Abstract
Objective:
Antimicrobial resistance is one of the serious threats in the world, including Ethiopia. Even though several studies were conducted to estimate common bacteria and their antibiotic-resistance profile in Ethiopia, it is difficult to estimate the overall resistant patterns due to the lack of a nationwide study. This systematic review aimed to determine the prevalence of gram-negative bacteria isolates and their antibiotic-resistance profile among pediatrics patients in Ethiopia.
Methods:
A web-based search using PubMed, EMBASE, Science Direct, the Cochrane Database for Systematic Reviews, Scopus, Hinari, Sci-Hub, African Journals Online Library, and free-text web searches using Google Scholar was conducted from August to September 16, 2021. Each of the original articles was searched by Boolean search technique using various keywords and was assessed using the Joanna Briggs Institute Critical Appraisal Checklist. The data were extracted using Microsoft Excel format and exported to STATA 14.0 for statistical analyses.
Results:
The database search delivered a total of 2,684 studies. After articles were removed by duplications, title, reading the abstract, and assessed for eligibility criteria, 19 articles were included in the systematic review. Of a total of 1372 (16.77%) culture-positive samples, 735 (53.57%) were gram-negative.
Conclusion:
Gram-negative bacteria were identified as the common pathogen causing infection in pediatrics and the level of resistance to commonly prescribed antibiotics was significantly higher in Ethiopia. Culture and susceptibility tests and well-designed infection control programs are important measures.
Introduction
Infectious diseases including bacterial infections are the leading causes of childhood mortality, particularly in developing countries.1,2 Even though the introduction of broad-spectrum antibiotics has been made over the last 70 years, the mortality and morbidity rate due to these life-threatening pathogens remains high. This is because of the increment of antibiotic resistance to gram-negative bacteria. 3
Antimicrobial resistance (AMR) is one of the serious threats to global public health; it affects the ability to treat the infection effectively and puts patients at risk of prolonged illness, complications, and death. It is estimated that 700,000 people die annually as a result of AMR, and this figure will rise to 10 million by 2050 if no action is taken. 4 AMR has drastically amplified due to the irrational prescribing and/or inappropriate use of antibiotics, and limited availability of reliable laboratories, particularly blood culture, in areas far from hospitals. These factors lead to compromised care, outdated antibiotic guidelines, and the emergence of resistance.3,5
Effective empirical therapy of bacterial diseases in the pediatrics population requires knowledge of local AMR patterns, acquired through up-to-date surveillance.6,7 Understanding the prevalence of common pathogens is essential for definitive antibiotic therapy and reduces exposure of non-involved bacterial pathogens to unnecessary antibiotics. 8 The World Health Organization (WHO) released a list of antibiotic-resistant priority pathogens to combat the AMR crisis, to promote the development of new antibiotics, and advocate rational use of the available ones. The majority of these microorganisms are gram-negative bacteria. 9 Gram-negative bacteria are common cause of different human infectious diseases, such as urinary tract infections, wound infections, gastroenteritis, ear infection, meningitis, septicemia, and pneumonia.10–14
Different studies conducted in the various areas of the world showed that there is a high burden of gram-negative resistance strains of bacteria to available antibiotics which compromise the success of various surgical procedures, cancer chemotherapy, and the treatment of many infections.15–18 In addition, due to the presence of the outer layer envelope, gram-negative bacteria are more resistant to antibiotics than gram-positive bacteria. 19 Currently, there is one published review on the patterns of antibiotic resistance to gram-negative bacteria in Ethiopia. 8 However, the study was conducted regardless of age classification and samples only from wound infection. Therefore, this systematic review aimed to synergize the previous work by determining the resistance patterns of gram-negative bacteria among pediatrics patients isolated from different clinical specimens in the country.
Objective
This systematic review aimed to determine gram-negative bacteria isolates and their antibiotic-resistance patterns among pediatrics patients in Ethiopia.
Method
Search strategy
This systematic review was conducted in compliance with Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA, 2009)
2
(Supplemental file 3). The comprehensive search for studies was done by two (BK and WY) of the authors from August to September 16, 2021. PubMed, EMBASE, Science Direct, the Cochrane Database for Systematic Reviews, Scopus, Hinari, Sci-Hub, African Journals Online Library, and free-text web searches using Google Scholar were searched for articles. First, articles were searched by examining the full title (“gram-negative bacteria isolates and their antibiotic-resistance patterns among pediatrics patients in Ethiopia”) and then keywords (bacterial infection, bacteria prevalence, drug resistance, antibiotic resistance, antimicrobial susceptibility, pediatric patients, Ethiopia). The specific name of each common gram-negative bacteria including
Study selection criteria
To minimize selection bias, all possible relevant articles were evaluated critically and that meet the inclusion criteria were selected. All available studies and data were incorporated based on the following predefined eligibility criteria.
Inclusion criteria
Exclusion criteria
We excluded studies limited to tuberculosis infections. Furthermore, reviews and systematic review articles, case reports, case series, and articles which were only available in abstract form were excluded. If multiple publications were reported with the same authors and findings; only the most recent or most complete publication for each data set for a specific outcome was selected. In addition, Intermediate susceptible strains were excluded from this study.
Data extraction
Essential data were extracted from eligible studies using Microsoft Excel spreadsheet format. Data were extracted by two of the authors (BK and WY) independently to enhance the quality and methodological validity. A standard extraction format adapted from the Joanna Briggs Institute (JBI) data extraction form was used to extract data. 20 The following information was identified for data extraction: The last name of the first author, year of publication, the region of the study conducted, study design and period, age of patients (years), site of clinical specimen obtained, total sample size, number of cultures positive sample, the number of gram-negative bacteria species isolated and antibiotic-resistance pattern. Any inconsistencies in the data extraction process were decided through discussion involving all authors.
Outcome measure
The main outcome of this systematic review was the patterns of antibiotic resistance for isolated gram-negative bacteria. The second outcome was the prevalence of isolates gram-negative bacteria.
Article quality assessment
The quality assessment was accompanied by two reviewers independently according to the critical appraisal checklist recommended by the JBI (20). Moreover, the disagreements were resolved by consensus and decided by taking the average score of the two reviewers. The JBI checklist was composed of 10 questions, the scores ranged from 0 to 10. The studies which obtained more than 60% were considered as good-quality studies. All of them had good quality and were included in the present systematic review.
Statistical analysis
The data were extracted using Microsoft Excel 2010 format and exported to STATA version 14.0 for further analysis. Frequency and percentages were computed using descriptive statics. Percent (%) of culture-positive = number of culture-positive divide by sample size from each specimen (N/n) ×100; percent (%) bacterial isolate = number of isolates bacteria divide by number of culture-positive (n) ×100. The extracted data were standardized to a prevalence of resistance, defined as the percentage of isolates being resistant out of the total number of isolates tested for the specific drug. Meta-analysis was not conducted because of the large variability in AMR methodology, geographical region, and a specimen was collected from a different site. A meta-analysis within each of the geographical regions was considered but rejected based on having too few studies for most of the pathogen–antimicrobial drug combinations per region. Since the number of studies from regions was small, they were combined and a percentage of resistance was generated.
Result
Characteristics of studies included in the analysis
In total, 2,684 articles were identified. After articles were removed by duplications, title, and reading the abstract, 114 studies were assessed for eligibility criteria. Consequently, 95 articles were excluded due to different reasons because they were irrelevant. Finally, a total of 19 studies met the inclusion criteria and were included in the final analysis (Figure 1). The articles were published between 2013 and 2021. Among the selected studies, two articles were unpublished (accepted article) which were obtained from the Addis Ababa University repository. Most of the studies (15/19, 78.95%) were cross-sectional studies, others were longitudinal or retrospective observational studies. Around one-third of the studies were conducted in Addis Ababa city (i = 7) followed by Amhara region (

Flow diagram of literature search and study selection.
Bacterial isolated
Overall, 1372 (16.77%) culture-positive samples were analyzed in the selected studies. Of these, 210/362 (58.01%), 153/300 (51.00%), and 643/2268 (28.41%) clinical specimens were taken from ear discharge, throat swab, and blood, respectively. The smallest culture-positive samples were obtained from the cerebrospinal fluid (CSF) analysis (177/4050, 4.37%). Of the total culture-positive sample, 735 were gram-negative (735/1372, 53.57%). Among gram-negative bacterial species,
Prevalence of isolated gram-negative bacterial.

CSF = Cerebrospinal fluid; K.P =Klebsiela pneumoniae; P.A =
Antibiotic resistance for gram-negative bacteria
More than 66.67% of isolates were resistant to ampicillin except
Resistance patterns of isolated organisms to all tested antibiotics.
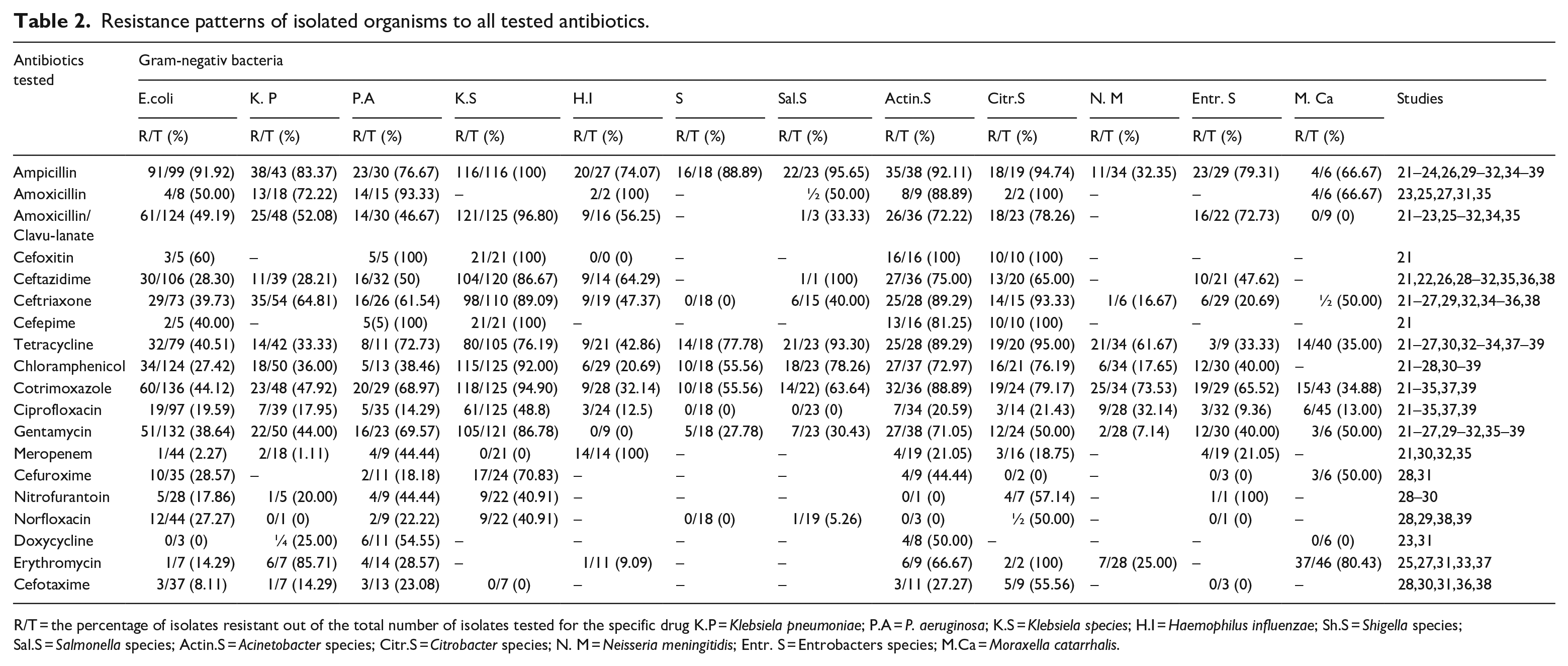
R/T = the percentage of isolates resistant out of the total number of isolates tested for the specific drug K.P =
Discussion
The overall prevalence of gram-negative bacteria isolated from various clinical samples (blood, urine, ear discharge, throat swab, CSF, and stool) from pediatrics patients was 53.75%. This is similar in studies from China, where 59.8% of the isolates were gram-negative bacteria,
40
and Pakistan, 55.7%.
41
However, it was lower than studies from low- and middle-income countries (63.9%), in Egypt (65.3%), and at Kigali, Rwanda (68.3%).42–44 The sources and numbers of the clinical samples collected, type of infections, types of patients, and geographical differences might be the cause of variation in the overall prevalence of gram-negative bacteria. The most frequent isolated gram-negative bacteria were
Our finding showed that the proportion of antibiotic resistance to gram-negative was ranging from 0% to 100%. In general, a high resistance rate was observed to the commonly prescribed antimicrobials. Except,
Tetracyclines, chloramphenicol, and cotrimoxazole are commonly used medications to treat
The limitations of this review; this systematic review provides useful information about the current status of antibiotic resistance among isolated bacteria among pediatrics patients. Besides, the review provides insight into the magnitude of the problem at the national level to help policymakers to design effective infection control programs and antibiotics use strategies to combat the antibiotics resistance burden in Ethiopia. The limitations of this review include there may be data not accessible by our search and therefore larger antibiotics resistance trends might have been missed. In addition, in Ethiopia antibiogram documentation habit is so poor, we may miss data related to drug resistance. Resistance data were obtained from different specimens, laboratory methodologies, disk diffusion methods, and different clinical and laboratory standard guidelines; these may have an impact on the variation in bacterial isolates and antibiotic resistance. Therefore, resistance rates shown in this review should be interpreted with caution due to the variable nature of the methodologies used for susceptibility testing in the publications that were combined to calculate the rates shown.
Conclusion
The current systematic review indicated that gram-negative bacteria were identified as the common pathogen causing infection in pediatrics patients.
The rate of AMR to gram-negative bacteria was significantly higher in Ethiopia.
Supplemental Material
sj-doc-3-smo-10.1177_20503121221094191 – Supplemental material for Gram-negative bacteria isolates and their antibiotic-resistance patterns among pediatrics patients in Ethiopia: A systematic review
Supplemental material, sj-doc-3-smo-10.1177_20503121221094191 for Gram-negative bacteria isolates and their antibiotic-resistance patterns among pediatrics patients in Ethiopia: A systematic review by Bekalu Kebede, Wubetu Yihunie, Dehnnet Abebe, Bantayehu Addis Tegegne and Anteneh Belayneh in SAGE Open Medicine
Supplemental Material
sj-xlsx-1-smo-10.1177_20503121221094191 – Supplemental material for Gram-negative bacteria isolates and their antibiotic-resistance patterns among pediatrics patients in Ethiopia: A systematic review
Supplemental material, sj-xlsx-1-smo-10.1177_20503121221094191 for Gram-negative bacteria isolates and their antibiotic-resistance patterns among pediatrics patients in Ethiopia: A systematic review by Bekalu Kebede, Wubetu Yihunie, Dehnnet Abebe, Bantayehu Addis Tegegne and Anteneh Belayneh in SAGE Open Medicine
Supplemental Material
sj-xlsx-2-smo-10.1177_20503121221094191 – Supplemental material for Gram-negative bacteria isolates and their antibiotic-resistance patterns among pediatrics patients in Ethiopia: A systematic review
Supplemental material, sj-xlsx-2-smo-10.1177_20503121221094191 for Gram-negative bacteria isolates and their antibiotic-resistance patterns among pediatrics patients in Ethiopia: A systematic review by Bekalu Kebede, Wubetu Yihunie, Dehnnet Abebe, Bantayehu Addis Tegegne and Anteneh Belayneh in SAGE Open Medicine
Footnotes
Acknowledgements
The authors thank the Debre Markos University Information Technology Directorate.
Author contributions
B.K. and W.Y. participated in data extraction, the database search, screening, and quality assessment: B.K., W.Y., D.A., B.A., and A.B. contributed to data analysis, drafting, or revising the article. All authors read and approved the final article.
Availability of data and materials
All relevant data are within the article and its supporting information files.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
