Abstract
The availability of whole-genome sequencing (WGS) as a tool for the diagnosis and clinical management of tuberculosis (TB) offers considerable promise in the fight against this stubborn epidemic. However, like other new technologies, the best application of WGS remains to be determined, for both conceptual and technical reasons. In this review, we consider the potential value of WGS in the clinical laboratory for the detection of
Introduction
Owing to advances in technology and reductions in cost, whole-genome sequencing (WGS) has been transformed from a centralized service used by a select few to interrogate single genomes into a relatively decentralized lab technique used by many to detect and track infectious pathogens [Long et al. 2014; Price et al. 2014; SenGupta et al. 2014; Snitkin et al. 2012; Quick et al. 2014, 2015]. This transformation has not spared the mycobacterial genus, with a number of papers presenting its application to the characterization of
A brief description of WGS
WGS begins at the bench, with the extraction and purification of genomic DNA. In very brief detail, this DNA is typically fragmented into shorter pieces, which are then sequenced in ‘reads’ of 100–500 base pairs (bps) for bench-top sequencers. There are a number of different sequencing platforms available [Loman et al. 2012a; Kwong et al. 2015; Heather and Chan, 2015]. The choice of platform depends largely on the question, which in turn is dictated by clinical needs. If the aim is to identify unknown organisms or to characterize a novel bacterium, one might prefer a sequencer that generates longer reads (such as the PacBio RS by Pacific Biosciences, Menlo Park, CA, USA), as such reads enable more accurate WGS workflow for 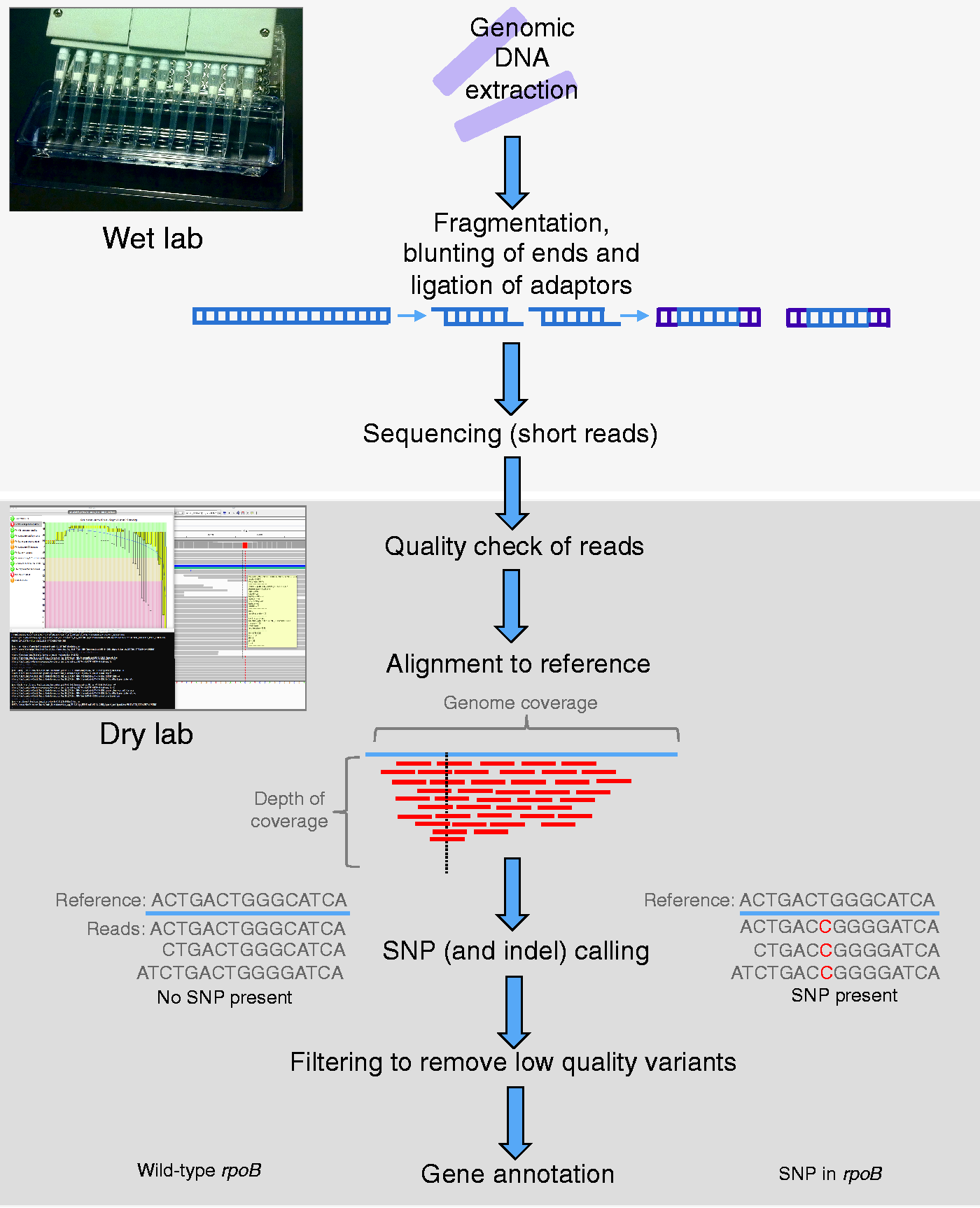
With the Illumina MiSeq platform, short reads of up to 300 bps in length are produced. To identify the microorganism in question based on these reads, a variety of tools can be utilized. The Basic Local Alignment Search Tool (BLAST [Altschul et al. 1990]) compares reads with existing microbial DNA databases and uses an algorithm to identify the most likely microorganism. Other methods include classifying the microorganism based on how well reads align to conserved coding sequences within phyla or species (‘clade-specific marker sequences’ [Segata et al. 2012]) or
Once reads have been assigned the identity ‘
Using a reference-based approach, single nucleotide polymorphisms (SNPs; i.e. a difference in a single base in the genome compared to the reference) and insertions/deletions (indels) present in the test isolate can be identified (‘called’) compared with the referent. This process, the quality control steps therein and the different tools used for identifying SNPs are reviewed in detail elsewhere in [Pabinger et al. 2014; Olson et al. 2015]. For the purposes of this work, we have focused on the utility of WGS for the clinician and, in particular, the use of these SNPs to predict drug resistance. In
Regardless of the application, the quality of WGS data depends on a number of factors, including the desired length of the sequencing reads and the cycle time [Quick et al. 2015]. These parameters in turn affect the turnaround time for results. Considering the most frequently used bench-top sequencers, raw sequencing results can be available in a clinically attractive span of just a few hours (for the Ion Torrent PGM) to as much as 39 hours with MiSeq for paired end 250 bp reads. By adjusting the sequencing protocol for MiSeq, it may be feasible to reduce this time frame without affecting key inferences, such as species and strain assignments [Quick et al. 2015]. An important consideration when making such adjustments is the ‘depth of coverage’; the more reads that span a position in the reference genome, the more support there is for the base identified. The optimal depth of coverage to detect clinically relevant variants needs to be determined.
Another factor influencing the time to obtain these data is whether samples are batched or run independently. According to Quick and colleagues [Quick et al. 2015], the MiSeq can sequence up to ∼100 isolates simultaneously. In our experience, the MiSeq 250-bp paired-end sequencing can generate a minimum of 10 million reads; if 20× coverage is desired, only ∼57 isolates of
WGS for detection of M. tuberculosis , including the prediction of drug resistance
In the clinical mycobacteriology lab, the goal is to secure a diagnosis of active TB and to provide clinicians with guidance on which antibiotics they should or should not prescribe for their patients. These two goals have classically been achieved with phenotypic tests, some dating to the 19th century. This begs the obvious question of whether WGS can help modernize the TB lab, with the goal of offering faster and more accurate results.
The current clinical workflow for detection of Clinical diagnostic workflow for 
Superimposed on this classic workflow (smear microscopy, culture, then DST), laboratories have overlaid molecular testing over the past two decades, using a variety of different platforms and clinical strategies. The first molecular tests approved were only licensed for the speciation of smear microscopy-positive samples [Parrish and Carroll, 2011], so their key role was in assigning a microbial name to such a sputum sample [Vuorinen et al. 1995; Carpentier et al. 1995]. Then, with time and experience, it became recognized that nucleic acid amplification testing could be offered on smear-negative samples where there was a high clinical suspicion of TB [Centers for Disease Control and Prevention (CDC) 2009]. To reduce costs of controls, these ‘rapid’ first generation tests were generally batched and as a result, might only have been done twice or three times per week, depending on laboratory volume. More recently, the GeneXpert (Cepheid Inc., Sunnydale, CA, USA) has offered a random-access real-time nucleic acid amplification test, which can be done on a single sample, without having to wait for samples from other patients. GeneXpert is conducted directly on the clinical specimen to detect both the presence of
WGS for diagnosis
Until recently, the utility of WGS for
If WGS is to be applied on the patient sample, the conceptual advantage is a more rapid result. However, the vast majority of samples are negative for
If instead WGS is applied on the positive culture, then the benefit of rapidity has been lost, as the patient should already be isolated and started on treatment, based on either smear microscopy, a nucleic acid amplification test or the Accuprobe result on the culture. In this case, WGS may offer a different opportunity, which is a more rapid identification of antibiotic resistance.
WGS for resistance
Examples of molecular diagnostics for drug resistance in

For meta-analyses: if available, ranges are shown in addition to pooled estimates, to indicate potential heterogeneity. All tests shown, with exception of GeneXpert, are line probe assays. Where no sensitivity is reported, no isolates were identified with resistance to the target drug. *Includes studies using MTBDR (first-generation). **Results not shown for second-line drugs, as only testing was only conducted on the 3 samples with resistance to first-line drugs. AID predicted 3/3 isolates to be susceptible to second-line, confirmed with phenotypic DST. ***Study did not report separate test results for positive RIF and INH resistance.
CI, confidence interval; CrI, credible interval; INH, isoniazid; RIF, rifampin.
As most rifampin-resistant isolates are also isoniazid-resistant, the GeneXpert uses
By sequencing the whole genome, in theory all resistance-connoting mutations that can guide clinical treatment can be identified by comparing the genome of the patient isolate with detailed databases of known resistance markers [Sandgren et al. 2009; Flandrois et al. 2014]. In practice, this will work, if (a) these markers accurately predict
In the same manuscript [Coll et al. 2015], Coll and colleagues also compared the performance of their database with KvarQ, a software that uses pre-specified ‘testsuites’ of known resistance-connoting mutations and other regions of interest to predict resistance [Steiner et al. 2014]. Using phenotypic data as the gold standard, sensitivity was substantially lower for nearly all drugs using the KvarQ method (though 95% CIs overlapped for all except EMB and KAN). Among first-line drugs, only RIF yielded similar point estimates to those obtained with Coll and colleagues’ mutation library, with sensitivity of 95.8% (95% CI 93.4–98.2%), while sensitivity for INH was only 86.9% (95% CI 83.1–90.7%). No results were available for ETH and CAP using the KvarQ software. Specificity was generally higher using KvarQ, though this difference was only significant for EMB and STR. Specificity for RIF was similar to that obtained with the mutation database, at 97.9% (95% CI 96.5–99.3%).
In a similar study [Walker et al. 2015], Walker and colleagues selected 23 candidate resistance-associated genes from the literature [Sandgren et al. 2009] and then used an algorithm to characterize mutations (SNPs and indels) within these genes and their promoter regions as resistance-connoting or benign. In a training dataset of 2099 isolates, 120 resistance-connoting mutations were identified, 772 were classified as benign and 101 could not be classified as either (called ‘uncharacterized’). The resistance-connoting and benign mutations identified in this training dataset were then used in a validation study on an additional 1552 genomes, 29% of which were resistant to at least one drug on drug susceptibility testing (DST). Using these mutations, authors were able to predict 89.2% of phenotypes as resistant or susceptible. 10.8% of phenotypes could not be predicted, as these contained mutations that had not been characterized. Among those where phenotype could be predicted and considering predictions for each drug independently, 112 of 6892 with drug-sensitive DST were predicted to be resistant based on WGS (1.6%), while 94 of 1221 with drug-resistant DST were erroneously predicted to be drug-sensitive (7.7%). The latter may be due to mutations with unknown function outside the 23 candidate genes interrogated. This is similar to Farhat and colleagues [Farhat et al. 2013]; in this study, authors performed targeted deep sequencing of known resistance genes to verify that resistance mutations were absent in subpopulations within isolates. They found that 13/47 isolates with phenotypic resistance had no previously known mutations. Unexplained resistance, wherein phenotypic resistance is present but known resistance-connoting mutations are absent has been most pronounced for PZA [Hewlett et al. 1995] and second-line drugs. For example, Farhat and colleagues [Farhat et al. 2013] found that, among isolates resistant to ciprofloxacin, KAN and CAP, 2/3, 6/18 and 1/6 isolates, respectively, had unexplained resistance. As the reliability of phenotypic testing is least well established for these drugs [Horne et al. 2013], this is where there is the greatest need for WGS, but presently also the greatest knowledge gap.
In clinical medicine, the physician wants to know whether the isolate has a resistance-connoting mutation or not, so that treatment can be tailored accordingly. Indeterminate test results offer little clinical guidance, and often steer clinicians to other antibiotics, where feasible. While it is logical to exclude isolates with uncharacterized mutations from a scientific paper that aims to understand resistance, in a clinical laboratory, these have to reported one way or the other. Analyses that classified such uncharacterized mutations as predictive of phenotypic susceptibility greatly affected test parameters; the sensitivity of WGS for INH and RIF resistance dropped from 94.2% (95% CI 91.1–96.5%) and 96.8% (95% CI 94.1–98.5%) with uncharacterized mutations excluded to 85.2% (95% CI 81.1–88.7%) and 91.7% (95% CI 87.9–94.5%) with uncharacterized mutations included, respectively. Sensitivity for PZA resistance in the latter analysis was the lowest overall, at only 24% (95% CI 17.9–30.9%). Until such mutations can be confidently assigned to the appropriate phenotype, it would seem that parallel, or at the least, sequential phenotypic testing should remain part of the diagnostic pathway.
Furthermore, these publications generally included biased samples, with relatively high proportions of drug-resistant isolates. As many clinical labs identify primarily drug-sensitive isolates, the operating parameters of WGS for this purpose may change when evaluated against more representative samples. While authors had generally high specificity for most drugs, the predictive value depends on the underlying prevalence of drug resistance. In a country such as Canada, which detected RIF resistance among only 17 of 1380
Despite these limitations, it is clear that WGS offers magnitudes more information than the molecular methods listed in Table 1, with the potential of greatly advancing clinical diagnostics for
Overall, these data support the great potential of WGS as a tool to predict resistance. However, databases of
Another issue for clinical application of WGS is timeliness of reporting. As of yet, two papers reported on the application of WGS in ‘real-time’ to clinical cases: a case report of a patient [Köser et al. 2012] with extremely drug-resistant (XDR) TB (defined as MDR TB plus resistance to an injectable second-line drug and a fluoroquinolone) and a prospective cohort of patients in the United Kingdom suspected of having XDR TB [Witney et al. 2015]. Köser and colleagues successfully obtained sequence data from a 3-day-old MGIT culture, identifying two concurrent but distinct strains of
The Witney and colleagues study also illustrated that for WGS data to be used clinically, the results need to be analyzed rapidly and presented in a clear, easily interpretable manner. Several groups have produced online tools (e.g. ‘PhyResSE’ [Feuerriegel et al. 2015] and ‘TB Profiler’ [Coll et al. 2015]) wherein raw sequencing data for an isolate can be uploaded and analyzed for resistance-connoting mutations. As mentioned previously, the KvarQ software can also predict resistance from raw sequencing data; in contrast to PhyResSE and TB Profiler, this can be done on a local server [Steiner et al. 2014]. Yet, despite efforts to make these reports accessible to the wider scientific community, a knowledge of genomics and/or bioinformatics is still required to interpret results. As an example, the quality of SNPs is provided with details such as depth of coverage, a parameter that most clinicians would be uncomfortable judging. Presently, PhyResSE and TB profiler are explicitly for research purposes only, which poses regulatory hurdles to the delivery of results destined for the clinical chart. Witney and colleagues [Witney et al. 2015] piloted a WGS report during the course of their study, but, unfortunately, clinician perception of this report and its interpretability was not assessed. Furthermore, though ‘best practices’ have been proposed for identifying SNPs [Olson et al. 2015], the current bioinformatics workflows used to analyze WGS data remain largely unstandardized. For implementation in the clinical lab, appropriate quality control measures [Clinical and Laboratory Standards Institute, 2014] and a standardized workflow need to be established. The lessons of the past five decades of emerging antibiotic resistance have demonstrated that even a simple dichotomous test result, i.e. resistant or susceptible, does not always predict appropriate care. Therefore, the application of WGS-based results to clinical care may benefit from evaluations done by experts in implementation science, rather than genomics or microbiology.
Conclusion
Offering increased resolution and substantially more data compared with conventional methods, WGS has revolutionized the arena of molecular epidemiology. Now, it seems poised to do the same for the clinical microbiology laboratory. The appeal of WGS for
The issues raised above are only further amplified when contemplating the countries of the world that suffer the greatest burden of TB and have the highest prevalence of drug-resistant strains. While it is clearly feasible to ship sequencing machines around the world, as has already been done with the GeneXpert platform, it is not as simple to distribute the technical and bioinformatic expertise required for next-generation sequencing where it is needed. A potential solution to the latter is open-source coding and online data treatment, but this is currently lacking for clinical use, even in settings with expertise in these methods. Ultimately, what is needed is an easy-to-use software complete with a graphical user interface that is capable of converting data-intense sequence files into a simple, concise clinical message. As done with GeneXpert [Theron et al. 2014b], these outputs then need to be field-tested in settings with a sufficient burden of drug-resistant TB to enable evaluation of whether test results altered treatment decisions and clinical outcomes. The relatively small number of MDR TB patients in countries such as Canada may preclude a formal evaluation of patient outcomes, simply due to sample size considerations. In order to assess its clinical utility for resource-rich countries where its use has been pioneered, we may need to first embed WGS in treatment studies conducted in the developing world, where the challenge posed by TB and drug resistance remains the greatest.
Footnotes
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article:
Conflict of interest statement
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
