Abstract
Background:
Metagenomic next-generation sequencing (mNGS) is a novel diagnostic tool increasingly used in the field of infectious diseases. Little guidance is available regarding its appropriate use in different patient populations and clinical syndromes. We aimed to review the clinical utility of mNGS in patients with a specific clinical syndrome and identify factors that may increase its utility.
Methods:
We retrospectively reviewed charts of 72 non-immunocompromised adults hospitalized with the clinical syndrome of ‘fever of undetermined origin’ and underwent mNGS testing. Standardized criteria from a previously published study were used to determine the clinical impact of mNGS testing. We applied logistic regression to identify factors associated with a positive clinical impact.
Results:
Of the 72 patients identified, 62.5% were males with a median age of 56. All patients had a fever at the time of evaluation. At least one organism was identified in 65.3% of cases; most commonly were Epstein–Barr virus (13.9%), cytomegalovirus (12.5%), and Rickettsia typhi (11.1%). Of those determined to have an infectious etiology of their febrile syndrome, 89.5% (n = 34/38) had a positive mNGS. Consistency between the organism(s) on mNGS and the clinically determined infectious etiology was 82.4%. mNGS had a positive clinical impact in 40.3% of cases, a negative impact in 2.8%, and no impact in 56.9% of cases. Besides age, we did not identify other factors associated with a higher likelihood of positive clinical impact.
Conclusion:
In our review, mNGS had a positive clinical impact in a large proportion of adults with fever of undetermined origin, with minimal negative impact. However, mNGS results should be interpreted carefully given the high rate of detection of pathogens of unclear clinical significance. Randomized clinical trials are needed to assess the clinical utility of this novel diagnostic tool.
Plain language summary
In this study, we evaluated the use of a new diagnostic tool, namely, metagenomic next generation sequencing (mNGS), in hospitalized, non-immunocompromised, adult patients with a fever that was otherwise unexplained. We reviewed the clinical utility of this tool in 72 patients and found that at least one organism was found in 65.3% of cases, with the 2 most common organisms being viruses. In patients who were found to have an infection as the cause of their fever, 89.5% had a positive mNGS study. In 82.4% of cases, the infectious organism(s) found on mNGS was the organism thought to be the cause of the fever. Based on definitions from another study, mNGS had a positive clinical impact in 40.3% of cases, a negative impact in 2.8%, and no impact in 56.9% of cases. This study suggests that mNGS has minimal negative impact and can be a useful tool in identifying a causative infectious organism in patients with unexplained fevers. Additional studies are needed to identify patients and clinical conditions that would most benefit from this tool.
Keywords
Introduction
Metagenomic next-generation sequencing (mNGS) is a relatively novel diagnostic tool that analyzes cell-free genetic material from a given sample without the need for pre-determined sequences. 1 Several reference libraries exist for these mNGS tests, the oldest of which is the Karius Test™ (Karius, Redwood City, CA, USA), which has a database of around 1250 organisms. mNGS first became commercially available around 2005 2 and has been increasingly employed in the field of infectious diseases (ID) since then, albeit with some reservations. 3 Its appeal comes from its ability to circumvent issues that are commonly faced when using traditional microbiologic testing. For instance, traditional methods fail to detect organisms in up to 60% of cases. 4 Organism identification through cultures can be time-consuming, taking days or even weeks for mycobacterial infections, and may necessitate invasive procedures like tissue biopsies in cases of local or deep-seated infections. Despite the promising potential of this tool, limitations include incomplete reference databases, particularly for rare pathogens; incomplete ability to differentiate between genetically similar pathogens 4 ; a high rate of detectability that may limit the interpretative utility of this tool 1 ; and limited ability to capture antimicrobial susceptibility data from these molecular tests. 5 Given its novelty and minimal prospective data, there is little guidance on the efficient and appropriate use of this tool in the clinical setting.
Several small-scale studies evaluating the utility of mNGS are emerging, including those conducted in specific patient populations (e.g. persons living with HIV), 6 and specific clinical syndromes (e.g. community-acquired pneumonia, central nervous system infections, and sepsis7–9), with varying results.
Fever is a commonly encountered symptom in emergency rooms and often leads to hospitalization, requiring a wide range of laboratory tests and consultations with specialists. Studies show that up to 51% of cases classified as fever of unknown origin (FUO) remain undiagnosed, and among the diagnosed cases, up to 38% are attributable to infectious causes. 10 To date, limited data are available regarding the use of mNGS for evaluating FUO, mostly consisting of case reports and case series. 11 While ‘fever of undetermined origin’ lacks a precise definition in the literature, it shares the same ICD10 code as ‘FUO’. It is typically employed in the inpatient setting for cases where the cause of fever is unclear, which aligns with the design of our study.
The objective of our study is to assess the clinical utility of mNGS in patients hospitalized with a ‘fever of undetermined origin’. We used previously published standardized criteria developed by Hogan et al. 12 to evaluate clinical impact.
Methods
This retrospective study was conducted at an academic, 881-bed institution in the Medical Center in Houston, TX. We identified patients with ‘fever of undetermined origin’ and mNGS testing at any time between 2017 up to and including the year 2021. Fever of undetermined origin was defined as at least two documented fevers (>38.3°C), 4 h apart without apparent source based on a careful history, physical examination, and initial diagnostic fever work-up, including laboratory and imaging. To ensure homogeneity within our study population, we excluded patients who had underlying immunocompromising conditions such as malignancy, HIV infection, neutropenia, or those who were undergoing transplantation or receiving immunosuppressive therapy. After Institutional Review Board (IRB) approval, we identified all hospitalized adults with a mNGS test performed at our institution between June 2017 and July 2021. The assay we used was the Karius Test™, a College of American Pathologists-accredited NGS laboratory. At our institution, test ordering is limited to ID specialists.
Based on a review of the literature, we evaluated some predetermined factors that may influence the clinical impact of mNGS. Those factors included antibiotic use prior to mNGS testing, age, medical comorbidities, pathogen type on mNGS, and the number of positive pathogens reported. We collected patients’ basic demographics and admission data, including microbiologic tests, lab work-up, imaging, treatment, clinical outcome, and follow-up. Two independent reviews (AMH and RI) of each case determined the clinical impact of mNGS using the previously published criteria by Hogan et al., 12 shown in Figure 1.

Standardized criteria for clinical impact of mNGS based on Hogan et al. 12 ’s study.
Our primary outcome was to determine the clinical impact of mNGS in our selected patient population. In addition, we aimed to examine the level of agreement between mNGS results and the clinically determined cause of fever; concordance between mNGS and traditional microbiologic findings; and to identify factors associated with a positive clinical impact. Statistical analysis was performed using Stata version 16 (StataCorp LLC, College Station, TX, USA). Patient characteristics were summarized by median with minimum and maximum value or frequency with percentage. The median was used instead of the mean due to a non-normally distributed patient population. Fisher’s exact test was used to determine whether the mNGS result was associated with the final etiology. Kappa statistics were also used to measure agreement with mNGS results. Logistic regression was used to determine whether any factors were associated with mNGS having a positive clinical impact (as opposed to a negative or no clinical impact). Regression models were also used to estimate the odds ratio (OR) with a 95% confidence interval (CI) for having a positive clinical impact. p Values < 0.05 were considered statistically significant.
The reporting of this study conforms to the Strengthening the Reporting of Observational Studies in Epidemiology guidelines (Supplemental Material).
Results
We identified 72 patients that met the inclusion criteria. In total, 62.5% were males, with a median age of 56 (range, 19–84), as seen in Table 1. The most common comorbidities were hypertension (43.1%) and diabetes mellitus (23.6%). The median length of hospital stay was 12 days (range, 0–201), and the median time from fever onset to ID consult was 1 day (range, 0–18). All patients had a fever at the time of evaluation. Other reported symptoms were cough (26.4%), abdominal pain (16.7%), and shortness of breath (15.3%). In all, 22 (30.6%) patients received antibiotics the week before mNGS testing. The median turnaround time of the test from the time of collection to the time of reporting was 26 h (range, 22–173). At least one organism was reported in 47 (65.3%) cases. The most commonly identified organisms were Epstein–Barr virus (EBV) (N = 10; 13.9%), cytomegalovirus (CMV) (9; 12.5%), and Rickettsia typhi (8; 11.1%) (Table 2). The median MPM (molecules per microliter) was 135 for EBV (range, 29–418) and 142 for CMV (range, 35–105,221). The most commonly identified bacteria were R. typhi (N = 8; 11.1%), Enterococcus faecalis (N = 6; 8.3%), and Klebsiella oxytoca (N = 4; 5.6%).
Patient demographics and admission data (N = 72).

Organisms identified on mNGS.
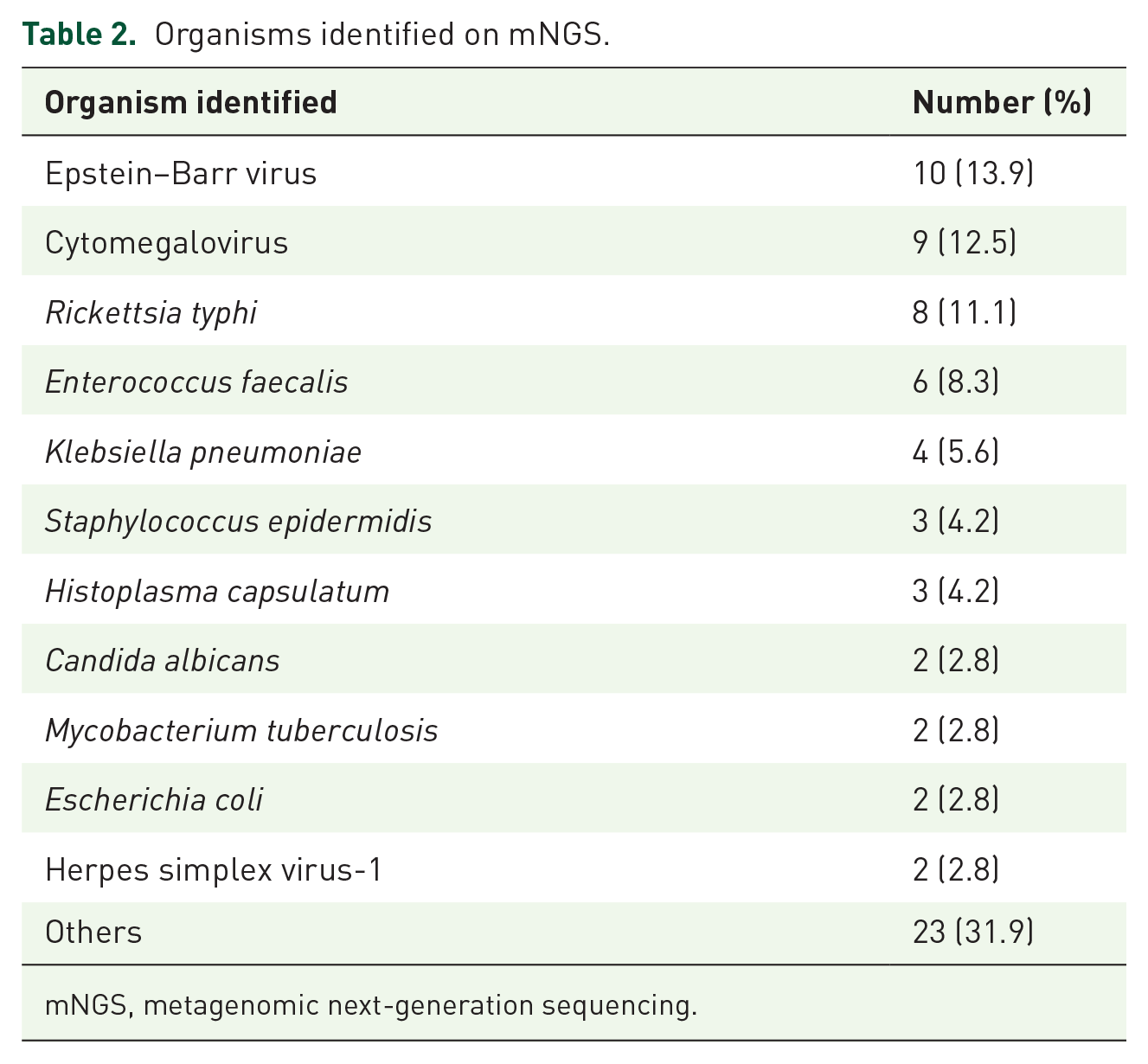
mNGS, metagenomic next-generation sequencing.
Of those determined to have a clinical infectious etiology of their fever (N = 38), 89.5% had a positive mNGS compared to 43.8% of those with non-infectious etiology (N = 16) (p = 0.002). A manual review of cases was done to evaluate the consistency between the organism(s) identified on mNGS and the clinically determined infectious etiology of fever. Based on this review, the consistency was 82.4% (28/34). In total, 14.7% (5/34) of patients had organism(s) on mNGS that were inconsistent with the final infectious diagnosis, and 1 patient did not have a clear etiology for his fever. mNGS had a positive clinical impact in 40.3% of cases (N = 29), a negative impact in 2.8% of cases (N = 2), and no impact in 56.9% of patients (N = 41) (Table 3).
Clinical impact of mNGS testing.

mNGS, metagenomic next-generation sequencing.
Among those with a positive clinical impact (N = 29), 12 (41.4%) were due to new diagnoses based on mNGS not confirmed by traditional microbiologic methods (Table 3). Among cases with a negative clinical impact, one was due to unnecessary diagnostic investigations due to mNGS results, and one was due to a longer hospital stay. Among those with no clinical impact (N = 41), the major reason was negative mNGS and not acted upon (N = 18; 43.9%), followed by mNGS identifying a new organism that was not acted upon (N = 15; 36.5%), and mNGS confirming the traditional microbiologic data and not acted upon (N = 8; 19.5%).
Based on logistic regression analysis, older age was the only factor associated with a higher odd of positive impact (10-year increase in age: OR: 1.37; 95% CI: 1.001–1.86; p = 0.049). Chief complaints (abdominal pain, cough, diaphoresis, diarrhea, headache, myalgias, sepsis, shortness of breath, other), the positivity of cultures, number of pathogens retrieved, and type of prior antibiotic use were not associated with a higher rate of positive clinical impact (Supplemental Table 2).
Discussion
We provide a case series describing the clinical utility of mNGS in hospitalized non-immunocompromised adult patients with fever of undetermined origin, using the standardized criteria for clinical impact outlined by Hogan et al. 12 In our study, at least one organism was identified in 65% of cases. Prior studies reporting specific clinical syndromes described similar rates ranging from 53% to 70%.9,12,13
Studies on the clinical impact of mNGS vary significantly, ranging from 7% to 56%. 14 In our study, mNGS had a positive clinical impact in 40.3% of cases, a negative impact in 2.8%, and no impact in 56.9% of cases. Compared to our study, the positive clinical impact rate in Hogan et al. 12 ’s study was only 7.3%, while most cases (86.6%) were determined to have no clinical impact. Another study using Hogan et al.’s standardized criteria assessed 80 patients without pre-defined clinical syndromes and found the rate of positive clinical impact to be 43%, with the majority of cases (55%) having no clinical impact. 15 Among those with a FUO, 50% had a positive clinical impact and 50% had no/uncertain clinical implications. Another common finding in several studies is that the positive clinical impact is primarily driven by the ability to de-escalate antimicrobial therapy when mNGS is negative.12,16,17 This reflects the importance of having a careful evaluation of the results by ID specialists to maximize the benefit from a stewardship standpoint and avoid unnecessary antimicrobial treatment.
Although rates of positive and neutral clinical impacts may vary between studies, there is a significantly smaller degree of negative clinical impacts in the aforementioned studies and our own. In our cases, the negative impact was due to additional diagnostic investigations being pursued and longer hospital lengths of stay. Both issues can contribute to increased healthcare costs as well as potential emotional distress to the patient and family. However, our study is consistent with others that highlight the significantly lower rates of negative clinical impacts of mNGS testing. Based on comparisons of our study, which focused on a particular clinical syndrome, and others that had more heterogeneous populations, it is likely that clinical impact varies based on patient population and specific clinical syndrome studied. We also believe that the rates of positive impact in our study were influenced by the fact that mNGS ordering is limited to ID specialists.
In our study, we show that mNGS has possible implications when used in cases with high suspicion of infection as the etiology of fever. The rate of a positive mNGS test was significantly higher when the etiology was infectious than when it was non-infectious. A similar pattern was noticed in Rossoff et al., 13 where mNGS was positive in 56% of cases that were infectious in etiology and only 14% of patients that were non-infectious in etiology. When comparing mNGS results and traditional microbiologic results, the literature reports a wide range of consistency, ranging from 26% to 100%. 14 This wide range is likely due to the heterogeneity of the studied patient populations, study designs, and clinical syndromes included in the different studies. In Rossoff et al. 13 ’s study, the sensitivity and specificity of mNGS for an infection were compared to conventional microbiological testing. Although sensitivity was higher for mNGS versus conventional testing (92% versus 72%, respectively; p < 0.01), specificity was lower (64% versus 89%, respectively; p < 0.01). The issue of specificity was addressed by one study on pediatric patients, where conventional testing was compared to mNGS. 18 They found that when a new organism was identified on mNGS that was not identified by conventional testing, antibiotic changes were made in 26% of cases, raising concerns for this tool’s potential misuse and the importance of clinical interpretation of the results. Literature shows that up to one-third of mNGS testing will result in two or more organisms.12,13,18,19 Thus, in addressing the limitations of this tool, it is critical to acknowledge its high detectability rates and its potential.
To date, factors influencing the utility of mNGS have not been fully and clearly delineated. In our study, we were unable to identify factors associated with higher rates of positive clinical impact other than older age. Chief complaint, type of antibiotic used, and comorbidities did not have an association with positive clinical effects. In Shishido et al. 15 ’s study, several factors were associated with the clinical utility of mNGS, including a clinical syndrome of sepsis, a specific patient population (solid organ transplant recipients), and fewer days of antibiotics before testing (<7 days). In Rossoff et al.’s study, mNGS had higher rates of positive clinical impact in immunocompromised patients than immunocompetent ones. In that study, mNGS testing had a higher yield than conventional methods in cases where invasive methods (e.g. biopsy) were required. 13
Our study was limited by its retrospective nature and smaller sample size. We did not perform a sample size calculation for power analysis. Furthermore, we were not able to investigate the ‘MPM’ titer and its possible impact on the significance of mNGS because there is no agreed-upon published data on what would be considered a high positive titer in the literature and the cutoff for positive reporting is different for each pathogen. In addition, the turnaround time of different microbiology tests differs according to each laboratory’s capacity and send-out procedures; thus, the utility of this tool may be influenced by the turnaround time at different institutions. Furthermore, the decision on the clinical impact of mNGS testing is not always clear and depends on the subjective review of patients’ charts and physicians’ documentation.
Conclusion
In this study, we showed that mNGS had a positive clinical impact in a large proportion of hospitalized non-immunocompromised adult patients with fever of undetermined origin, with minimal negative impact. Besides age, we did not identify other factors associated with a higher likelihood of positive clinical impact. mNGS may be a valuable tool in many cases with a potential benefit from a stewardship perspective, but we believe the test should be carefully utilized with the assistance of an ID specialist to avoid oversensitivity and treatment of unnecessary positive ‘background noise’ organisms. Randomized clinical trials are needed to delineate specific clinical syndromes, patient populations, and other factors that increase the likelihood of a positive clinical impact of mNGS testing and to guide clinicians toward the most efficient use of this novel tool.
Supplemental Material
sj-docx-1-tai-10.1177_20499361241244969 – Supplemental material for Clinical utility of metagenomic next-generation sequencing in fever of undetermined origin
Supplemental material, sj-docx-1-tai-10.1177_20499361241244969 for Clinical utility of metagenomic next-generation sequencing in fever of undetermined origin by Marilyne Daher, Roumen Iordanov, Mayar Al Mohajer, M. Rizwan Sohail, Kristen Andrews Staggers and Ahmed Mufeed Hamdi in Therapeutic Advances in Infectious Disease
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
