Abstract
Background:
Diarrhea causes significant morbidity among people living with human immunodeficiency virus. Compared to conventional diagnostic tests, multiplex polymerase chain reaction gastrointestinal panels provide faster and more accurate pathogen detection.
Objectives:
This study aimed to characterize diarrheal pathogens in people living with human immunodeficiency virus using multiplex gastrointestinal polymerase chain reaction, evaluate their impact on antibiotic use, and synthesize global evidence through a systematic review.
Design:
We conducted a retrospective, cross-sectional study at a hospital in the Caribbean region of Colombia from 2020 to 2024. Concurrently, a systematic review was performed following Preferred Reporting Items for Systematic Reviews and Meta-Analyses guidelines.
Methods:
Adult people living with human immunodeficiency virus hospitalized with diarrhea who underwent testing using a multiplex gastrointestinal polymerase chain reaction panel were included in the cross-sectional study. A systematic review was conducted by searching major electronic databases without geographic or language restrictions. Clinical, microbiological, and treatment-related data were extracted and analyzed.
Results:
The retrospective study included 49 people living with human immunodeficiency virus hospitalized with diarrhea, while the systematic review analyzed 679 cases across 8 studies. Compared to conventional methods, multiplex polymerase chain reaction panels showed higher overall pathogen detection rates. At least one pathogen was detected in 79.5% of cases in the retrospective study. Bacterial pathogens were the most frequent in both the retrospective (70.1%) and the systematic review studies (58%–80%), with diarrheagenic
Conclusion:
Multiplex gastrointestinal polymerase chain reaction panels improve pathogen detection and help guide antimicrobial decision-making in people living with human immunodeficiency virus with diarrhea. These findings support the use of molecular diagnostics to enhance clinical management and epidemiological surveillance in this population.
Registration:
PROSPERO (CRD42025644820).
Keywords
Introduction
Human immunodeficiency virus (HIV) infection remains a major global public health concern, associated with significant morbidity, mortality, and economic impact. Although antiretroviral therapy (ART) has markedly improved survival and quality of life and reduced the incidence of opportunistic infections in people living with HIV (PLWH), diarrhea continues to pose a frequent and debilitating complication in this population. 1 Diarrhea affects more than 50% of PLWH and may be caused by opportunistic pathogens specific to immunocompromised hosts, as well as organisms commonly affecting the general population. 2 Identifying the causative agent of diarrhea in PLWH is often difficult due to the limited diagnostic sensitivity performance of conventional methods. 3 Consequently, a considerable proportion of cases (30%–60%) remain without a definitive etiological diagnosis. 4
The rise of antimicrobial resistance among diarrheal pathogens has become a major public health problem. Growing resistance to commonly used antibiotics, such as fluoroquinolones and third-generation cephalosporins, has progressively reduced effective treatment options in both pediatric and adult populations. 5 Current clinical guidelines recommend the use of antibiotics in immunosuppressed patients with diarrhea, especially in cases of severe or invasive infection, to improve outcomes and reduce complications.6,7 Multiplex gastrointestinal polymerase chain reaction (PCR) panels offer a rapid, sensitive, and comprehensive method for pathogen detection infectious diarrhea. Their use has been associated with improved clinical outcomes, including reduced hospital length of stay, more targeted antimicrobial therapy, and increased rates of discharge without antibiotics.8,9 However, to date, information regarding the implementation and performance of multiplex gastrointestinal PCR panels in PLWH is scarce. Moreover, no systematic reviews have synthesized the available evidence on the use of these molecular diagnosis tests in PLWH.
The aim of this study was to characterize diarrheal pathogens in PLWH using multiplex gastrointestinal PCR panels at a tertiary care center in Colombia, evaluate their impact on antibiotic use, and synthesize global evidence through a systematic review.
Methods
Cross-sectional study
The observational, descriptive, retrospective cross-sectional study was conducted at Hospital Universidad del Norte, a tertiary care center located in the Caribbean region of Colombia. The inclusion criteria included all PLHIV aged 18 years or older who were hospitalized with diarrhea and underwent multiplex gastrointestinal PCR testing between 2020 and 2024. No specific exclusion criteria were applied, as the study aimed to include all consecutive patients who met the predefined clinical and diagnostic criteria during the study period. Diarrhea was defined as the occurrence of ⩾3 loose or watery stools in a 24-h period. The diagnosis of HIV infection was made according to the national diagnostic algorithm and guidelines established by the Colombian Ministry of Health. The study was approved by the Research Committee of Hospital Universidad del Norte and the Ethics Committee of Universidad del Norte (No. 325/2024). Due to the retrospective design, which involved the review of electronic medical records and the use of anonymized clinical data, the requirement for written informed consent was waived by the Ethics Committee.
Clinical and laboratory data were extracted from electronic medical records. Variables collected included pathogen identification based on multiplex PCR panel results, sociodemographic data (age and sex), immunological status (CD4+ T-cell count and HIV viral load), clinical features of diarrhea (duration and consistency), antibiotic use before and after testing, length of hospital stay, intensive care unit (ICU) admission, and in-hospital mortality. Data accuracy was verified by cross-referencing medical and laboratory sources.
Stool samples were analyzed using the FilmArray® Gastrointestinal Panel (BioFire Diagnostics, Salt Lake City, Utah), following the manufacturer’s instructions. In parallel, samples were also processed using conventional diagnostic methods, including modified Ziehl–Neelsen staining for the detection of intestinal coccidian oocysts (e.g.,
Statistical analysis
Data were analyzed using RStudio (Posit Software, PBC, Boston, MA, USA). Descriptive statistics were used to summarize patient characteristics. Continuous variables were reported as medians with interquartile ranges (IQR) and categorical variables as frequencies and percentages. Patients with missing data were excluded from analyses involving the corresponding variables. For continuous variables with values below the limit of detection (LOD), observations were excluded from analysis when fewer than 50% of the values fell below the LOD threshold. 10
Systematic review
A systematic review was conducted in accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines. 11 We included studies reporting pathogens causing diarrhea identified through multiplex gastrointestinal PCR panels in PLWH. The review protocol was registered in PROSPERO (CRD42025644820). Literature research was performed in MEDLINE via PubMed, EMBASE, LILACS via Virtual Health Library, and Google Scholar. Additional sources included ClinicalTrials.gov, International Clinical Trials Registry Platform, Cochrane Central Register of Controlled Trials, The Kings Fund (England), The New York Academy of Medicine (United States), TESEO (Spain) and LAReferencia (Latin America). Full search strategies and database-specific URLs are provided in the Supplemental Table 1.
We included studies published in English, Spanish, or Portuguese. Eligible study designs comprised case reports, randomized controlled trials, case–control studies, and cohort studies. No restrictions were applied regarding publication date, participant age, or geographic location. Studies were excluded if they did not report the diagnostic method used, did not confirm HIV status, or did not involve participants with diarrhea. Studies using molecular tests limited to a single pathogen or a restricted group of pathogens (e.g., only bacteria, viruses, or parasites) were also excluded. Review articles and publications without original data were not eligible. All records were managed in Rayyan software, where screening decisions were documented.
Titles and abstracts were first screened for eligibility, followed by full-text review, both conducted independently by two reviewers. Disagreements at any stage were resolved by consultation with a third reviewer. When necessary, corresponding authors were contacted to obtain missing information. Data extraction and synthesis were performed from the final set of included studies. Extracted variables included sample size, patient age and sex, country of study or case, pathogens identified, CD4+ T-cell count, ART status, antibiotic treatment, and clinical outcomes. Information was synthesized, and proportion and distribution patterns were calculated across studies. The quality and risk of bias of included studies were assessed according to Murad et al. criteria for case series and case reports, 12 The Risk Of Bias in Nonrandomized Studies (ROBINS-I) for nonrandomized studies, 13 and the Cochrane risk of bias tool for randomized controlled trials. 14
Results
Cross-sectional study
This retrospective cross-sectional study included 49 hospitalized PLWH with diarrhea (Supplemental Figure 1). The median age was 37 years (IQR 33–47), and 77.6% were male. Diarrheal symptoms had lasted a median of 7 days before admission. Most patients (79.6%) were aware of their HIV status, and 51% were on ART. The median CD4+ T cell count was 100 cells/µL (IQR 21–285), with 87.5% having counts below 200. Median HIV viral load was 65,919 copies/mL (IQR 172–240,000). Coinfections were identified in 63.3% of patients, with oral candidiasis, syphilis, and tuberculosis being the most prevalent. Comorbidities were present in 24.5% of patients, mainly chronic kidney disease, psychiatric disorders, and hypertension (Table 1).
Characteristics of 49 PLWH and diarrhea from a tertiary care center in Colombia.

HIV: human immunodeficiency virus; ICU: intensive care unit; PLWH: people living with human immunodeficiency virus; IQR: interquartile range.
Other coinfections include
Other comorbidities include urolithiasis, heart failure, hypothyroidism, and epilepsy (two cases each), asthma (one case).
Diarrheal pathogens identified
Multiplex gastrointestinal PCR panels detected at least one pathogen in 79.5% of cases. Bacteria were most frequently identified (70.1%), particularly enteropathogenic and enterotoxigenic
Pathogens identified by multiplex PCR in PLWH and diarrhea from the present study and the systematic review.

Data are presented as numbers (%). EAEC: enteroaggregative
Conventional diagnostic methods identified pathogens in only 13 cases. Blood cultures (
Antimicrobial therapy and outcomes
Antibiotic use and adjustments are described in Table 3. Empiric antimicrobial therapy was initiated in 39 patients (79.6%), most commonly with ceftriaxone plus metronidazole in 22 patients. Following multiplex PCR results, antimicrobial therapy was de-escalated in 17 cases (43.6%). In one case, empiric therapy was completely discontinued, while in 16 cases targeted de-escalation was performed by discontinuing a single agent in patients receiving combination therapy. Metronidazole was the most frequently withdrawn antibiotic, primarily due to the absence of parasitic pathogens. In contrast, new antimicrobial agents were started in 18 patients (36.7%), most often ceftriaxone or nitazoxanide.
Antibiotic use and adjustments in 49 PLWH and diarrhea from a tertiary care center in Colombia.

The percentages have been calculated from the total number of patients (
The median time to resolution of diarrheal symptoms after starting antibiotics was 3 days (IQR 2–7), and the median hospital stay was 9.5 days (IQR 5.75–17.25). Ten patients (18.4%) required ICU admission. There were three in-hospital deaths, only one of which was attributed to gastrointestinal sepsis.
Systematic review
From 2047 records identified, 8 studies met inclusion criteria and were analyzed in the systematic review.15–22 The PRISMA flow diagram is presented in Supplemental Figure 2, and study characteristics are summarized in Table 4. Risk of bias in the included studies was low (Supplemental Table 2). Most studies originated from North America18–20 and Asia,16,17,22 with additional reports from Latin America 15 and Africa 21 ; no studies were identified from Europe or Oceania. Study designs included four cross-sectional studies,15,16,19,21 three retrospective cohorts,17,18,20 and one case report. 22
Characteristics of studies evaluating PLWH and diarrhea included in the systematic review.
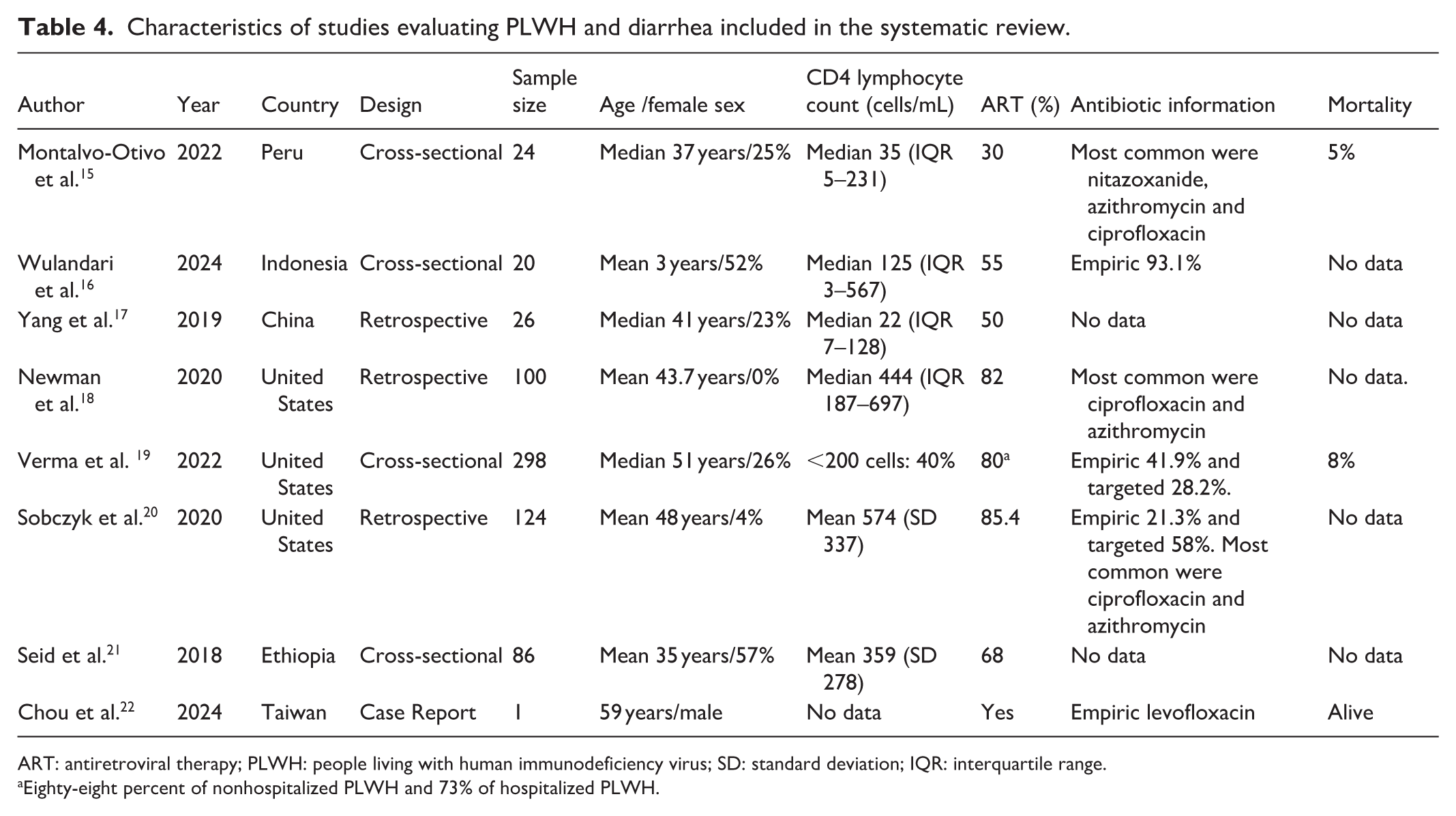
ART: antiretroviral therapy; PLWH: people living with human immunodeficiency virus; SD: standard deviation; IQR: interquartile range.
Eighty-eight percent of nonhospitalized PLWH and 73% of hospitalized PLWH.
A total of 679 PLWH with diarrhea were analyzed across the included studies, including 78 children (11.4%). The proportion of male patients ranged from 47% to 100%. Clinical settings were reported in four studies,16,18,19,22 with most patients hospitalized and a smaller number managed as outpatients. Diarrhea duration before consultation ranged from 2 to 20 days. Median CD4+ T cell counts ranged from 22 to 574 cells/mm3. HIV viral load data were available in three studies, with values ranging from <50 to 50,220 copies/mL.18–20 ART coverage varied between 30% and 88%. Opportunistic infections were reported in only one study, which documented syphilis in 54 of 268 patients. 18
Diarrheal pathogens identified
Multiplex gastrointestinal PCR platforms used in the included studies were primarily FilmArray (BioFire Diagnostics) in six studies15,17–20,22 and QIAGEN-based platforms in two.16,21 The diarrheagenic pathogens identified in the studies included in the systematic review are presented in Table 2. Bacterial pathogens were the most frequently detected, found in 58–80% of cases, with
One pediatric study found higher viral infection rates among PLWH (57.1%) compared to HIV-negative children, who had more frequent bacterial infections (70.6%).
16
Another study reported increased risk of infectious diarrhea due to
Multiplex PCR panels demonstrated higher detection rates than conventional methods. In one study, microscopy identified 28% of parasitic infections, mainly
Antimicrobial therapy and outcomes
Prior antibiotic use or recent hospitalization was reported in 20%–30% of patients (Table 2). Empiric treatment commonly included ciprofloxacin15,18,20,22 and azithromycin.16,18,20 Antimicrobial therapy was modified to target treatment based on multiplex PCR results in 28.2%–58% of cases,19,20 with durations typically ranging from 3 to 7 days.16,18,19
Clinical outcomes were variably reported. One study documented a 7.7% mortality rate. 19 Symptom resolution ranged from 30% 17 to 74%. 18 Severe immunosuppression (CD4+ <200 cells/μL) was associated with increased hospitalization and a lower likelihood of pathogen detection by PCR, although not with overall infection rates. 19 Early testing, within 24 h of admission, was linked to a higher probability of identifying a pathogen. 19
Discussion
This retrospective study, in conjunction with the systematic review, characterizes etiological pathogens, patterns of antimicrobial use, and clinical outcomes in PLWH who underwent multiplex gastrointestinal PCR panel testing for diarrhea. Most patients were adults, predominantly male, and with CD4+ T lymphocyte counts below 200 cells/µL. Bacterial pathogens were the most frequently identified, with
Studies using conventional stool testing in PLWH and diarrhea have reported different pathogen profiles compared to those identified by multiplex gastrointestinal PCR panels in our study and the accompanying systematic review. A large cohort study conducted in the United States between 1992 and 2002 assessed bacterial diarrhea in PLWH, identifying
Since their regulatory approval, multiplex gastrointestinal PCR panels have been widely adopted in clinical practice, due to their rapid turnaround time and broad pathogen detection capabilities.
26
While traditional stool cultures detect pathogens in approximately 6% of cases, multiplex gastrointestinal PCR panels can identify pathogens in 32% or 50% of cases.4,27 significantly improving diagnostic yield especially for organisms difficult to isolate using conventional methods.
26
Current guidelines recommend multiplex gastrointestinal PCR testing for patients with immunosuppression, severe or prolonged illness, or in suspected outbreaks, and as a complement to traditional diagnostic approaches.6,7 These assays typically demonstrate high specificity (>98%) and sensitivity (90%–100%).
3
However, lower sensitivity has been reported for pathogens such as
Beyond improving diagnostic accuracy, multiplex gastrointestinal PCR panels have demonstrated a measurable impact in clinical decision-making. In a retrospective analysis of over 9402 patients, those tested with a multiplex gastrointestinal PCR panel were 8.4% less likely to undergo endoscopy, 29.4% less likely to receive abdominal imaging, and 36.2% less likely to be prescribed antibiotics compared to those who underwent conventional stool testing. 32 In our study, empirical antibiotics were discontinued in 34% of patients following result availability, most often after the exclusion of parasitic infections. Similarly, two studies included in the present systematic review reported initiation of targeted therapy in 28.2% 19 and 58% 20 of cases. These findings suggest that multiplex gastrointestinal PCR testing facilitates more appropriate therapeutic decisions and contributes to a meaningful reduction in antimicrobials used in empirical therapy, supporting its role in antimicrobial stewardship programs.4,9 The implementation of this panel facilitates the mitigation of negative consequences associated with the prolonged use of empiric broad-spectrum regimens reducing unnecessary drug exposure, microbiota disruption, and the risk of antimicrobial resistance.
This study has several limitations. Given its retrospective design of the cross-sectional study, no formal sample size calculation was performed, and the relatively small cohort may have limited statistical power and generalizability. Importantly, we included all consecutive PLWH who met the predefined inclusion criteria at our institution during the 4-year study period. Moreover, the patient population included in our retrospective cross-sectional study may not be representative of other settings, as most patients had advanced immunosuppression, which likely influenced the high pathogen detection rate and the predominance of opportunistic organisms. Therefore, these findings may not be fully generalizable to PLWH with higher CD4+ counts or those managed in outpatient settings. Furthermore, its retrospective design required the use of medical records, which could have resulted in incomplete data for some variables. Among the strengths of the study is the inclusion of PLWH with documented diarrhea, increasing the likelihood that the identified pathogens were causative rather than colonizing. In addition, clinical outcome data were collected, including antibiotic use and patient prognosis. Regarding the systematic review, limitations include methodological heterogeneity among included studies, many of which were retrospective and cross-sectional in nature. Most of the included studies also lacked detailed reporting of patient characteristics and contextual factors that could influence the prevalence of diarrhea-causing pathogens. Nevertheless, all included studies were assessed as having a low risk of bias, which lends confidence to the overall findings.
Conclusions
This study synthesizes current evidence on the use of multiplex gastrointestinal PCR panels in PLWH presenting with diarrhea, enhancing our understanding of the epidemiology of diarrheal pathogens in this population. These assays not only increase diagnostic yield, particularly for difficult-to-culture organisms, but also inform clinical decision-making by enabling targeted antimicrobial therapy and reducing unnecessary interventions. The findings support the adoption of multiplex PCR panels as a valuable diagnostic tool to improve accuracy and guide antimicrobial management in PLWH. Importantly, in settings with limited access to multiplex PCR, the epidemiological patterns identified through its implementation may help guide locally adapted empiric treatment strategies and optimize resource utilization. Nonetheless, the number of studies evaluating these assays in this population remains limited, and data from several geographic regions are lacking. Further research is warranted, specifically prospective, controlled studies directly comparing a multiplex PCR-guided diagnostic pathway versus a conventional pathway. Such studies should focus on cost-effectiveness and clinically meaningful outcomes, including time to targeted therapy, complication rates, and mortality in PLWH presenting with diarrhea.
Supplemental Material
sj-docx-1-smo-10.1177_20503121261436154 – Supplemental material for Multiplex polymerase chain reaction for pathogen identification and antibiotic stewardship in human immunodeficiency virus-associated diarrhea: A retrospective study and systematic review
Supplemental material, sj-docx-1-smo-10.1177_20503121261436154 for Multiplex polymerase chain reaction for pathogen identification and antibiotic stewardship in human immunodeficiency virus-associated diarrhea: A retrospective study and systematic review by Santiago Gómez-Jordan, Samuel Jarava, Francisco Villadiego, Valentina Lara, David Saavedra-Segura, Vanessa Sarmiento and Diego Viasus in SAGE Open Medicine
Supplemental Material
sj-docx-2-smo-10.1177_20503121261436154 – Supplemental material for Multiplex polymerase chain reaction for pathogen identification and antibiotic stewardship in human immunodeficiency virus-associated diarrhea: A retrospective study and systematic review
Supplemental material, sj-docx-2-smo-10.1177_20503121261436154 for Multiplex polymerase chain reaction for pathogen identification and antibiotic stewardship in human immunodeficiency virus-associated diarrhea: A retrospective study and systematic review by Santiago Gómez-Jordan, Samuel Jarava, Francisco Villadiego, Valentina Lara, David Saavedra-Segura, Vanessa Sarmiento and Diego Viasus in SAGE Open Medicine
Supplemental Material
sj-docx-3-smo-10.1177_20503121261436154 – Supplemental material for Multiplex polymerase chain reaction for pathogen identification and antibiotic stewardship in human immunodeficiency virus-associated diarrhea: A retrospective study and systematic review
Supplemental material, sj-docx-3-smo-10.1177_20503121261436154 for Multiplex polymerase chain reaction for pathogen identification and antibiotic stewardship in human immunodeficiency virus-associated diarrhea: A retrospective study and systematic review by Santiago Gómez-Jordan, Samuel Jarava, Francisco Villadiego, Valentina Lara, David Saavedra-Segura, Vanessa Sarmiento and Diego Viasus in SAGE Open Medicine
Supplemental Material
sj-docx-4-smo-10.1177_20503121261436154 – Supplemental material for Multiplex polymerase chain reaction for pathogen identification and antibiotic stewardship in human immunodeficiency virus-associated diarrhea: A retrospective study and systematic review
Supplemental material, sj-docx-4-smo-10.1177_20503121261436154 for Multiplex polymerase chain reaction for pathogen identification and antibiotic stewardship in human immunodeficiency virus-associated diarrhea: A retrospective study and systematic review by Santiago Gómez-Jordan, Samuel Jarava, Francisco Villadiego, Valentina Lara, David Saavedra-Segura, Vanessa Sarmiento and Diego Viasus in SAGE Open Medicine
Supplemental Material
sj-docx-5-smo-10.1177_20503121261436154 – Supplemental material for Multiplex polymerase chain reaction for pathogen identification and antibiotic stewardship in human immunodeficiency virus-associated diarrhea: A retrospective study and systematic review
Supplemental material, sj-docx-5-smo-10.1177_20503121261436154 for Multiplex polymerase chain reaction for pathogen identification and antibiotic stewardship in human immunodeficiency virus-associated diarrhea: A retrospective study and systematic review by Santiago Gómez-Jordan, Samuel Jarava, Francisco Villadiego, Valentina Lara, David Saavedra-Segura, Vanessa Sarmiento and Diego Viasus in SAGE Open Medicine
Footnotes
Ethical approval
The study was approved by the Research Committee of Hospital Universidad del Norte and the Ethics Committee of Universidad del Norte (No. 325/2024).
Consent to participate
The requirement for written informed consent was waived by the Ethics Committee.
Consent for publication
All authors give consent for publication.
Author contributions
All of the authors make substantial contributions to the conception. All authors approve the final version, and all agree to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. SGJ, SJ: design, conception, acquisition of information, manuscript draft, analysis, review, editing. FV: design, acquisition of information, conception, editing. VL: acquisition of information, statistical analysis, manuscript draft. DS: design, acquisition of information, manuscript draft. VS: acquisition of information, review. DV: supervision, design, conception, manuscript draft, analysis, review, editing.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
All information is available on request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
