification dermpathology images, and 148 high-magnification dermpathology images for validation. To prevent potential data leakage across patients, we implemented secure data storage protocols and anonymized patient information, ensuring confidentiality. Access to the data was restricted to authorized personnel only, and all necessary precautions were taken to protect patient privacy.
Data augmentation
Data augmentation is a method of increasing the number and diversity of samples. A category-balanced dataset is essential for training, and an uneven distribution of samples may lead to bias. Considering our dataset is unbalanced and exhibits a considerable variation in the number of samples for each disease category (e.g. there are 441 macroscopic images of Nv, but only 65 macroscopic images of LP), we enhanced the dataset used for model training using horizontal flip, vertical flip, rotation, and luminance shift. As shown in Figure 3, after enhancement, the number distribution of each disease is relatively balanced.

Image distributions of the dataset before and after augmentation. Abbreviations: LP: lichen planus; Ecz: eczema; Pso: psoriasis; SK: seborrhoeic keratosis; Nv: nevus; LM: low magnification; HM: high magnification.
Model construction
Constructing unimodal models
To determine the most appropriate model structure for the three types of images, we train and validate the convolution-based neural networks Alexnet, VGG16, ResNet50, and the transformer-based neural network ViT on their respective unimodal datasets. In comparison to convolutional neural networks, the ViT is a novel model architecture introduced by Google researchers that utilizes the attention mechanism. It divides the input image into patches, applies position encoding, and subsequently feeds it into the Transformer Encoder (TE) to determine the interdependence among pixels. The attention mechanism possesses stronger global modeling capabilities and is capable of capturing global information more effectively. Additionally, due to the ability to perform parallel calculations, the output of each position can be computed independently, resulting in a significant improvement in efficiency.
During the training process, to accelerate convergence and prevent overfitting, we adopt the transfer learning technology by initializing the model with pre-trained weights from ImageNet. We utilize Adam as the optimizer with an initial learning rate of 0.0001. Additionally, we employ an early termination strategy to prevent overtraining. The training process is terminated when the training loss continues to decrease or remains unchanged, but the validation loss increases. We assigned the names model 1, model 2, and model 3 to the trained macroscopic image model, low-magnification dermatopathology image model, and high-magnification dermatopathology image model, respectively.
Construct multimodal model using soft voting strategy
Multimodal models can process multiple types of images simultaneously, enabling richer information representation. Data from different modalities can complement each other, providing more comprehensive and accurate information. Additionally, the multimodal model closely aligns with the diagnostic behavior of clinicians and offers better interpretability.
36
In this context, we utilized the soft voting method in ensemble learning to construct the multimodal DL model for verifying its diagnostic assistance to clinicians of varying expertise levels during subsequent testing. We selected the voting method because it closely approximates the decision-making process of clinicians, prioritizing more important image types. Soft voting, unlike hard voting, utilizes the soft labels outputted by the base learners, providing more informative data. The structure of the designed model is depicted in Figure 4. The base learners 1, 2, and 3 represent model 1, model 2, and model 3, which were trained in the previous section. They are used to extract features from unimodal data, respectively. The output probabilities of the base learners are utilized as soft labels, containing more information regarding disease categories. Subsequently, these labels are fused using a weighted average method (as depicted in Equation (1)). Finally, a softmax layer will be employed to obtain the final result.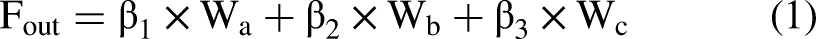
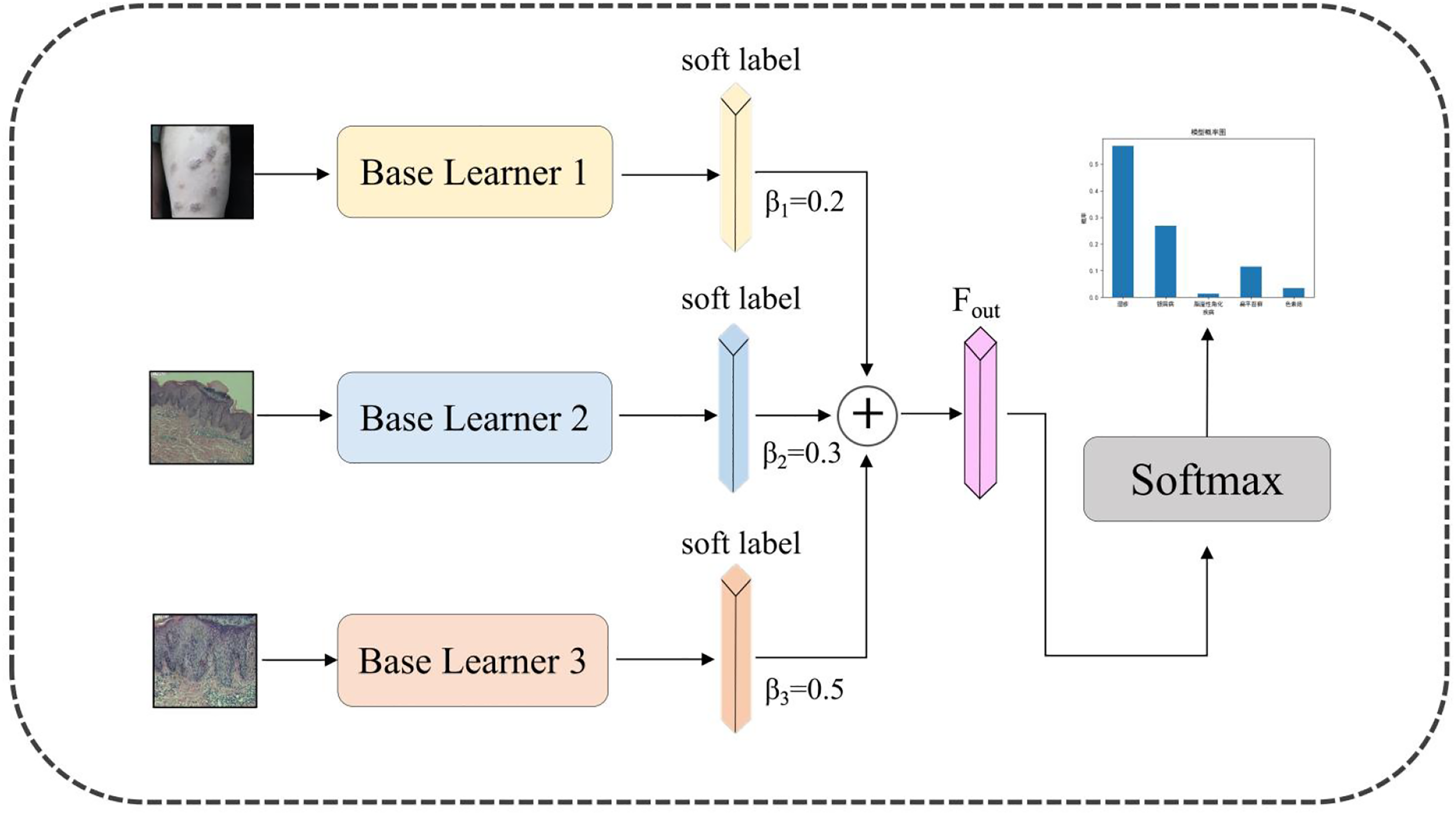
Structure of the proposed multimodal deep learning model.
Clinician study
A total of 12 clinicians participants certified by the health administrative departments under the State Council of China participated in this study, including three director dermatologists (>15 years of dermatology practice), three dermatologists-in-charge (3–5 years of dermatology practice), three resident dermatogists (<3 years of dermatology practice, in the middle of residency training), and three general practitioners (3–5 years of general practice). They were divided into three groups: A, B, and C. Each group contained one director dermatologist, one dermatologist-in-charge, one resident dermatologist, and one general practitioner. The participants included in the three groups were not duplicated.
Next, to compare the impact of multimodal model and unimodal models on doctors’ diagnostic decisions, we established seven different experimental arms. The detailed description of these experimental arms is shown in Table 2. For experimental arms (e, f, g) with model assistance (including unimodal models and a multimodal model), we applied the respective models to the images in the testset to generate the probability distribution histograms. The relative size of the probability of each disease in the histogram indicated the diagnostic confidence of the model, and they sum up to 1. For macroscopic images, model 1 output the corresponding histograms; for low and high-magnification dermatopathological images, model 2 and model 3 output the histograms, respectively. For macroscopic images, low-magnification dermatopathological images, and high-magnification dermatopathological images, the model 4 output the corresponding histograms.
Table of samples clusters.

Finally, the groups of doctors mentioned above would read the patient information in the testset across seven different experimental arms and provide diagnostic results. Specifically, experimental arms a and e were assigned to group A participants to study the effectiveness of model 1 as a clinical aid. Experimental arms b and f were assigned to group B participants to study the effectiveness of model 2 and model 3 as clinical aids. Experimental arms c, d, and g were assigned to group C participants to investigate the effectiveness of the metadata and multimodal model as clinical aids, respectively. The allocation of image to experimental arms was counterbalanced across participants, so that each image had approximately the same number of participants for each arm. It is not strictly the same because each image was read 12 times (once per participant) across seven experimental arms. Specifically, for participants in groups A and B, the readings of individual images were evenly distributed across the arms because each image was read two times on four experimental arms (once per participant). However, for participants in group C, each image was read four times across the remaining three experimental arms (c, d, g). Consequently, each image was read by one reader for two conditions and by two readers for the third condition.
Before the clinicians’ test began, each participant was given an introduction on how to perform the specific test. Participants were asked to diagnose the samples under various experimental conditions. In addition, they were told that the diagnostic accuracy was used as a measure of the outcome, but not the time of diagnosis. Following the completion of the test by the clinicians, we collected their diagnostic accuracy and diagnostic time for further statistical analysis.
Statistical analyses
We used Stata 16.0 statistical software to construct the logit regression models 37 in order to compare and explore the impact of the model on clinicians’ decision-making. The dependent variable in the logit models was the participant's judgment of the case as true (T) or false (F), represented as T = 1 and F = 0. Whether the participants used assistance was selected as an independent variable. The auxiliary methods consist of model assistance and metadata assistance. Based on this, we established logit regression models for two conditions: controlling doctor level or not. Ultimately, we chose the Logit model result with the highest prediction accuracy under the two conditions as the final outcome for detailed analysis. Additionally, we utilized linear models to analyze the effect of different auxiliary methods on diagnosis time. In linear model, the dependent variable was the subject's time to diagnosis, while the usage of assistance by the participants was chosen as an independent variable.
Results
Performance of unimodal model on validation dataset
The performance indicators of the constructed unimodal model on the validation set are shown in Table 3. ViT achieved higher Top-1 accuracy rates than all other models used for comparison on the macroscopic image, low-magnification dermatopathology image, and high-magnification dermatopathology image datasets. The accuracy rates were 0.8636, 0.9545, and 0.9673 respectively. ViT also outperformed other models on the AUC indicator. ViT achieved AUC scores of 0.9952 and 0.9989 on the low-magnification dermatopathology image and high-magnification dermatopathology image datasets, respectively. These scores were 0.18% and 0.30% higher than Resnet50, the best model in the convolutional neural network. Additionally, the ViT achieved satisfactory precision and recall indicators. This demonstrates that the attention-based model has powerful long-distance modeling capabilities and can effectively extract image features compared to CNNs. Therefore, we chose ViT as the base learner to build the multimodal model.
The performance of unimodal models on validation dataset.

Performance of the models on the testset
Table 4 summarizes the diagnostic accuracy of the unimodal models (model 1, model 2, model 3) and the multimodal model (model 4) on the testing set. It showed that model 4 achieved the highest 98% accuracy, followed by model 3, model 2, and model 1. The result indicates that the multimodal model was overall better than the unimodal model at identifying the five skin disorders. The intuitive reason is that multimodal learning can aggregate information from multiple data sources, enabling the model to learn a more comprehensive representation. Furthermore, the unimodal models all show some degradation compared to the metrics on the training set. This indicates that during the training process, the models may have learned the characteristics of some specific datasets, indicating overfitting. However, multimodal models can learn more general data features, effectively alleviating this situation and achieving better performance.
Different models’ diagnostic performance in the five-category classification task.

Abbreviations: LP: lichen planus; Ecz: eczema; Pso: psoriasis; SK: seborrhoeic keratosis, Nv: nevus.
Specifically, model 1's overall performance significantly differed from that of model 4, with the lowest overall accuracy. To investigate the specific reasons for the classification errors, we created a confusion matrix for both models. According to Figure 5, model 1 primarily misclassified Ecz as LP or Pso. Additionally, it misclassified three Pso images as LP. This is likely because these three diseases have very similar features and can be easily misdiagnosed, even by dermatologists. Furthermore, a few SKs were misclassified as Nv, which also had an impact on the model's accuracy. For model 4, only one case of Pso was misclassified as LP. This result demonstrates that combining information from multiple modal images can yield a model with enhanced feature representation capabilities.

Confusion matrices of the models and five skin disorders. Figures a and b are the confusion matrices of model 1 and model 4 in classifying the test set, respectively. The x-axes are the predicted labels, which are the diagnoses made by the model. The y-axes are the true labels, which are the pathological results. The number in each small square represents the corresponding number of images with the same predicted true label. Five categories of skin disorders are abbreviated as: LP: lichen planus; Ecz: eczema; Pso: psoriasis; SK: seborrhoeic keratosis; Nv: nevus.
Effect of model assistance on the diagnostic accuracy of clinicians
The logit model, which controls the type of clinician, had a higher accuracy of 86.2% compared to the model that does not control the type of clinician (as provided in Supplemental Table S1, with an accuracy of 84.3%). Hence, we chose this model for further analysis. Table 5 shows that all auxiliary methods of the model are beneficial for clinicians’ diagnosis (OR > 1), except for metadata, which does not provide significant improvement (OR < 1). Among all auxiliary models, the multimodal model 4 had the highest Logit Coefficient Estimate (2.972) and OR (19.525), making it the most effective auxiliary method for clinicians. Dermatopathology models 2 and 3 are also beneficial for clinicians, although not as effective as the multimodal model. Macroscopic image model 1 is not as evidently helpful to clinicians compared to the first two ways, but it still shows statistical significance (P < 0.05). However, metadata is not beneficial for clinicians’ diagnosis and may even interfere with their decision-making (Logit Coefficient Estimate < 0). This may be because our metadata only includes general information such as age and gender.
Logit regression analysis of diagnostic accuracy rates for participants (control clinicians’ type).

Effect of model assistance on the diagnostic time of clinicians
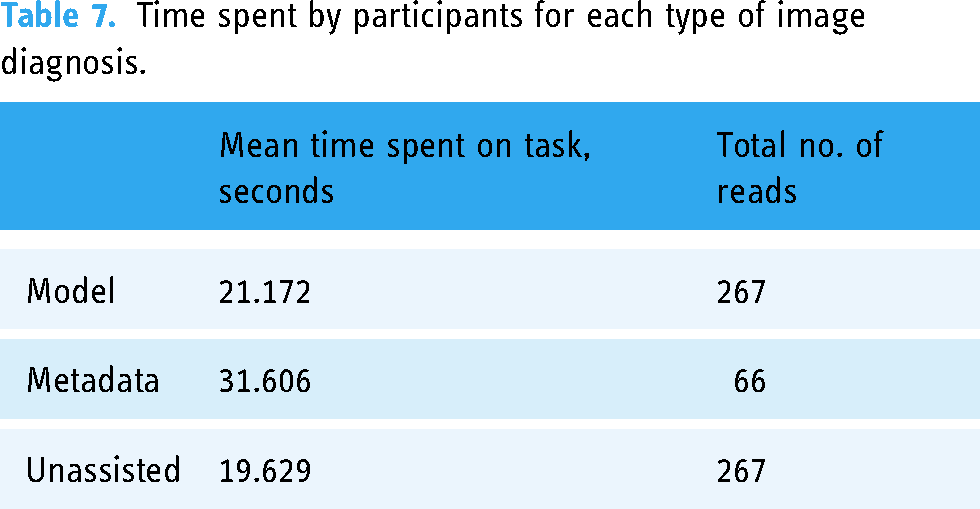
Table 6 presents the impact of various auxiliary methods on the diagnosis time of clinicians. Despite the model-assisted approach providing an additional percentage of diagnostic confidence for the five skin disorders during diagnosis, the extra information does not significantly enhance the clinician's diagnostic time (P > 0.05). The inclusion of patient metadata has resulted in an increased diagnostic time for clinicians (P< 0.05). This indicates that metadata may not provide valuable information. Although it cannot significantly enhance clinicians’ diagnostic abilities, it adds to their workload by requiring them to read this information. Furthermore, we calculated the average diagnostic time for clinicians. According to Table 7, clinicians’ average diagnostic time per patient increased by less than 2 seconds with model assistance. However, with metadata assistance, it increased by nearly 12 seconds. In conclusion, the multimodal DL model offers the most remarkable and efficient improvement for clinicians when considering the above information. Moreover, this improvement will not significantly impose additional burden on clinicians.
Analysis of participants’ diagnostic time using linear regression.

Time spent by participants for each type of image diagnosis.
Benefit from assistance varies with clinician's background level
The analysis above revealed that only the dermatopathology model 2&3 and the multimodal model 4 were statistically significant in improving clinicians’ diagnoses. Still, the effect of these models on improving doctors at different levels may vary. As shown in Figure 6, the director dermatologists could reach 98% diagnostic accuracy without the aid of dermatopathology model 2&3. In contrast, the resident dermatologists only achieved 82%. When assisted by the model, the dermatologists-in-charge can attain a level comparable to that of director dermatologist, with general practitioners surpassing even the resident dermatologists.

Bar graphs showing varying experience level of clinicians’ diagnositic accuracy for cases with five-class skin diseases under (left) unassisted and (right) model-assisted conditions. Panel a illustrates readers reading LM&HM dermatopathological images, while Panel b represents reading macroscopic skin lesion images alongside LM&HM dermatopathological images. Each plot breaks down performance by the reader's previous experience with skin images: director dermatologists (DD; n = 3), dermatologists-in-charge (DIC; n = 3), resident dermatologists (RD; n = 3), general practitioners(GP; n = 3). LM: low magnification; HM: high magnification.
Likewise, the diagnostic accuracy of dermatologists-in-charge and general practitioners was very low in the absence of the multimodal model. However, general practitioners improved significantly with the multimodal model and even outperformed director dermatologists. This could be due to general practitioners relying heavily on models. In our test set, general practitioners, aided by the multimodal model, outperformed even the performance of the director dermatologist. While this does not conclusively prove that clinicians of lower seniority can reach the expert level with the model's assistance, it does confirm that the model has a more notable impact on advancing clinicians with lower seniority.
Discussion
In this study, we quantitatively explored the impact of different auxiliary methods on clinicians’ decision-making by building models and conducting clinical tests. We found that among the auxiliary testing methods, multimodal models can provide clinicians with the greatest assistance, improving diagnostic accuracy without significantly increasing the diagnostic time, which is consistent with the expected results. This trial provides quantitative validation for the superiority of a multimodal CAD model.
In the medical field, most of the existing networks based on multimodal fusion are focused on medical images such as CT, MRI, and ultrasound. For instance, Hao et al. 38 proposed a novel multimodal neuroimaging feature selection method with consistent metric constraints (MFCC) for Alzheimer's disease diagnosis based on two types of multimodal image data: VBM-MRI and FDG-PET. Liu et al. 39 use a deep learning model to diagnose significant liver fibrosis in chronic hepatitis B (CHB) patients by integrating ultrasound contrast-enhanced micro-flow (CEMF) cines, B-mode images, and clinical parameters. These multimodal data differ greatly from the skin disease modal data, making it difficult to directly migrate the models to skin disease diagnosis. Thus, an increasing number of tasks involve multimodal input in the field of dermatology. Similar to the findings of Yap et al., 25 Bi et al., 27 Cai et al., 40 Tang et al., 28 and Tian et al., 29 we all discovered that the completed multimodal data fusion CAD systems achieved greater diagnostic accuracy than single-modality CAD systems. However, Yap and Bi et al.'s multimodal CAD systems fused patient clinical images and dermatoscopic images25,27; Cai et al. proposed a multimodal Transformer that fuses two modalities: macroscopic skin lesion images and metadata. 40 Tang et al.'s system fused patient clinical images, dermatoscopic images, and patient information metadata 28 ; and Tian et al. combined clinical images captured under different light sources with nine distinct views 29 ; To date, there is no multimodal fusion CAD system that fuses modal information such as skin pathology and other laboratory examination data. For the diagnosis of skin diseases, pathological images can provide characteristic manifestations of diseases, which are vital for the diagnosis of skin diseases, and can be independent of and complementary to the modal information provided by clinical images and dermoscopic images. Because skin pathology images provide information about disease characteristics that other modalities cannot, we included them in the multimodal CAD model we developed for this study. This, we believe, is an important reason for our multimodal CAD model's excellent performance.
More importantly, after constructing the models, we further performed clinical quantitative validation of the models. Previous studies have mostly pitted models and doctors against each other, with the models and doctors competing to reflect the models’ diagnostic performance. According to Hekler et al., 41 a diagnostic model obtained by fusing human and AI models’ diagnoses could achieve better image classification than classification by only dermatologists or only CNN models. When dermatologists collaborated with the model, the average accuracy increased by 1.36%. 41 This implies that figuring out a good way to combine humans and artificial intelligence could help the model perform better. In fact, studies have shown that when various types of models are applied in real-world scenarios, they often confront both over-reliance (repeating the model's errors) and under-reliance (ignoring the predictions of accurate algorithms).42–45 While numerous models with outstanding performance have been developed and some are currently in use, 46 only a limited number have undergone testing in real-world settings.10,30–32 Therefore, more studies are necessary to evaluate the extent of improvement that can be achieved in clinicians’ diagnostic results through the utilization of these models.
There are also some limitations to our research. First, the multimodal model in this study used a weighted fusion approach. However, there are several other fusion approaches to the multimodal model, and different methods may also influence the model's performance. The second limitation is that, due to insufficient data for some rare diseases, our database currently only includes five skin disorders. However, currently there is no multimodal dataset available in public databases of skin diseases that includes both macroscopic images and dermatopathological images, so our proposed model cannot be verified by an external dataset. The third limitation is that the vast majority of the data we use for training comes from Asian populations, and the predictions of the algorithms are heavily dependent on the features of the training data. 34 Therefore, they may exhibit uncertainty in different settings, so the accuracy of our deep learning algorithms and their usefulness in assisting clinicians cannot be generalized to non-Asian populations.
For future research: (1) Conduct a multicenter study to increase the size of the dataset by combining patient data from other hospitals and supplementing data from new patient visits to include more skin disease categories. (2) Consider adding further large-scale prospective validation of the model in future research.
Conclusions
In this study, we collected and constructed a multimodal dataset consisting of macroscopic images, dermatopathological images, and metadata for five skin disorders. We then developed a new multimodal DL model for the diagnosis of skin conditions, performed quantitative validation of the model-assisted effects, and experimentally demonstrated that the developed model does have better assisted effects for clinicians. On the basis of these findings, we conclude that the multimodal model is superior to the unimodal model when used for both independent and assisted diagnosis of skin diseases, providing evidence of the model's clinical applicability.
Supplemental Material
sj-doc-1-dhj-10.1177_20552076241257087 - Supplemental material for Exploring the influence of transformer-based multimodal modeling on clinicians’ diagnosis of skin diseases: A quantitative analysis
Supplemental material, sj-doc-1-dhj-10.1177_20552076241257087 for Exploring the influence of transformer-based multimodal modeling on clinicians’ diagnosis of skin diseases: A quantitative analysis by Yujiao Zhang, Yunfeng Hu, Ke Li, Xiangjun Pan, Xiaoling Mo and Hong Zhang in DIGITAL HEALTH
Footnotes
Acknowledgments
We would like to thank all the doctors and nurses at the Department of Dermatology, the First Hospital of Jinan University and the clinicians who participated in our study.
Author contributions
(I) Conception and design:YZ and HZ; (II) Administrative support: HZ; (III) Provision of study materials or patients:YH; (IV) Collection and assembly of data: XM and XP; (V) Data analysis and interpretation:YZ and KL; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors.
Availability of data and materials
The code of model and the priviate dataset used in the current study can be available from the corresponding author with reasonable request.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics approval and consent to participate
The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by the Ethics Committees of the First Affiliated Hospital of Jinan University and individual consent for this retrospective analysis was waived (Approval number: KY-2023-130; Approval Date: 2023-03-23).
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Clinical Frontier Technology Program of the First Affiliated Hospital of Jinan University, China (No. JNU1AF-CFTP-2022-a01211); and Science and Technology Projects in Guangzhou (2023A03J1027).
Guarantor
HZ.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
