Abstract
Background:
Antimicrobial resistance is one of the most significant challenges to global public health and the risk factors in severe pneumonia are constantly growing. Therefore, this study aimed to identify factors associated with antimicrobial resistance and conduct survival analysis of severe pneumonia patients with single and multiple pathogens in the National Referral Hospital, Jakarta, Indonesia.
Methods:
A retrospective method was used, and secondary data were collected from severe pneumonia patients admitted to the intensive care unit at Cipto Mangunkusumo National Referral Hospital, Jakarta, Indonesia, from January 2016 to December 2022. Respiratory specimens were collected through bronchial washing. Furthermore, univariate and multivariate analyses were performed to analyze factors associated with antimicrobial resistance. Kaplan‒Meier survival curves were generated with the log-rank test to compare 30-day mortality between patients infected with single, dual, and multiple pathogens.
Results:
The results showed that a total of 333 patients from 415 enrolled were analyzed. Klebsiella pneumoniae (35.4%), Acinetobacter baumannii (29.3%), and Pseudomonas aeruginosa (15.4%) were the most frequently isolated Gram-negative pathogens. Factors associated with resistance to aminoglycoside, carbapenem, and quinolone were sepsis, cerebrovascular disease, and ventilator-associated pneumonia, as indicated by p < 0.05. In addition, the Kaplan–Meier curves showed that multiple pathogens influenced the survival rate of severe pneumonia patients (p < 0.05).
Conclusions:
Sepsis, cerebrovascular disease, and ventilator-associated pneumonia were associated with antimicrobial resistance in severe pneumonia patients. The survival rate of patients infected with multiple pathogens was low. This suggests the importance of further awareness regarding empirical antibiotic stewardship and mortality assessment in severe pneumonia patients.
Background
Pneumonia is one of the most common causes of hospitalization along with septicemia, heart failure, and osteoarthritis, accounting for more than 600,000 cases annually. 1 This condition is the second most common nosocomial infectious illness after urinary tract infection and the first leading community-acquired infection. Severe pneumonia is associated with a high mortality rate, with 16%–36% of patients eventually dying within a short period, despite effective antibiotic therapy. 2 Streptococcus pneumoniae is the most frequent pathogen of community-acquired pneumonia (CAP), while Gram-negative organisms are the major pathogen of hospital-acquired pneumonia (HAP). 3 Previous studies reported that Klebsiella pneumoniae, Acinetobacter baumannii, and Pseudomonas aeruginosa were identified among patients with CAP.4–6 These top three Gram-negative pathogens were also included in the most common multidrug-resistant Gram-negative bacteria and the World Health Organization’s (WHO) critical priority list for antibiotic-resistant bacteria investigation and development, posing a significant threat to global public health.7,8 According to the previous studies, there were no significant differences in mortality rates between patients having severe pneumonia with bacteria infection only, single viral infection, and with coinfections.9,10
Excessive and irrational use of antibiotics leads to antimicrobial resistance (AMR), specifically in pneumonia patients. 7 AMR is a growing and serious global threat to public health. 11 Bacteria AMR or antibiotic resistance reportedly caused 1.27 million deaths worldwide in 2019. 12 A previous study stated that the risk factors for antibiotic resistance in nosocomial pneumonia include home infusion therapy, chronic dialysis within 30 days, home wound care, antimicrobial therapy in the 90 days before, living in a nursing home or extended-care facility, and family history. 13
There are no available studies conducted globally and in Indonesia focusing on the three dominant Gram-negative bacteria. These include K. pneumoniae, A. baumannii, and P. aeruginosa using bronchial washing specimen. Therefore, this study aimed to identify factors associated with AMR and to perform survival analysis on severe pneumonia patients with single and multiple pathogens.
Methods
Study population
A retrospective method was used, and secondary data in the form of medical records were collected from all severe pneumonia patients with antibiotic resistance. Data collection was carried out at the intensive care unit (ICU) of Cipto Mangunkusumo National Referral Hospital, Jakarta, Indonesia, between January 2016 and December 2022. This location was selected because it served as a national referral hospital and the only center where bronchoscopy was performed for severe pneumonia patients to collect respiratory specimens. The sample size calculation was carried out using the logistic regression sample size calculation formula (Supplemental material File 3). 14 From the results of the sample size calculation, the minimum sample to be taken is 207 patients. The inclusion criteria include individuals aged 18–75 years old, who agreed to participate and had overall suitable conditions for bronchoscopy. On the other hand, the exclusion criteria include pregnant women and patients whose families declined bronchoscopy.
The bronchoscopy procedure was performed according to local guidelines applied in the hospital. In accordance with the policy, all pneumonia patients should have culture specimens upon admission. Sputum or nasopharyngeal samples were collected for nonintubated patients as well as tracheal aspirate samples for intubated patients. Bronchoscopy was performed after 3 days when there was no improvement with empirical antibiotic therapy. Before the procedure, both written and verbal informed consent were obtained from the families of patients, along with approval for potential future data usage. The consent was not directly received from patients due to the severe conditions. Chest radiography was initially performed followed by bronchoscopy. Bronchoscopy was carried out more than 3 days after the onset of pneumonia diagnosis for an average of 30 min depending on the overall condition of patients. Intravenous midazolam and propofol were given by the anesthesiologist for optimum sedation, then the patients were observed 1 h postprocedure.
During the pandemic starting in March 2020, routine COVID-19 antibody/antigen tests for bronchoscopy teams were carried out every 2 weeks, and reverse transcription polymerase chain reaction swab tests were conducted once a month as standard hospital operational procedures. All personnel engaged in aerosol-generating procedures such as bronchoscopy were required to comply with the personal protective equipment (PPE) protocol, requiring the use of PPE level 3. 15 These PPE include face shields, N95 masks with goggles, or a Powered Air-Purifying Respirator, along with gowns and gloves. Before donning and doffing the PPE, all personnel must adhere to standard hand hygiene. 16
Ethical considerations
This study was approved by the ethical committee of the Medical Faculty of Universitas Indonesia (Approval number: KET-303/UN2.F1/ETIK/PPM.00.02/2021) and by the hospital review committee. All processes in this study were conducted in accordance with the Declaration of Helsinki.
Subject definitions
CAP is a respiratory infection affecting the alveoli, which occurs during outpatient care or within the initial 48 h of hospital admission. 17 Severe community-acquired pneumonia (SCAP) was diagnosed based on the 2019 American Thoracic Society guideline, which included either one major criterion or three or more minor criteria. 18 HAP patients had pneumonia occurring ⩾48 h after hospital or healthcare facility admission. Ventilator-associated pneumonia (VAP) diagnosis required mechanical ventilation for >2 consecutive calendar days, starting from day 1 when the ventilator was first placed or the day before.4,19
The WHO defined a confirmed case of COVID-19 as a positive nucleic acid amplification test regardless of clinical or epidemiological criteria (option A). It was also defined as a person meeting clinical and/or epidemiological criteria with a positive professional-use or self-test severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) Antigen (option B). 20 Single pathogen infection in this study were defined as pneumonia patient infected with one of the three predominant causative Gram-negative bacteria including K. pneumoniae, A. baumannii, and P. aeruginosa. Dual pathogens infection referred to pneumonia patients infected with two of the three predominant causative Gram-negative bacteria. In this study, multiple pathogens infection was defined as pneumonia patients infected with one or more of the three predominant causative Gram-negative bacteria along with SARS-CoV2 or Candida coinfections.
Diagnostic and drug susceptibility tests
Respiratory specimens used in this study were only collected through bronchial washing. The specimens were analyzed to confirm the etiology using the Vitek 2 Compact system (Biomerieux). Inoculation was carried out on Blood Agar (211307-BD, France) and Mac-Conkey Agar (CM0115B-Oxoid, United Kingdom) followed by incubation at a temperature of 35°C–37°C lasting 18–24 h. Subsequently, the isolated specimens were transferred to the Vitek system for etiology identification. The antibiotic susceptibility test (listed in Supplemental material Table 1, File 1) was performed using the disk diffusion method based on the Clinical Laboratory Standards Institute M100 guideline. 21 Antimicrobial sensitivity and resistance profile for each pathogen are shown in Supplemental material Figures 1–3, File 1. The reference bacterial strains used for quality control in this study were Escherichia coli, P. aeruginosa, Staphylococcus aureus, and Candida albicans (ATCC). Disk diffusion was adopted for resistance testing instead of Vitek Antibiotic Susceptibility Testing, primarily because of the protocol policy and financial considerations within the center during data collection. Moreover, gene testing results were not obtained because, during the study period, the procedure was not yet available in Indonesia.
Statistical analysis
Categorical variables were reported as n (%) and bivariate analysis was performed to analyze the association of subject characteristics with antibiotic susceptibility by a 2 × 2 table using chi-square or Fisher’s exact tests. Variables with p values <0.25 were further analyzed with multivariate logistic regression and all p values <0.05 were considered statistically significant. Kaplan‒Meier survival analysis was performed with the log-rank test to analyze the 30-day mortality between severe pneumonia patients infected with single and multiple pathogens. All recorded data were analyzed using the Statistical Package for the Social Sciences version 26 (IBM Corporation, Armonk, NY, USA).
Results
Clinical and demographic characteristics of severe pneumonia patients
The majority of patients were male (60.2%) with a mean age of 54.6 years, had CAP (38.1%) and neurologic/cerebrovascular diseases as the most common comorbidity (41.4%) (Table 1). The most common Gram-negative pathogen of pneumonia was K. pneumoniae, while the leading comorbidity in CAP, HAP, and VAP groups was cerebrovascular disease (Table 2).
Baseline characteristics of patients with severe pneumonia.
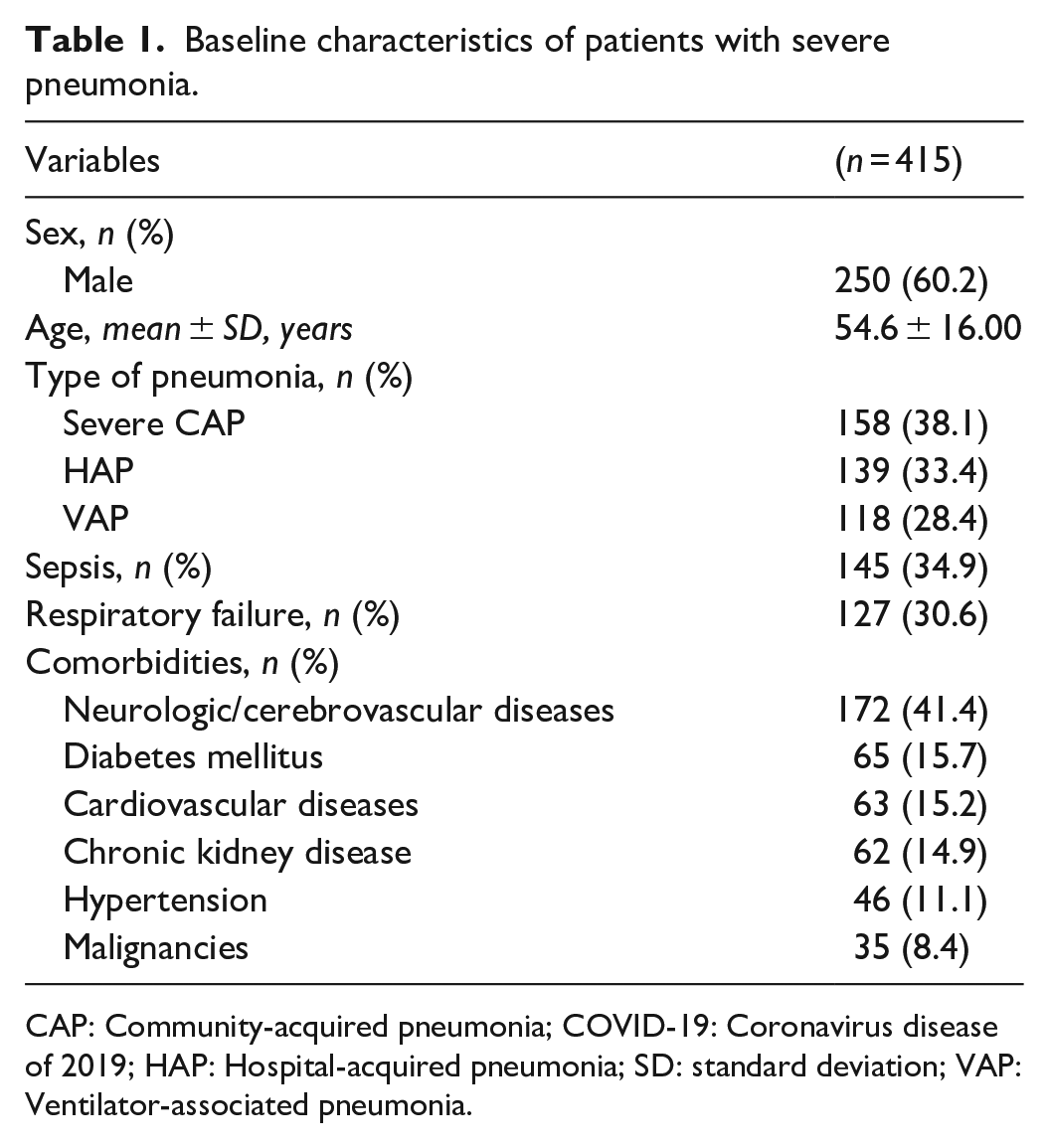
CAP: Community-acquired pneumonia; COVID-19: Coronavirus disease of 2019; HAP: Hospital-acquired pneumonia; SD: standard deviation; VAP: Ventilator-associated pneumonia.
Characteristics based on the type of pneumonia.

CAP: Community-acquired pneumonia; COVID-19: Coronavirus disease of 2019; HAP: Hospital-acquired pneumonia; VAP: Ventilator-associated pneumonia.
K. pneumoniae, A. baumannii, and P. aeruginosa were the most common Gram-negative identified pathogens (Figure 1). As reported in Figure 1 and Table 2, other causative pathogens of pneumonia include Pseudomonas fluorescens, S. aureus, Staphylococcus epidermidis, Streptococcus viridans, and Serratia marcescense. During the study period, culture medium for S. pneumoniae was not available resulting in no data for this pathogen.

Pathogen distribution of severe pneumonia patients.
Among the 415 patients enrolled, the analysis focused on 333 who had severe pneumonia and showed antibiotic resistance with one or more of the top three predominant Gram-negative pathogens (K. pneumoniae, A. baumannii, and P. aeruginosa). Factors of antibiotic resistance included in the univariate analysis were sex, age, type of pneumonia (SCAP, HAP, VAP, COVID-19), sepsis, respiratory failure, comorbidities such as neurologic disorders, diabetes mellitus, cardiovascular diseases, chronic kidney disease, hypertension, and malignancies. Variables with p values <0.25 were further analyzed with multivariate logistic regression.
Factors associated with antibiotic resistance in K. pneumoniae, A. baumannii, and P. aeruginosa
In the multivariate regression model for K. pneumoniae pathogen, a statistically significant association was found between sepsis and chloramphenicol (p = 0.11), gentamicin (p = 0.45), amikacin (p = 0.03), amoxicillin+clavulanic acid (p = 0.029), cefoperazone/sulbactam (p = 0.01), doripenem (p ⩽ 0.01), meropenem (p = 0.001), and imipenem (p < 0.01) resistance. Severe CAP was significantly associated with doripenem (p = 0.02) and imipenem (p = 0.029) resistance (Table 3).
Factors associated with antibiotic resistance in K. pneumoniae.

p < 0.05 (statistically significant); ‘—’: no comorbidity with the specific antibiotic; CAP: Community-acquired pneumonia; CI: confidence interval; OR: odds ratio; VAP: Ventilator-associated pneumonia.
Based on the multivariate regression model for the A. baumannii pathogen, there was a statistically significant association between malignancies and amikacin resistance (p = 0.029). Sex, particularly male, was significantly associated with ceftazidime resistance (p = 0.032), while VAP was substantially related to levofloxacin resistance (p = 0.019) (Table 4).
Factors associated with antibiotic resistance in A. baumannii.

p < 0.05 (statistically significant); ‘—’ : no comorbidity with the specific antibiotic; CI: confidence interval; OR: odds ratio; VAP: Ventilator-associated pneumonia.
In the multivariate regression model for the P. aeruginosa pathogen, there was a statistically significant relation between VAP with amikacin (p = 0.02), aztreonam (p = 0.012), and ciprofloxacin (p = 0.042) resistance. Cerebrovascular disease was significantly associated with doripenem (p = 0.013), meropenem (p = 0.024), and imipenem (p = 0.012) resistance, while sepsis was substantially related to cefoperazone/sulbactam (p = 0.47), imipenem (p = 0.06), and levofloxacin (p = 0.07) resistance. Respiratory failure was significantly associated with doripenem (p = 0.01), meropenem (p = 0.032), and imipenem (p = 0.035) resistance (Tables 5a and 5b).
Factors associated with antibiotic resistance in P. aeruginosa.
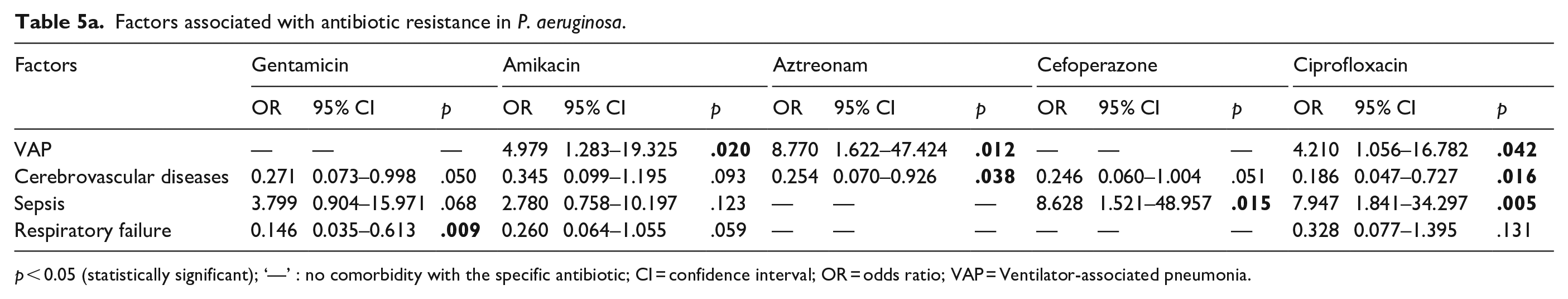
p < 0.05 (statistically significant); ‘—’ : no comorbidity with the specific antibiotic; CI = confidence interval; OR = odds ratio; VAP = Ventilator-associated pneumonia.
Factors associated with antibiotic resistance in P. aeruginosa.

p < 0.05 (statistically significant); ‘—’ : no comorbidity with the specific antibiotic; CAP = Community-acquired Pneumonia; CI = confidence interval; OR = odds ratio.
Survival analysis of patients infected with single and multiple pathogens
Among the 333 patients infected with at least one of the top three pathogens, the 30-day survival rate was compared between 305 (91.5%) infected with a single pathogen, 4 (1.2%) with dual pathogens infection, and 24 (7.2%) who had coinfections of SARS-CoV-2 or Candida (multiple pathogens infection). For the 24 severe pneumonia patients infected with multiple pathogens, 21 were coinfected with SARS-CoV-2, while the remaining three had coinfections with Candida tropicalis, Candida famata, as well as both C. tropicalis and C. famata, respectively. There was a significant difference in the 30-day mortality between the three groups (p = 0.002), with patients infected with dual and multiple pathogens having a lower survival rate. Kaplan‒Meier survival curves between the three groups are shown in Figure 2, while more details are presented in Supplemental material Table 2a–2c, File 2.

Kaplan–Meier curves for severe pneumonia patients infected with single, dual, and multiple pathogens.
Discussion
This is the first study conducted globally to elaborate on factors associated with antibiotic resistance in the three most commonly found Gram-negative bacteria and conduct an analysis of 30-day mortality among severe pneumonia patients infected with single and multiple pathogens. During the COVID-19 pandemic in March 2020, based on hospital protocols all pneumonia patients had to be screened for SARS-CoV-2, so that several of the study subjects were confirmed to have the disease, as showed by a positive bronchial washing polymerase chain reaction for severe acute respiratory syndrome coronavirus 2. However, most of the patients could not be included in the analysis due to a lack of bacteria culture results. 22 Additionally, the results of this study may help the management of severe pneumonia patients admitted to the ICU in selecting proper empiric antibiotics. All patients received initial antibiotics according to the guidelines of the in-house AMR control program. The empirical antibiotics most frequently administered for severe pneumonia patients are carbapenems, third-generation cephalosporins, and fluoroquinolones.
In this study, males were found to be the most frequently affected by severe pneumonia (60.2%), with a mean age of 54.6 years. Similar results were found in previous studies, with the frequency of male pneumonia patients being 65.4% and 64.6%, respectively.23,24 Males were reported to have poorer health habits and more other health issues, which make them more likely to get severe pneumonia. 25 We found that CAP was the most common type of severe pneumonia in this study (38.1%). K. pneumoniae (33.5%), A. baumannii (29.7%), and P. aeruginosa (10.7%) were the most frequently isolated Gram-negative bacteria among CAP patients. A previous study yielded identical results, in which Klebsiella pneumonia, A. baumannii, and P. aeruginosa were the top three most isolated microorganisms among adult patients with CAP. 6 The most common pathogenic cause of CAP worldwide, specifically in Western European countries is reportedly S. pneumoniae. However, several studies in Southeast Asia reported that Gram-negative bacteria were the most common cause of CAP, including studies from Thailand, 26 Singapore, 27 and Indonesia. 6 Differences in demographics might be a potential explanation for variations in the etiology of CAP. Comorbidities and patient conditions might also play a significant role. Previous studies reported that in immunocompromized patients and the elderly, the predominant etiology of CAP was Gram-negative bacteria.26,28 Similar to the results in this study, K. pneumoniae was identified as the most common etiology of SCAP in elderly patients treated in the ICU.26,27 In this study, K. pneumoniae, A. baumannii, and P. aeruginosa were also the most common pathogen for both HAP and VAP patients. Previous studies have reported similar findings, indicating that in patients with HAP or VAP, the predominant bacterial pathogens were Gram-negative bacteria, consistent with the top three pathogens identified in our study.29,30 These three Gram-negative bacteria are listed among the most significantly antibiotic-resistant bacteria and are classified as pan-drug-resistant, thereby requiring special attention due to the threat to global health. 31
This study identified underlying cerebrovascular disease, sepsis, and VAP as the most frequent factors associated with the acquisition of nonsusceptibility to carbapenem, fluoroquinolones, and aminoglycosides in K. pneumoniae, A. baumannii, and P. aeruginosa isolates. Carbapenem resistance to Gram-negative bacilli is listed as the second most urgent AMR threat by the Center for Disease Control and Prevention, and also considered a pathogen of critical priority by the WHO. Lima et al. 32 reported that patients with longer hospital stays and previous bacteria infections, specifically hospital-acquired types, and were treated using carbapenem supported with mechanical ventilation had an increased risk of sepsis caused by nonsusceptible Gram-negative bacteria. Frequent use of antibiotics and extended stays in the ICU results in aminoglycoside-resistant Gram-negative bacteria. 2 Underlying cerebrovascular disease, healthcare-associated pneumonia, and previous exposure to fluoroquinolone within 3 months were associated with fluoroquinolone resistance.32,33
Severe pneumonia patients admitted to the ICU have a high risk of developing antibiotic resistance. Therefore, improving antibiotic stewardship programs is important and recommended to optimize the utilization and management of antibiotics for better outcomes. 34 In this study, antibiotic susceptibility testing found that the majority of the patients had a high resistance rate. For K. pneumoniae isolates, the highest resistance was against cefotaxime (46.9%). K. pneumoniae is a type of Enterobacterales that shows a high frequency of antibiotic resistance. In a previous study, this bacteria showed high resistance against cefotaxime (79.2%) and ceftazidime (75.7%). 35 Furthermore, we found that A. baumannii isolates were resistant to ceftriaxone (39.6%) and cefotaxime (39.2%). Previous study reported similar results where the highest resistance of A. baumannii strains was to the cephalosporin group, specifically ceftriaxone (83%) and ceftazidime (75%). This bacteria is known to be the predominant cause of nosocomial infection and is considered a highly drug-resistant strain worldwide with a high mortality rate. 36 In our findings, the resistance of A. baumannii to carbapenem was 35.5% against doripenem, 25.9% against meropenem, and 37% against imipenem. Carbapenem is commonly prescribed, but recent reports have proven that A. baumannii can increase resistance to the drug ranging from 62% to 85%.36,37 This bacteria also was reported to show high resistance to aminoglycoside, penicillin, and fluoroquinolone, with rates of 86% for gentamicin, 38 90% for piperacillin/tazobactam, 39 and 71% for levofloxacin, 40 compared to our findings of 38.8%, 38.8%, and 34.1%, respectively. This study showed P. aeruginosa isolates were 13.9% resistant to gentamicin, aztreonam, and meropenem, 18% resistant to tigecycline, 12.1% to ciprofloxacin, and 9.5% to piperacillin/tazobactam. This findings align with results from other studies, reporting P. aeruginosa high resistance rate: 97% to aminoglycoside, 41 21.5% to aztreonam, 42 19.3% to meropenem, 42 90% to tigecycline, 43 13.4% to ciprofloxacin, 42 and 10.8% to piperacillin. 42 Awareness of factors associated with AMR can guide future interventions and reduce the risk of developing multidrug resistance infections, thereby reducing the mortality of patients with multidrug resistance pathogens.
A low survival rate was observed in the group of patients infected with dual pathogens and those who had SARS-CoV-2 or Candida coinfections (multiple pathogens). These results are in line with previous studies on pneumonia patients having virus coinfection with other pathogens such as C. tropicalis and C. albicans, leading to increased in-hospital mortality.44,45 Previous study reported high mortality in subjects with Gram-negative K. pneumoniae or A. baumannii bacterial pneumonia with SARS-CoV-2 co-infection. 46 Hedberg et al. 47 also reported that when compared with influenza virus and respiratory syncytial virus (RSV) co-infection, bacterial pneumonia patients with SARS-CoV-2 co-infection had the highest percentage of 30-day mortality. In the analysis, the presence of another Gram-negative pathogen and SARS-CoV-2 or Candida significantly increased the 30-day mortality.
This study had several limitations, first, the pathogens were not analyzed within 72 h of hospitalization. Second, some subjects were not enrolled in this study due to a lack of documentation. There was also no culture medium or rapid antigen testing for the identification of other possible pathogens such as S. pneumoniae and Legionella pneumophilia. Gene testing for (OXA-48, aac(6’)-Ib, aac(3)-II, aph(3’)-Ia, rmtA (3), rmtB, rmtD) was not performed due to the unavailability of the procedure in Indonesia (2016–2022). 48 Third, data on antibiotic use 90 days before treatment was not included due to the unavailability. Considering the study’s duration amid the COVID-19 pandemic, SARS-CoV-2 might have affected mortality rates observed in this study.
The strength of this study lies in the provision of insight into the factors associated with AMR and a comparison of 30-day survival rates in severe pneumonia patients infected with single, dual, and multiple pathogens. Microbial pattern and antibiotic resistance type data may provide additional important information for the selection of empirical antibiotics in at-risk patients admitted to the ICU. However, these results should be used with caution due to the lack of data on previous antibiotic use. This study also collected respiratory specimens through bronchoscopy within 7 years, including during the COVID-19 pandemic.
Conclusions
In conclusion, this study found high rates of K. pneumoniae, A. baumannii, and P. aeruginosa isolates in severe pneumonia patients admitted to the ICU. Based on the results, sepsis, VAP, and underlying cerebrovascular disease were the most common factors associated with AMR, mainly to carbapenems, quinolones, and aminoglycosides. The 30-day survival rate was lower among patients infected with multiple pathogens. Therefore, early identification of multiple pathogens is an important issue to improve outcomes in severe pneumonia patients with AMR. Further prospective multicenter studies in severe pneumonia patients are needed to gather information on differences between various demographics. Drug resistance gene detection is required to validate a causal relationship between these factors and nonsusceptibility to carbapenems, quinolones, and aminoglycosides.
Supplemental Material
sj-docx-1-smo-10.1177_20503121241264097 – Supplemental material for Factors associated with antibiotic resistance and survival analysis of severe pneumonia patients infected with Klebsiella pneumoniae, Acinetobacter baumannii, and Pseudomonas aeruginosa: A retrospective cohort study in Jakarta, Indonesia
Supplemental material, sj-docx-1-smo-10.1177_20503121241264097 for Factors associated with antibiotic resistance and survival analysis of severe pneumonia patients infected with Klebsiella pneumoniae, Acinetobacter baumannii, and Pseudomonas aeruginosa: A retrospective cohort study in Jakarta, Indonesia by Gurmeet Singh, Tonny Loho, Mira Yulianti, Dita Aditianingsih, Laila Fakhriyatuz Zakiyah, Sudirman Fakhruddin Masse and Muhammad Rizki Triono in SAGE Open Medicine
Supplemental Material
sj-docx-2-smo-10.1177_20503121241264097 – Supplemental material for Factors associated with antibiotic resistance and survival analysis of severe pneumonia patients infected with Klebsiella pneumoniae, Acinetobacter baumannii, and Pseudomonas aeruginosa: A retrospective cohort study in Jakarta, Indonesia
Supplemental material, sj-docx-2-smo-10.1177_20503121241264097 for Factors associated with antibiotic resistance and survival analysis of severe pneumonia patients infected with Klebsiella pneumoniae, Acinetobacter baumannii, and Pseudomonas aeruginosa: A retrospective cohort study in Jakarta, Indonesia by Gurmeet Singh, Tonny Loho, Mira Yulianti, Dita Aditianingsih, Laila Fakhriyatuz Zakiyah, Sudirman Fakhruddin Masse and Muhammad Rizki Triono in SAGE Open Medicine
Supplemental Material
sj-docx-3-smo-10.1177_20503121241264097 – Supplemental material for Factors associated with antibiotic resistance and survival analysis of severe pneumonia patients infected with Klebsiella pneumoniae, Acinetobacter baumannii, and Pseudomonas aeruginosa: A retrospective cohort study in Jakarta, Indonesia
Supplemental material, sj-docx-3-smo-10.1177_20503121241264097 for Factors associated with antibiotic resistance and survival analysis of severe pneumonia patients infected with Klebsiella pneumoniae, Acinetobacter baumannii, and Pseudomonas aeruginosa: A retrospective cohort study in Jakarta, Indonesia by Gurmeet Singh, Tonny Loho, Mira Yulianti, Dita Aditianingsih, Laila Fakhriyatuz Zakiyah, Sudirman Fakhruddin Masse and Muhammad Rizki Triono in SAGE Open Medicine
Footnotes
Acknowledgements
The authors are grateful to all patients and families, as well as the healthcare workers (doctors and nursing staff) for the contributions in conducting this study. The authors are also grateful to the scientific community for the valuable feedback on the preprint version of this manuscript, which greatly contributed to improve the quality.
Author contributions
GS, MY, TL, and DA prepared the materials and collected and analyzed the data. GS and TL contributed to the study conception and design. GS, LFZ, SFM, and MRT wrote the first draft of the manuscript. All authors have read and approved the final manuscript.
Availability of data and materials
Data for this study are available upon request from the corresponding author (G.S.). The data are not publicly available to avoid compromising the privacy of the patients.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethics approval and consent to participate
This study was approved by the ethical committee of the Medical Faculty of Universitas Indonesia (Approval number: KET-303/UN2.F1/ETIK/PPM.00.02/2021) and by the hospital review committee.
Informed consent was obtained from all legally authorized representatives who participated in this study, according to the local ethics and hospital review committee.
Consent for publication
Not applicable.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
