Abstract
Sleep is vital to many processes involved in the well-being and health of children; however, it is estimated that 80% of children with Rett syndrome suffer from sleep disorders. Caregiver reports and questionnaires, which are the current method of studying sleep, are prone to observer bias and missed information. Polysomnography is considered the gold standard for sleep analysis but is labor and cost-intensive and limits the frequency of data collection for sleep disorder studies. Wearable digital health technologies, such as actigraphy devices, have shown potential and feasibility as a method for sleep analysis in Rett syndrome, but have not been validated against polysomnography. Furthermore, the collected accelerometer data has limitations due to the rigidity, periodic limb movement, and involuntary muscle contractions prevalent in Rett syndrome. Heart rate and electrodermal activity, along with other physiological signals, have been linked to sleep stages and can be utilized with machine learning to provide better resistance to noise and false positives than actigraphy. This research aims to address the gap in Rett syndrome sleep analysis by comparing the performance of a machine learning model utilizing both accelerometer data and physiological data features to the gold-standard polysomnography for sleep analysis in Rett syndrome. Our analytical validation pilot study (
Introduction
At an estimated prevalence of 1 in 10,000 females, Rett syndrome (RTT) accounts for up to 10% of genetically linked severe intellectual disabilities in females.1–3 RTT is associated with a spontaneous mutation in the methyl CPG binding protein 2 (MeCP2) gene, 3 located on the X-chromosome at Xq28. RTT is characterized by regression with loss of acquired spoken language and volitional hand use, disrupted or absent ambulation, and repetitive hand movements. 4 Associated clinical features include seizures, autonomic and breathing abnormalities, growth failure, scoliosis, gastrointestinal and nutritional symptoms, and impaired sleep.5,6
Sleep is regarded as essential for the well-being and health of children and is critical to many somatic, psychological, and cognitive processes.7,8 However, sleep problems are highly prevalent in RTT, with studies showing that around 80% of children with RTT suffer from sleep disorders.9,10 Disturbances, such as night-time laughing and screaming, sleep walking, and night terrors 11 adversely impact the quality of life for both the child with RTT and their families.12,13 Current methods of sleep evaluation in the home are limited to sleep diaries and caregiver-completed questionnaires,14,10 however, these methods are subjected to a number of biases, including recall and observer bias. Caregivers could miss night wakings 15 or misinterpret a child laying with their eyes closed as the child being asleep. Due to the severe communication impairments that accompany RTT, self-reports of sleep disturbances are not possible. Caregiver-independent, objective methods of sleep assessment are necessary to further explore sleep trends and the progression of sleep disturbances in this population. The gold standard for obtaining objective sleep measures is polysomnography (PSG), a procedure that uses a number of sensors to measure and record brain waves, oxygen levels, respiration rate, and heart rate, along with leg and eye movement to determine and classify sleep stages. 16 While highly effective in measuring sleep quality, PSG is resource intensive and impractical for longitudinal assessments. The unreliability of caregiver reports and the resource requirements of PSG highlights the need for new methods of effective sleep assessment that are suitable for longitudinal use in RTT research.
Recent research has explored the use of digital health technologies (DHTs), such as wearable accelerometers to measure body movements from the wrist (actigraphy), as a method of sleep analysis. 17 Actigraphy devices have shown promise when used with children with neurodevelopmental disorders (NDDs), such as Down syndrome 18 and autism spectrum disorders, 19 due to their ease of use and ability to capture data within the home environment for a prolonged period of time. Researchers have determined the feasibility of actigraphy for sleep analysis in RTT, 20 however, analytical validation of actigraphy against gold standard PSG has yet to be examined in RTT literature. In addition, individuals with RTT have a higher prevalence of involuntary muscle contractions, rigidity, 21 and increased prevalence of periodic limb movements during sleep, 22 which could potentially limit the interpretability of actigraphy alone. Currently, available models for sleep analysis and sleep scoring with wearable devices are not applicable for children with RTT due to the fact that the models are designed with data from typically developing adults 23 and greater sensitivity and specificity are required for this population. These limitations can be addressed with the addition of physiological data collection and a model specifically trained on data from children with RTT.
Distinct hormonal patterns and the underlying sub-cortical network of brain structures that govern sleep significantly influence physiology,24,25 meaning that changes in physiological signals can be correlated to sleep stages. As early as 1968, fluctuations in electrodermal activity (EDA) were found to increase during the late stages of non-rapid eye movement (NREM) sleep and decrease during rapid eye movement (REM) sleep. 26 In 1973 Aldredge et al. determined that heart rate averages trended higher in REM sleep and lower in NREM sleep with the variance in heart rate decreasing with the depth of sleep. 27 Other researchers found that heart rate variability (HRV) changes during sleep are highly individualized and vary based on the basal autonomic activity of each individual. 28 Sleep quality has been associated with both HR and HRV. 29 These works show that meaningful and distinct sleep characteristics can be determined by collecting both heart rate and inter-beat interval data. More recent studies have found that during NREM, many of the measurable physiological processes decrease when compared to being awake. These processes include brain activity, respiration, body temperature, and blood pressure. Alternatively, these signals show an uptick in measured values during REM sleep. 25 Wearable devices, such as the Empatica E4, make it possible to collect many of these physiological measures in a non-invasive way. As discussed previously, wearable devices that measure accelerometer data have been able to distinguish sleep vs wake periods, 30 however, these sensors struggle to differentiate between NREM and REM sleep. By combining accelerometer data with physiological data, it is possible to train a machine learning algorithm to predict sleep stages in children with RTT.
Work in the field of machine learning and sleep state analysis has focused on automating the labeling process of PSG 31 or reducing the need for a full PSG sleep study by combining in-home monitoring with machine learning.32,33 In their work, Mikkelsen et al. 34 evaluated the performance of a machine-learning algorithm based on input from a mobile around the ear electroencephalography (EEG) when compared to an actigraphy, using PSG as the ground truth. It was found that the EEG alone outperformed the actigraphy and was acceptable compared to the PSG, however, 85% of the participants reported that the around the ear EEG negatively influenced their sleep, to some degree. More often, sleep quality is considered to be the target measure. Studies have explored machine learning for sleep quality both with commercial smartwatches and clinical actigraphy with positive results.35,36 Commercial wearable devices, such as FitBit, claim to track sleep using their integrated sensors. However, studies have shown that the FitBit algorithms tend to overestimate total sleep time and struggle to correctly estimate light and deep sleep.37,38 It has been shown that adding HRV data and additional body movement measures did increase the accuracy of FitBit’s algorithm. 38 While these studies show the promise of combining machine learning and physiological data for sleep analysis, current work is based on typically developed adults38,35 and does not translate to the sleep patterns of children. The existing techniques and data for automated sleep analysis are even less applicable to children with special needs, such as RTT. While other works have shown promise using automated algorithms with physiology and accelerometer data to differentiate between autism 39 and to classify high severity and low severity RTT, 40 to our knowledge, there exist no studies that utilize automated algorithms with physiological and accelerometer data for sleep analysis of children with RTT.
This paper fills a gap in the research literature by examining the analytical validation of a wearable sensor-based sleep analysis against gold-standard PSG in RTT. We validate the combination of physiological and accelerometer data by training a machine learning algorithm on extracted features in order to output sleep metrics. This process follows the best practices for analytical validation as presented by Goldsack et al. 41 Our work also considers the impact of feature selection and parameters on the accuracy of machine learning algorithms for sleep analysis.
Methods
Participants
Seven participants (age range 4–16 years, mean age 7.22 years, standard deviation
Data collection
Physiological data were collected using the Empatica E4 device.43,44 The device was shipped to participants, and they wore the device on their wrist continuously for two days. On the third night, overnight PSG was performed through the Vanderbilt Sleep Core while the participant concurrently wore the E4 device. A standard PSG protocol with monitoring of respiratory effort, blood oxygen saturation, nasal airflow, heart rate, electromyography, EEG, and electrooculography was conducted using Nihon Kohden Polysmith Sleep Systems. 45 The PSG studies were scored visually in 30-second epochs with analysis and interpretation performed by a board-certified sleep medicine neurologist with expertise in sleep measures for NDDs at the Vanderbilt Sleep Research Core.
The E4 collects data from four main sensors: A photoplethysmography (PPG) sensor, an EDA sensor, a 3-axis accelerometer, and an infrared thermopile. The PPG measures volumetric variations of blood circulation using red and green light.46,47 The lights are oriented towards the wrist skin, which allows the light to be absorbed and reflected. A photodetector then measures the reflected light. The reflection measurements during green light exposure are generally a sequence of valleys caused by high light absorption during a heartbeat. The measured valleys are correlated to heartbeats and are used to estimate heart rate. The red light provides a reference light level for canceling out motion artifacts and allowing for maximization of pulse wave detection. 47 Empatica uses a proprietary algorithm in order to extract the blood volume pulse (BVP) from the PPG signal. The resultant BVP output is stored in a CSV file with a sampling rate of 64 Hz. Interbeat interval (IBI) and heart rate are computed from the BVP and output to CSV files. The IBI data are output intermittently with 1/64 second resolution while the heart rate file contains the average heart rate values over the span of 10 seconds, sampled at 1 Hz.
Innervating signals from the brain cause changes in the permeability of sweat glands on the skin, which can be measured as changes in electrical conductance on the skin surface.
48
The E4 uses a minuscule amount of current between two electrodes to measure these changes as the pores on the wrist fill with sweat.
49
The EDA data, measured in the conductance unit of microSiemens (
An onboard 3-axis micromachined microelectromechanical system accelerometer is used to measure linear motion without a fixed reference.
51
The E4 provides a measurement of acceleration in the unit of 1/64 g at 32 Hz by measuring the continuous gravitational force (g) that is applied in each of the three spatial dimensions (
Temperature data are sampled at 4 Hz using an infrared thermopile on a scale of
All of the generated CSV files from each sensor are zipped and downloaded from the Empatica Data Manager before preprocessing.
Data processing
The sleep data generated from the PSG are stored in a CSV file in 30 s epochs with six possible labels. The stages of sleep include lights on awake (L), lights off awake (W), sleep stage N1 (N1), sleep stage N2 (N2), sleep stage N3 (N3), and REM (R). The timestamps along with the labels are imported into a Jupyter Notebook and the labels are converted into numerical labels, starting with zero and ending with five, so that they are compatible with the scikit-learn. 52 After initial testing, it was determined that without eye movement data, differentiation between all six labels was beyond the capabilities of the current work. Therefore, the six classes are consolidated into three broader classes. Lights on awake and lights off awake are combined into an awake category, designated with the label 100. N1-N3 are combined into a non-REM sleep category, labeled 010. REM sleep remains as its own category, labeled with 001 following the rules of one-hot-encoding, used for categorical data. 53 The consolidation of sleep stages into 3 classes follows the procedure set in previous works by Korkalainen et al. 54 and Sun et al. 55 The distribution of labels can be seen in Figure 2.
Each 30 s epoch is resorted into the 3 resultant labels and stored in a data frame with the timestamp.
After unzipping the physiological data, each CSV file is loaded into the Juypyter Notebook.56,57 The CSV files given begin with a Unix timestamp that is converted into Universal Time Coordinated time and the initial timestamp along with the sampling rates are used to generate timestamps for the length of the collected data. Using the generated timestamps, the labels and physiological data are concatenated into a data frame. Features based on prior work in physiological data research are then extracted from the physiological data.58,59 Using filters, the SCL and SCR are extracted from the EDA data. Following the method presented in Bian et al., 58 a low-pass filter with a 0.5 Hz cutoff frequency removes noisy data. A high pass filter with a 0.05 Hz cutoff frequency is then used to isolate the SCL baseline which is stored in a data frame. The isolated SCL level is subtracted from the filtered data to find the SCRs, which are also stored in a data frame. To match the standard PSG, 30-second epochs without overlapping windows are applied to the physiological data. Interpolation is used to account for any missing data caused by the different sampling rates of the sensors on the E4, as detailed in the “Data collection section. Once all the data are synced and interpolated, the standard deviation and mean of each window is calculated. Multiple features were derived from each sensor in order to explore the full extent of changes to physiology during sleep, which sets the stage for future work that evaluates sleep quality along with sleep stages in children with RTT. The initial features extracted are presented in Table 2 and a graphical overview of the data collection and data processing procedure is depicted in Figure 1.
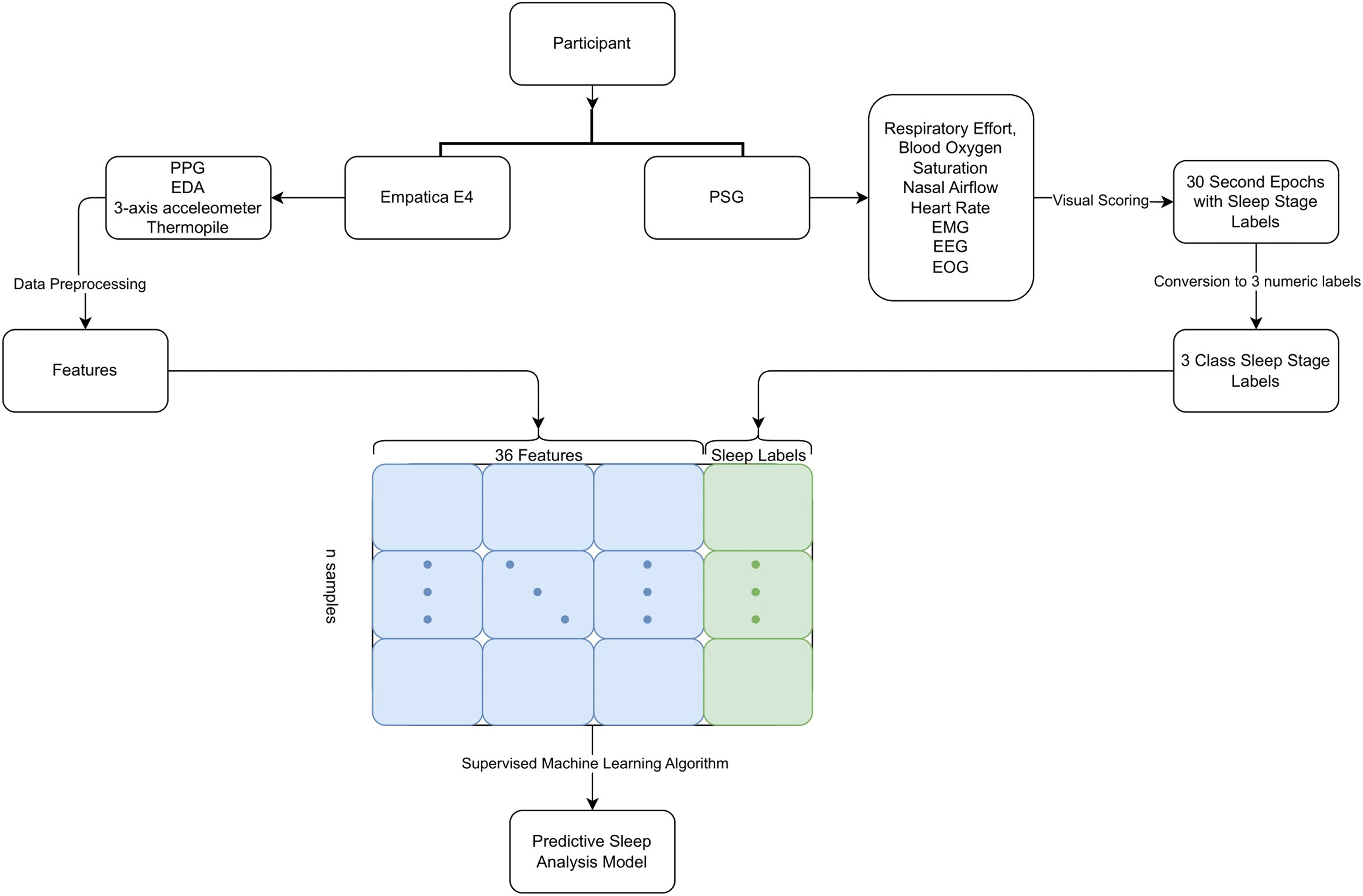
Process of collecting data and creating a predictive sleep analysis model.

Distribution of sleep stages shown with all six labels and how they are recategorized into 3 classes.
R-MBA clinical severity scores of each participant.

ID: Identification; R-MBA: Revised Motor-Behavioral Inventory
Physiological features extracted and how many times each feature was used after feature selection.

PPG: photoplethysmography; EDA: electrodermal activity; BVP: blood volume pulse; IBI: interbeat interval; SCL: skin conductance level; SCR: skin conductance response.
Feature selection
Feature selection in machine learning is used to eliminate redundant features or features that may be unnecessary. By reducing the number of features, the resultant models are less likely to over-fit and the training time is optimized. 60 Model accuracies are also increased as the model no longer has to parse through noisy data. When training models that lack a large amount of training data, feature selection reduces the search space, making the resulting model more accurate. 61
To begin with the feature selection, a dataset of all the available data from the participants was created, and the permutation importance of each feature was generated. Knowing that physiological features are often colinear, we utilized hierarchical clustering on the Spearman rank-order correlations of the features, as detailed in the Permutation Importance page 62 on scikit-learn. 52 This allowed us to determine if feature reduction was possible without loss of information with the available features. The correlation between features was found and plotted and a distance matrix was created. The distance matrix was used to create a dendrogram using Ward’s linkage 63 for hierarchical clustering. The resulting dendrogram (seen in Figure 3) allowed us to choose a feature from each cluster and create a new training set using only the selected features. A comparison of the accuracy between the model generated from all the features and the model generated from the selected features showed a 2% drop in accuracy, indicating that the reduction of features would not negatively affect the performance of the model.

Dendrogram of hierarchical clustering and heatmap of feature correlation used to determine starting point for feature selection.
The features chosen from the dendrogram included heart rate standard deviation and mean, BVP standard deviation and mean, the mean of acceleration in the
Machine learning
Machine learning uses computational algorithms in order to build models that can represent a given dataset. 64 The most commonly used method of machine learning for practical applications is supervised machine learning. Supervised machine learning algorithms produce hypotheses and predictions by learning the general pattern found in data and correlating the patterns to the provided labels. 65 For supervised machine learning, a labeled dataset has to be provided. A subset of the data, the training set, is used to train the model using both the features and the labels. The remaining data, the test set, is used to evaluate the model. The test set labels are removed and the unlabeled features are predicted on by the model. The predicted labels are then compared to the ground truth labels. 66 Variation in physiological signals is expected and can be attributed to a variety of factors, including age, activity level, and, in the case of this work, neurological disorders. When features are varied but possess fundamental qualities that can distinguish the different classes, i.e., physiological signals, supervised machine learning is especially applicable. 67 For this reason, supervised machine learning was used to create our predictive sleep analysis model.
The term ’individual model’ refers to a model that is trained only on data from the participant that the model will be predicting on. One well-known method of evaluating individual models is K-fold cross-validation, which prevents over-fitting and increases the robustness of the evaluation of the model.
68
K-fold cross-validation works by splitting the data into K-equal folds. One fold is held out as the test set each time and the remaining folds are used to train the model. When dealing with unbalanced classes within the data, literature suggests the use of stratified K-fold cross-validation. Stratified K-fold validation uses the same basic method as K-fold cross-validation but maintains the class ratio throughout the K folds of the original dataset.
30
For our work, the value of K was determined by the following equation:
In order to evaluate the significance of the addition of physiological features, as opposed to using only accelerometer features as seen in previous studies with actigraphy, individual models using an SVM and stratified K-fold were developed using only features derived from the accelerometer data. This allowed for a direct comparison of individual models with and without physiological features.
Results and discussion
In order to ensure that the resultant models were correctly predicting each class, the fold with the best F1 score from the stratified K-fold was extracted and stored for each subject. This resulted in the predictions given by each of the seven models having an accuracy of 85.1% and an F1 score of 84.4 when compared to the ground truth labels from the PSG. Each participant’s model predicted with an accuracy between 72.6% and 96.7% as can be seen in Figure 4. By extracting the highest F1 models, we were able to address the overfitting issue we saw when looking at the models with the best accuracies.The confusion matrix in Figure 5 shows the confusion matrix for all the predictions given by the seven individual models when compared to the PSG labels. This shows that the individual models were able to predict all three sleep stages at above 50% accuracy.
Because we do not collect eye movement data, the transition between awake and Non-REM as well as the transition from Non-REM to REM sleep is particularly difficult to differentiate, 72 as can be seen by the lower accuracy of the awake sleep state.

Accuracy of best F1 scored model for each participant.

Confusion matrix for all predictions given by individual models.
While our seven participants showed a wide range of R-BMA severity scores, there were no noticeable trends or correlations between the disease severity and accuracy of the models. We did not explore clinical severity as part of this pilot study, since this initial work aims not to create a model for severity, but to provide an analytical validation of a proxy for sleep analysis against gold-standard PSG. The use of wearable sensors and machine learning for sleep analysis as a predictor for disease severity in RTT is beyond the scope of this current work, especially given the small sample size and preliminary nature of this work. Using information gleaned from the dendrogram of hierarchical clustering and K best feature selection, the final model was trained using 23 input features. The final features for each model relied heavily on temperature and accelerometer data measures, with seven out of seven models using temperature and accelerometer features. Features derived from PPG were consistently used for five of the seven models. These features being used is consistent with other studies in RTT that explore HRV, temperature, and accelerometer measures.20,39,40,73
While the overall accuracy of the predictions given by the model utilizing only accelerometer data is comparable to the accuracy of the model with both physiological features and accelerometer features, when analyzing the distribution of predictions, it is clear that accelerometer data alone is unable to provide sufficient information for the model to predict the REM stage of sleep accurately. This can be seen in Figure 6, where the accelerometer-only model predicted REM sleep with 28.5% accuracy in contrast to 74% accuracy with the addition of physiological features.
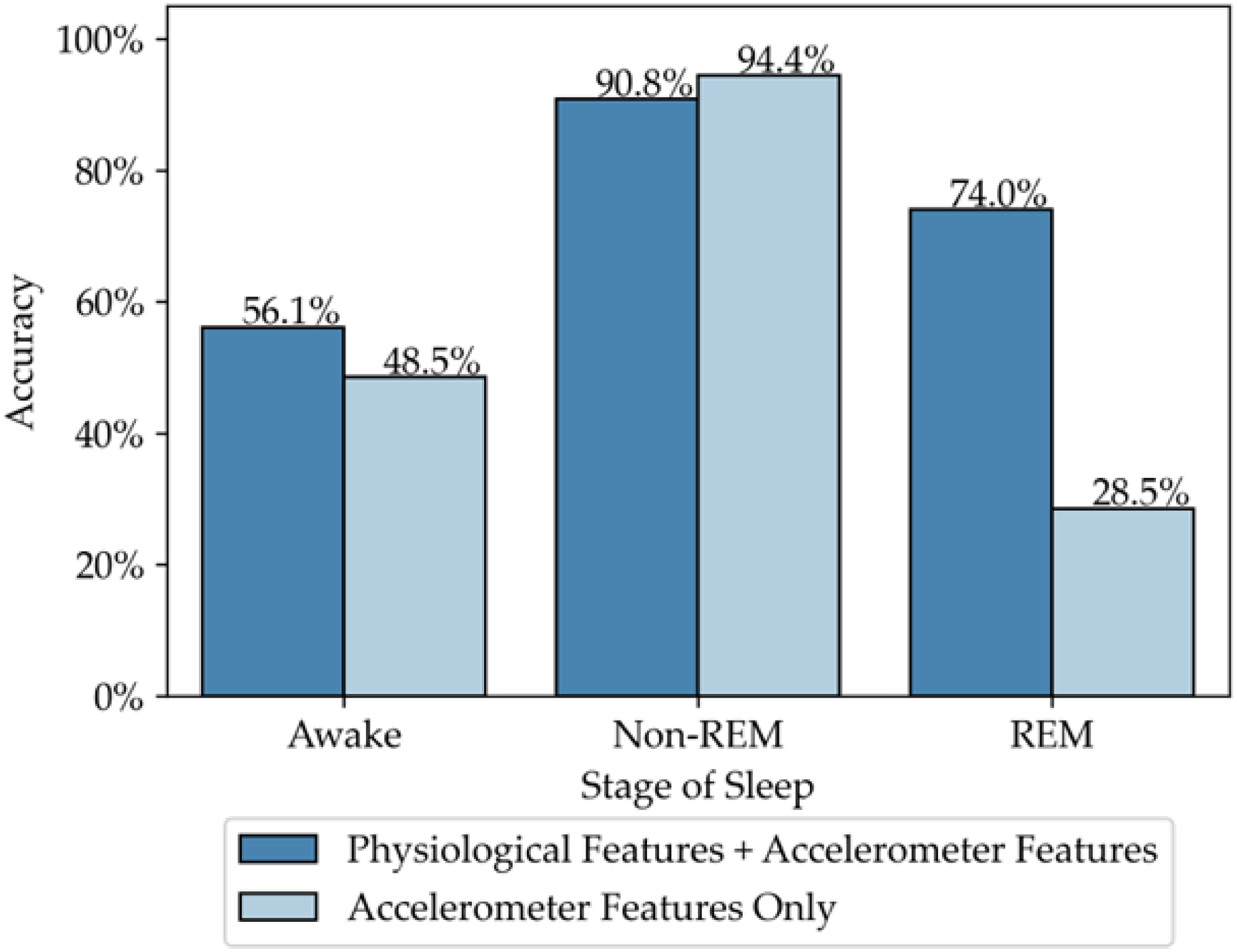
Comparison of accuracy for each stage of sleep predictions using physiological features and accelerometer features vs only using accelerometer features.
Overall, the individual models with physiological features and accelerometer features perform well when compared to commercial products, which tend to have accuracies varying between 60% and 90% when doing epoch-to-epoch comparisons to PSG, depending on the class (awake, Non-REM, REM) being predicted.72,37 Our work shows that the addition of physiological data improves the model fit and should be considered for future research in NDDs such as RTT. While actigraphy-based sleep analysis has been done, 20 in light of clinical features, related to temperature dysregulation 73 and HRV 74 in RTT, wearable devices that incorporate physiological data should be considered in future studies. The present work provides evidence for the feasibility of using DHTs, such as wearable sensors, in RTT, even at young ages, as there was no data loss due to non-compliance with wearing the device. Although some previous studies have questioned the feasibility of DHTs in RTT due to repetitive hand movements and interference with accelerometer readings, the incorporation of physiological data in our work circumvented these concerns. It is also important to note that models developed for commercially developed devices rely on data from a large group of people. In contrast, our current sample size only consists of 7 individuals. The sample size of the current work is a limitation, however, more participants are being recruited and further analysis and validations are planned. The variations in accuracies for the individual participants may be due to the placement of the E4 during data collection, as it is on the wrist and the device not being flush against the skin can result in noisy data. The placement of the E4, as well as other wearable devices, may be adjusted to reduce the probability of data loss and reduce noise in the data for future studies. Future work will focus on the creation of group models and the possible addition of non-invasive methods for obtaining muscle activation near the eyes for REM detection. The expansion of the model to accommodate more classes, such as being able to differentiate between N1, N2, and N3 sleep stages, as well as being able to detect sleep apnea, is pending the collection of additional data.
Conclusion
Our current work paves the way for sleep analysis from wearable sensors for children with Rett by providing evidence for the feasibility of DHTs in RTT for continuous monitoring and demonstrating the benefits of incorporating physiological features. This will expand the ability of clinicians to monitor and analyze how sleep patterns for children with Rett differ from other children, which may allow for new interventions to be explored which can better inform sleep-based interventions and support families. The individual models developed and validated are clinically significant as they allow for progression of sleep disturbances to be tracked longitudinally with more frequent data points. Compared to the current standard of care, where sleep is assessed via PSG far less frequently, our method also allows for sleep monitoring to occur in the child’s natural environment. Our work establishes the viability, through analytical validation, of using wearable devices and machine learning for sleep analysis, which paves the way for the establishment of group models in future work to expand the reach of sleep analysis through wearable sensors.
This initial work analytically validates the use of wearable physiological sensors and accelerometer data as a method of sleep analysis in children with RTT as compared to gold-standard PSG. This addresses the gap in the literature for affordable and non-invasive methods of sleep analysis for children with RTT. Our models are able to predict 3 sleep stages, awake, non-REM, and REM sleep, with around 85% accuracy with individual models when compared to the gold-standard PSG. During our model creation, we explored feature selection to reduce the search space and training time without reducing accuracy and F1 and reduced our features from 36 to 23, which we hope will inform decisions regarding the use of different types of DHT for NDDs such as RTT. However, we acknowledge that this research is in its preliminary stage, featuring a limited sample size. Therefore, it is important to exercise caution when interpreting the results and we emphasize the necessity for additional data before generalization of the results. Future work will expand the validation to other classes of sleep and additional sleep metrics. In addition, future studies will examine the clinical validation of these models to the most used clinical outcome measures in RTT with the goal of also demonstrating both analytical and clinical validation for use in future clinical trials in RTT.
Footnotes
Acknowledgments
We graciously acknowledge the children and families who participated in this study.
Author Contributions
Conceptualization was done by M.M., A.U., C.F.,S.U.P., and N.S.; methodology was done by M.M., A.U., and N.S.; software was handled by M.M. and A.U.; validation was done by M.M., S.U.P., and N.S.; formal analysis was done by M.M.; investigation was carried out by C.F. and S.U.P.; resources were handled by S.U.P. and N.S.; data curation was done by M.M., A.U., C.F., and S.U.P.; writing—original draft preparation was done by M.M.; writing—review and editing was done by A.U., C.F., S.U.P., and N.S.; visualization was done by M.M.; supervision was done by S.U.P. and N.S.; project administration was done by C.F. and S.U.P.; funding acquisition was done by S.U.P. All authors have agreed to the submission of this manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by 1R21TR003942-01 to S.U.P.
Guarantor
M.M.
Ethical approval
This study was approved by the IRB (IRB Number 210217).
Informed consent
Informed consent was obtained from all participants.
