Abstract
Background
Early detection and treatment are crucial for reducing gastrointestinal tumour-related mortality. The diagnostic efficiency of the most commonly used diagnostic markers for gastric cancer (GC) is not very high. A single laboratory test cannot meet the requirements of early screening, and machine learning methods are needed to aid the early diagnosis of GC by combining multiple indicators.
Methods
Based on the XGBoost algorithm, a new model was developed to distinguish between GC and precancerous lesions in newly admitted patients between 2018 and 2023 using multiple laboratory tests. We evaluated the ability of the prediction score derived from this model to predict early GC. In addition, we investigated the efficacy of the model in correctly screening for GC given negative protein tumour marker results.
Results
The XHGC20 model constructed using the XGBoost algorithm could distinguish GC from precancerous disease well (area under the receiver operating characteristic curve [AUC] = 0.901), with a sensitivity, specificity and cut-off value of 0.830, 0.806 and 0.265, respectively. The prediction score was very effective in the diagnosis of early GC. When the cut-off value was 0.27, and the AUC was 0.888, the sensitivity and specificity were 0.797 and 0.807, respectively. The model was also effective at evaluating GC given negative conventional markers (AUC = 0.970), with the sensitivity and specificity of 0.941 and 0.906, respectively, which helped to reduce the rate of missed diagnoses.
Conclusions
The XHGC20 model established by the XGBoost algorithm integrates information from 20 clinical laboratory tests and can aid in the early screening of GC, providing a useful new method for auxiliary laboratory diagnosis.
Introduction
Gastric cancer (GC) is the third most common cancer in China, accounting for 10.5% of cases according to the latest global cancer burden estimate released by the International Agency for Research on Cancer. The mortality rate of GC is 12.4% in China, and GC is the third leading cause of cancer death.1,2 The five-year overall survival rate of patients with early-stage localised GC is more than 60%, whereas that of patients with distant metastasis is less than 5%. Early detection and treatment are crucial for reducing the mortality rate of GC patients.3,4 Nevertheless, more than 70% of patients are often diagnosed with advanced GC because early symptoms are not obvious. 5 In recent years, with the continuous development of endoscopic technology and the popularisation of endoscopic screening, the detection rate of early-stage GC has been increasing globally, and endoscopic therapy has also greatly improved the survival rate.6,7 However, large-scale endoscopic screening of early-stage GC has not yet been carried out in China. The intestinal preparation process for endoscopic and pathological examination is complex and invasive, and patient compliance is poor, which is not suitable for routine GC screening. 8 Endoscopic screening is currently performed mainly in symptomatic patients, resulting in a significantly lower detection rate of early GC than in developed countries. 9 Due to its technical limitations, including manual sampling, the diagnostic effect of endoscopic techniques on GC, especially early-stage GC, is not perfect. 10 The ability of traditional endoscopy to observe small lesions has long been unable to meet current clinical needs. The missed diagnosis rate of confocal laser endoscopy in chronic atrophic gastritis patients is 8–15% due to the limited field of view and the inability to observe the entire oesophagus or stomach cavity.11,12 Therefore, there is an urgent need to develop non-invasive, convenient and valuable early screening and diagnostic techniques for GC as an auxiliary means of endoscopic diagnosis.
Non-invasive markers that can detect cancer in its early or precancerous stages are promising for clinical development.13–15 Some conventional protein markers of digestive tract tumours, such as carcinoembryonic antigen (CEA), carbohydrate antigen 19-9 (CA 19-9), CA 242 and CA 72-4, are used for routine screening, efficacy monitoring and prognosis judgement. 16 Unfortunately, for GC patients, these methods are not effective at early screening and diagnosis because of their poor specificity and sensitivity. 17 Gastric cancer diagnostic markers such as pepsinogen (PG) and gastrin-17 (G17) have not been used as first-line screening markers because they are not fully covered by national medical insurance.18,19 Another emerging blood-based biomarker derives from the liquid biopsy, which involves obtaining samples of blood or other body fluids and is used to analyse cancer-derived molecules or cancer cells. 20 However, liquid biopsy cannot be commonly carried out in the laboratory due to its high cost, low quantity of circulating tumour cells and circulating cell-free nucleic acids, complex operation and high requirements for personnel and instruments.21,22 While clinical laboratories report most test results as individual numbers, findings or observations, clinical diagnosis usually relies on the results of multiple tests. The multiple routine tests for general surgery admission testing are commonly inexpensive, covered by medical insurance and easy to obtain. The routine laboratory indicators of each patient need to be fully utilised, artificial intelligence needs to be used to determine the associations between the indicators, an early screening model needs to be established for GC, and, more importantly, the diagnostic efficiency of early GC needs to be improved. In recent years, several studies have combined test indicators with machine learning (ML) to develop disease prediction models based on large databases; these models not only improve the performance of non-invasive serological tests but are also easy to apply in routine examinations of patients at most medical institutions.23,24
ML focuses on how to improve the performance of specific algorithms in experiential learning.25,26 The method can discover and utilise the interaction effect and non-linear relationship between related factors and balance the influence of sample error by randomly selecting a large number of samples.27,28 The resulting models trained with large amounts of data are more reliable than logistic regression models that fit only simple test samples. Extreme gradient boosting (XGBoost) is an ML system developed by Chen and Guestrin and is available as an open-source package.29,30 It is widely used in the Kaggle competition and many other ML competitions and has enabled great achievements, even better than deep neural nets. The XGBoost algorithm is increasingly used in cancer diagnosis and treatment areas, such as surgical intervention, image interpretation, drug development and personalised treatment.31–33
In this study, we included a large number of clinical laboratory test results from thousands of patients, used the XGBoost algorithm to establish a GC prediction model and evaluated the diagnostic efficacy of this model, especially for the early diagnosis of GC. We also demonstrated that the model was effective in the diagnosis of GC patients with negative conventional protein marker results. Therefore, this model provides accurate and convenient decision support for early GC diagnosis and reasonably interprets a large amount of test data, greatly enhancing user trust.
Patients and methods
Subjects
The subjects of this study were mainly newly admitted patients registered in the pathology laboratory of Xinhua Hospital from March 2018 to June 2023. New admissions between July 2023 and December 2023 were included in the external validation set. We collected clinical, pathological and laboratory test data from those patients who met the inclusion and exclusion criteria. The patient's admission number was the unique sample identification number. All disease names in the databases were obtained from the
All patients were categorised into two groups: precancerous disease and GC. The inclusion criteria for patients were as follows: (1) had a precancerous disease, including patients with gastroscopic or pathological results, and were clinically diagnosed with diseases, including atrophic gastritis, gastrointestinal metaplasia and gastric dysplasia. (2) Gastric cancer patients were included if they met the clinical and histopathological diagnostic criteria. The exclusion criteria for patients were as follows: unclear clinical diagnosis, repeated examination or treatment and coexisting malignancies. For patients with precancerous disease or GC who were admitted multiple times, only the data for the first diagnosis without treatment (including surgical treatment and drug therapy) were considered to minimise bias. To ensure the compatibility of different versions of TNM staging in the database, the pTNM stage was reworked according to the 8th edition Cancer Staging Manual issued by AJCC. 34 The Clinical Guidelines for the Diagnosis and Treatment of GC released by The Chinese Society of Clinical Oncology (CSCO) in 2023 defined early-stage GC as characterised by confinement within the mucosa and submucosa, regardless of evidence of regional lymph node metastasis. 10 Therefore, this study defined with pTNM-stage 0 and pTNM-stage IA cancer as early GC. In addition, no further restrictions were imposed on the completeness or accuracy of the data, and the data selected from the database could be considered as real-world big data.
Feature correlation screening
Feature correlation screening was designed to reduce the redundancy among features and retain features that were highly correlated with the dependent variable. In our method, the threshold for feature correlation analysis was 0.9. Features that had a high linear correlation with the dependent variable were preferentially retained. We also used L1 regularisation (also known as LASSO) as a feature screening method to further optimise feature selection. We set the regularisation coefficient to 1 to ensure rigor in feature selection, helping us choose the most predictive features to establish the model.
XGBoost algorithm
Parameter settings
XGBoost is a powerful gradient lift tree algorithm for processing complex datasets and predictive modelling. In our approach, we used the XGBoost model based on the scikit-learn (sklearn) library for modelling. The booster parameter specifies the type of base learner. We chose a tree-based gradient lifting method, namely, gbtree. In the modelling process, we used the training set for model training, and the validation set for hyperparameter tuning and model selection.
The objective function of XGBoost
The objective function of XGBoost is depicted as follows:
Taylor's expansion:


The core algorithm of XGBoost
Trees should be constantly added, and characteristic divisions should be performed to grow a tree. By adding a tree, a new function was learned to fit the residuals of the last prediction. When we finish training and obtain
Our goal was to ensure that the predicted values of the tree group were close to the true values to the greatest extent possible and had the greatest generalisation ability. In each iteration, a tree is added to the existing tree to fit the residual difference between the predicted result of the previous tree and the true value. The iteration process is depicted as follows. It starts from constant prediction and then adds a new function each time:




Testing the performance of prediction model
In the model, the importance of each feature was measured by characteristic coefficients and relative weights. The performance of the ML model based on XGBoost was evaluated by sensitivity [TP/(TP + FN)], specificity [TN/(TN + FP)], accuracy [TP + TN/(TP + FP + TN + FN)], positive predictive value [TP/(TP + FP)] and negative predictive value [TN/(TN + FN)]. The matrices of the true and predicted conditions are shown in Table S1. In addition, the prediction ability of the model was evaluated by using the area under the receiver operating characteristic curve (AUC), accuracy rate (ACC), recall rate and F1 score. The prediction score directly represents the quantitative index of the prediction ability of the model. Validation sets and test sets were used to evaluate the diagnostic efficiency of the model.
Statistical analysis
The Deepwise and Beckman Coulter DxAI platforms (https://dxonline.deepwise.com/login) were used to perform the XGBoost algorithm. This platform was based on scikit-learn 1.2.2 for packaging modelling. A detailed introduction to the algorithm and the original code can be found on the following website (https://xgboost.readthedocs.io/en/latest/python/sklearn_estimator.html#). The distributed variables are presented as the means ± SDs, and the significance of the differences was determined with Student's
Results
Random splitting of the data
Based on the inclusion and exclusion criteria, 4400 patients were selected, among whom 1673 had GC and 2727 had precancerous disease. The Z score standardised method was used to normalise the quantitative data to ensure comparability under various conditions before the analysis. To perform reliable model evaluation, we used a random splitting method to divide the complete set of subjects, with a total data size of 4400, into a training set, a validation set and a test set. Among them, the training set accounted for 60% of the total data (2640 samples), and the validation set and test set each accounted for 20% of the total data (1760 samples) (Figure 1 and Table 1). The training set was used to determine the model hyperparameters, and the validation set and the test set were used to evaluate the performance of the trained optimal function. 35 Patients with early-stage GC (pTNM stage 0–IA) accounted for 19.19% of all GC patients. Among the precancerous disease patients, 46.09%, 38.50% and 15.40% had atrophic gastritis, gastrointestinal metaplasia and gastric dysplasia, respectively. The feature results of each item were taken based on the inpatient number as the unique identifier. If one patient was tested multiple times for the same item immediately after admission, the results were averaged.

Schematic representation of case enrollment process. PD: precancerous disease; GC: gastric cancer.
Clinical features of the subjects.

Feature filtering
In the process of feature cleaning and feature substitution for model parameter setting, features with more than 30% missing values were automatically deleted and not included in model training, and features with more than 30% missing values were filled with median data. After extraction of the database information, 860 laboratory routine test items were obtained, 52 of which had missing values of less than 30%. The clinical guidelines for the diagnosis and treatment of GC released by the CSCO indicate that age and sex are closely related to the occurrence of GC. 34 Together with the two variables of age and sex, a total of 54 variables were included in this study for feature selection. Finally, 20 of the 54 candidate features were selected, and a prediction model named “XHGC20” was established using the XGBoost algorithm. We listed and ranked the 20 model features in Table 2 according to their relative weights. The missing values of these 54 candidate features in the model are shown in Table S4. Except for sex, the other 19 haematological indicators were divided into four main laboratory test categories: (1) five indicators of blood cells; (2) three indicators of coagulation function; (3) nine biochemistry indicators; and (4) two indicators of protein-related tumour markers. Among the 20 indicators, the characteristic coefficient of total protein was the most significant (0.1662), followed by fibrin (pro) degradation products (FDPs), C-reactive protein (CRP) and red blood cell volume distribution width (RDW).
Twenty features and relative coefficients.

Comparison of prediction scores among different groups
A total of 4400 patients were categorised into GC and precancerous disease groups and used to establish the XGBoost model based on the 20 indicators mentioned above. The XGBoost model “XHGC20” was used to determine the prediction score to measure the risk of GC. A comparison of the prediction scores of the two groups revealed that the prediction score of the GC group was significantly greater than that of the precancerous disease group [0.974 (95% CI 0.068–1.000) vs. 0.013 (95% CI 0.000–0.667),
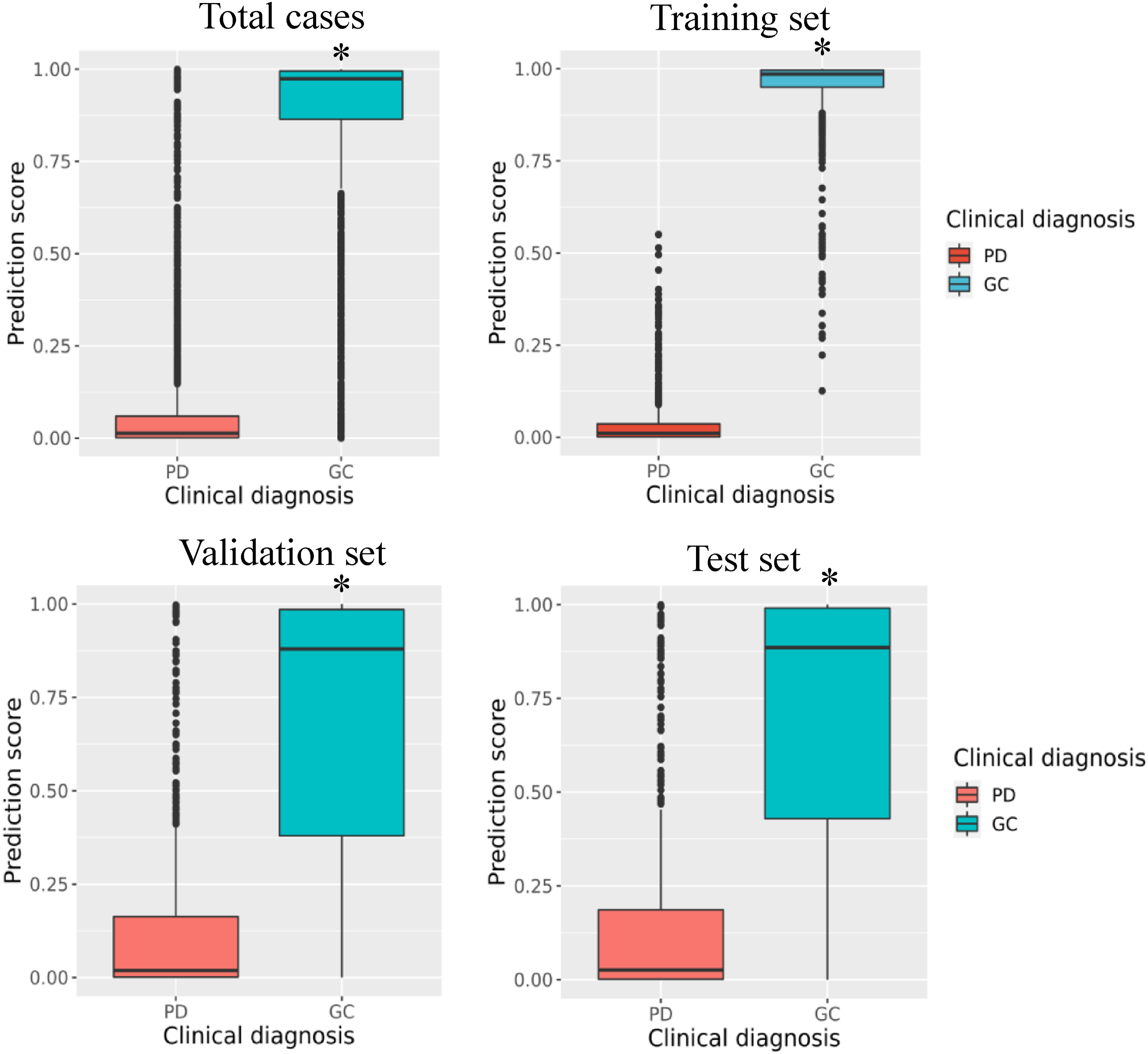
Comparison of prediction scores between precancerous disease and gastric cancer in the training, validation and test set, respectively. *
Diagnostic efficiency of the XGBoost model
The fitting results were also satisfactory. The model showed great diagnostic performance in the training cohort, with an AUC of 0.999 and an ACC of 0.989 (sensitivity = 0.988, specificity = 0.992) (Table 3 and Figure 3(a)). With respect to the validation set, the XHGC20 model (AUC = 0.901, ACC = 0.818) also had good diagnostic efficiency, with high specificity (0.830) and sensitivity (0.806) in distinguishing GC from precancerous lesions when the cut-off value was set to 0.265. The diagnostic performance in the test set was also good (AUC = 0.907, ACC = 0.827, sensitivity = 0.875, specificity = 0.771, cut-off value = 0.216). According to the precision–recall curve, the XHGC20 model also demonstrated good diagnostic efficiency (Figure 3(b)). The precisions of the validation set and test set were 0.811 and 0.802, respectively. The results showed that the XHGC20 model established by the XGBoost algorithm could distinguish between GC and precancerous lesions well, and its diagnostic efficiency was very good.
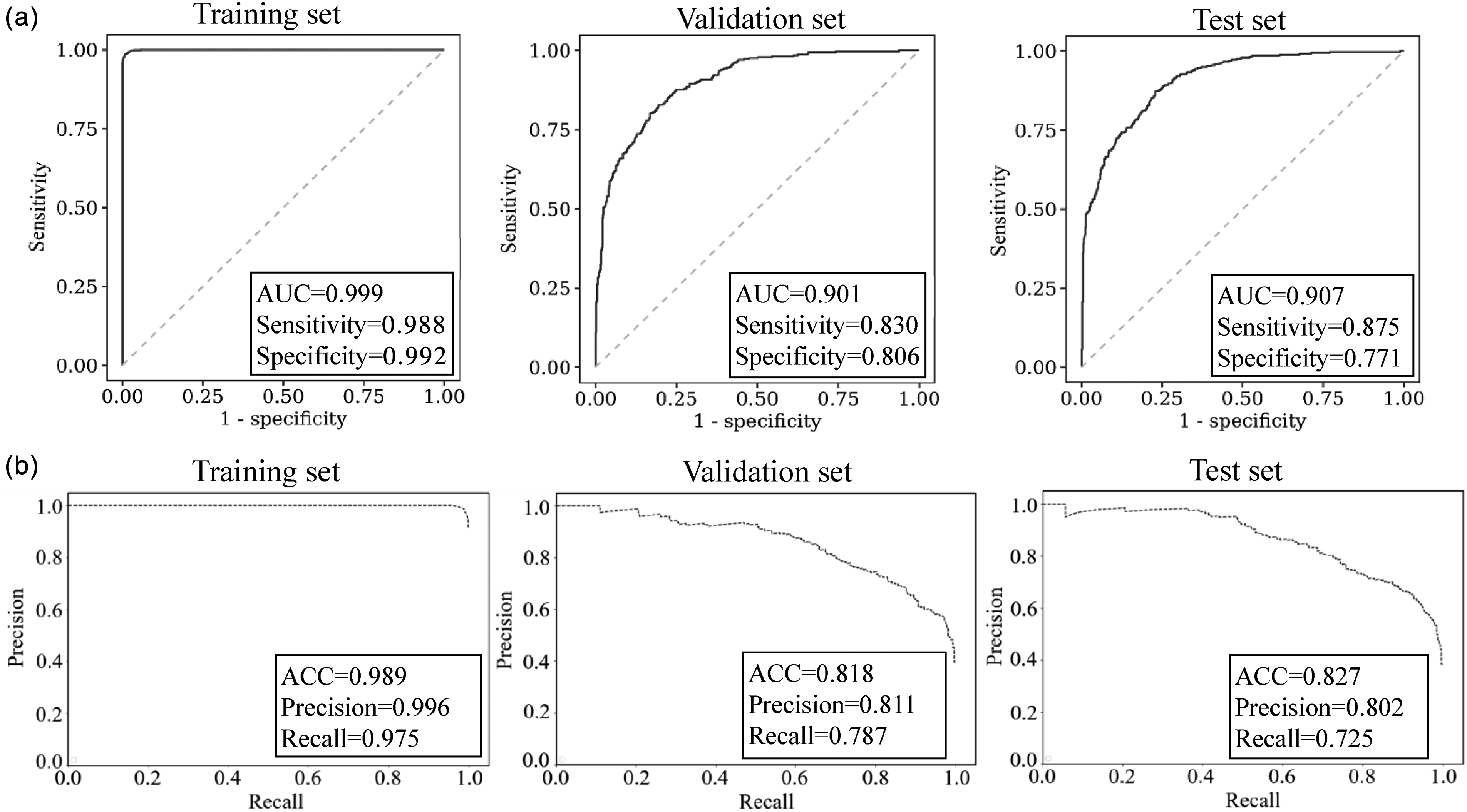
Diagnostic efficiency of the XHGC20 model for distinguishing between gastric cancer and precancerous disease. (a) ROC curve of the XHGC20 model in the training, validation and test set, respectively. (b) PR curve of the XHGC20 model in the training, validation and test set, respectively. PR: precision–recall.
Diagnostic efficiency of the XHGC20 model in gastric cancer.

AUC: area under the receiver operating characteristic curve; CI: confidence interval; ACC: accuracy; PPV: positive predictive value; NPV: negative predictive value; Sens.:sensitivity; Spec.:specificity.
The consistency between the XGBoost model and clinical diagnosis
The agreement between the prediction behaviour of the XHGC20 model and the actual diagnosis was further analysed. In all the training, validation and test sets, the patients’ risk scores were consistent with the clinical diagnosis results (Figure 4(a)). The upper segment of the histogram above the zero score line showed that some of the GC cases predicted by the model actually involved pathologically benign disease (false-positives: 25/1003 in the training set, 107/335 in the validation set and 92/335 in the test set). The lower segment of the histogram beneath the zero score line indicates that some patients with a pathological diagnosis of GC were predicted to have benign disease (false-negatives: 4/1637 in the training set, 53/545 in the validation set and 60/545 in the test set). As seen from the calibration curve (Figure 4(b)), the prediction value of the model was close to the actual diagnosis probability. The calibration curve was an evaluation index suitable for probabilistic models such as XGBoost. The curve was constructed with the predicted value as the abscissa and the real value as the ordinate. The closer the calibration curve to the diagonal, the better the performance of the model. The Brier scores of the training set, the validation set and the test set were 0.011, 0.182 and 0.173, respectively. The Brier score ranged from 0 to 1, with a smaller value indicating a higher accuracy of the prediction model. A Brier score of 0.0–0.1 represents very good prediction accuracy, and 0.1–0.2 represents good prediction accuracy. The consistency of the total number of patients in the GC and precancerous disease groups predicted by the training and validation sets was compared with the pathological diagnosis results, and the Chi-square test showed good consistency (χ2 = 340.521,

Consistency analysis between the model risk score and clinical diagnosis based on the XHGC20 model. (a) Risk score in the training, validation and test set, respectively. (b) Calibration curve of the XHGC20 model in the training, validation and test set, respectively. FP: false-positive rate; FN: false-negative rate.
Chi-square test of the XGBoost model for gastric cancer.

*
Diagnostic efficiency of the XGBoost model for early-stage GC
To evaluate the diagnostic effect of the XHGC20 model for early-stage GC, 64 patients in the validation cohort (accounting for 19.10%) and 65 patients in the test cohort (accounting for 19.40%) were included in the next analysis. The XHGC20 model had a high diagnostic efficiency in distinguishing between early-stage GC and precancerous disease. With respect to the validation cohort, the AUC was 0.888, the specificity was 0.807, and the sensitivity was 0.797 (cut-off value = 0.270). The efficacy in the test set was also good, with an AUC of 0.904, specificity of 0.881, sensitivity of 0.781 and cut-off value of 0.216 (Figure 5(a), Table 5). In addition, in the validation and test sets, the AUC of the XHGC20 model for distinguishing advanced GC from precancerous lesions was 0.904 and 0.907, respectively (Figure 5(b)). According to the pathological diagnosis results of the patients, the total number of precancerous lesions and early GC patients predicted by the training set and the validation set were analysed, and the Chi-square test revealed good consistency (χ2 = 109.737,

ROC curve of theXHGC20 model for diagnosis of early or advanced gastric cancer in the validation and test set. (a) ROC curve of the XHGC20 model for diagnosis of early gastric cancer. (b) ROC curve of the XHGC20 model for diagnosis of advanced gastric cancer.
Diagnostic efficiency of the XHGC20 model in early gastric cancer.
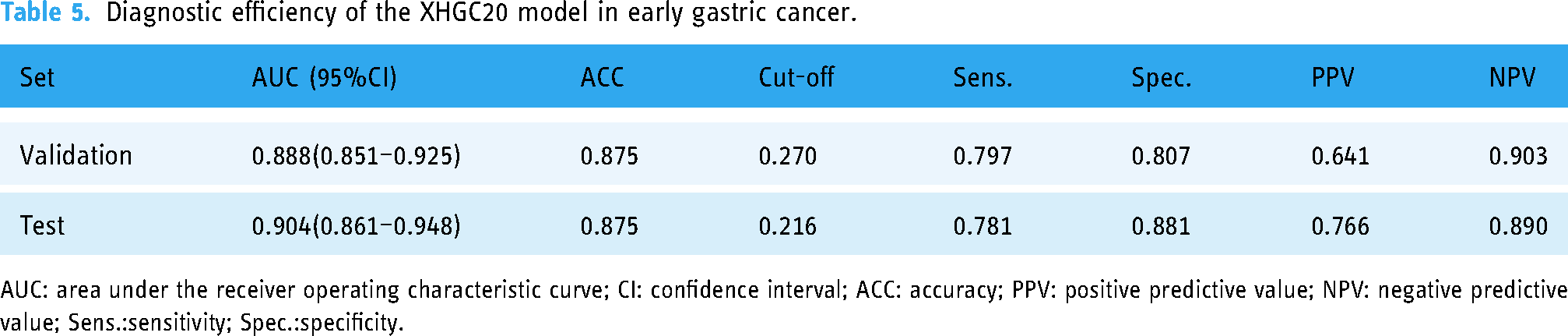
AUC: area under the receiver operating characteristic curve; CI: confidence interval; ACC: accuracy; PPV: positive predictive value; NPV: negative predictive value; Sens.:sensitivity; Spec.:specificity.
Diagnostic effect of the XGBoost model on patients with GC negative for tumour protein markers
We also analysed the diagnostic efficacy of four protein tumour markers commonly used in the diagnosis of gastrointestinal tumours for distinguishing GC from precancerous lesions in enrolled patients. According to the Roche Elecsys electrochemiluminescence instruction manual, the diagnostic thresholds for CEA, CA 19-9, CA 242 and CA 72-4 were 5.2 ng/mL, 39 U/mL, 20 U/mL and 6.9 U/mL, respectively. The AUC of the XHGC20 model was 0.9011, which was much greater than the AUCs of CEA, CA 19-9, CA 242 and CA 72-4 (0.647, 0.567, 0.557 and 0.583, respectively) (Table S4). The specificity and sensitivity of the combined use of four protein tumour markers for distinguishing GC from precancerous lesions were 0.414 and 0.764, respectively, which were also inferior to the prediction scores. The results showed that the diagnostic efficiency of the XHGC20 model for GC was better than that of protein tumour markers commonly used in the clinic.
Among the 1673 GC patients in our study, 1368 patients were tested for at least one conventional protein tumour marker, 1301 patients (95.10%) were negative for at least one marker, and 665 patients were negative for all four markers (false-negative rate = 0.566) (Table 6, Figure 6). When the established XHGC20 model was used to diagnose GC patients with negative results for the four markers alone or in combination, 933, 1009, 987, 877 and 593 patients were diagnosed with GC, accounting for approximately 90% of the patients. The AUCs were 0.968 (95% CI 0.961–0.975), 0.968 (95% CI 0.961–0.975), 0.975 (95% CI 0.969–0.981), 0.971 (95% CI 0.964–0.978) and 0.970 (95% CI 0.963–0.978) (Table 6, Figure 6). The established XHGC20 model could be used for identifying GC patients with negative gastrointestinal tumour marker data, possibly helping to reduce the rate of missed diagnoses of GC in a certain population.
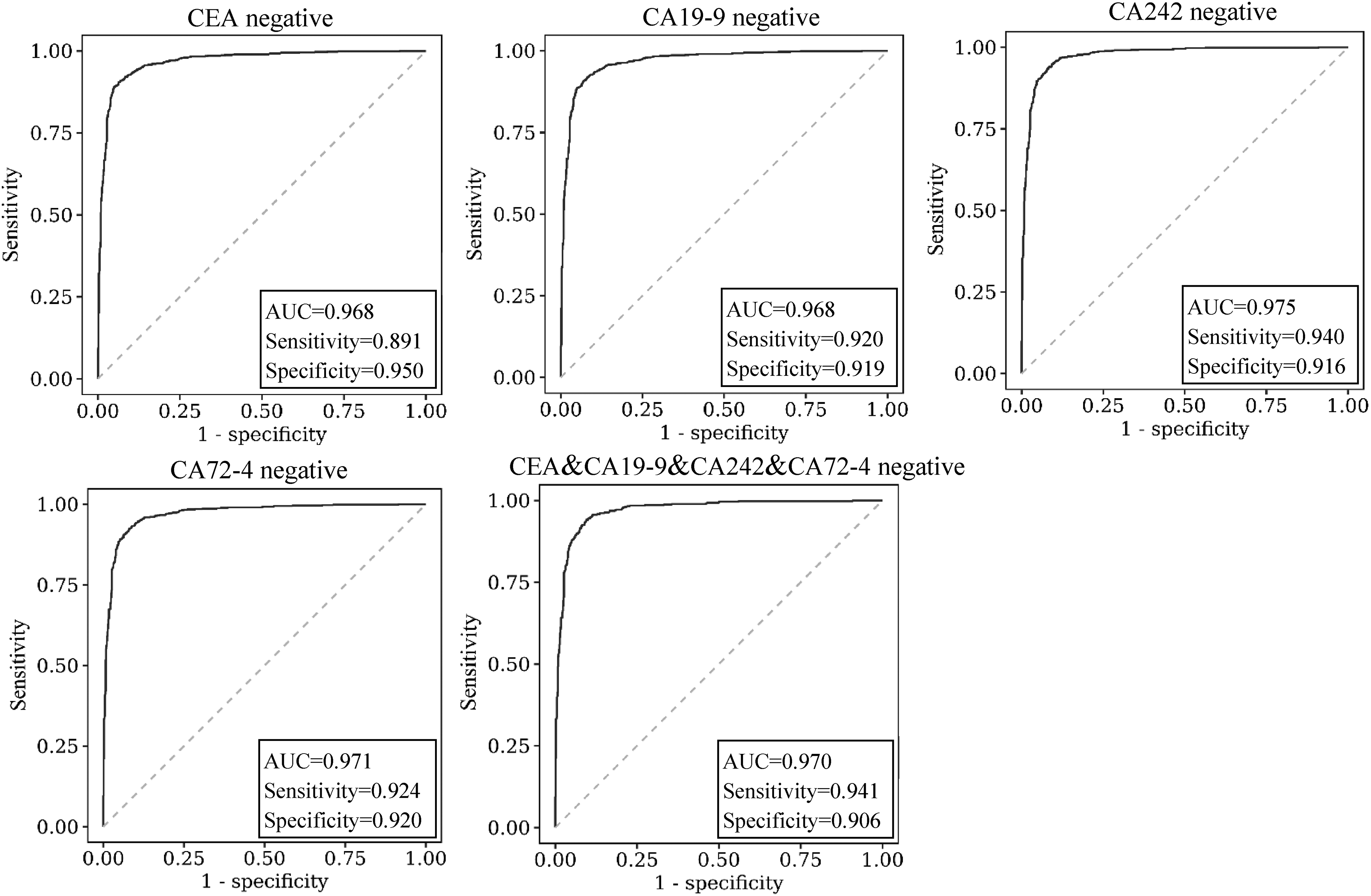
ROC curve of the XHGC20 model in distinguishing between tumour marker-negative gastric cancer and precancerous disease.
Diagnostic efficiency of the XHGC20 model in protein marker-negative gastric cancer.

AUC: area under the receiver operating characteristic curve; CI: confidence interval; ACC: accuracy; PPV: positive predictive value; NPV: negative predictive value; Sens.:sensitivity; Spec.:specificity; CEA: carcinoembryonic antigen; CA: carbohydrate antigen.
Diagnostic efficiency of the XGBoost model in the external validation set
To extrapolate the performance results of our model to other populations, we verified the model's diagnostic efficiency for GC on an external validation set. According to the inclusion and exclusion criteria for patients with GC and precancerous diseases, a total of 306 patients were included in the external validation set, and their clinical and pathological characteristics are shown in Table S6. According to the external validation set, the prediction score of the GC group was greater than that of the precancerous disease group [0.689 (95% CI 0.345–1.000) vs. 0.179 (95% CI 0.000–0.428),
Discussion
Early diagnosis of GC can significantly improve the survival rate and reduce the mortality rate. 36 At present, GC is mainly diagnosed by endoscopy and pathological biopsy combined with clinical symptoms. 37 Although the results are intuitive and accurate, these methods are invasive, and patient compliance is poor, making the approaches unsuitable for large-scale screening. 38 The results of endoscopic examination are easily affected by instrument performance and the visual field, and there is a certain rate of missed early-stage GC detections.39,40 In addition, the sensitivity and specificity of traditional laboratory markers for the diagnosis of early GC are not very good in clinical application, as using these markers can easily lead to missed diagnoses and unsatisfactory medical experiences. 41 Detection or diagnostic tools developed using ML methods have a wide range of clinical applications and are being rapidly appearing because they can positively impact doctors’ diagnosis and treatment activities.42–50 Using ML technology to investigate an ideal multi-indicator combined diagnosis method could be the basis of a most promising breakthrough in the early detection of GC.51,52
The XGBoost algorithm based on gradient boosting decision trees has been widely used in the medical field. Pan et al. constructed an eight-factor XGBoost model to predict the risk of death in ICU patients with COVID-19 that demonstrated good predictive ability.
53
Jiang et al. developed a predictive model using XGBoost and CT image-based deep learning to predict preoperative microvascular invasion in patients with hepatocellular carcinoma.
54
The XGBoost algorithm has also been widely used in cancer risk prediction and prognosis analysis. Leung et al. evaluated the properties of seven different ML models for predicting GC risk after
The XHGC20 model included 20 individual variables with potential contributions, which were divided into four main categories, excluding sex. Of these 20 variables, the most significant feature was total protein (characteristic coefficient = 0.1662). The decrease in total protein in GC was due to the rapid growth of malignant tumour cells, which consume a large amount of nutrients in the body, resulting in a decrease in hepatocyte synthesis. Malnutrition is a serious problem in GC patients and may lead to hypoproteinaemia, impaired organ function and decreased immunity.58,59 The second most important feature was FDPs, the characteristic coefficient of which was 0.0832. Hypercoagulability is associated with tumour development and may play a role in metastasis. Zhang et al. analysed the clinical characteristics of patients in three tertiary referral centres and suggested that FDP was the most effective indicator for predicting GC peritoneal metastasis, liver metastasis or bone metastasis, followed by
Compared with traditional tumour markers, which were useful for diagnosing GC, this model has great diagnostic efficacy for total GC. The AUCs of the validation set and test set were 0.901 and 0.907, respectively. When the cut-off value was 0.265, the sensitivity and specificity of the XHGC20 model were 0.830 and 0.806, respectively, in the validation cohort. When the XHGC20-derived prediction score was used to assist in the diagnosis of early GC, the AUCs of the validation set and test set were 0.888 and 0.904, respectively. When the cut-off value of the validation set was 0.270, the sensitivity and specificity were 0.797 and 0.807, respectively. In addition, the calibration curve showed that the prediction results of the XHGC20 model were in good agreement with the actual diagnosis in both general and early GC patients. These results indicated that this model could be used as an auxiliary tool for the diagnosis of early-stage GC and could provide reliable guidance for clinical practice. To a certain extent, this approach has solved the difficult problem of early diagnosis of GC by traditional methods and has a positive impact on the treatment and prognosis of these patients.
Another purpose of this study was to analyse the diagnostic efficacy of the XHGC20 model in GC patients with negative detection results according to conventional tumour markers. The sensitivity and specificity of the conventional protein tumour markers CEA, CA19-9, CA242 and CA72-4 in patients with GC were low, often leading to misdiagnosis of GC. In this study, among 1673 GC patients, 95.10% were negative for at least one tumour protein marker and 56.55% were negative for all four tumour protein markers. The XHGC20 model was effective at diagnosing patients with negative results for four tumour markers (AUC = 0.970), and the false-negative rate was only 0.108. Notably, the efficacy of this model in GC patients negative for four tumour markers was lower than that in GC patients negative for a single tumour marker, exhibiting consistency with the actual situation and proving that the detection of XHGC20-related markers has a good auxiliary role in the diagnosis of GC. Furthermore, the external validation population we selected was independent of the modelling population, and we found that the performance of the established predictive model could be extrapolated to the external validation population. The model was also very effective in the diagnosis of early GC and protein marker negative GC. Although PG and G17 play a guiding role in the monitoring of atrophic gastritis and GC, they are not widely used due to incomplete coverage by national medical insurance. Since more than 30% of the PGI and PGII data were missing, these data were filtered out during the data screening process. Our XHGC20 model overcomes the shortcomings of patients’ low compliance with invasive methods, barium meal and other imaging examinations and fully integrates routine laboratory physical examination screening indicators.
The population included in the model included newly admitted patients from Xinhua Hospital. This hospital receives a large number of tumour patients from all over the country every day, and a large number of GC patients are diagnosed, which aided in the development of this study. In this study, patient information and laboratory test results were obtained from the database after ethical review. The laboratory indicators included in the XHGC20 model were non-invasive blood test results, involving general requirements for testing personnel and equipment, low cost and good patient compliance. The XGBoost model can be used to improve the accuracy of the combined diagnosis of multiple serum markers. A prediction score combining 20 serum markers can distinguish not only GC patients but also early-stage patients from precancerous lesions. This strong evidence proves that the XHGC20 model is a promising tool for assisting in the diagnosis and screening of early-stage GC. Based on the good accuracy and low cost of the XHGC20 model, it is expected to be useful as an intelligent tool for screening high-risk patients for primary prevention. Large prospective cohort studies can further determine whether individuals identified as part of the high-risk group by the XHGC20 model will be diagnosed with GC in subsequent years.
Conclusion
We used the XGBoost algorithm to establish a GC diagnosis model XHGC20 which integrates 20 non-invasive detection indicators. The diagnostic efficiency of this model was satisfactory, and it could significantly improve the ability to recognise GC and distinguish early-stage GC from precancerous lesions. These modelling data are easy to obtain and convenient for clinical application. The diagnostic model integrates a variety of clinical laboratory data and helps to improve the ability of laboratory auxiliary diagnosis and prediction. Future studies need to include more non-invasive markers and larger populations to improve the accuracy and reliability of the model.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076241240905 - Supplemental material for Predicting early gastric cancer risk using machine learning: A population-based retrospective study
Supplemental material, sj-docx-1-dhj-10.1177_20552076241240905 for Predicting early gastric cancer risk using machine learning: A population-based retrospective study by Xing Ke, Xinyu Cai, Bingxian Bian, Yuanheng Shen, Yunlan Zhou, Wei Liu, Xu Wang, Lisong Shen and Junyao Yang in DIGITAL HEALTH
Footnotes
Acknowledgements
The authors thank the teams of Beckman Coulter Commercial Enterprise (China) Co., Ltd and Beijing Deepwise & League of PHD Technology Co., Ltd for their technical assistance.
Contributorship
Ke X, Cai X and Bian B designed the study and wrote the main manuscript. Shen Y, Zhou Y and Liu W helped to collect data and revise the manuscript. Wang X, Shen L and Yang J reviewed and edited the main manuscript. All authors reviewed the manuscript.
Data availability statement
The datasets used during the current study are available from the corresponding author on reasonable request.
Declaration of conflicting interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: All authors have reviewed the final version of the manuscript and approved it for publication.
Ethics approval
The ethical approval of this research was obtained from the Ethics Committee of Xinhua Hospital, Shanghai Jiao Tong University School of Medicine (Approval No. XHEC-D-2023-163). All patients had signed informed consent.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported in part by the National Natural Science Foundation of China (grant number 81802082 and 81672363); and the Natural Science Foundation of Shanghai Science and Technology Innovation Action Plan (grant number 21ZR1441500); and the Clinical Research Project of Shanghai Municipal Health Commission (grant number 202340054); and the Science and Technology Commission of Shanghai Municipality (grant number 20JC1410100); and the Hangzhou Qianjiang Plan Program.
Guarantor
LS
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
