Abstract
Introduction:
Increasing occurrence of infections caused by multidrug-resistant Gram-negative bacteria resulted in colistin being the last agent for treatment. Apart from plasmid-mediated
Methodology:
A total of 100 carbapenem-resistant
Results:
Among 100 carbapenem-resistant
Conclusions:
The performance of cBMD was excellent, whereas the E-test was unacceptable. VITEK-2 and RPNP performed better but remained unreliable due to high error rates. Multiple mutations in the target proteins involving lipopolysaccharide formation, modification, and regulation were seen, resulting in colistin resistance.
Introduction
The increasing incidence of multidrug-resistant (MDR) Gram-negative bacilli (GNB) such as
As part of the routine antibiogram panel, the susceptibility of colistin must be tested in settings where these MDR GNBs are prevalent to avoid delays in reporting. For appropriate therapeutic decision-making, rapid and reliable colistin susceptibility testing methods is the need of the hour in routine diagnostic microbiology laboratories. To date, few studies around the globe have elaborately studied the performance of different colistin susceptibility methods, showing opposing results; therefore, more studies are needed to establish the most accurate method. 6
The addition of L-Ara4-N and phosphoethanolamine (PEtN) molecules causing an alteration in the lipopolysaccharide region of bacterial outer membrane is the most common way leading to colistin resistance.
7
Mutations involving the two-component regulatory systems like
Disk diffusion is the most prevalent antimicrobial susceptibility testing method used in routine diagnostic microbiology laboratories. But this method is not recommended for detection of colistin resistance like other antibiotics because of high error rates leading to discordant and unreliable results compared to minimum inhibition concentration (MIC) based methods.11,12
Some studies have demonstrated excellent correlations between the results of the E-test and broth microdilution (BMD) methods for colistin,12–16 while others questioned its reliability.17,18 Although E-test strips are convenient in determining MIC levels but because of conflicting performance results, routine colistin susceptibility testing using E-test is not recommended. In addition, limited studies have tested the performance of automated methods like VITEK-2 for colistin susceptibility and could not reach any conclusive decision.12,15,18
BMD is mostly considered the gold standard method for determining antibiotic MICs, although not convenient for routine clinical laboratories. 19 It is sometimes becomes impractical to include BMD as a part of the routine antibiogram panel for laboratories specifically in resource-poor settings due to its extensive requirement of human resources, high cost, and lengthy test timing. Many laboratories perform BMD either due to specific requests for colistin susceptibility or after detecting resistance to other drugs like carbapenems, causing a delay in reporting.20,21 In addition, Colistin shows a property of adherence of varying degrees to different surfaces used to make BMD plates, which may cause a reduction in antibiotic concentrations affecting colistin susceptibility testing. 22 Surfactant polysorbate 80’s addition to BMD tray minimized colistin adhesion leading to a significant decrease in colistin MICs, affecting mainly bacteria with relatively low MICs.17,20,23 Nevertheless, surfactant polysorbate 80 is not recommended by the Clinical and Laboratory Standards Institute (CLSI) for colistin susceptibility by BMD. 24
‘Colistin Heteroresistance’ is defined as colistin‑resistant subpopulations that emerge from a colistin‑susceptible population under colistin pressure and may be evidenced by the presence of skip wells in BMD. 7 Due to all these issues, commercial BMD (cBMD) methods have come up recently. These methods do not need hectic standardization steps, do not have cumbersome logistic problems and show excellent correlation with BMD. 25 Apart from cBMD platforms, two other methods, namely colistin broth disk elution method and colistin agar test, have performed at par with BMD and are now recommended as standard tests for colistin susceptibility testing by CLSI.24,26
The breakpoints for colistin have changed every year, and different organizations have differences among them, which ultimately complicates the interpretation of colistin susceptibility results.24,27 In this study, the performance of various colistin MIC testing methods, such as cBMD, E-test, and VITEK-2, along with the colorimetric test, that is, RPNP, were tested against BMD using a collection of non-repeating carbapenem-resistant
Material and methods
Isolate collection
A total of 100 CRE isolates comprising only
Colistin susceptibility
Various phenotypic methods used to determine colistin MICs are cBMD (Mikrolatest® MIC colistin), E-test (Himedia, India), and VITEK-2 (bioMérieux, Marcy-l'Étoile, France) following which comparison of the results were made with BMD results according to CLSI breakpoints (intermediate, ⩽ 2 µg /mL; resistant, ⩾ 4 µg/mL).
24
For the purpose of this study, the ‘Intermediate’ category mentioned in CLSI guideline is considered as ‘Susceptible’. In the interpretation of colistin MIC, as EUCAST breakpoints (resistant > 2µg/mL; susceptible ⩽ 2 µg/mL) were similar to that of CLSI, we used both the guidelines in CLSI/EUCAST format for the study.
27
In case of disagreement, the BMD method was used for reporting susceptibility. cBMD is a ready-to-use BMD test kit coated with different colistin concentrations to determine colistin MIC. Rapid Polymyxin NP (RPNP) (ELITech Group, Puteaux, France) is a colorimetry-based method detecting bacterial growth indicated by glucose metabolism in the presence of 2 μg/mL of colistin. Colistin resistance is indicated by pH shift due to the formation of acid metabolites resulting from glucose metabolism, which in turn causes a color shift of phenol red indicator from orange to yellow.
28
VITEK-2 AST susceptibility cards AST- GN-280 (bioMérieux, Marcy-l'Étoile, France) were used as per the instructions mentioned by the manufacturer, where colistin MIC is reported in the range between ⩽ 0.5 to ⩾ 16 μg/mL.
Evaluation of minimum inhibitory concentration correlations
Essential agreement (EA) was defined as an E-test/ cBMD/ VITEK-2 MIC equal to or within ± 1 dilution of the MIC result of BMD. Categorical agreement (CA) was met when E-test/ cBMD/ RPNP/ VITEK- 2 interpretive criteria agreed (susceptible /resistant) with BMD interpretive criteria. A major error (ME) occurred when cBMD/ RPNP/ VITEK-2 results were resistant, and BMD was susceptible and was calculated only for susceptible isolates. A very major error (VME) occurred when cBMD/ RPNP/ VITEK-2 results were susceptible, and BMD was resistant and was calculated only for resistant isolates. 30 Colistin susceptibility performance was considered acceptable if both the CA and EA were ⩾ 90%. In addition, MEs and VMEs rates were benchmarked as ⩽ 10% and ⩽ 3%, respectively, for acceptable performance. 31
All the discordant results (MEs and VMEs) were retested in triplicate to reconfirm the results. In the repeat testing, if the error was resolved, the repeat results were kept as final. If repeat MIC values were within ± 2 log dilution and categorical error in the form of MEs and VMEs remained, the categorical error was accepted. Skip well phenomenon was defined as the absence of growth of an isolate at a lower antimicrobial concentration(s). A single skip well did not affect the MIC interpretation, while multiple skip wells were considered uninterpretable according to CLSI guidelines. 19 For the skip well phenomenon in BMD panels, the isolates were retested.
Modified rapid polymyxin NP
RPNP solution was prepared by adding 6.25 g of cation-adjusted Mueller-Hinton II broth powder and 0.0125 g of phenol red (Sigma-Aldrich) in 225 mL of distilled water. 32 The pH of the resultant solution was adjusted at 6.7 with 1 mol/L HCl, following which sterilization was carried out by autoclaving the solution at 121°C for 15 min at 15 lbs. After sterilization, the RPNP solution was cooled at room temperature, and 25 mL of filter-sterilized 10% anhydrous D(+)-glucose (Sigma-Aldrich) was added.
The RPNP test was modified by incorporating two additional wells, one containing colistin-free RPNP solution plus EDTA (80 µg/mL) [EDTA is an inhibitor of PEtN transferase] and the other having RPNP solution containing colistin (2 µg/mL) along with EDTA (80 µg/mL). In the modified scheme (Figure 3), row A wells were filled with 150 µL of colistin-free RPNP solution. In row B wells, 150 µL of RPNP solution containing colistin sulfate was filled. Next, row C wells were added with 150 µL of colistin-free RPNP solution plus EDTA only. Finally, row D wells were added with 150 µL of RPNP solution containing both colistin sulfate and EDTA. In the next step, to all wells of Column 1, 50 µL of 0.85% NaCl (negative sterility control) was added. In other respective columns for each strain, 50 µL of a 3.0 to 3.5 McFarland bacterial suspension corresponding to ~109 CFU/mL was added and adequately mixed with 150 µL reaction solution in each well. Incubation of the 96 -well plate was done at 35 ± 2°C under aerobic conditions for 4 h.
At every 1 h, change of color of the 200 µL reaction mixture in each well was monitored. Any color change from orange to yellow in wells containing colistin sulfate was considered positive for colistin resistance. When the wells with colistin-containing solution supplemented with EDTA remained orange, signifying the absence of glucose metabolism due to inhibition of
DNA extraction and conventional multiplex PCR for the mcr-1, mcr-2, mcr-3, mcr-4, and mcr-5 genes
Extraction of the genomic DNA of the 100 CRE and 10 intrinsically colistin-resistant
Whole genome sequencing, sequence assembly, annotation, and analysis
The only two colistin-resistant XDR [non-susceptibility to at least one agent in all but two or fewer antimicrobial categories]
30
non-
Nucleotide sequence accession numbers
The whole-genome sequences of
Statistical analyses
SPSS software v.20.0 (SPSS Inc., Chicago, IL) was used to analyze the data, and
Results
Out of 100 CRE isolates most common organism to be isolated was
Colistin susceptibility by different methods
Among these 100 CRE isolates, according to CLSI/EUCAST guidelines, total colistin resistance by standard BMD method was observed to be 15%. Species wise,
Colistin resistance profile among different species of CRE by different phenotypic methods along with their MIC50/90 values.

BMD, Broth microdilution; CLSI, Clinical and Laboratory Standards Institute;
Evaluation of minimum inhibitory concentration correlations and error rates
Colistin had the same MIC50 and MIC90 (0.5 µg/mL) in the colistin susceptible strain. In colistin-resistant strains, overall MIC50 of 8 µg/mL and MIC90 of 16 µg/mL were observed. The MIC50/90 rates of cBMD were similar to BMD, whereas both E-test and VITEK-2 had higher MIC90 values (Table 1). The rate of colistin resistance was identical for all the methods with respect to
The overall sensitivity (SN) for cBMD was 100%, whereas specificity (SP) was 98.8%.VITEK-2 showed SN ad SP, of 86.7% and 88.2%, respectively. In comparison, RPNP had SN similar to VITEK-2 but with a higher SP of 91.8%. Among all the phenotypic tests compared in the study, E-test fared poorly with SN and SP of 60% and 90.6%, respectively (Table 2).
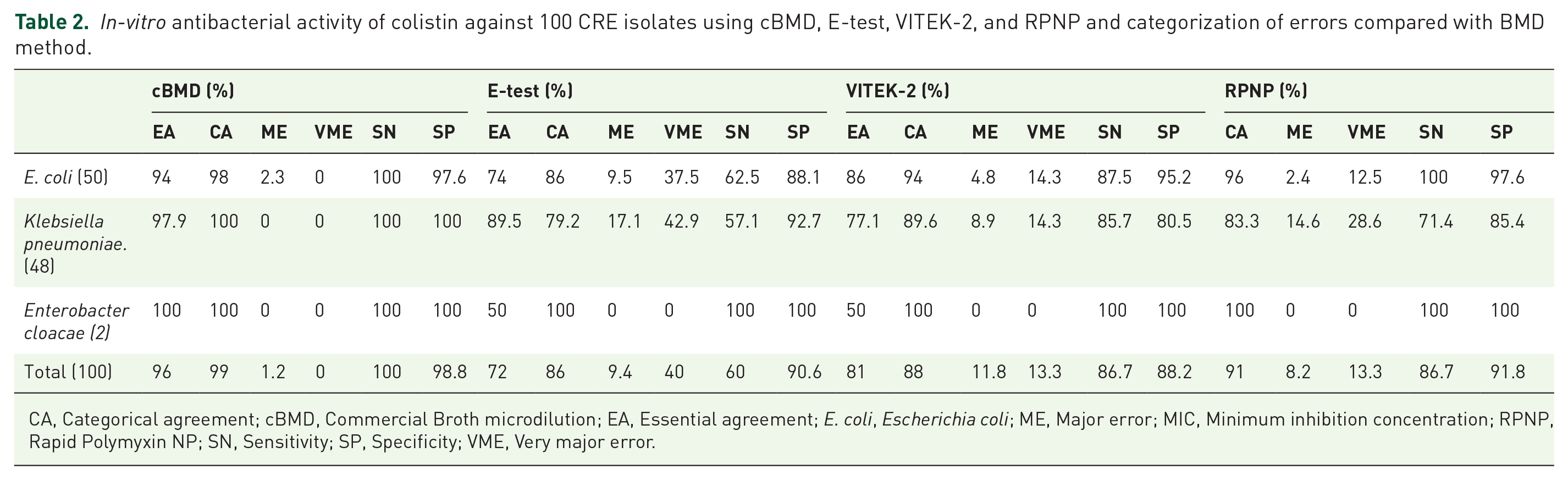
CA, Categorical agreement; cBMD, Commercial Broth microdilution; EA, Essential agreement;
According to CLSI/ EUCAST breakpoints, EA, CA, ME, and VME for cBMD/E-test/VITEK-2 were 96%/72%/81%; 99%/86%/88%, 1.2%/9.4%/11.8% and 0%/40%/13.3%, respectively. In RPNP, CA, ME, and VME rates were 91%, 8.2%, and 13.3%, respectively (Table 2).
The performance of cBMD for
A Scatter diagram was plotted using colistin MIC results of 100 isolates by cBMD, VITEK-2, E-test, and RPNP and compared with the BMD test result. Ninety-five isolates showed an excellent correlation in the case of cBMD when compared with BMD results (Figure 1(a)). VITEK-2 showed excellent correlation with 79% isolate lying with ± 1 log2 dilution of BMD (Figure 1(b)). However, a relatively poor correlation was seen between the E-test and BMD results with 77% isolates having E-test MIC lying between ± 1 log2 dilution of BMD MIC. Sixteen percent of isolates having BMD MIC of ⩽ 0.5 µg/mL had E-test MIC of ⩾ 2 µg/mL (Figure 1(c)). Comparing the RPNP method results with BMD, a good categorical agreement was observed with isolates having MIC value ⩽ 2 µg/mL, showing 78/85(91%) isolates susceptible to colistin. Few isolates, that is, 2 out of 15 (13.3%) with MIC > 2 µg/mL by BMD, were wrongly classified as susceptible by RPNP (Figure 1(d)). The correlation plot between MIC results of cBMD and BMD values was drawn for the comparative analysis. Pearson R2 value was found at 0.8736, which indicates excellent concordance between both methods. On the other hand, both VITEK-2 and E-test showed fair concordance compared to BMD (Figure 2).

Scatter diagram of colistin MICs obtained by BMD compared with a) cBMD, b) VITEK-2, c) E-test, and d) CNP for 100 isolates. Essential agreement is highlighted in green (perfect agreement) and yellow (±1 log2 dilution). The solid horizontal and vertical lines represent the clinical breakpoint value established by CLSI/ EUCAST. Blue color signifies susceptible MIC, whereas Red color signifies resistance MIC range.
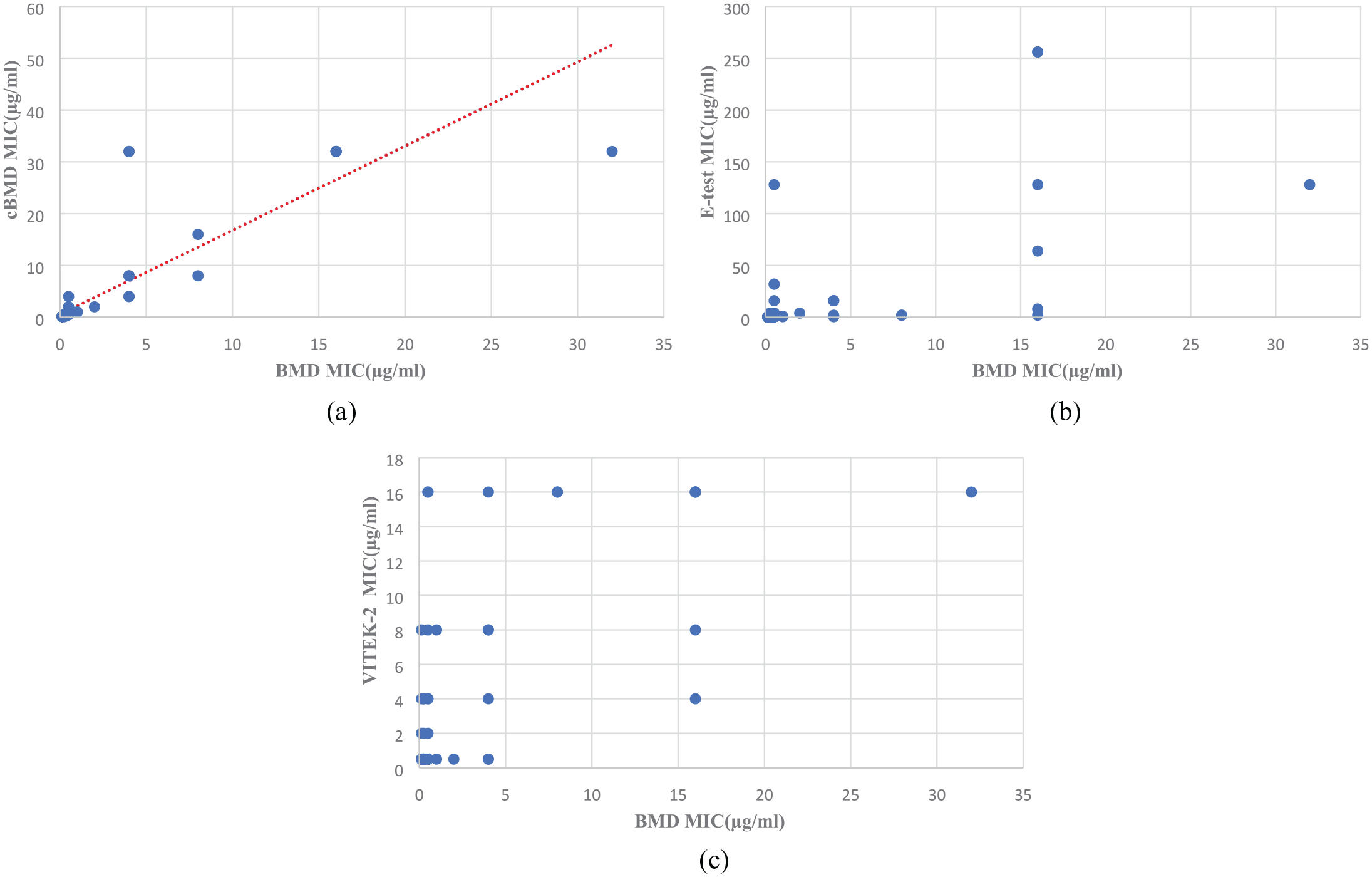
Correlation plot of colistin MICs obtained by BMD compared with a) cBMD, b) E-test, c) VITEK-2 for 100 isolates. Pearson correlation values show an excellent correlation between BMD and cBMD, whereas it shows a fair correlation for E-test and VITEK-2.

Modified rapid polymyxin NP test (mRPNP). Wells A1-A6 were filled with colistin-free RPNP solution. Wells B1 to B6 were filled with RPNP solution + colistin sulfate. Wells C1 to C6 were filled with colistin-free NP + EDTA. Wells D1 to D6 were added with RPNP solution + colistin sulfate + EDTA. Wells in column 1 were filled with 0.85% NaCl (negative sterility control), whereas Columns 2 to 7 represent the mRPNP test performed for
Modified rapid polymyxin NP and conventional multiplex PCR for mcr 1-5 genes
Only one
Whole-genome sequencing and analysis
The WGS analysis revealed that the closest neighbor of H-53 and H-62 strains is
Cumulative results of amino acid substitutions in various genes contributing to colistin resistance found upon WGS analysis of two non-MCR colistin-resistant XDR

PROVEAN score is a measure of the change in protein structure: if the score is equal or below to the predefined threshold (cut off = −2.5), the variant is predicted to have a ‘deleterious’ effect; if above, the variant is predicted to have a ‘neutral’ effect. In the latter column, there is a prediction of the mutation effect on the protein functionality.
Discussion
Colistin therapy is one of the last viable options while treating serious life-threatening infections caused by MDR GNBs, especially CRE. 5 However, a worldwide trend of increasing colistin MIC levels has been noted,37,38 underlining the need for a robust and precise colistin susceptibility testing method.
The final verdict for an optimal method for colistin susceptibility testing is an ongoing debatable and developing field. Therefore, the performances of four colistin susceptibility methods against CRE isolates were evaluated in this study.
While false colistin susceptible result is considered as a serious error, false colistin resistant results should be viewed as an equally gravely. Therefore, excellent essential agreement regarding colistin susceptibility testing is needed.
In our study, overall colistin resistance among CRE was 15% with standard BMD which is similar to findings presented in other studies worldwide, with colistin resistance among
MIC50/90 of
In this study, cBMD meet all the CLSI recommendations regarding VMEs and MEs when compared with the MIC results of BMD with EA and CA lying around 96% and 99%, respectively, with a high concordance with BMD (R2, 0.8736; Figure 2a). Similar results were found in Singh
All the phenotypic methods did appear to perform well against
In the case of VITEK-2, previous studies have a controversial and ambiguous conclusion for
RPNP showed high CA (91%), but it also suffered from high ME (8.2%) and VME (13.3%), which is similar to previously done studies.28,49 There are several possible explanations for the higher VME rates observed in general for the RPNP method. Primary reasons for higher error rates may be due to difficulty in interpreting the color changes, inoculum effect, and borderline MICs. 49 Previous studies have noted this same limitation, with both in-house methods and the commercial kits.50,51
BMD is a very laborious and expensive method for MIC detection. However, the cBMD method in the present study is easy to use without the requirement of additional equipment or great technical expertise. Therefore, after internal validation, cBMD can be utilized for colistin MIC testing in clinical microbiology laboratories. Furthermore, as per cost comparison, cBMD and standard BMD costs are comparable, while the costs of both RPNP and VITEK-2 are higher.
This present study also looked for the presence of plasmid-mediated
In this study, extensive analysis of the genome of both the XDR
This study has several limitations. The isolates were taken from a single center with fewer numbers contributed by
Conclusion
In conclusion, the present study's findings showed that cBMD performed well with excellent correlation compared to BMD, whereas the performance of gradient tests like the E-test was unacceptable. This unacceptable performance of the E-test is probably related to the poor and unpredictable diffusion of colistin sulfate. VITEK-2 and RPNP performed better than E‑test but remained unreliable due to high ME and VME. Further studies correlating MICs with the clinical outcome are needed to determine the precise breakpoint to lead patient management. The present study highlights the importance of optimal colistin susceptibility amid CRE isolates. Although there is a substantial colistin resistance among the CRE isolates mediated mainly by the accumulation of mutation in lipopolysaccharide formation and regulation pathway, only one isolate with a plasmid-mediated
Supplemental Material
sj-docx-1-tai-10.1177_20499361221080650 – Supplemental material for Analysis of colistin resistance in carbapenem-resistant Enterobacterales and XDR Klebsiella pneumoniae
Supplemental material, sj-docx-1-tai-10.1177_20499361221080650 for Analysis of colistin resistance in carbapenem-resistant Enterobacterales and XDR Klebsiella pneumoniae by Raunak Bir, Hitender Gautam, Nazneen Arif, Priyanka Chakravarti, Jyoti Verma, Sayantan Banerjee, Sonu Tyagi, Sarita Mohapatra, Seema Sood, Benu Dhawan, Rama Chaudhry, Arti Kapil, Bimal Kumar Das and Bhabatosh Das in Therapeutic Advances in Infectious Disease
Footnotes
Acknowledgements
We thank the Laboratory staff, Bacteriology Section, All India Institute of Medical Sciences, New Delhi, India, and Translational Health Science and Technology Institute, Faridabad, for helping us carry out this study.
Author contributions
Conflict of interest statement
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Intramural Research Grant from All India Institute of Medical Sciences, New Delhi with Project Code No.: A-597.
Ethics statement
Ethical approval for this study has been granted by the Institutional Ethical Committee, AIIMS, New Delhi (Ref. No.: Institutional Ethical Committee-283/01.06.2018), and all the protocols were followed as per the ethical standards. As the inclusion criteria was CRE isolates from routinely sent samples, the requirement of informed consent was exempted.
Data transparency
All relevant data related to this scientific work is available.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
