Abstract
Objective
The goal of this work is to show how to implement a mixed reality application (app) for neurosurgery planning based on neuroimaging data, highlighting the strengths and weaknesses of its design.
Methods
Our workflow explains how to handle neuroimaging data, including how to load morphological, functional and diffusion tensor imaging data into a mixed reality environment, thus creating a first guide of this kind. Brain magnetic resonance imaging data from a paediatric patient were acquired using a 3 T Siemens Magnetom Skyra scanner. Initially, this raw data underwent specific software pre-processing and were subsequently transformed to ensure seamless integration with the mixed reality app. After that, we created three-dimensional models of brain structures and the mixed reality environment using Unity™ engine together with Microsoft® HoloLens 2™ device. To get an evaluation of the app we submitted a questionnaire to four neurosurgeons. To collect data concerning the performance of a user session we used Unity Performance Profiler.
Results
The use of the interactive features, such as rotating, scaling and moving models and browsing through menus, provided by the app had high scores in the questionnaire, and their use can still be improved as suggested by the performance data collected. The questionnaire's average scores were high, so the overall experiences of using our mixed reality app were positive.
Conclusion
We have successfully created a valuable and easy-to-use neuroimaging data mixed reality app, laying the foundation for more future clinical uses, as more models and data derived from various biomedical images can be imported.
Introduction
Images from computed tomography (CT) and magnetic resonance imaging (MRI) are a cornerstone of diagnostic clinical practice and decision-making for surgical planning, as they allow to reproduce patient brain anatomy, offering important information about anatomic-functional organization in brain areas that are damaged or near tumour mass. 1 Medical data visualization is routinely done using dedicated software, where images are manipulated using a computer keyboard and mouse pad, which require surgeons to mentally reconstruct the two-dimensional information in three dimensions to better understand the cerebral structures anatomy. In this context, mixed reality (MR) technologies could represent a helpful tool for healthcare professionals, since they provide an immersive experience to the user, where three-dimensional (3D) anatomical structures are superimposed on the real-world environment.
Among available MR technologies, Microsoft® HoloLens 2™ (hereafter called HoloLens 2) is Microsoft's second generation of visors, a device conceived and built for industrial, design and healthcare use. 2 HoloLens 2 device offers an immersive experience to users, where they can interact with virtual 3D objects, i.e., holograms, surrounded by real-world environment. Holograms manipulation, (e.g., placement, movement, rotation, and scaling) is ensured by instinctive and intuitive hand movements through specific recognition systems. Compared with the first-generation Microsoft® HoloLens™ (hereafter called HoloLens), the HoloLens 2 is designed with numerous improvements in order to optimize the user experience (see the ‘Device and App development’ section. Several examples of HoloLens devices applications were reported in literature, ranging from surgical planning3–6 to neuro-rehabilitation purposes.7–11 The use of HoloLens has been associated with significant time reduction for tasks requiring spatial understanding of the liver anatomy. 12 More in general, thanks to HoloLens 2 healthcare providers reduced training time by 30%, at an average savings of $63 per labour hour. 13 Moreover, MR technologies represent effective tools during surgery to provide proper visualization of the operative field. The alteration of the visual-motor axis caused by monitor position in laparoscopic surgery can lead to decreased ergonomics and surgical performance, spatial disorientation, and increased risk of iatrogenic injury. 14 The shift from on-screen visualization to MR devices paves the way for new opportunities. By placing virtual objects in the real environment, the MR technology inserts the users in a real-time and immersive interaction, thus allowing treatments and procedures to be planned in more detail and with greater accuracy. 15
Previous MR workflows for medical practice included first-generation HoloLens applied to laparoscopic liver resection and congenital heart surgery. 3 In this paper, we propose and describe the first original MR workflow using the second generation of head-mounted display (HMD) HoloLens 2 in neurosurgery. Moreover, we present a dedicated MR app created to guide neurosurgery planning on paediatric patients with epilepsy. In the case of patients with drug-resistant focal epilepsy, MRI techniques are pivotal tools in preoperative planning. The characterization of eloquent cortices surrounding the epileptic focus is crucial to achieve. 16 In this context, morphologic MRI, together with functional MRI (fMRI) and diffusion tensor imaging (DTI) provide important anatomical information about epilepsy, locating the epileptic focus and performing the functional mapping. By loading 3D structures from morphologic, fMRI, and DTI images on HoloLens 2, surgeons can better visualize brain structures and estimate the damage location and extension, thus improving surgical planning.
The application we present in this article is designed to be an important tool that can help and guide neurosurgery planning, enabling surgeons to explore in an immersive and interactive way the MR environment created from morphologic, fMRI, and DTI images. This provides them the possibility to better visualize brain structures and estimate the damage location and extension, which is fundamental in patients with epilepsy.
Methods
The nature of this study is to outline the steps leading to the creation of an MR app based on MRI data, presenting in detail how MR technology can be used to improve neurosurgical planning, and highlighting its current and potential benefits and clinical applications.
Since typical neuroimaging file formats cannot be directly used on Unity™ (hereafter called Unity), the first step of our workflow addresses compatibility. An automated process was implemented using the Python programming language to make conversions from Neuroimaging Informatics Technology Initiative (NIfTI) and Visualization Toolkit (VTK) formats to formats readable by Unity (see the ‘Data processing’ section). Secondly, we coded the actual app for HoloLens 2 with the models derived from these conversions using the C# programming language. The steps we carried out are summarized in Figure 1. The volumetric, fMRI, DTI processing (Figure 1(a to c)) was conducted using Python, and then the app creation and its features implementations (Figure 1(d to f)) were carried out using C#, which is supported in the Unity software development environment.

Summary scheme of implemented pipelines. (a) Volumetric data processing: From the NIfTI T1w image to the correctly positioned 3D models (see 'Data processing' section); (b) fMRI data processing in Python: from the NIfTI activation maps to the projected activation values (see 'Data processing' section); (c) DTI data processing in Python, detail of the AF: from the VTK points-lines model to the OBJ points-surface model with less filaments and changed coordinate system (see 'Data processing' section); (d–f) screens from the HoloLens 2 point of view of some features introduced in the MR app (see 'Device and app development' section); (d) possibility to visualize brain thickness values over the cortex; (e) possibility to visualize brain activation values corresponding to right-hand movement over the cortex; (f) possibility to make the cerebral hemispheres transparent for the CST and AF visualization. AF: arcuate fasciculus; CST: corticospinal tract; fMRI: functional magnetic resonance imaging; 3D: three-dimensional; DTI: diffusion tensor imaging; NIfTI: Neuroimaging Informatics Technology Initiative; VTK: Visualization Toolkit.
Data acquisition
Written informed consent from patients was acquired before data acquisition and processing. The MRI data from an epileptic patient (15 years old, female) was acquired in accordance with the standardized preoperative study acquisition as performed in patients undergoing surgical treatment. The brain MRI data from the patient were obtained at Bambino Gesù Children's Hospital (Rome) on a 3 T Siemens Magnetom Skyra scanner (Siemens Medical Systems, Erlangen, Germany) equipped with 32 channels head-coil (coil dimensions L-W-H: 440 mm × 330 mm × 370 mm). Image protocol consisted of: T1-weighted sagittal 3D magnetization-prepared rapid gradient-echo sequence (TR = 770 ms, TE = 2.27 ms, TI = 1040 ms, FA = 9°, ST = 0.8 mm) with in-plane acquisition matrix of 288 × 288 pixels, transversal bold sequence (TR = 2900 ms, TE = 30 ms, FA = 90°, ST = 4 mm) with acquisition matrix 370 × 370, transversal bold sequence (TR = 3000 ms, TE = 30 ms, FA = 90°, ST = 4 mm) with acquisition matrix 370 × 370, transversal dynamic bold sequence (TR = 2700 ms, TE = 30 ms, FA = 90°, ST = 4 mm) with acquisition matrix 456 × 456, transversal DTI sequence (TR = 9100 ms, TE = 99 ms, FA = 90°, ST = 2 mm) with acquisition matrix 896 × 896. Particularly, in this work we used a T1w image, four fMRI images related to cognitive and motor tasks, and two DTI models representing white matter (WM) fibres as source data. The T1w image in NIfTI format was obtained by the T1-weighted sequence. The following cognitive and motor-related fMRI tasks in NIfTI format were acquired: (1) imagination of toys, acquired using the first bold sequence of the protocol, (2) listening to a fairy tale, acquired using the second bold sequence of the protocol, (3) right-hand movement and (4) left-hand movement both acquired using the dynamic bold sequence of the protocol. Those task activations were spatially co-registered to the morphological T1w image. From the DTI sequence, we derived two VTK-format WM tracts in the left hemisphere: Corticospinal tract (CST) and arcuate fasciculus (AF). The CST extends from the cerebral cortex to lower motor neurons and interneurons in the spinal cord and controls limb and trunk movements. The AF is a bundle of axons that connects two speech areas, e.g., Broca's area of the frontal lobe and Wernicke's area of the temporal lobe, respectively responsible for language production and language comprehension (both written and spoken).
Data processing
Raw data from MRI sequences were firstly pre-processed with specific software and then converted to be properly loaded on the MR app. Specifically, after the said pre-processing performed with FreeSurfer and MRtrix3, a conversion pipeline was entirely developed in Python programming language and allowed to automatically perform conversion of volumetric, functional, and diffusion data into a specific format suitable for the MR app.
Volumetric data of the T1w image were firstly pre-processed with FreeSurfer 7.3.2 software, 17 using a standard automatic pipeline (i.e., recon-all) that sequentially performed skull stripping, intensity correction, and transformation to Talairach-Tournoux space to produce grey matter (GM) and WM segmentation. Combining information from tissue intensity and neighbourhood constraints, the GM–WM boundary was first determined and then tessellated to generate the inner cortical surface (white surface). The outer surface (pial surface) was then generated through the expansion of the white surface with a point-to-point correspondence, thus obtaining a 3D model for both left and right hemispheres. Based on brain surfaces, several cortical parameters were extracted, including cortical thickness and sulcal depth. Finally, we applied the implemented Python pipeline section dedicated to the volumetric data processing, that consisted of (1) switching of FreeSurfer outputs orientation, (2) translation of those 3D models, and (3) storing 3D models’ vertices and faces in txt format files. In particular, for visualization purposes, FreeSurfer outputs were switched from their left-posterior-superior (LPS) orientation to the right-anterior-superior (RAS) orientation of 3D Slicer (i.e., software we used for visualization of neuroimaging data). Due to a shift between the model origin (0, 0, 0) and the image centre, a translation was performed on 3D structures to produce models properly overlapping the image from which they were extracted. In the end, recon-all outputs were converted in txt format to be used as 3D models in the MR app.
DTI data was pre-processed using MRtrix3. 18 Data was denoised via PCA algorithm (dwidenoise) and corrected for distortion due to eddy currents via ACID toolbox. 19 We identified CST and AF using constrained spherical deconvolution (CSD), as provided in MRtrix3. 20 To perform automatic tracking and minimize manual inputs, different regions of interest (ROIs) were manually drawn on the high-resolution anatomical template. 21 Fibre tracking was performed through the CSD streamlines algorithm, computing the pathways between a pair of ROIs. Finally, we applied the implemented Python pipeline section dedicated to the DTI data processing, consisting of: (1) random extraction of 10% of CST and AF filaments, (2) surfaces creation around CST and AF filaments lines, (3) CST and AF coordinate system conversion and (4) 3D models conversion to Wavefront OBJ format (hereafter called OBJ). In particular, the number of fibres composing the CST and AF 3D models was very large, preventing a clear interpretation of the bundles shape. To improve shape interpretability, we decided to randomly extract only 10% of the total filaments from the VTK files of CST and AF, but they can also be 100% loaded in the MR project. Then, in order to produce mesh elements (composed by faces and vertices), MRTrix3 output was firstly manipulated creating a tubular surface around the lines. Subsequently, coordinate system conversion (from LPS to RAS system) and format conversion (from VTK to OBJ) were performed, similarly to volumetric data.
Since cognitive and motor-related tasks files were 3D activation maps of the brain in NIfTI format, we needed to perform surface projection in order to visualize them on the cortex and store the projection values in a usable file format. We applied the implemented Python pipeline section dedicated to the fMRI data processing, through which the following actions were performed: (1) volume activation values at the vertices of the cortical meshes were computed by sampling points around each vertex, (2) activation at these points was sampled, (3) average samples were associated with each vertex, and (4) the projection values were saved in txt format.
Development tools for mixed reality
Device and app development
The MR device selected for our project was HoloLens 2, produced and developed by Microsoft®. This is a HMD mounting Windows 10 operating system, and represents an improvement of the first type of HoloLens. As reported in Table 1, HoloLens 2 has several upgrades compared to the previous model,22–24 including improved resolution and an increased field of view, that result in less eye fatigue with longer use of the headset. Both HoloLens and HoloLens2 have voice input recognition, but the last model has one more microphone array with respect to the first version, ensuring better performance for voice command. Also, gesture input was improved, allowing both hands control. Additionally, HoloLens 2 implements eye-tracking, which enables useful features such as selecting a button by simply blinking or enlarging a text by looking at it. Although both versions have a custom-made Microsoft® Holographic Processing Unit (HPU), the one in the HoloLens 2 has been improved to better interpret measurements from the gyroscope, accelerometer, magnetometer and better handle voice and gesture inputs.
Main technical features of HoloLens and HoloLens 2.

FOV: field of view; HPU: holographic processing unit.
We decided to develop our app in Unity, a cross-platform graphics engine (or game engine) developed by Unity Technologies© and released in 2005. The Unity development environment enables the creation of interactive content (e.g., video games or mobile applications), with graphics processed in real time. Unity is also a cross-platform graphics engine, as it ensures final product compatibility with multiple different platforms, such as PC, smartphones, PlayStation, Xbox, and virtual, augmented, and MR glasses. Since Unity supports scripting in C#, we decided to use Visual Studio® 2019 as an integrated development environment. Our MR application was developed with the OpenXR™ standard, which provides an application programming interface for virtual, augmented or MR hardware. To accelerate MR app development in Unity we used the Mixed Reality Toolkit (MRTK) that provides a set of components for spatial interactions and user interface.
Gestures included touch, touch and hold, two-finger pinch (e.g., to move a slider), one-handed manipulation to move a hologram, or two-handed manipulation to rotate and magnify it. Finally, spatial mapping allowed surfaces in the environment where the user moves (e.g., walls, floors and desks) to be detected as meshes. Specifically, HoloLens 2 automatically builds triangular face meshes on surfaces, that can be set as visible or not by the user. In order to get a realistic view of the holograms, we set the holograms to be obscured by surfaces that maybe be interposed with the user.
Files used within Unity project, i.e., assets, can be imported or directly created on Unity. Since the goal of our project was the visualization of brain characteristics and activations from a paediatric patient with epilepsy, it was important that the reconstructed 3D models preserved the number and position of vertices within the mesh. For this purpose, the 3D models were recreated directly in Unity, in this way we avoided Unity automatic adjustments that would have taken place by directly importing an external 3D model. In order to recreate the cerebral 3D models directly on Unity, txt files created with the Python pipeline containing mesh vertices and faces of the cerebral cortex were read and processed through a C# script. We used 3D Slicer to verify the correct placement of the 3D models within the Unity project.
We associated a set of MRTK scripts to each 3D model, thus allowing user interaction (grab, rotate, enlarge, and shrink) with objects, controlled by hand gestures and their relative inputs via holographic buttons. We decided to attach the CST and AF meshes to the 3D model of the left hemisphere, so they can move and enlarge together as one block. To set the visibility (i.e., transparency) of the CST and AF models, a function has been implemented and a shader was assigned to the meshes through a script computing the colours of each rendered pixel. By moving a slider hologram, the user can set the value of the Alpha colour parameter of the shader assigned to the meshes as shown in Figure 2.

Screen captures of the app as seen by the HoloLens 2. (a) Slider set to its maximum value, corresponding to fully opaque meshes; (b) Slider set to its minimum value, corresponding to fully transparent meshes.
Moreover, 3D models were shown or hidden by a specific function based on changing a Boolean variable when the user presses the corresponding button. Brain activation and cortex features’ values saved in txt were converted to colours and displayed on the cortex meshes. To do this, a function with multiple methods was created, which changed the colours of the meshes in accordance with the txt values by pressing the button holograms. To improve the speed of the application we saved this meshes colours information and loaded it already set, so the HoloLens 2 did not have to perform in real-time the values-to-colours conversion and vertex association, avoiding slow for loops. The chosen colour scale ranges from blue to red, where blue is related to lower values while red is associated with higher values.
User interface design
When HoloLens 2 was worn and the MR app was launched, 3D models (holograms) and a graphical interface consisting of various holographic menus appeared in front of the user (see Figure 3). The displayed menus are three and consist of various buttons to show/hide the meshes (Figure 3(a), top left), a slider to control the transparency of the cerebral cortex (Figure 3(a), top right), and various buttons used to select the brain feature or activation to be displayed on the cortex (Figure 3(a), bottom). In addition, a small menu was set up to follow the user during movement, allowing him to hide/show the other menus or to reload the scene to put all the models back to their original position (Figure 3(b)).

Screen captures of the app project in Unity (left) and the app as seen by the HoloLens 2 (right). (a) Scene containing the three menus and the three-dimensional (3D) brain and nerves models; (b) Menu that follows the user.
The app created on Unity can be used either from the HoloLens 2 or from a pc via the Unity game window, which acts as a simulator. The use of the app from the HoloLens 2 can be streamed in real-time by a user not wearing the HMD via pc with the Microsoft HoloLens™ application.
Users’ evaluation
To get an evaluation of the MR app, we submitted the questionnaire proposed by Kumar et al. 3 to four neurosurgeons who had no prior experience using HoloLens 2. The statements in the questionnaire regarded the rating of (1) comfort level while wearing the HoloLens 2, (2) 3D depth perception of the models, (3) screen size of the HoloLens 2, (4) visibility when using HoloLens 2, (5) understanding of the morphology with HoloLens 2, (6) ability to walk around the model, (7) ability to rotate the model, (8) ability to scale the model, (9) ability to move the model, (10) ability to browse through the menus, (11) likelihood to recommend HoloLens 2 to others in their profession (see Table 2). For each of the questionnaire's statements, the neurosurgeons had to circle the response that best characterized how they feel about the statement. For statements 1–10 the responses could range from Extremely dissatisfied to Extremely satisfied, while for statement 11 the response could range from Extremely unlikely to Extremely likely.
Results
Users’ experience
The four neurosurgeons’ experience in using HoloLens 2 consisted of wearing the glasses, moving around the models, grasping, moving, enlarging and shrinking the models, and pressing menu buttons to activate the various features. The questionnaire's results are shown in Table 2.
Questionnaire's average scores on 6-point Likert scale for neurosurgeons (n = 4).

After an initial period in which the neurosurgeons had to figure out how to spatially relate to the holograms, interaction with the app was easier and button selection was faster. Menus with checkboxes immediately allowed the neurosurgeons to understand which meshes were being hidden/shown and which activation or feature was being observed relative to the cerebral cortex. The left and right hemispheres can be grasped individually or simultaneously to be moved and zoomed in or out. Zooming was particularly useful for better-observing areas of activation. By setting the transparency of the hemispheres through the slider, it was possible to observe the relationship between the activated cortical areas with respect to the AF and CST bundles beneath it. Since all scores were greater than or equal to 4.75, it can be concluded that the neurosurgeons’ overall experiences of using HoloLens 2 were positive.
Performance testing
In order to test how our MR app runs on HoloLens 2 and to get performance information, we used the Unity Performance Profiler. In particular, we marked up our C# code block methods by using BeginSample and EndSample static methods of the Profiler class. 25 We collected data concerning the performance of a user session, focusing on those tasks more responsible for application slowdown, which are the colouring of the cortical mesh according to brain activity and morphological characteristics and the restarting of the application itself. Each menu button involves the execution of a marked script in addition to those that are executed by default to run the application. During the test, an engineer with experience using HoloLens 2 assessed the performance indices for each enable/disable the buttons and sliders associated with brain morphological information and meshes as well as functional activations (Figures 1 to 3).
Tables 3 and 4 report performance information for each activity. The performances are expressed in terms of CPU (total time for the central processing unit to execute the instruction), TUM (total memory used by the app during the execution of the command), Time (time the app spends on running the script block only), GCA (bytes allocated by the garbage collector (GC) in a specific frame when the script block runs) and GCA T (GC allocating time taken by the script block only). In particular, Table 3 shows that after an activity or characteristic button is pressed, the app takes an average time of 129 ms to produce the output, where the 27% (35 ms) is for the code block. When the CPU exceeds 66 ms, the app goes under 15 frames per second (FPS). Command execution causes an average memory usage of 252 MB. When there is not enough free memory to make an allocation, Unity runs the GC, a memory recovery feature that automatically frees up memory space allocated to objects no longer needed by the program. Table 3 reports both GCA and GCA T of the code blocks, with average values respectively of 3251376 bytes (3.25 MB) and 0.37 ms. The GCA causes a major spike compared to the memory allocated by the GC when no button command is running, which is always on the order of 104 bytes (10−2 MB) (see Table 5). As it is possible to deduce from Table 4, when the user pushes the Reset button the app only takes 82.96 ms to produce the output, but the complete task (subsequent execution of Pink and Slider codes) requires an overall time of 1381.04 ms, with Pink command responsible for all the GCA. Moreover, Reset by itself does not cause an allocated memory spike, but the Pink command call does (see Table 6).
Values of app performances when an activity or characteristic button is pressed.

CPU: total time for the central processing unit to execute the instruction; TUM: total memory used; GCA: bytes allocated by the garbage collector.
Values of app performances when Reset button is pressed; values from Slider command are the same of Pink because they are called together

CPU: total time for the central processing unit to execute the instruction; TUM: total memory used; GCA: bytes allocated by the garbage collector.
Total bytes allocated by the garbage collector (GC) before and after a command execution
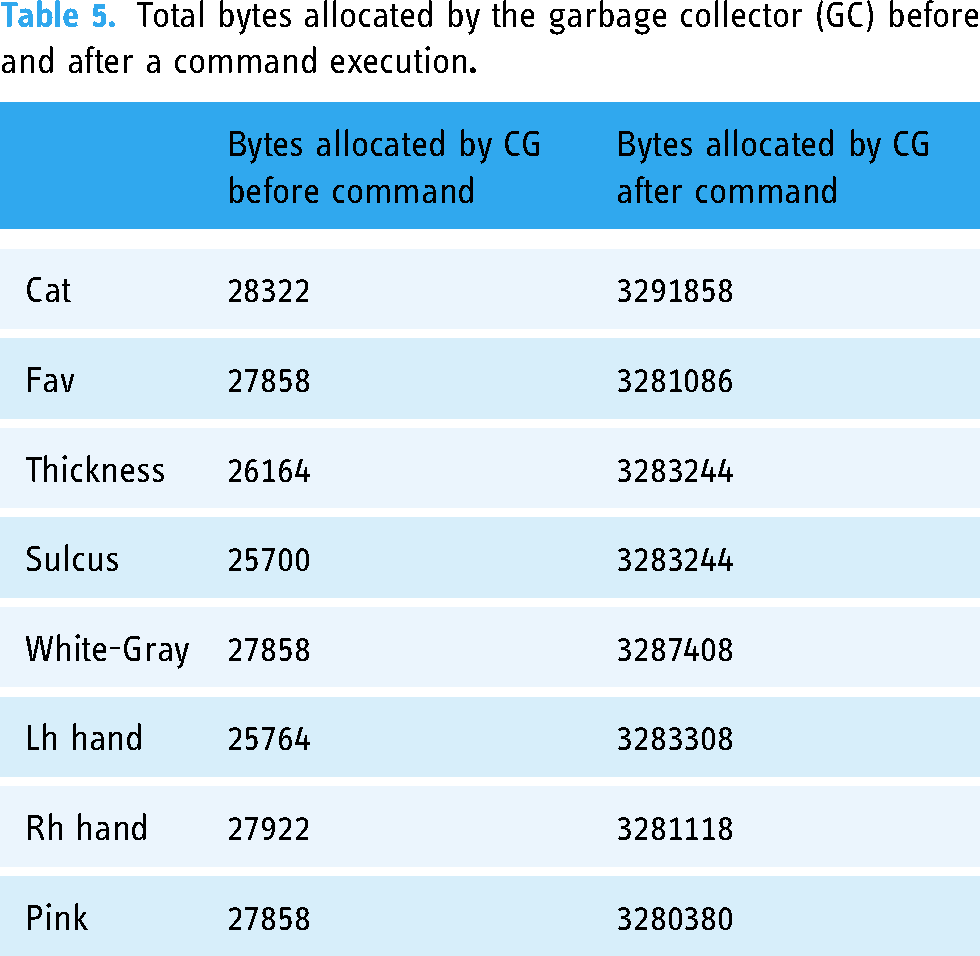
Total bytes allocated by the garbage collector (GC) before and after Reset button is pressed and after Pink button is triggered

Discussion
The purpose of this study was to create an MR app with practical applications in neurosurgical field, representing a first step toward the fruition of a technology that is expected to have a major impact on the medical field in the coming years. In fact, previous studies reported surgeons experience with the HoloLens app, which was perceived as a better tool for visualizing and analyzing neuroimaging data than classical visualization techniques3,26–29 In neurosurgery, the use of 3D printing was discussed. 30
The skull and its structures can be segmented with standard CT and MRI and successively printed with a 3D printer. In this context, Filho et al. have created an anatomical model that can provide reliable neuroendoscopic training, 31 thus demonstrating that 3D printed models can result in powerful neurosurgical training simulators. Furthermore, 3D printed anatomy integrated with an augmented reality system is an important addition to neurosurgery training, as shown for basic and advanced procedures of skull base approaches by Lee et al.. 32 This could bring further future potential to our work, which by integrating MR application with 3D printed structures could not only improve surgical planning but also train and enhance surgeons’ skills.
As reviewed by Cole et al., 33 there are many imaging modalities currently available to the practicing neurosurgeon, each with its own strengths. For this reason, we extended our application to include not only MRI but also computerized tomographic (CT) images. This choice was also made based on what has been reported in literature regarding the application of augmented reality in support of stereotactic radiosurgery (SRS), a specific treatment recognized as the most effective method for the management of high-grade cerebral arteriovenous malformations.34,35 Indeed, the use of time-lapsed computerized tomography angiography (4D-CTA), based on computerized tomography angiography (CTA) techniques, can support radiosurgery plans by providing less invasive, 3D techniques. 36 In this regard, we provide an example in Figure 4, where blood vessel models, reconstructed from CT, can be explored, examined, and manipulated. 37
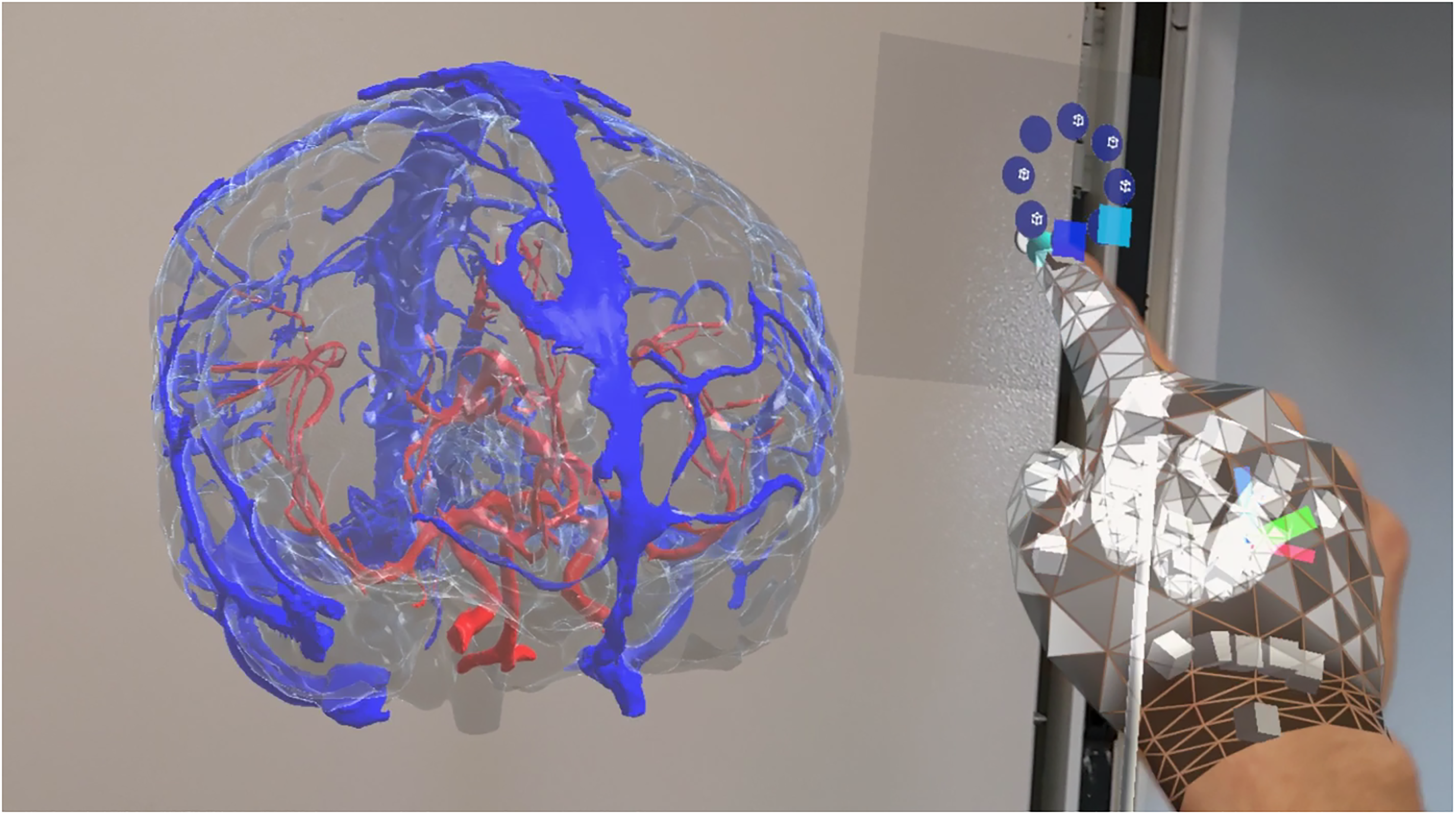
Screen capture of the app as seen by the HoloLens 2. In this scene, the user can visualize and interact with blood vessels reconstructed from computed tomographic (CT) scans.
The combination of SRS and augmented reality can help optimize SRS case planning by creating more efficient treatment planning workflows and ultimately optimizing the efficiency and safety of radiosurgical treatment, with the ultimate goal of improving patient outcomes and quality of care. 35
This app was indeed designed with the aim of supporting clinicians in the pre-operative study phase. It is also an important informative tool, as it provides an immersive experience to the patients or relatives (in the case of paediatric patient), who are consequently informed about the pathological condition and the surgical procedure by means of an intuitive and comfortable technology. In this context, we implemented an MR app as simple and intuitive as possible, even for those who have never used MR devices. Simple and well-designed menus immediately give an idea of what are the possible commands. As previously reported in a recent review, depth perception in real environments allows the user to get a better understanding of the patient anatomy, 3 and the interpretation of organ spatial organization (and the related tasks) requires less time than neuroimaging visualization with regular pc monitors. 12 This project provides an optimal workflow to convert the standard formats of neuroimaging data into format readable by Unity, and ultimately displayable on the HoloLens 2.
This MR study paves the way for further clinical application, as several 3D models from different biomedical images can be loaded within the MR app in the future. Moreover, spatial location can be configured to be shared among multiple users, in order to provide shared experiences where users interact with the same holograms in the same environment. The hand gesture feature, together with the voice command option made the HoloLens 2 a suitable device for usage in sterile environments, like the operating room. If necessary, additional morphological characteristics and brain activation can be loaded into the environment.
Our study has a limitation, specifically the very large number of vertices that compose each hemisphere could be responsible for the low FPS (CPU = 129 ms) of the app when executing brain activity and morphological characteristics visualization commands. Indeed, left hemisphere mesh has 93303 vertices (5.0 MB) and 186602 triangular faces (2.1 MB), while right hemisphere has 109904 vertices (5.9 MB) and 219804 triangular faces (2.1 MB). In this context, one possible way to speed up the application could require vertices reduction and consequently mediate the values of various brain activities and morphological characteristics in the areas around the newly reduced number of vertices.
Conclusions
In this paper, we provided an accurate description of each step required for the creation of an MR app for paediatric neuroimaging data visualization. We have effectively succeeded in creating a realistic holographic experience from MRI, allowing the surgeon to move freely around patient anatomy, and to visualize brain characteristics and activations on the cerebral cortex with colour maps.
Footnotes
Acknowledgements
The authors gratefully acknowledge the financial support of Fondazione Enrico ed Enrica Sovena.
Contributorship
A.N., M.A., M.L., and C.P. researched the literature and conceived the study. M.A., C.P., A.N. set up and used the software. M.A. and C.P. took care of coding. M.A., A.N., M.L., C.P., and A.D.B. were involved in data analysis. M.A. and A.N. administered the questionnaire. M.A. prepared the original draft of the manuscript. M.A., A.N., M.L., C.P., M.C.R.-E., L.F.T., C.G., A.S., L.P., E.P., L.D.P., C.E.M., A.M., and F.G. reviewed and edited the manuscript. All authors approved the final version of the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Informed consent statement
All subjects involved in the study agreed to participate in it and to be aware of what this study involves by signing a written informed consent prior to study initiation.
Ethical approval
The study was conducted according to the guidelines of the Declaration of Helsinki and approved by the ethical committee of Bambino Gesù Children's Hospital (protocol number 1867/2019).
