Abstract
Objectives:
Legionellosis, primarily caused by Legionella pneumophila serogroup 1, remains a public health concern in Thailand, particularly in tourism-driven regions. Conventional culture-based surveillance is hindered by limited sensitivity and prolonged turnaround times. This study aimed to develop and validate an optimized multiplex TaqMan real-time PCR assay for simultaneous detection of Legionella spp., Legionella pneumophila, and Legionella pneumophila serogroup 1 in environmental water, while also investigating antimicrobial resistance genes.
Methods:
A modified primer–probe set targeting the 23S–5S rRNA, mip, and wzm genes was combined with optimized DNA extraction and PCR conditions. Analytical sensitivity was assessed using colony suspensions and spiked water, and specificity was tested against 24 bacterial strains. The assay was validated on 202 environmental water samples collected from four provinces in southern Thailand. Samples positive by real-time PCR were further screened for antimicrobial resistance genes (lpeAB and tet56) using conventional PCR.
Results:
The optimized assay achieved a detection limit of 1 CFU/mL in suspensions and 1 CFU/L in filtered water, with 100% specificity. Among 202 field samples, PCR detected Legionella in 48.51%, significantly higher than culture (24.26%; p < 0.001). Detection was particularly enhanced in cold-water systems from premise plumbing (41.61% versus 9.49%; p < 0.001). Both methods identified Legionella pneumophila serogroup 1 in 14.29% of samples. Spatial mapping identified Patong subdistrict, Phuket, as a contamination hotspot. Antimicrobial resistance surveillance showed lpeAB in 6.1% of samples and tet56 in one Legionella pneumophila isolate, representing the first report of antimicrobial resistance in environmental Legionella from Thailand.
Conclusions:
This optimized multiplex real-time PCR assay offers a rapid, sensitive, specific, and cost-effective alternative to culture for Legionella surveillance. The detection of antimicrobial resistance genes highlights potential emerging resistance risks, underscoring the need for integrated molecular and resistance monitoring. Implementation of this approach can strengthen outbreak preparedness and improve water safety management in high-risk settings.
Introduction
In Thailand, Legionellosis has been mostly associated with European foreign tourists visiting famous provinces like Phuket, Phang Nga, and Krabi. The first case was documented in 1984. Reports from the Bureau of Epidemiology, Department of Disease Control, Ministry of Public Health, indicate that from 2010 to 2017, there were 62 cases in the upper southern region, while from 2010 to 2023, there were 55 cases in Phuket province. 1 Despite the low incidence of Legionellosis, the economic impact of cases reported by travelers to Thailand is substantial.
Legionella spp. are aerobic Gram-negative rods and nonspore-forming bacteria which are currently classified into 67 validly published species with correct names, according to the List of Prokaryotic names with Standing in Nomenclature. 2 Only 24 species were recognized as human pathogens. 3 Legionella pneumophila (L. pneumophila) is the most common cause of Legionellosis and can be classified into 16 serogroups. Of these, serogroup 1 is the most prevalent and is associated with clinical human cases, accounting for 80%–90% of confirmed cases. 4 Other non-pneumophila species being etiological agents of Legionellosis are Legionella longbeachae, Legionella anisa, Legionella micdadei, Legionella bozemanii, and Legionella dumoffii.4,5 Legionella are naturally found in environmental water (e.g. lakes, ponds, and rivers) and the human-made water distribution system (e.g. cooling tower, water heater tanks, fountains, and spa pools). 6 Exposure to aerosols from water contaminated with Legionella can lead to infections in the respiratory tract. There are two distinct clinical patterns of Legionellosis: a self-limited flu-like illness known as Pontiac fever, which accounts for over 90% of cases, and a more severe form characterized by pneumonia or multisystem disease referred to as Legionnaires’ disease. 6
The detection of Legionella from environmental water samples is routinely done as a surveillance policy of the public health organization. Both cultural methods and noncultural methods can be used as tools for Legionella detection. Like other pathogen detection methods, the higher sensitivity and specificity of molecular testing overcome the constraints of culture-based techniques. A concurrent identification of human pathogen species or serogroup of Legionella with multiplex PCR has been previously documented in both clinical and environmental water samples.7–10 In Thailand, routine surveillance for Legionella contamination in environmental water samples predominantly relies on the gold standard culture method. This study aimed to optimize and validate a multiplex TaqMan real-time PCR assay for the simultaneous detection of Legionella spp., L. pneumophila, and L. pneumophila serogroup 1 in environmental water samples collected from southern Thailand, along with the characterization of antimicrobial resistance (AMR) markers (lpeAB and tet56) identified in Legionella-positive samples to establish baseline information for future surveillance of resistance in environmental isolates.
Methods
Bacteria and culture
A list of bacterial isolates used in this study is shown in Table 1. Legionella strains were subcultured on buffered charcoal yeast extract (BCYE) agar and incubated at 35 °C in 5% CO2 for 48 h. Colonies were identified based on morphology—convex, round, and cut-glass appearance with central white zones and colored edges—and serogrouped into L. pneumophila serogroup 1–15 and other Legionella spp. using serogroup antisera (Denka Company Limited, Tokyo, Japan). Non-Legionella bacterial strains were grown on tryptic soy agar at 37 °C for 16–18 h and identified using Gram staining and biochemical methods. All procedures were approved by the Thammasat University Institutional Biosafety Committee (029/2567), with safety practices aligned to the biological risk group involved.
Lists of microorganisms used in this study and the modified primer set specificity determination.
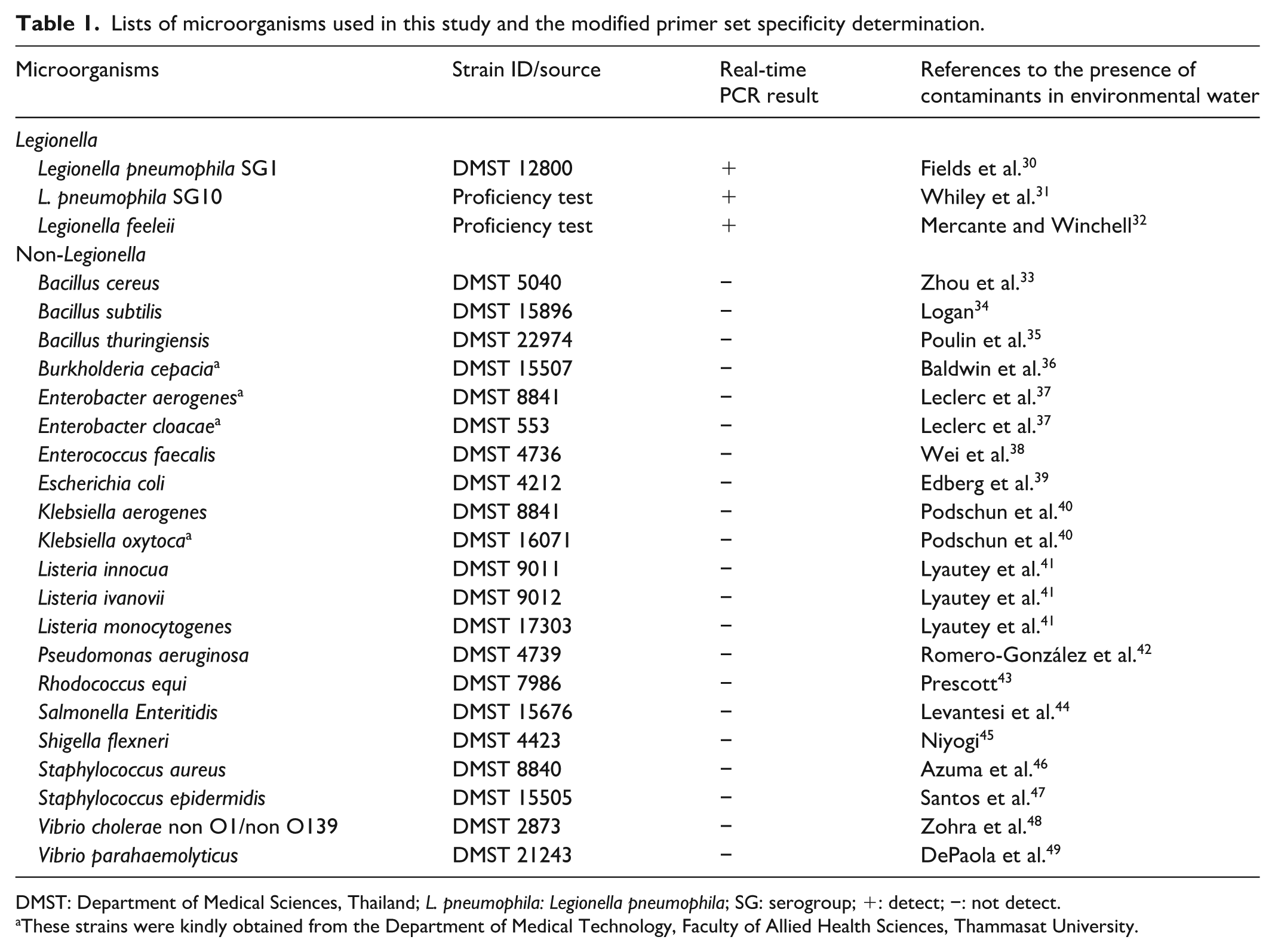
DMST: Department of Medical Sciences, Thailand; L. pneumophila: Legionella pneumophila; SG: serogroup; +: detect; −: not detect.
These strains were kindly obtained from the Department of Medical Technology, Faculty of Allied Health Sciences, Thammasat University.
Optimization process
A fresh colony of Legionella was suspended in sterile normal saline to a concentration of 10 8 CFU/mL, adjusted to the McFarland standard no. 0.5. This suspension was used to compare quantification cycle (Cq) values obtained from different DNA extraction protocols and real-time PCR master mixes.
DNA extraction
DNA extraction was performed using three protocols: (1) Zybio EXM 3000 automated extractor with Zybio extraction kit (Zybio, Inc., Chongqing, China); (2) MagPurix 12A system with Zinexts Viral/Pathogen Kit (Zinexts Life Science Corp., New Taipei City, Taiwan); and (3) QIAamp DNA Mini Kit (Qiagen GmbH, Hilden, Germany).
Real-time PCR
Equal volumes of extracted DNA were used as templates in two commercial reagents: qPCRBIO Probe Mix Lo-ROX (PCR Biosystems Ltd, London, UK) and iTaqTM Fast Supermix with ROX (Bio-Rad Laboratories, Inc., Hercules, CA, USA). A duplex primer–probe set targeting Legionella spp. and L. pneumophila, adapted from a previously published report, 7 was employed. Primer sequences, reaction compositions, and thermal cycling conditions are provided in Supplemental File S1. All tests were conducted in duplicate.
Based on the comparison of Cq values, the combination of the MagPurix 12A system with Zinexts Viral/Pathogen Kit (Zinexts Life Science Corp.) and qPCRBIO Probe Mix Lo-ROX (PCR Biosystems Ltd) yielded the best performance. This optimized protocol was subsequently used in all downstream experiments.
Comparison of duplex and multiplex sets
Given the added diagnostic value of a multiplex primer set targeting Legionella spp., L. pneumophila, and L. pneumophila serogroup 1, the multiplex set—adapted from a previously published report 11 —was compared with the duplex primer–probe set.
LOD experiments
As shown in Figure 1, the limit of detection (LOD) was determined using two experimental models: (1) a colony suspension model and (2) a spiked water model.

Schematic representation of the experimental design for determining the LOD.
In the colony suspension model, 10-fold serial dilutions (108–100 CFU/mL) were prepared from pure Legionella cultures. Three types of PCR templates were evaluated: genomic DNA extracted from the undiluted 108 CFU/mL colony suspension and subsequently diluted 10-fold; extracted DNA obtained from 10-fold serially diluted (108–100 CFU/mL) bacterial suspensions prior to extraction; and direct bacterial suspensions used without DNA extraction. All template types were tested to assess detection sensitivity.
In the spiked water model, 1 mL of each bacterial dilution was inoculated into 1 L of reverse osmosis (RO) or tap water to simulate environmental samples. DNA was subsequently obtained from three template preparations: extracted DNA from non-filtrated water, extracted DNA from filtrated water, and direct unprocessed water samples.
Environmental water sample processing
A total of 202 residual environmental samples were sourced from the Regional Medical Sciences Center, 11/1 Phuket, collected across four provinces (Phuket, Krabi, Phang Nga, and Nakhon Si Thammarat) as illustrated in Figure 2(a) and stored at 2 °C–8 °C for under 1 month. The processing of water samples varied according to their respective sources. 12 Each 1 L water sample was dechlorinated with 0.5 mL of 0.1 N sodium thiosulfate (Merck KGaA, Darmstadt, Germany) and filtered through 0.2 µm polycarbonate membranes. Filters were resuspended in 5 mL sterile water. Samples were either plated directly onto BCYE; PCV medium (BCYE supplemented with polymyxin B, cycloheximide, and vancomycin); GPCV medium (BCYE supplemented with glycine, polymyxin B, cycloheximide, and vancomycin); and PCV⁻ medium (PCV without L-cysteine), or acid-treated (if turbid) before plating. Plates were incubated at 35 °C ± 2 °C for 7 days. Nonpotable water (e.g. from cooling systems) followed a similar procedure, excluding filtration.
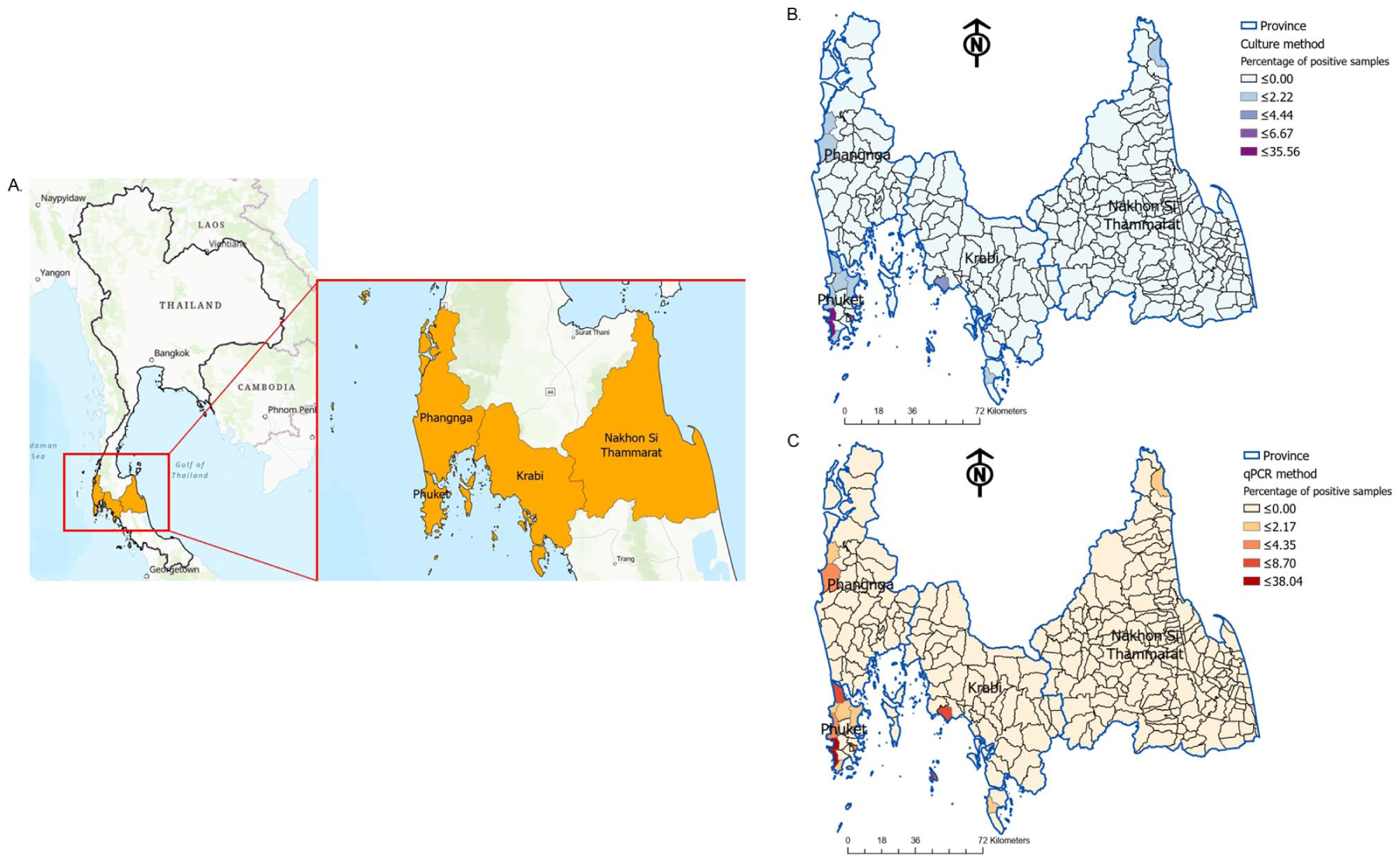
Geographical distribution of Legionella detection in environmental water samples across southern Thailand. (a) Overview map of Thailand highlighting the study area encompassing Phuket, Krabi, Phang Nga, and Nakhon Si Thammarat provinces. (b) Choropleth map illustrating the percentage of positive Legionella detections at the subdistrict level using the culture method. The color gradient represents increasing percentages of positive samples, as indicated in the legend. (c) Choropleth map illustrating the percentage of positive Legionella detections at the subdistrict level using the real-time PCR method. The color gradient represents increasing percentages of positive samples, as indicated in the legend.
Two hundred microliters of each processed sample were extracted using the Magpurix 12 A System and Zinexts Kit. Extracted DNA was analyzed using the optimized real-time PCR protocol with the modified primer set.
Detection of lpeAB and tet56 antimicrobial resistance genes by conventional PCR
DNA samples that tested positive for Legionella by real-time PCR were further subjected to conventional PCR for the detection of the lpeAB and tet56 AMR genes. Primer sequences were adopted from previously published studies as follows: lpeAB—Fw (5′-GTG ATG ATT GTC TTA TTG GTG CGA-3′) and lpeAB—Rw (5′-ATG GCG TTT AAG ATG ATG GTG ATT-3′), yielding a 359 bp amplicon 13 ; and tet56—Fw (5′-ATG TCT AAA AAT ATC AAA ATT CTC GTC-3′) and tet56—Rw (5′-CTA TGA TGA TTC ATA TTG AGG TAA GG-3′), yielding a 1170 bp amplicon. 14 PCRs were performed in a total volume of 25 µL containing 1× PCRBIO Taq Mix Red buffer (cat. no. PB10.13-02; PCR Biosystems Ltd), 0.4 µM of each forward and reverse primer, and 2 µL of DNA template (<500 ng), with PCR-grade water added to the volume. Each amplification set included a no-template control to monitor potential contamination. Thermal cycling was carried out on a Bio-Rad T100 thermal cycler with an initial denaturation at 95 °C for 1 min, followed by 40 cycles of denaturation at 95 °C for 15 s, annealing at 57.5 °C (lpeAB) or 52 °C (tet56) for 15 s, and extension at 72 °C for 10 s (lpeAB) or 30 s (tet56). PCR products were resolved on 1.5% (w/v) agarose gels (HIMEDIA, Mumbai, India) in 1× TBE buffer prepared in-house from analytical-grade reagents (Sigma–Aldrich, St. Louis, MO, USA), stained with SYBR Safe DNA gel stain (Thermo Fisher Scientific, Waltham, MA, USA), and visualized under UV transillumination.
Statistical analysis and data visualization
Descriptive statistics as mean (standard deviation) were calculated using Microsoft® Excel for Mac (version 16.96). Group comparisons were performed using two-proportion z-tests for sample sizes ⩾30 or Fisher’s exact tests for sample sizes <30, with p < 0.05 considered statistically significant. Statistical tests were run using IBM SPSS Statistics for Windows, version 25.0 (IBM Corp., Armonk, NY, USA). Spatial distribution of Legionella-positive samples was mapped by subdistrict using ArcGIS Pro (v2.6.0; Esri, Redlands, CA, USA), employing Natural Breaks (Jenks) classification.15,16 Administrative shapefiles were sourced from the Humanitarian Data Exchange platform. 17
Results
Optimization protocol
As shown in Table 2, the combination of the MagPurix extractor and the qPCRBIO Probe Mix reagent yielded the lowest Cq values among all tested protocols and was selected as the optimized approach for subsequent analyses. To further verify assay performance, the sensitivity of the multiplex primer set was compared with that of the duplex primer set using DNA templates from L. pneumophila serogroup 1, L. pneumophila serogroup 10, and Legionella feeleii. Table 3 presents the comparative Cq values obtained from both primer–probe sets. During the preliminary evaluation of the original multiplex primer set (Supplemental File S2), false-negative amplification was observed for L. feeleii, and occasional nonspecific or false-positive signals were detected in several non-Legionella species. These limitations, resulting from suboptimal probe–primer complementarity and threshold behavior under single-threshold mode, affected the diagnostic reliability of the original design. Consequently, a modified primer set was generated by combining the Legionella spp. component from the duplex assay with the L. pneumophila and serogroup 1 components from the original multiplex assay. When tested with equivalent template volumes, the modified primer set successfully detected all intended targets with Cq values comparable to the original assays. Further improvement was achieved by increasing the DNA template volume, which enhanced the detection sensitivity across all targets. Accordingly, the modified primer set and optimized protocol were adopted for the remainder of this study.
Cq values for the detection of Legionella spp. and L. pneumophila targets using various protocols.

Duplex primer set was used with 2 µl of DNA template. Data were presented as mean (SD) from duplication.
L. pneumophila: Legionella pneumophila; SD: standard deviation.
Comparison of Cq values for target detection using the duplex, original multiplex, and modified multiplex primer sets under optimized conditions.

Data were presented as mean (SD) from duplication.
L. feeleii: Legionella feeleii; L. pneumophila: Legionella pneumophila; ND: not detect (Cq > 45 cycles); SD: standard deviation; SG: serogroup; –: not done.
Real-time PCR results varied depending on the determination mode used for interpretation, as presented in Supplemental File S2. Analysis using the regression mode for each target yielded negative results. When applying the regression mode to all targets simultaneously, Cq values were observed without characteristic S-shaped amplification curves for the Legionella spp. target. Additionally, the single-threshold mode—whether applied to individual targets or all targets combined—consistently produced Cq values lacking S-shaped amplification curves and led to false-positive signals with non-Legionella species.
LOD in experimental settings and specificity of the modified primer set
The LOD of the real-time PCR assay varied depending on sample type and water matrix, as summarized in Figure 1 and Table 4. For colony suspensions, the LOD obtained by real-time PCR was comparable to that of the culture method, and direct PCR was able to detect concentrations as low as 101 CFU/mL. In spiked RO and tap water samples, filtration markedly enhanced detection sensitivity for both culture and real-time PCR, achieving LOD values within the range of 100–101 CFU/L. Non-filtrated water samples also yielded lower LODs than those observed with direct PCR. Comparable LOD values were obtained across all target genes in the modified primer set. The modified primer set demonstrated 100% specificity for Legionella, with no amplification observed in any non-Legionella species listed in Table 1, all of which are commonly present in environmental water.
A comparison of the LOD values between culture and real-time PCR under different conditions.

gDNA: genomic DNA; LOD: limit of detection; L. pneumophila: Legionella pneumophila; RO: reverse osmosis; SG: serogroup; –: not applicable.
LOD of colony suspension samples and artificially spiked RO/tap water was reported in unit of CFU/mL and CFU/L, respectively.
Legionella detection in real-world environmental water samples
A total of 202 leftover environmental water samples from the Regional Medical Sciences Center, 11/1 Phuket were utilized for Legionella detection using our optimized real-time PCR protocol. Full details are provided in Supplemental File S3, and a summary is shown in Table 5. With the exception of a single natural sample collected from the sea, the majority of samples were obtained from human-made water distribution systems. Table 5 illustrates that the real-time PCR assay achieved a detection rate of 48.51% (98/202), significantly outperforming the traditional culture method, which showed a detection rate of 24.26% (49/202; p < 0.001). Forty-five samples yielded positive results with consistent identification across both methods.
Summary of Legionella detection, species identification, AMR genes, and statistical significance (n = 202).

Detailed sample-level data are provided in Supplemental File S3.
AMR: antimicrobial resistance; CT: cooling towers; DF: decorative fountains; DK: drinking water; L. pneumophila: Legionella pneumophila; PPS: premise plumbing systems; SP: swimming pool.
It is important to note that the temperature data for the samples were derived from recorded observations during collection and did not involve exact temperature measurements.
Statistical significance between PCR and culture detection rates was evaluated using the two-proportion Z-test or Fisher’s exact test as appropriate.
The prevalence of Legionella spp., as determined by both culture and real-time PCR methodologies, was found to be the most predominant among the detected species, with comparable detection rates of 63.27% and 68.37%, respectively (p = 0.536). Specifically, L. pneumophila non-serogroup 1 was identified via culture in 22.45% of samples and by real-time PCR in 17.35% of samples (p = 0.426). Furthermore, L. pneumophila serogroup 1 exhibited identical detection rates of 14.29% with both culture and real-time PCR methods (p = 1.000). The positive detection rate of Legionella in hot water samples did not substantially differ between culture (55.38%, 36/49) and real-time PCR (63.08%, 41/98; p = 0.373). Conversely, a statistically significant increase in Legionella detection was observed in cold-water samples from premise plumbing systems when employing real-time PCR compared to culture methods (41.61% versus 9.49%, p < 0.001). Although the overall detection rate of Legionella by real-time PCR was higher than that of the culture method, two samples yielded positive results by culture but were negative by PCR (Supplemental File S3). These discrepancies can be attributed to the presence of non-amplifiable DNA due to degradation during sample processing.
Environmental water samples yielding positive results by culture or real-time PCR, with known sampling locations (Supplemental File S4), were utilized in a spatial distribution analysis to compare pathogen prevalence at the subdistrict level as shown in Figure 2(b) and (c). The highest Legionella prevalence was observed in Patong subdistrict, Kathu District, Phuket, with detection rates of 35.56% (16/45) by culture and 38.04% (35/92) by real-time PCR.
Antimicrobial resistance gene detection
Conventional PCR screening of 98 environmental water samples that tested positive for Legionella by real-time PCR identified two AMR genes, lpeAB and tet56. The lpeAB gene was detected in 6.1% (6/98) of samples, occurring predominantly in Legionella spp. (7.4%, 5/68) and in a single isolate of L. pneumophila serogroup 1 (1.0%, 1/14), while no L. pneumophila non-serogroup 1 isolates were positive. In contrast, the tet56 gene was detected in only one L. pneumophila isolate, as summarized in Table 5.
Discussion
This present study introduced an optimized real-time PCR assay leveraging a modified multiplex primer set for the simultaneous detection of Legionella spp., L. pneumophila, and L. pneumophila serogroup 1 in environmental water samples collected from the southern provinces of Thailand. This assay demonstrated considerable improvements in both analytical performance and field applicability compared to conventional culture methods, which are inherently limited by low sensitivity, prolonged incubation periods, and dependence on bacterial viability. The real-time PCR assay developed in this study achieved a detection rate of 48.51%, significantly higher than the 24.26% observed with culture-based diagnostics (p < 0.001), aligning with global trends that advocate for molecular diagnostics in Legionella surveillance.7,9,18
Despite the superior sensitivity and rapid turnover of real-time PCR for Legionella detection, the occurrence of culture-positive yet PCR-negative results, as revealed in this study, underscores critical methodological nuances. DNA degradation from non-intact cells or suboptimal extraction, genetic variability or mutation in the target genes, sample heterogeneity, or PCR inhibition by environmental substances are among the potential causes of these discrepancies. 19 Consequently, PCR offers substantial advantages; however, its application as an independent instrument in environmental surveillance necessitates caution; complementary culture methods are still required to confirm viability and facilitate outbreak investigations.
The superior sensitivity of the modified primer set was largely attributed to the inclusion of three distinct genetic targets: the 23S–5S rRNA gene for Legionella spp., the mip gene for L. pneumophila, and the wzm gene specific to L. pneumophila serogroup 1. This tri-target design ensured comprehensive coverage of clinically relevant Legionella species and serogroups, addressing a major limitation of the original multiplex set, which either failed or lacked sufficient PCR efficiency to detect non-pneumophila species such as L. feeleii. While L. feeleii detection was documented in the original study, 7 its absence in the present analysis was hypothesized to stem from a mutation in the tested strain that precluded effective primer binding. Notably, the modified assay was capable of detecting as few as 1 CFU/L in spiked environmental samples that were filtrated, a detection threshold that surpasses most standardized methods and rivals or exceeds sensitivity benchmarks reported in the literature.20,21 The analytical specificity of the assay was validated against 24 bacterial strains, including common waterborne contaminants and non-target species. During the early evaluation phase, the original multiplex primer set exhibited sporadic cross-reactivity with a few non-Legionella species (e.g. Staphylococcus aureus, Rhodococcus equi, and Vibrio parahaemolyticus), as presented in Tables S3 and S4 of Supplemental File S2. However, these false-positive signals were completely eliminated following primer modification. The final optimized primer–probe set developed and validated in this study achieved 100% analytical specificity for Legionella detection, confirming its robustness and reliability for environmental testing where microbial diversity is high.
L. pneumophila serogroup 1 remains the predominant cause of Legionnaires’ disease worldwide, and its identification is crucial for both clinical diagnosis and outbreak prevention. In this study, L. pneumophila serogroup 1 was detected in 14.29% of samples by both culture and PCR, reinforcing its epidemiological relevance. Interestingly, a substantial proportion of samples tested positive exclusively by PCR, particularly those from cold-water sources, suggesting that culture may underrepresent environmental risk due to its inability to detect viable but non-culturable cells. 6 This limitation highlights the need for future studies to incorporate viability-based molecular assays, such as propidium monoazide or ethidium monoazide coupled with quantitative polymerase chain reaction (qPCR), which can discriminate between viable and non-viable Legionella cells.22–24 Such approaches would enhance the accuracy of environmental surveillance and provide a clearer assessment of the infectious risk posed by viable Legionella populations. This finding underscores the necessity of incorporating molecular diagnostics to achieve comprehensive environmental risk assessments. Moreover, the real-time PCR assay enabled more nuanced detection across temperature regimes, with significant detection increases in cold-water samples (41.61% by PCR versus 9.49% by culture, p < 0.001). This is particularly relevant for countries like Thailand, where water systems exposed to tropical climates may serve as reservoirs for Legionella growth and dissemination.
Spatial distribution mapping revealed Patong subdistrict in Phuket as a critical hotspot, with the highest positivity rates across both detection methods. This high-resolution geospatial data is instrumental for public health agencies to prioritize interventions, especially in high-risk regions frequented by international tourists. Such targeted surveillance is aligned with WHO’s risk-based management frameworks for water safety.
Macrolides and fluoroquinolones, most commonly azithromycin and levofloxacin, are recommended as first-line therapies for moderate to severe Legionnaires’ disease. 25 Although AMR has historically not been regarded as a major concern in Legionella infections, recent evidence indicates emerging resistance in clinical strains. Documented cases include in vivo selection of fluoroquinolone resistance associated with treatment failure 26 and azithromycin resistance mediated by efflux mechanisms in L. pneumophila serogroup 1. 27 These reports highlight the potential for reduced efficacy of standard therapies and underscore the importance of ongoing resistance surveillance.
Within this context, the present study constitutes, to the best of our knowledge, the first report of AMR gene detection in environmental Legionella isolates from Thailand. Although the prevalence observed was lower than that reported in the international investigation, 28 this comparatively reduced detection rate suggests that resistance may currently be less pervasive in Thailand. Such variation may reflect regional differences in antimicrobial usage patterns, environmental conditions, or water system management practices that shape selective pressures on Legionella populations. It may also indicate that Thailand is at an earlier stage in the emergence and dissemination of AMR genes, thereby offering an important window of opportunity to strengthen surveillance and implement preventive strategies before resistance becomes more widespread.
Of particular significance was the identification of the lpeAB efflux pump gene, previously associated with reduced macrolide susceptibility, and the unexpected detection of the tet56 gene in a L. pneumophila isolate. The tet56 gene encodes a ribosomal protection protein that confers tetracycline resistance, and to date, it has only been described in non-pneumophila species such as L. longbeachae and L. bozemanae, 28 making our detection the first report in L. pneumophila. Interestingly, this tet56-positive sample also exhibited a discrepancy between culture (which identified Legionella spp.) and real-time PCR (which identified L. pneumophila). This discordance may reflect mixed-species contamination within the sample, limitations in the species-level resolution of culture methods, or the detection of non-viable L. pneumophila cells by PCR. Such findings highlight the complexity of interpreting AMR gene data in environmental contexts and reinforce the value of applying complementary methods to improve resolution and reliability.
Collectively, these discoveries enhance the current comprehension of AMR genes in Legionella and underscore the significance of integrating AMR gene monitoring into environmental surveillance frameworks. Even resistance determinants that are not explicitly associated with first-line therapies, such as tet56, can function as early markers of emerging resistance and indicators of genetic exchange in clinically significant species. However, the findings necessitate cautious interpretation. This study’s primary limitation is the absence of antimicrobial susceptibility testing. Consequently, the phenotypic significance of the detected AMR genes has yet to be determined.
Upon detecting Legionella, Thailand’s public health authorities initiate epidemiological investigations, 29 with key control strategies encompassing chlorine management, hot water system regulation, and cooling tower disinfection. To ensure effective chlorine management, maintaining a residual chlorine concentration of at least 0.2 mg/L throughout the water distribution system is paramount; deviations from this standard necessitate immediate corrective actions. Hot water system regulation focuses on inhibiting Legionella growth by maintaining water temperatures above 60 °C in storage and distribution, with a minimum of 50 °C at distal points to prevent biofilm formation. Cooling tower disinfection involves routine cleaning and periodic shock treatment with chlorine to eliminate Legionella and other microorganisms. These control strategies, when implemented effectively, are essential for minimizing the risk of Legionnaires’ disease, particularly in susceptible individuals and healthcare environments. Furthermore, proactive prevention through the development and implementation of comprehensive water safety plans is strongly emphasized. These plans should incorporate regular monitoring and microbiological surveillance, including systematic testing for Legionella, to guarantee continuous water safety and prevent future colonization. Suspected outbreaks necessitate prompt and detailed case reporting, coupled with thorough environmental investigations to identify the source of contamination and guide appropriate public health interventions.
Operationalizing molecular surveillance requires balancing technical performance with economic and logistical constraints. The cost analysis presented in this study shows that real-time PCR incurs a cost of $7.95/sample, compared to $34.66 for culture (Supplemental File S5). This fourfold reduction in cost, coupled with a significantly shorter turnaround time, reinforces the assay’s suitability for routine surveillance, especially in low- and middle-income countries where diagnostic resources are limited. Furthermore, the ability to process previously collected, refrigerated water samples up to 1 month old without loss of diagnostic performance enhances its utility in real-world public health workflows.
Conclusion
This study developed an optimized multiplex real-time PCR assay for rapid and sensitive detection of Legionella spp., L. pneumophila, and L. pneumophila serogroup 1 in environmental water. The assay outperformed culture in sensitivity, reduced costs, and provided results within a day, making it suitable for routine surveillance. Detection of AMR genes (lpeAB and tet56) highlights emerging resistance risks. These findings support integrating molecular diagnostics with culture methods to strengthen public health monitoring and outbreak preparedness.
Supplemental Material
sj-docx-1-smo-10.1177_20503121251403893 – Supplemental material for Two-stage surveillance of Legionella in environmental water: Optimized multiplex TaqMan qPCR for pathogen detection followed by antimicrobial resistance gene identification
Supplemental material, sj-docx-1-smo-10.1177_20503121251403893 for Two-stage surveillance of Legionella in environmental water: Optimized multiplex TaqMan qPCR for pathogen detection followed by antimicrobial resistance gene identification by Watcharee Thongkhaw, Iriyaporn Kongthap, Watcharaporn Kamjumphol, Hiranya Sritart and Nattamon Niyomdecha in SAGE Open Medicine
Supplemental Material
sj-docx-2-smo-10.1177_20503121251403893 – Supplemental material for Two-stage surveillance of Legionella in environmental water: Optimized multiplex TaqMan qPCR for pathogen detection followed by antimicrobial resistance gene identification
Supplemental material, sj-docx-2-smo-10.1177_20503121251403893 for Two-stage surveillance of Legionella in environmental water: Optimized multiplex TaqMan qPCR for pathogen detection followed by antimicrobial resistance gene identification by Watcharee Thongkhaw, Iriyaporn Kongthap, Watcharaporn Kamjumphol, Hiranya Sritart and Nattamon Niyomdecha in SAGE Open Medicine
Supplemental Material
sj-docx-3-smo-10.1177_20503121251403893 – Supplemental material for Two-stage surveillance of Legionella in environmental water: Optimized multiplex TaqMan qPCR for pathogen detection followed by antimicrobial resistance gene identification
Supplemental material, sj-docx-3-smo-10.1177_20503121251403893 for Two-stage surveillance of Legionella in environmental water: Optimized multiplex TaqMan qPCR for pathogen detection followed by antimicrobial resistance gene identification by Watcharee Thongkhaw, Iriyaporn Kongthap, Watcharaporn Kamjumphol, Hiranya Sritart and Nattamon Niyomdecha in SAGE Open Medicine
Supplemental Material
sj-docx-4-smo-10.1177_20503121251403893 – Supplemental material for Two-stage surveillance of Legionella in environmental water: Optimized multiplex TaqMan qPCR for pathogen detection followed by antimicrobial resistance gene identification
Supplemental material, sj-docx-4-smo-10.1177_20503121251403893 for Two-stage surveillance of Legionella in environmental water: Optimized multiplex TaqMan qPCR for pathogen detection followed by antimicrobial resistance gene identification by Watcharee Thongkhaw, Iriyaporn Kongthap, Watcharaporn Kamjumphol, Hiranya Sritart and Nattamon Niyomdecha in SAGE Open Medicine
Supplemental Material
sj-docx-5-smo-10.1177_20503121251403893 – Supplemental material for Two-stage surveillance of Legionella in environmental water: Optimized multiplex TaqMan qPCR for pathogen detection followed by antimicrobial resistance gene identification
Supplemental material, sj-docx-5-smo-10.1177_20503121251403893 for Two-stage surveillance of Legionella in environmental water: Optimized multiplex TaqMan qPCR for pathogen detection followed by antimicrobial resistance gene identification by Watcharee Thongkhaw, Iriyaporn Kongthap, Watcharaporn Kamjumphol, Hiranya Sritart and Nattamon Niyomdecha in SAGE Open Medicine
Footnotes
Acknowledgements
We thank Dr. Sirinart Chumean for kindly providing PCR reagents, which were essential for the successful completion of this research. We also extend our gratitude to Ms. Madtika Fungkrajai and Ms. Thanawan Soimanee for their contributions to the AMR gene detection process. In addition, the authors acknowledge the use of QuillBot Premium as an AI-assisted language editing tool to enhance the clarity, grammar, and readability of the article.
Ethical considerations
This study did not involve human participants, animal subjects, or personal data, and therefore did not require ethical approval.
Author contributions
Writing the article: N.N.; conceiving and planning the experiments: N.N., W.T.; data analysis: N.N., W.T., H.S.; data curation and visualization: W.T., H.S.; resources and review of the article: I.K., W.K.; funding acquisition: W.T., N.N.; supervision and project administration: N.N.; reading and approving the final version of the article: all authors; revision: N.N.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by Thammasat University Research Fund, contract no. TUFT 048/2568. The funder had no role in study design, data collection and analysis decision to publish, or preparation of the article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
All data supporting the findings of this study are available within the article and its Supplemental Information.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
