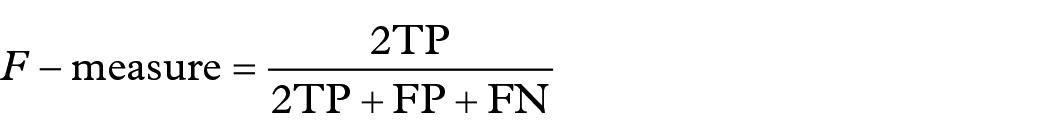Abstract
Background:
Drug–drug interactions (DDI) are an important cause of adverse drug reactions (ADRs). Could large language models (LLMs) serve as valuable tools for pharmacovigilance specialists in detecting DDIs that lead to ADR notifications?
Objective:
To compare the performance of three LLMs (ChatGPT, Gemini, and Claude) in detecting and explaining clinically significant DDIs that have led to an ADR.
Design:
Observational cross-sectional study.
Methods:
We used the French National Pharmacovigilance Database to randomly extract Individual Case Safety Reports (ICSRs) of ADRs with DDI (positive controls) and ICSRs of ADRs without DDI (negative controls) registered in 2022. Interaction cases were classified by difficulty level (level-1 DDI being the easiest and level-2 DDI being the most difficult). We give each LLM (ChatGPT, Gemini, and Claude) the same prompt and case summary. Sensitivity, specificity, and F-measure were calculated for each LLM in detecting DDIs in the case summaries.
Results:
We assessed 82 ICSRs with DDIs and 22 ICSRs without DDIs. Among ICSRs with DDIs, 37 involved level-1 DDIs, and 45 involved level-2 DDIs. Correct responses were more frequent for level-1 DDIs than for level-2 DDIs. Regardless of difficulty level, ChatGPT detected 99% of DDI cases, and Claude and Gemini detected 95%. The percentage of correct answers to all DDI-related questions was 66% for ChatGPT, 68% for Claude, and 33% for Gemini. ChatGPT and Claude produced comparable results and outperformed Gemini (F-measure between 0.83 and 0.85 for ChatGPT and Claude and 0.63–0.68 for Gemini) to detect drugs involved in DDI. All exhibited low specificity (ChatGPT 0.68, Claude 0.64, and Gemini 0.36) and reported nonexistent DDIs for negative controls.
Conclusion:
LLMs can detect DDIs leading to pharmacovigilance cases, but cannot reliably exclude DDIs in cases without interactions. Pharmacologists are crucial for assessing whether a DDI is implicated in an ADR.
Plain language summary
Background:
Drug-drug interactions (DDIs) are reactions between two (or more) drugs that can cause harmful side effects (unwanted or unexpected effects that comes along with the beneficial effect of a drug), known as adverse drug reactions (ADRs). We wanted to assess the capacity of Large Language Models (LLMs), a type of machine learning designed to generate language and presented as chatbots, to detect these drug interactions.
Goal of this study:
To compare how well three LLMs—ChatGPT, Claude, and Gemini—could identify and explain DDIs that led to an ADR and a pharmacovigilance report.
Methods:
The researchers used reports from the French pharmacovigilance database and tested the LLMs on 82 cases where DDIs were present and 22 cases without DDIs.
Results:
ChatGPT detected 99% of DDI cases, while Claude and Gemini detected 95%. However, all the LLMs struggled to exclude the lack of interaction in ADR cases, often detecting non-existent DDIs. Overall, ChatGPT and Claude performed better than Gemini.
Conclusion:
While LLMs can help detect DDIs leading to an ADR, they are unreliable for confirming the absence of interactions. Human expertise, from pharmacologists, remains essential for assessing drug interactions in pharmacovigilance cases.
Keywords
Introduction
Since the release of ChatGPT in 2022, there has been a massive dissemination and utilization of large language models (LLMs). These models are constructed using neural networks and are mainly designed for natural language processing tasks. 1 They function as next-word predictors, generating text based on the sequence of previous words. 2 LLMs are now increasingly used for many tasks, including tasks for which they have not been specifically designed, trained, and validated (e.g., problem solving and analysis). In medicine, their use is increasing. 3
In pharmacovigilance, studies evaluating the utility of LLMs are also on the rise. According to Matheny et al., LLMs offer opportunities to support signal-identification activities. 4 LLMs were assessed as tools for literature screening to categorize medical publications as relevant or not for safety signal (e.g., GPT-3.5, GPT-4, Claude2). 5 LLMs could also identify and categorize adverse drug reactions (ADRs) in medical terms from the Medical Dictionary for Regulatory Activities from unstructured text (GPT-3.5). 6 In causality assessment, ChatGPT (version unknown) produced ambiguous results in a case report of toxic epidermal necrolysis. 7 Finally, in a previous study published in 2023, we highlighted the limitations of ChatGPT (version GPT-4.0), one of the most well-known LLMs, in responding to queries directed to our drug information service. 8 These results were corroborated by Pariente et al., with ChatGPT v3.5. 9
Regarding drug interactions, few studies are currently available. Two studies evaluating the capacity of LLMs to identify drug–drug interactions (DDIs) for different selected drug pairs were published. According to Juhi et al., ChatGPT (version unknown) was effective in identifying DDIs between two drugs but was partially effective in explaining them, based on an analysis of 40 DDI pairs. 10 ChatGPT provided inconclusive answers and incomplete guidance regarding DDIs. The authors concluded that there was a need for further improvement to identify and explain DDIs accurately. In 2024, Al-Ashwal et al. compared the accuracy of ChatGPT-3.5, ChatGPT-4, Microsoft Bing AI, and Bard in predicting DDI with clinical reference tools. 11 The authors investigated DDIs between sodium/glucose cotransporter 2 inhibitors or macrolides and the most frequently prescribed drugs listed in the DrugStats Database. They identified discrepancies among different LLMs chatbots regarding their accuracy in detecting DDIs. To date, only one study, published in September 2024, has evaluated ChatGPT’s ability to detect DDIs in real-world hospitalized patient settings. 12 The authors found that ChatGPT-3.5 exhibited low sensitivity but good specificity in detecting DDIs from prescribed medications for 120 hospitalized patients. According to a recent systematic review, in healthcare research performed with LLMs, only 5% of studies have utilized real patient care data. 13
To our knowledge, no studies in the field of pharmacovigilance have evaluated LLMs for the detection of clinically significant DDIs, while these interactions remain a critical aspect of pharmacovigilance. 14 To address this question, we compared the performance of three LLMs—Claude, Gemini, and ChatGPT—in their ability to detect and explain clinically significant DDIs that have led to an ADR.
Methods
French National Pharmacovigilance Database
We used the French National Pharmacovigilance Database (FNPV) to extract Individual Case Safety Reports (ICSRs) of ADRs induced by DDIs. FNPV gathers spontaneous ICSRs from French healthcare professionals or patients. Each report is assessed by clinical pharmacologists in the relevant regional pharmacovigilance center (30 regional pharmacovigilance centers are present nationwide)—according to the updated French method for the causality assessment of ADRs—before being recorded in the database. 15 ICSRs are classified according to the type of case: ADRs, medication error with or without ADRs, drug interactions, pregnancy, breastfeeding, drug dependence, withdrawal, overdose (accidental or intentional), and occupational exposure. The updated French causality assessment method ensures satisfactory reproducibility among users. 16
ICSRs selection and classification
ICSRs involving ADRs induced by DDIs (positive controls) and registered in the FNPV between January 1st, 2022 and December 31st, 2022 were extracted. This time frame was selected to ensure that all LLMs were trained using data covering this period. In addition, to evaluate the specificity of LLMs, we also extracted ICSRs of ADRs without DDIs during the same period. These ICSRs were considered negative controls. The selected ICSRs used as negative controls had to include a suspected drug and at least one other concomitant or suspected medication. We randomly selected cases from these two datasets for both the positive and negative controls.
Each case of interaction was classified according to the type of interaction: pharmacokinetic (PK) or pharmacodynamic (PD). A difficulty level score was then assigned to each case, ranging from 1 to 2. Level-1 DDI was the easiest (e.g., interactions clearly mentioned in the Summary of Product Characteristics), which corresponded to level B4 of the French method for assessing ADRs. 17 Level-2 DDI was the most difficult (requiring crossing information from two Summary of Product Characteristics or requiring more in-depth bibliographic research to identify the interaction), which corresponded to levels B2 and B3 of the French method for assessing ADRs. 17
After selecting the cases, to avoid redundancy, duplicated reports (defined as reports involving the same drugs or the same type of DDIs) were excluded from the analysis.
LLMs choice
For each case, three LLMs were used: ChatGPT-4o, 18 Gemini 1.5 Flash, 19 and Claude 3.5 Sonnet. 20
We used ChatGPT as we had previously worked with this chatbot, and it was also evaluated in two studies on DDIs.8,10,11 Furthermore, according to the literature, ChatGPT-4 surpasses ChatGPT-3.5 in drug information queries. 21
As we wanted to test other LLMs, we chose to evaluate Gemini and Claude, while these LLMs are under-evaluated in medicine, they are considered two of the latest major LLMs. 22
Prompts and information given
We conducted a zero-shot prompting approach, where LLMs relied entirely on their pre-trained knowledge. This method was chosen as it could reflect the routine use of LLMs in pharmacovigilance practice, where pharmacologists would typically issue a single prompt for each case.
For the study, a case (positive or negative control) was randomly selected from the ICSRs extracted from the FNPV. For each LLM, a new conversation was initiated, and the same standardized introductory text was provided, followed by the case summary. This process was repeated for every case to ensure that a new conversation was opened for each instance, thereby eliminating potential biases within the LLMs. The introductory text was the following one (provided in French as the ICSRs are written in French): “You are a pharmacologist. I will give you pharmacovigilance cases, and you will tell me if (1) there are drug interactions involved in the case. If so, (2) what type of interaction (pharmacokinetic, pharmacodynamic), (3) which drugs are involved, and (4) what is the interaction mechanism?”
Regarding the case summary, based on the information obtained from the ICSRs, we provided the following details to the LLMs (when available): the patient’s gender and age, medical history, medications, a summary of clinical symptoms, relevant biological data for analysis, and the outcome following any potential intervention. The case summary did not include any indication of whether an interaction was present or not.
The LLMs were interrogated between July and September 2024.
Sensitivity, specificity, and F-measure
Based on the responses provided by the LLMs, we calculated the percentage of correct answers and their 95% confidence interval (95% CI) for the four questions asked: (1) are there drug interactions involved in the case? If so, (2) what type of interaction (pharmacokinetic, pharmacodynamic), (3) which drugs are involved, and (4) what is the interaction mechanism?
For negative controls, if the LLM identified an interaction, we considered it a false positive for all the questions asked, and conversely, if no interaction was found, it was a true negative. We calculated the sensitivity and specificity of each LLM regarding the drugs involved, the type of interaction, and the interaction mechanism.
To measure the predictive performance of LLMs, we also calculated the F-measure that varies from 0 to 1 (the highest possible value being 1, indicating perfect precision and sensitivity). For positive controls, we also calculated the percentage of correct answers according to the bibliographic score. The following formulas were utilized:
where FP is false positive, FN is false negative, NPV is negative predictive value, PPV is positive predictive value, TP is true positive, and TN is true negative. TP represents the number of correct answers for positive controls.
Results
Out of 137 ICSRs extracted that implicated DDI, 55 were excluded (due to redundancy in interacting drugs and/or interacting mechanisms), and 82 were analyzed. We also processed 22 ICSRs as negative controls. The 82 ICSRs included 31 PK interactions and 51 PD interactions. Thirty-seven ICSRs implicated level-1 DDI (29 ICSRs with PD interactions and 8 ICSRs with PK interactions) and 45 ICSRs implicated level-2 DDI (22 ICSRs with PD interactions and 23 with PK interactions). ChatGPT detected 99% (95% CI (98–99)) of DDIs cases. Claude and Gemini detected 95% (95% CI (90–99)) of DDI cases (Figure 1). ChatGPT and Claude performed comparably across all questions (including: are there drug interactions involved in the case; if so, what type of interaction (pharmacokinetic, pharmacodynamic); which drugs are involved; what is the interaction mechanism?) and better than Gemini. ChatGPT, Claude, and Gemini provided correct answers for all questions in 66% (95% CI (56–76)), 68% (95% CI (58–78)), and 33% (95% CI (23–43)) of ICSRs involving DDIs (Figure 1). ChatGPT and Claude showed higher performance (F-measure between 0.83 and 0.85 for the three questions) than Gemini (F-measure between 0.63 and 0.68 for the three questions; Table 1). Sensitivity was higher for ChatGPT and Claude than for Gemini. All chatbots presented a low specificity (with values of 0.68 for ChatGPT, 0.64 for Claude, and 0.36 for Gemini).

Percentage of correct answers by chatbot based on the type of question for ICSRs implicating drug-drug interaction.
Sensitivity, specificity, and F-measure based on the type of question and by chatbots.

FN, false negative; FP, false positive; NPV, negative predictive value; PPV, positive predictive value; TN, true negative; TP, true positive.
For all questions, the percentage of correct answers was higher for ICSRs with level-1 DDI than for ICSRs with level-2 DDI (Table 2). Furthermore, for level-2 difficulty, the percentage of correct answers regarding the detection of drugs involved in DDI, the type of interaction, and its mechanism was higher for PK interactions than PD interactions (Table 2). The number of PK interactions for level-1 DDI was too small to compare with PD interactions.
Correct answers by level of difficulty and type of interaction.

DDI, drug–drug interactions; PD, pharmacodynamic; PK, pharmacokinetic.
Discussion
This study is the first to evaluate the efficacy of LLMs to identify DDIs for pharmacovigilance purposes based on zero-shot prompting. By asking LLMs about the existence of an interaction, the drugs involved in the interaction, the type of interaction, and the mechanisms involved, we sought to assess their ability to analyze the interaction like a pharmacologist. We found that while LLMs are relatively effective at detecting the existence of DDIs leading to ADRs, their performance declines when analyzing these interactions. ChatGPT, Claude, and Gemini provided correct answers for all questions in 66%, 68%, and 33% of ICRs involving DDIs. ChatGPT and Claude were more accurate, sensitive, and specific than Gemini. All the LLMs presented a low specificity with 0.68 for ChatGPT, 0.64 for Claude, and 0.36 for Gemini. The percentage of correct answers decreased as the difficulty of identifying DDIs increased.
While ChatGPT, Claude, and Gemini showed relatively high sensitivity to detect clinically significant DDIs that led to an ADR, all LLMs, notably Gemini, showed poor specificity to exclude the involvement of a DDI in a case of ADR. Our results are in line with those of Al-Ashwal et al., which suggest high sensitivity but poor specificity to detect clinically significant DDIs compared with clinical tools (Micromedex and Drugs.com). 11 However, a recent study 12 contrasts with our results and those of Al-Ashwal et al. The authors found that ChatGPT-3.5 had a low sensitivity but a good specificity to detect DDI from prescribed medicines for 120 hospitalized patients. This difference could be explained by the fact that we used pharmacovigilance data containing only clinically significant DDIs that led to an ADR, and Al-Ashwal et al. used highly selective drug pairs that could induce clinically significant DDIs. The data used by Radha et al. concerned hospitalized patients in whom DDIs are not all clinically relevant.
In addition, from a pharmacologist’s perspective, the percentage of correct answers to all DDI-related questions was insufficient for all LLMs (below 70% for ChatGPT and Claude, and around 30% for Gemini). Contradictory information was also sometimes provided, particularly regarding the type of interaction (pharmacodynamic vs pharmacokinetic) and the mechanisms proposed by the LLMs. ChatGPT and Gemini performed better at explaining the interaction mechanisms than at identifying the type of interaction, despite the latter being a much simpler question. These findings highlight the lack of scientific reasoning in LLMs and the fact that the answers provided by the chatbots are likely independent of each other. These results are not unexpected, as LLMs function as next-word predictors, generating outputs based on the probability of which words are likely to follow, 23 and are not specifically designed to detect DDIs.
Finally, the percentage of correct answers decreased as the difficulty of identifying DDIs increased. LLMs answered better for difficult DDIs (level-2) regarding PK interactions. Our recent publication showed that ChatGPT’s performance declines with increasing task difficulty in drug information services. 8 We can speculate that PK interactions are better reported in the literature or easier to identify than PD interactions.
Our result also illustrates that chatbots in a zero-shot prompting condition are prone to “hallucinations,” meaning they may generate incorrect or fabricated information—in our case, inventing drug interactions to explain ADRs. Andrikyan et al. highlighted that chatbots could potentially provide harmful answers to patients’ questions about drug treatment in zero-shot prompting conditions. 24 In our specific field, hallucinations by LLMs could lead to misclassification of a pharmacovigilance case if LLMs are used without proper verification by a pharmacologist.
In conclusion, we believe that LLMs (in a zero-shot prompting condition) cannot yet be relied upon as tools for pharmacovigilance cases where humans make the final judgment on detected DDIs. LLM response will still require verification of the implicated drugs, the type of interaction, and the specific mechanism involved. These verifications could be time-consuming for a pharmacologist in daily practice. Future studies should test different prompting methods and their effect on the efficacy of LLMs in the field of DDIs. The development of small language models specifically trained in pharmacovigilance could also be relevant. 25 Finally, the daily and unreasonable use of LLMs raises the question of their environmental impact on electricity consumption and water.26,27
One of the main strengths of this study is the use of the FNPV. We analyzed over 100 ICSRs from real-world clinical scenarios, all of which had been previously validated and assessed by pharmacologists. Furthermore, we used several chatbots and demonstrated that they do not all have the same effectiveness. Finally, we used four different questions (are there drug interactions involved in the case; if so, what type of interaction; which drugs are involved; what is the interaction mechanism) to better characterize the performance of LLMs in detecting DDIs from pharmacovigilance cases.
Limitations
This study has several inherent limitations. We only evaluated three AI platforms—ChatGPT, Claude, and Gemini. Future studies should evaluate other chatbots, such as MedPalm or other open-source models like DeepSeek or Perplexity, in detecting DDIs. We used zero-shot prompting and not few-shot prompting, chain of thought, or reflection of thoughts prompting, while they could increase the results. 28 Furthermore, we did not use several prompts to assess how they can influence the results. We know that LLMs may generate different answers to the same question and can be inconsistent. 28 In this study, we wanted to evaluate the utility of LLMs for daily routine use in pharmacovigilance practice. We thus considered that zero-shot prompting was a reasonable option. As this is a pharmacovigilance study, we only evaluated the efficacy of chatbots on clinically significant DDIs. Also, with the constantly growing performance of LLMs, these results may no longer be valid in the coming years, notably if LLMs are specifically trained with reference textbooks.
Conclusion
LLMs may detect DDIs leading to pharmacovigilance cases. Nevertheless, they cannot rule out their involvement in pharmacovigilance cases without DDIs. Pharmacologists are crucial for assessing whether a DDI is implicated in an ADR.
Footnotes
Acknowledgements
The authors are indebted to the National Pharmacovigilance Centers that contributed data. The opinions and conclusions in this study are not necessarily those of the various centers or the ANSM (Agence Nationale de Sécurité du Médicament et des produits de santé, France).