Abstract
Background
Colorectal cancer is one of the highly malignant cancers with a poor prognosis. The exact mechanism of colorectal cancer progression is not completely known. Recently, microRNAs (miRNAs, miRs) were suggested to participate in the regulation of multiple cancer development, including colorectal cancer.
Methods
MiR-4319 expression in colorectal cancer patient samples was detected by real-time polymerase chain reaction. MiR-4319 was knocked down in the colorectal cancer cells by siRNA transfection to study the role of miR-4319 in the cell cycle and proliferation of colorectal cancer cells.
Results
MiR-4319 expression was found to be inverse correlated with survival in colorectal cancer patients. Overexpression of miR-4319 markedly reduced the proliferation of colorectal cancer cells and altered cell cycle distribution. A further experiment showed that ABTB1 is the target gene of miR-4319. MiR-4319 was regulated by PLZF.
Conclusion
Our studies indicated that reduced expression of miR-4319 was correlated with poor prognosis in colorectal cancer patients; miR-4319 also suppressed colorectal cancer cell proliferation by targeting ABTB1. ABTB1 might become an excellent therapeutic target for colorectal cancer treatment.
Key summary
Established knowledge on this subject
Colorectal cancer is a highly malignant cancer with a poor prognosis. MicroRNAs (miRNAs, miRs) participate in the regulation of multiple cancer development, including colorectal cancer. The dysregulation of miR-4319 and its target genes are involved with inflammation, autoimmune diseases, as well as cancers, such as prostate cancer and acute myeloid leukemia.
Significant/new findings of this study
MiR-4319 expression was inversely correlative with colorectal cancer patients' survival rate. Overexpression of miR-4319 significantly reduced colorectal cancer cell growth and altered cell cycle distribution. ABTB1 is a target gene of miR-4319, which is further regulated by PLZF. ABTB1 might become an excellent therapeutic target for colorectal cancer treatment.
Introduction
Colorectal cancer (CRC), as one of the highly malignant cancers with a poor prognosis, leads to a large number of deaths worldwide, and millions of people get CRC every year.1, 2 CRC mostly affects the rectum, the sigmoid colon and the distal part of the descending colon. 3 Colorectal tumorigenesis begins with a progressive transformation of epithelial cells. 4 At the early stage, the cancer is within the mucosal layer, which is made up of epithelium, lamina propria and muscularis mucosae (stage 0–1). With the development of the CRC, tumour cells grow through or into the wall of the colon and rectum (stage 2), and spread further to the nearby lymph nodes, such as the mesenteric or inguinal lymph nodes (stage 3). At the late stage of CRC, the tumour cells metastasise to the other parts of the body through the blood, such as the abdominal cavity, ovaries, lung, liver and kidneys (stage 4).5, 6 An increasing number of risk factors for CRC have been reported recently, such as aging, unhealthy eating habits, smoking, obesity, insufficient physical exercise, Streptococcus gallolyticus bacteremia, inflammatory bowel disease and genetic factors. 7 The treatment for CRC includes surgery, chemotherapy, radiation therapy and ablation. 8 However, as the detailed mechanisms of CRC development are not completely understood, the 5-year survival rate for CRC is low, especially in the case of advanced stages. 5 Thus a better understanding of CRC pathogenesis is a high priority for providing CRC patients with effective diagnostic and prognostic strategies.
MicroRNAs (miRNAs), as a group of small non-coding RNA, contain about 19–22 nucleotides. They bind to complementary mRNA sequences to regulate gene expression, resulting in enhanced degradation of their target genes. MiRNAs have great promise as potential biomarkers in CRC because of their altered expression levels, high tissue specificity and stability in tumour development. 9 Moreover, they are easily detected in the stool, tissue and blood samples of CRC patients. 9 MiR-143 and miR-145 are the initial confirmed RNAs involved in CRC development. Both miR-145 and miR-143 display significantly decreased levels of mature miRNA in colorectal cancer tissues in comparison to normal mucosa specimens. 10 Furthermore, miR-1 has been reported to act as a tumour suppressor in CRC, which could directly target the oncogene mesenchymal-epithelial transition factor (MET). MiR-1 is downregulated during CRC progression.11, 12 In addition, miR-21 could be significantly upregulated in chemotherapy-resistant colon cancer cells, suggesting an important role in the regulation of colon cancer cell differentiation.13, 14 Besides, various reports have demonstrated different expressions of miRNAs in the tumour tissues, blood and stool of CRC patients, including miR-7, miR15, miR-16, miR-19a, miR-20a, miR21, miR-22, miR-29a, miR-31, miR-92a, miR-93, miR-101, miR-155, let-7c, miR-126, miR-132, miR-139-3p, miR-181a, miR-183, miR-375 and miR-378, highlighting the potential of miRNAs as prognostic and diagnostic biomarkers in CRC.15–18
The dysregulation of miR-4319 and its target genes are involved with inflammation, autoimmune diseases, as well as cancers, such as prostate cancer and acute myeloid leukemia (AML).19, 20 In this paper, we demonstrate that miR-4319 is expressed at a low level in CRC tissues, which predicts the poor clinical outcome of CRC. MiR-4319 regulates the proliferation, cell cycle, invasion and migration of CRC cells. Mechanically, miR-4319 is found to be transcriptionally regulated by PLZF, and acts as a tumour suppressor in CRC by targeting ABTB1. Our data suggest that miR-4319 might be a potential biomarker for CRC, providing opportunities to develop better diagnostic and therapeutic strategies.
Methods
Real-time polymerase chain reaction
The total RNAs of CRC tissues and normal tissues were extracted by Trizol reagent (Sigma, St Louis, MO, USA). Real-time polymerase chain reaction (RT–PCR) was conducted using a 7500 real-time PCR system (Applied Biosystems Corp., Foster City, CA, USA), SYBR Green Real-time PCR Master Mix (Sigma) was used according to the manufacturer's instructions. 27 In this study, the primer for miR-4319 is: 5′–GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACGTGGCT–3′. For miR-4319 RT–PCR: 5′–CACCCAGAGCAAAGCCAC–3′ (forward) and 5′–GTGCAGGGTCCGAGGT–3′ (reverse); for control gene (U6) RT primer: 5′–GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACAAAATATGGAA–3′; for control gene (U6) RT–PCR: 5′–TGCGGGTGCTCGCTTCGGCAGC–3′ (forward) and 5′–GTGCAGGGTCCGAGGT–3′ (reverse). For ABTB1 RT–PCR: 5′–CCGTCACCCTGCATGGCATCTC–3′ (forward) and 5′–CACCCACCACAGTGTCCTCGTCT–3′ (reverse). For GAPDH RT–PCR: 5′–ACAACTTTGGTATCGTGGAAGG–3′ (forward) and 5′–GCCATCACGCCACAGTTTC–3′ (reverse). For PLZF RT–PCR: 5′–TGCGGCTGAGAATGCATTA–3′ (forward) and 5′–ACACAGCAGACAGAAGACGG–3′ (reverse). For ChIP–qPCR: 5′–GAACCAACCCAAATGTCCAA–3′ (forward) and 5′–TCCAGCTTCATCCATGTCCC–3′ (reverse).
Cell culture and transfection
HCT116 and SW480 cells were ordered from the ATCC. Cells were cultured in Dulbecco's modified Eagle's medium (Invitrogen, Pleasanton, CA, USA) supplemented with 10% fetal bovine serum (Invitrogen). MiR-4319 inhibitor, miR-4319 mimics and ABTB1 siRNA were ordered from Sigma. For transfection experiments, cells were plated in a six-well plate with 5 × 105 cells per well. Subsequently, cells were transfected by Lipofectamine2000 (Thermo Fisher, CA, USA). Cells were collected for further analysis 48–72 hours after transfection.
Luciferase assay
SW480 and HCT116 cells were plated in a 24-well plate one day before transfection. PsiCHECK2, psiCHECK2-ABTB1, psiCHECK2-ABTB1-mut and pRL-TK plasmids were used to transfect HEK293T cells. The luciferase signal was measured through the Dual Luciferase Assay Kit (Promega, Madison, WI, USA) after transfection for 48 hours. The primer for ABTB1 3′UTR: 5′–CTACTCGAGGCCCCTGGCTGGGCAGCCCCAG–3′ (forward), 5′–CTAGCGGCCGC TCATGCTCCCACGGTGCCTTCAAGCTT–3′ (reverse). The primer for mutant 3′UTR: 5′–TGGGGGGGAACACCAGTGACGAA–3′ (forward), 5′–TTCGTCACTGGTGTTCCCCCCCA–3′ (reverse).
RNA immunoprecipitation
SW480 and HCT116 cells were transfected with pSL-MS2-miR-4319 and pSL-MS2-12x vectors (Addgene Cambridge, MA, USA). Following 48 hours' transfection, cells were harvested to perform RNA immunoprecipitation (RIP) experiments using Magna RIP RNA-Binding Protein Immunoprecipitation Kit (Millipore, Bedford, MA, USA). Human anti-Ago2 antibody or IgG control (Millipore) was added in cell lysates. MiRNA expression was detected by RT–PCR using the TaqMan microRNA assay (Applied Biosystems).
Western blot analysis
Antibodies against ABTB1 (ab99547; Abcam, Cambridge, MA, USA), PLZF (sc-28319; Santa Cruz, Dallas, TX, USA), actin (ab8227; Abcam) were used. Western blot was performed in procedures as previously reported. 28
Cell viability examination
Cell viability was evaluated using 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT). HCT116 and SW480 cells were plated at 1000 cells for each well in a 96-well plate. After 3 days' incubation at 37℃, 20 µl of MTT (2.5 mg/ml; Sigma) was added to the well followed for 3 hours by incubation at 37℃. Subsequently, 150 µl/well of dimethyl sulphoxide was added to solubilise the water-insoluble purple precipitate. Absorbance measurements were performed at the 595 nm wavelength using a microplate reader (BioTek Synergy, Winooski, VT, USA).
Colony formation assay
One to two × 103 cells were seeded in a six-well plate, then were fixed by 4% PFA for 10–20 minutes at room temperature after 2 weeks, followed by staining with crystal violet (0.05%) for 10–20 minutes. Colonies were counted through a microscope.
Migration assay
A wound was created though a sterile pipette tip scratch against the cell culture plate. Twenty-four hours later, the width of the wound was measured through an inverted phase microscope with DP controller software (Olympus, Center Valley, PA, USA).
Transwell assay
A million of HCT116 or SW480 cells were seeded in an invasion chamber with serum-free media on top of extracellular matrix. The cells in the bottom of the chamber were photographed quantitatively using a standard microplate reader (at 570 nm) after incubation for 24 hours.
Cell cycle analysis
Cells were collected and stained through a cell cycle detection kit (BD Biosciences, Bedford, MA, USA), and then analysed using FACS Aria III (BD Biosciences) as previously described. 29 The percentages of the cells at the S, G2/M and G0/G1 phases were analysed by way of a cell-cycle program (Modfit, USA).
Statistical analysis
Statistical significance was calculated using Graph Prism 7 software. Data were shown as mean ± SD. Data are derived from three independent experiments. In all cases, P < 0.05 was thought to be statistically significant.
Results
Low miR-4319 levels are associated with poor prognosis in CRC patients
To test whether miR-4319 has a potential function in colon cancer development, we first determined miR-4319 expression in different colon cancer cell lines. As demonstrated in Figure 1(a), compared with HIEC, a normal human intestinal epithelial cell line, miR-4319 expression was obviously decreased in colon cancer cell lines, which indicated that miR-4319 expression was decreased in colon cancer. To reveal further the potential pathological relevance between miR-4319 and CRC, we measured the level of miR-4319 in normal intestinal tissue and CRC tissue. Consistently, miR-4319 was expressed at a much lower level in CRC tissues (Figure 1(b)). Furthermore, the clinical data also showed that the expression level of miR-4319 was decreased with advanced clinical stage (Figure 1(c)). By analysing the lymph node metastatic state, we found that miR-4319 was also expressed at a lower level in metastatic patients than metastatic-free patients (Figure 1(d)). A reduced level of miR-4319 was highly associated with a poor prognosis in CRC (Figure 1(e)) and patients with low miR-4319 expression levels were also associated with poor distant metastasis-free survival (Figure 1(f)).
The level of miR-4319 was downregulated in colorectal cancer (CRC) tissues and correlated with poor prognosis. (a) The expressions of miR-4319 in CRC cell lines (HCT116, Caco2, HT29, SW620 and SW480) were lower than the normal human intestinal epithelial cell line (HIEC). (b) The expression of miR-4319 was decreased in CRC tissues (n = 100) compared to normal human intestinal tissues (n = 50). The relative expression was shown as 2−ΔCT. (c) The expression of miR-4319 was significantly decreased with advanced clinical stage. (d) The expression of miR-4319 was significantly decreased in CRC tissues of lymph node metastatic (LNM+) patients compared to lymph node metastatic-free (LNM-) counterparts. (e) Kaplan–Meier analysis of the correlation between miR-4319 expression and the overall survival of CRC patients. (f) Kaplan–Meier analysis of the correlation between miR-4319 expression and the distant metastasis-free survival of CRC patients. *P < 0.05; **P < 0.01 (Student's t-test).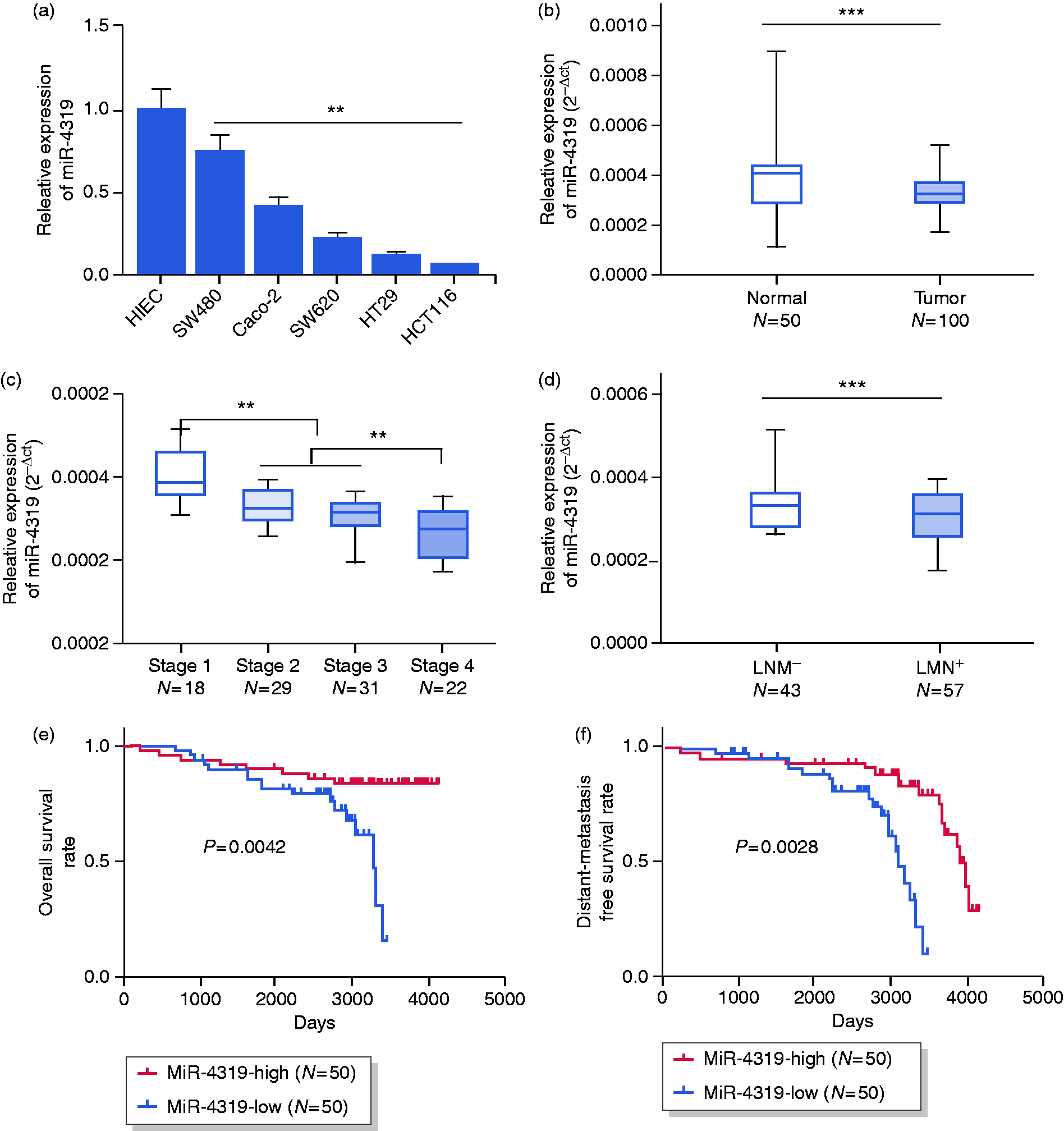
MiR-4319 inhibits proliferation and regulates cell cycle distribution
To investigate further the biological functions of miR-4319 in CRC progression, we transfected miR-4319 in SW480 and HCT116 cells. We examined miR-4319 expression after the transfection in two CRC cells (Figure 2(a)), and then tested the cell viability by MTT assay. It was demonstrated that the growth of miR-4319 overexpressed cells was much lower than that of control cells (Figure 2(b) and (c)). Colony formation data showed that miR-4319 decreased the cell colony formation capacity (Figure 2(d)). Furthermore, we tested whether miR-4319 plays a role in regulating cell cycle distribution. It was found that miR-4319 altered the cell cycle distribution, resulting in decreased population in the S phase and increased population in the G1 phase (Figure 2(e) and (f)), which suggested that miR-4319 might downregulate cell proliferation by regulating cell cycle distribution. To confirm further the function of miR-4319 in cell proliferation, we treated SW480 and HCT116 cells with miR-4319 inhibitor. MiR-4319 expression was significantly downregulated after inhibitor treatment in both HCT116 and SW480 cells (Figure 2(g)). Consistently, miR-4319 inhibitor treatment increased cell proliferation (Figure 2(h) and (i)) and colony formation capacity (Figure 2(j)). The cells after miR-4319 inhibitor treatment also showed different cell cycle distribution by comparing with control cells (Figure 2(k) and (l)). Based on the above results, we concluded that miR-4319 inhibited cell proliferation and regulated cell cycle distribution in CRC cells.
MiR-4319 inhibited the proliferation and regulated cell cycle distribution of colorectal cancer (CRC) cells. (a) The real time polymerase chain reaction (RT–PCR) analysis of miR-4319 levels in HCT116 and SW480 cells transfected with miR-4319 mimics or the negative controls (miR-NC) at 48 hours post-transfection. (b) and (c) The 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay showed that the cell viabilities of miR-4319-transfected HCT116 and SW480 cells were decreased compared to the control groups. (d) Colony formation abilities of miR-4319-transfected HCT116 and SW480 cells were decreased compared to miR-NC-transfected cells. (e) and (f) The effect of miR-4319 on the cell cycle of HCT116 and SW480 cells was detected by flow cytometry. (g) Quantitative RT–PCR analysis of miR-4319 levels in HCT116 and SW480 cells transfected with miR-4319 inhibitor or the negative controls (NC-inhibitor) at 48 hours post-transfection. (h) and (i) The MTT assay showed the cell viabilities of miR-4319 inhibitor-transfected HCT116 and SW480 cell lines were increased. (j) The colony formation abilities of miR-4319 inhibitor-transfected HCT116 and SW480 cells were increased. (k) and (l) The effect of miR-4319 inhibitor on the cell cycle of HCT116 and SW480 cells was detected by flow cytometry. The data represent the mean ± SD from three independent experiments. *P < 0.05; **P < 0.01.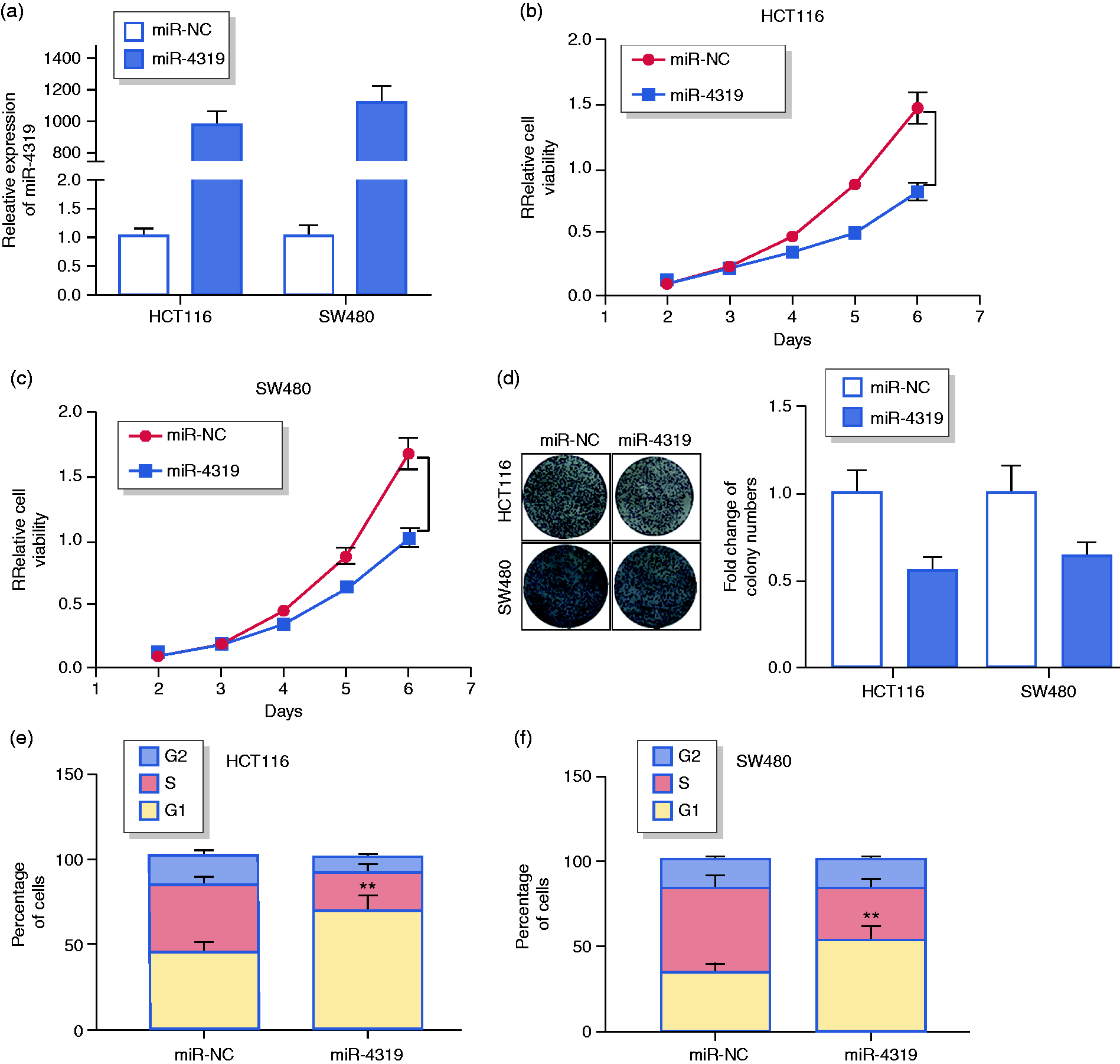
MiR-4319 suppresses CRC cell migration and invasion
Considering that miR-4319 was highly associated with distant metastasis-free survival in CRC patients, we next detected whether miR-4319 inhibited cell migration and invasion in CRC cell lines. A wound healing assay showed that miR-4319 overexpression inhibited cell migration, whereas miR-4319 inhibitor treatment promoted migration in both SW480 and HCT116 cells (Figure 3(a) and (b)). Then we performed a transwell assay to determine whether miR-4319 affected the invasive abilities of CRC cells. The results demonstrated that the invasive ability was decreased in miTR-4319 transfected cells, whereas the miR-4319 inhibitor could rescue this phenomenon (Figure 3(c)). These results indicated that miR-4319 suppressed CRC cell migration and invasion.
MiR-4319 suppressed the migration and invasion of colorectal cancer (CRC) cells. (a) and (b) The migration abilities of HCT116 and SW480 cells were significantly reduced by miR-4319 overexpression, while they were increased by the miR-4319-inhibitor. (c) The transwell assays were performed to determine the invasive abilities of HCT116 and SW480 cells transfected with miR-4319 mimics, miR-NC, miR-4319 inhibitor and NC-inhibitor. The data represent the mean ± SD from three independent experiments. *P < 0.05; **P < 0.01 (Student's t-test).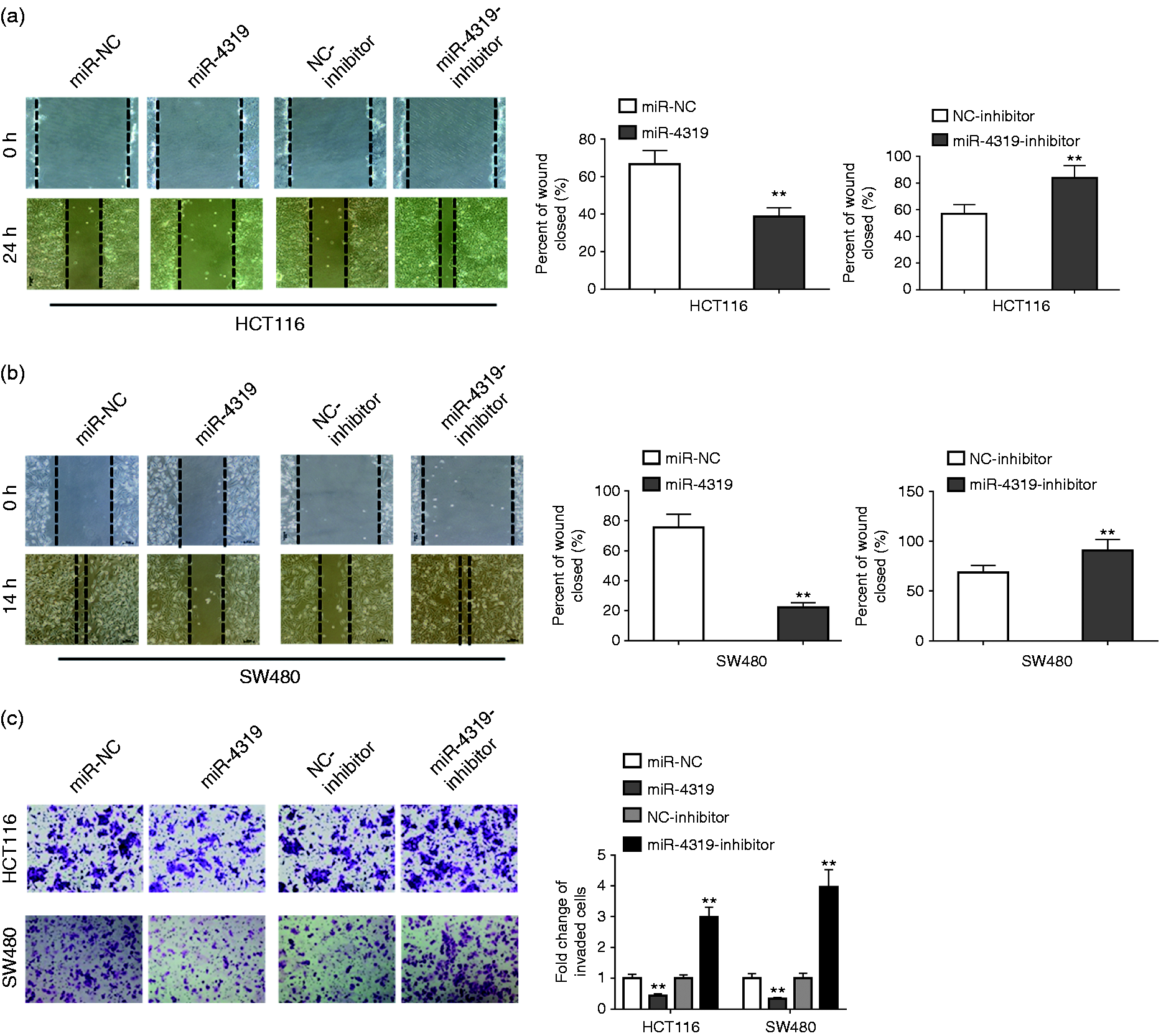
ABTB1 is the target of miR-4319
To identify the target gene of miR-4319, we searched potential targets through online miRNA prediction websites. Interestingly, ABTB1 showed strong potential to be the target of miR-4319, as there exist multiple binding sites on ABTB1 (Figure 4(a)). The indicated mutations and different binding sites within the ABTB1 sequences were shown in Figure 4(a). To analyse further whether miR-4319 targets ABTB1 and regulates its expression in CRC cells we generated the luciferase reporter plasmids including specific mutations and potential binding sites within the sequence of ABTB1. The mutated ABTB1 expressed luciferase reporter constructs and control vectors were transfected into HEK293 cells in the presence of miR-4319. Significant suppression was found when miR-4319 was transfected together with the wild-type ABTB1, while luciferase activity of mutated ABTB1 was not significantly affected compared to the control group (Figure 4(b) and (c)). In addition, both the mRNA level and protein level of ABTB1 were clearly downregulated by miR4319, whereas they were upregulated by the miR-4319 inhibitor (Figure 4(d) and (e)). IP assay showed that both miR-4319 and ABTB1 could be enriched by Ago2 immunoprecipitates from CRC cell lysis (Figure 4(f) and (g)), suggesting that miR-4319 interacts with ABTB1 in CRC cells. ABTB1 level was negatively correlated with miR-4319 in the CRC samples (Figure 4(h)). Our results indicated that ABTB1 should be a direct target of miR-4319 in CRC cells.
ABTB1 was a direct target of miR-4319 in colorectal cancer (CRC) cells. (a) Schematic diagram of predicted wild-type or mutated miR-4319 binding sites in the 3′UTR of ABTB1. (b) and (c) HCT116 and SW480 cells were co-transfected with constructed luciferase reporter vectors (ABTB1 3′UTR, ABTB1 3′UTR-mut, psi-check2) and miR-4319 or control mimics (miR-NC) and performed luciferase reporter assays. (d) The mRNA expressions of ABTB1 in HCT116 and SW480 cells transfected with miR-4319, miR-4319-inhibitor or their negative control mimics (miR-NC, NC-inhibitor) were detected by quantitative real time polymerase chain reaction (RT–PCR). (e) Protein expressions of ABTB1 in HCT116 and SW480 cells transfected with miR-4319 and miR-4319-inhibitor or their negative control mimics (miR-NC, NC-inhibitor) were detected by western blot. (f) and (g) RT–PCR analysis of miR-4319 and ABTB1 levels isolated from Ago2 and IgG immunoprecipitates derived from HCT116 and SW480 cells. (h) Pearson's analysis showed the correlation between miR-4319 and ABTB1 levels in CRC samples (n = 100). The data represent the mean ± SD from three independent experiments. *P < 0.05; **P < 0.01.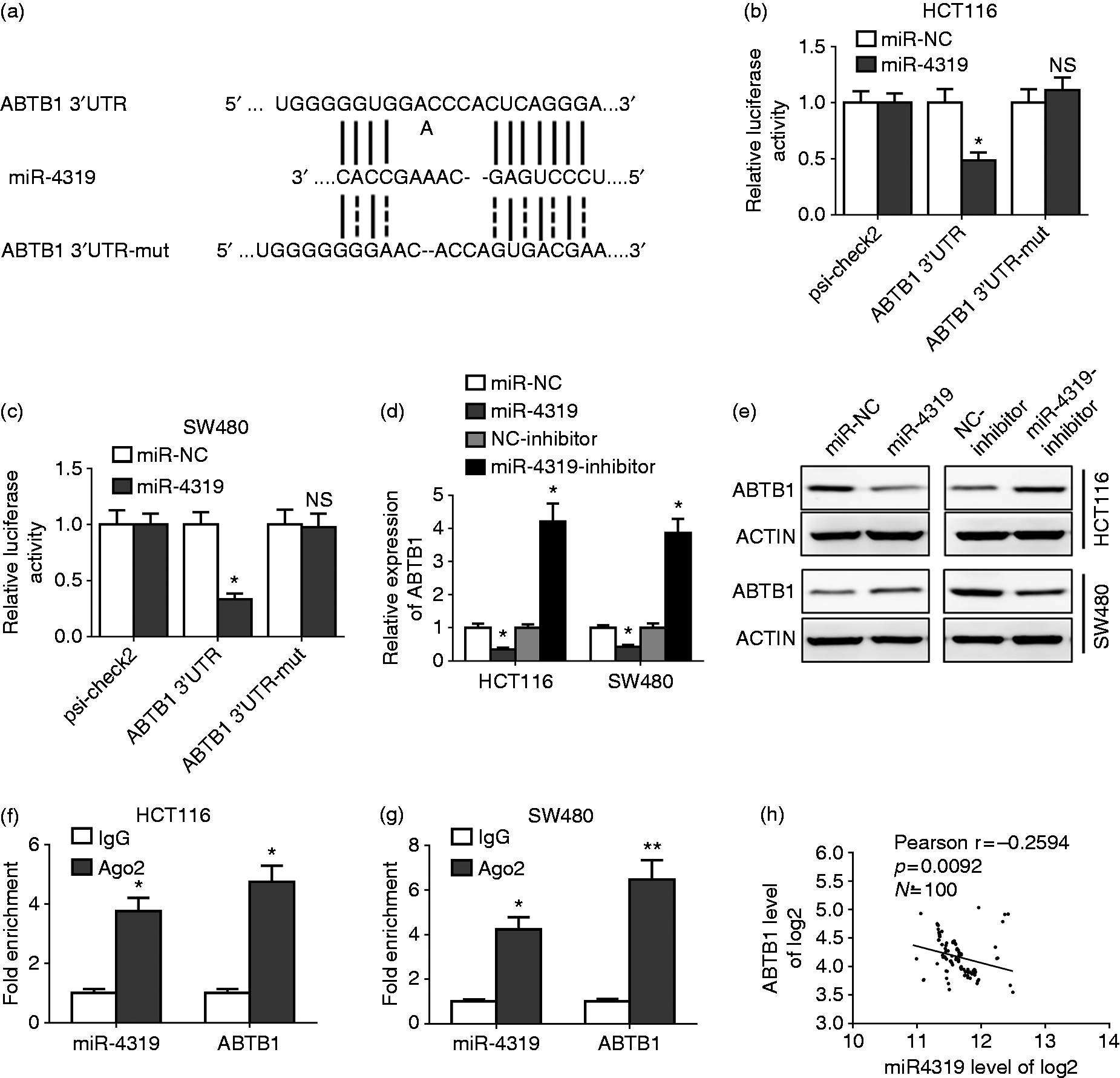
MiR-4319 mediates progression of CRC cells by targeting ABTB1
To investigate whether miR-4319-mediated regulation of CRC cell progression was dependent on targeting ABTB1, we used ABTB1 siRNA to knock down endogenous expression level of ABTB1 in HCT116 cells. Both ABTB1 mRNA and protein level was significantly decreased after siRNA transfection, and ABTB1 siRNA transfection also downregulated ABTB1 mRNA and protein level on miR-4319 inhibitor treatment (Figure 5(a) and (b)). In addition, knockdown of ABTB1 resulted in decreased cell viability and colony formation capacity in both control and miR-4319 inhibitor treated cells (Figure 5(c) and (d)). Knockdown of ABTB1 also dramatically enhanced cell population in the G1 phase (Figure 5(e)). The cell migration and invasion capacity was inhibited when ABTB1 was knocked down by siRNA transfection in either control or miR-4319 inhibitor treated HCT116 cells (Figure 5(f) and (g)). Taken together, our results suggested that ABTB1 plays a critical role in the regulation of CRC cell progression through targeting by miR-4319.
MiR-4319 mediated the progression of colorectal cancer (CRC) cells by targeting ABTB1. (a) and (b) The expression levels of ABTB1 were examined in four groups of HCT116 cells, treated with NC-inhibitor + si-NC, NC-inhibitor + si-ABTB1, miR-4319-inhibitor + si-NC, miR-4319-inhibitor + si-ABTB1, by real time polymerase chain reaction (RT–PCR) and western blot. (c) and (d) The cell survival in HCT116 cells treated as indication were detected by 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) and colony formation assays. (e) The cell cycle of HCT116 cells treated as indication was detected by flow cytometry. (f) and (g) Scratch wound and transwell assays were performed to determine the migratory and invasive abilities in HCT116 cells. The data represent the mean ± SD from three independent experiments. *P < 0.05; **P < 0.01.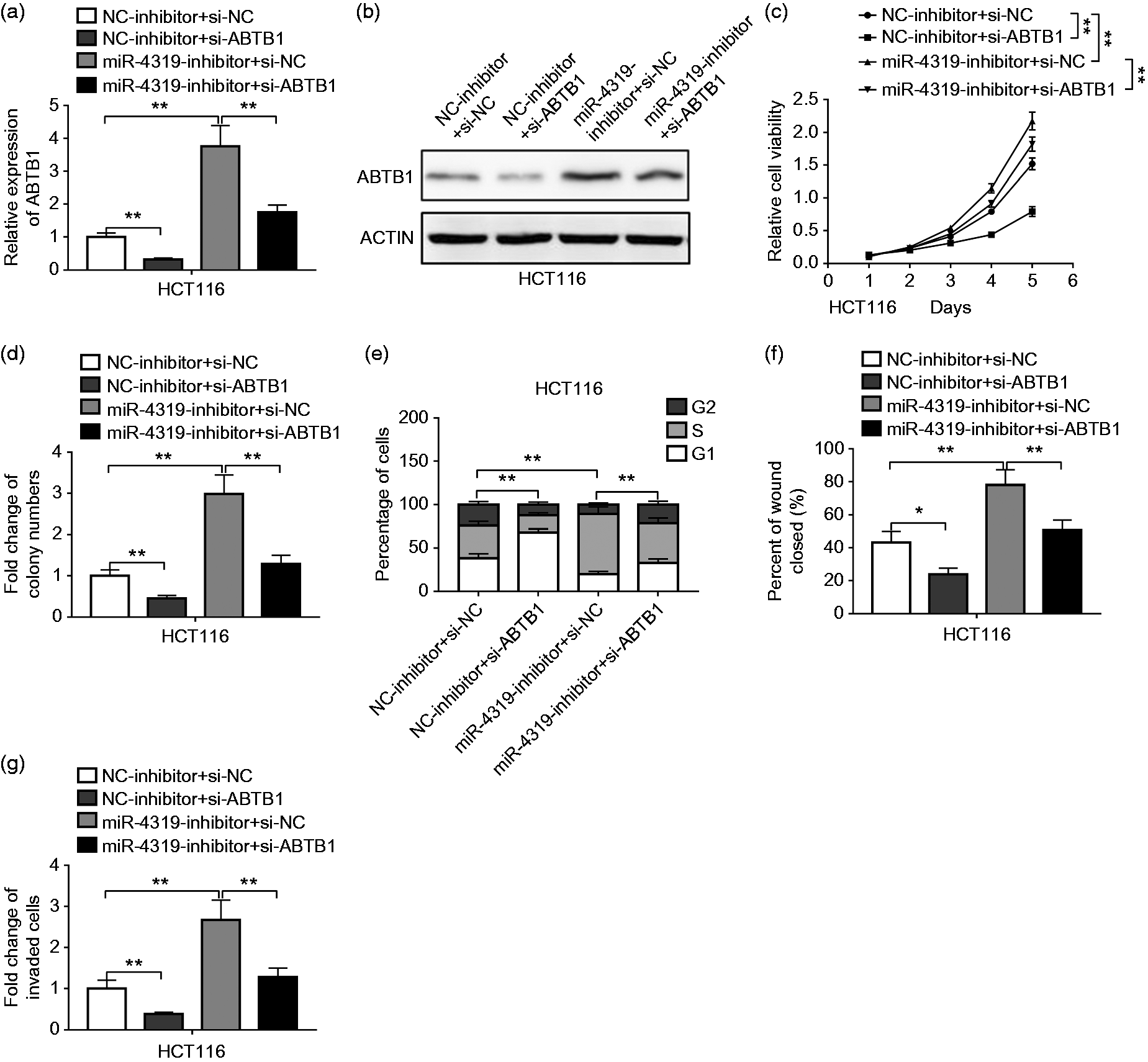
MiR-4319 is transcriptionally regulated by PLZF
To find which gene was the upstream regulated gene for miR-4319, we predicted through rVista 2.0 and found that PLZF was a candidate regulated gene for miR-4319. To confirm this prediction, we transfected the HCT116 and SW480 cells with two snRNA targeting PLZF. A significant decrease of the PLZF mRNA level and protein level was observed when siPLZF-1 or siPLZF-2 was transfected (Figure 6(a) and (b)). Indeed, miR-4319 expression was inhibited in PLZF knockdown cells (Figure 6(c)). Compared with the non-specific promoter region, knockdown of PLZF could specifically inhibit miR-4319 promoter activity (Figure 6(d) and (e)). ChIP assay showed that PLZF binds to miR-4319 promoter DNA, suggesting that PLZF is a transcriptional regulator of miR-4319 (Figure 6(f)).
MiR-4319 was transcriptionally regulated by PLZF in colorectal cancer (CRC) cells. (a) and (b) The mRNA and protein levels of PLZF were determined by real time polymerase chain reaction (RT–PCR) and western blot in HCT116 and SW480 cells transfected with PLZF siRNAs (si-PLZF-1 and si-PLZF-2) or negative control (si-NC). (c) The expression level of miR-4319 was determined by RT–PCR in HCT116 and SW480 cells transfected with PLZF siRNAs (si-PLZF-1 and si-PLZF-2) or negative control (si-NC). (d) and (e) PLZF knockdown reduced the activity of the miR-4319 promoter in HCT116 and SW480 cells. (f) CHIP–PCR analysis of HCT116 and SW480 cells using PLZF antibody. The data represent the mean ± SD from three independent experiments. *P < 0.05; **P < 0.01.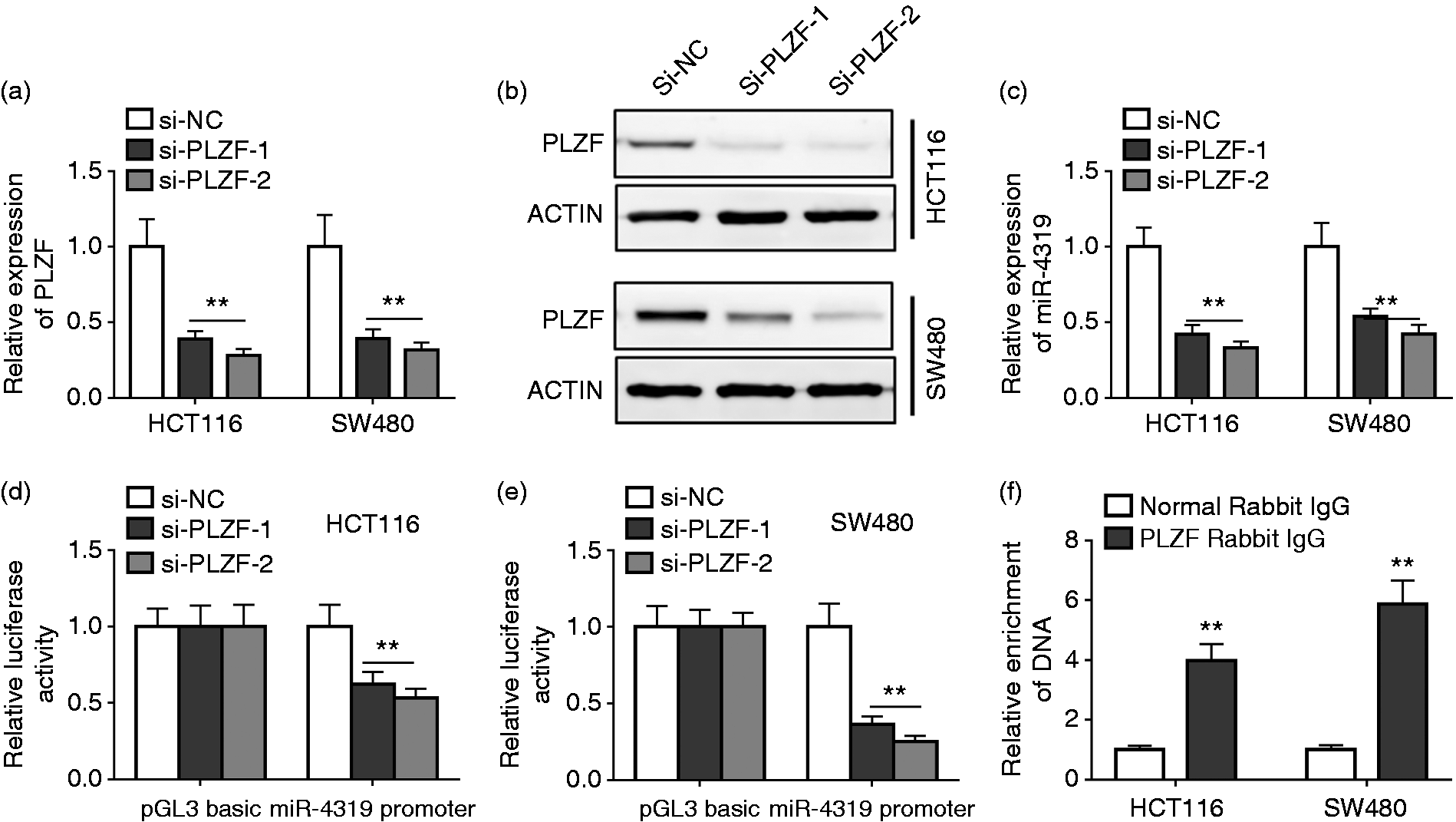
Discussion
Various studies have recently suggested that miRs could be important regulators of tumorigenesis and progression, and the dysfunction or aberrant expression of miRs is associated with cancers.18, 21, 22 Thus the studies of the cellular and molecular mechanisms of miRs during CRC tumorigenesis will lead to the development of new biomarkers for diagnosis and prognosis. Our results have provided evidence suggesting that miR-4319 expression was downregulated in tumour tissues of CRC patients. MiR-4319 expression is found to be positively correlated with CRC patients' survival. Moreover, miR-4319 overexpression could suppress the proliferation and colony formation capability of CRC cell lines, whereas the inhibition of miR-4319 arrested CRC cell lines at the S phase of the cell cycle. Furthermore, the abilities of CRC cell migration and invasion were increased by the inhibition of miR-4319. Our data are also supported by previous studies indicating the role of miR-4319 in tumour progression. A recent study indicates that miR-4319 may be involved in the regulation of prostate cancer pathogenesis. 19 The proliferation of primary prostate epithelial cells was increased by the depletion of miR-4319, whereas the re-expression of miR-4319 suppressed cell growth and their resistance to chemotherapeutic drug estramustine-induced apoptosis. 19 Further studies indicated that the regulatory role of miR-4319 in prostate epithelial cells was dependent on Her-2 (human epidermal growth factor receptor 2) signalling pathway. 19 Moreover, the decrease of the intronic miR-4319 was found in the bone marrow sample of the AML patient. 20 While its host gene SETBP1 (sterol regulatory element binding protein 1) was overexpressed in the patient's sample, which demonstrated a possible molecular mechanism underlying the primary myelofibrosis progression to AML. 20 Therefore, our study suggests that miR-4319 exhibits an effect on tumour progression, supporting its role as a promising therapeutic candidate. Nevertheless, further animal studies are needed to test the therapeutic role of miR-4319 against CRC. For example, transferring miR-4319-expressing plasmid, mimics or virally encoded constructs to tumour-bearing mice to analyse its anti-tumour efficacy and anti-metastasis activity.
PLZF is also recognised as a zinc finger and BTB domain containing 16, and it acts as a tumour-suppressor gene in many types of cancers including colon cancer, prostate cancer, melanoma, leukemia, hepatocellular carcinoma and gallbladder cancer.23,24 The decreased expression of PLZF was found in malignant cell lines and various solid tumours. For instance, hypermethylation in the PLZF promoter and downregulated level of PLZF was found in the majority of pancreatic cancer specimens. 23 Furthermore, the expression of PLZF was dramatically downregulated in hepatocellular carcinoma samples at both the RNA and protein levels in comparison to those of adjacent normal tissues. 25 Besides, PLZF expression was found to be substantially lower in high-risk melanomas than that in low-risk melanomas. 26 Although PLZF has emerged as an promising functional component during CRC progression, the underlying mechanism of tumorigenesis mediated by PLZF is not fully understood. Here we showed that PLZF transcriptionally regulated miR-4319, leading to miR-4319-mediated progression of CRC cells. PLZF knockdown reduced the expression of miR-4319 in HCT116 and SW480 cells. Besides, the CHIP assay demonstrated that PLZF directly bound to the promoter of miR-4319 and regulated its activity. Taken together, miR-4319 inhibits the malignancy of CRC cells by targeting ABTB1, which is transcriptionally regulated by upstream PLZF. Our results suggested that miR-4319, PLZF and ABTB1 could be potential targets for developing clinic diagnosis and treatment strategies for CRC.
In conclusion, our current findings demonstrate a previously unrecognised role of miR-4319 in the suppression of CRC, and suggest it may be a promising biomarker for CRC prognosis. Using the CRC cell lines, we showed that miR-4319 impeded cell proliferation, invasion and migration in vitro. We found that miR-4319 mediated the progression of CRC cells by directly targeting ABTB1, and miR-4319 was transcriptionally regulated by PLZF. Our current study improves our understanding of the role of miRNA in CRC, and suggests novel targets for therapeutic intervention in human CRC.
Footnotes
Author contribution
Longchang Huang, Ye Zhang, Zengyao Li, Xiaoqian Zhao, Zhong Xi, Hang Chen, Haoze Shi, Taojian Xin, Renhui Shen: performed the experiments, analyzed the data. Tong Wang: Conceived and supervised this study, wrote the paper.
Declaration of conflicting interests
The authors declare that they have no competing interests.
Funding
This study was supported by the National Science Foundation (81371683 and 81702592), Wuxi Medical Emphasis Disciplines Funding (syxzdxk-pwk), Wuxi Science and Technology Development Plan (CZE02H1701), Clinical Medicine Project of Jiangsu Province (BL2014023) and National Sciences Foundation of Jiangsu Province (BK20160198).
Ethics approval
All human studies have been approved by the appropriate ethics committee of Wuxi People’s Hospital on 05/07/2008, and have therefore been performed in accordance with the ethical standards laid down in the 1964 Declaration of Helsinki and its later amendments.
Informed consent
Informed consent was obtained from all the participants.
*The first two authors contributed equally to this work.
