Abstract
Nonalcoholic fatty liver disease (NAFLD) is becoming the most frequent chronic liver disease worldwide. Metabolic (dysfunction) associated fatty liver disease (MAFLD) is suggested to replace the nomenclature of NAFLD. For individuals with metabolic dysfunction, multiple NAFLD-related factors also contribute to the development and progression of MAFLD including genetics and epigenetics. The application of genome-wide association study (GWAS) and exome-wide association study (EWAS) uncovers single-nucleotide polymorphisms (SNPs) in MAFLD. In addition to the classic SNPs in PNPLA3, TM6SF2, and GCKR, some new SNPs have been found recently to contribute to the pathogenesis of liver steatosis. Epigenetic factors involving DNA methylation, histone modifications, non-coding RNAs regulations, and RNA methylation also play a critical role in MAFLD. DNA methylation is the most reported epigenetic modification. Developing a non-invasion biomarker to distinguish metabolic steatohepatitis (MASH) or liver fibrosis is ongoing. In this review, we summarized and discussed the latest progress in genetic and epigenetic factors of NAFLD/MAFLD, in order to provide potential clues for MAFLD treatment.
Keywords
Introduction
Nonalcoholic fatty liver disease (NAFLD), first named in 1980, is to describe fatty liver disease arising without significant alcohol consumption. 1 The spectrum of NAFLD ranges from simple steatosis (also called nonalcoholic fatty liver, NAFL) to nonalcoholic steatohepatitis (NASH) and even NAFLD-related cirrhosis, which can further develop into liver carcinoma.2,3 At present, NAFLD has become one of the most common causes of chronic liver disease around the world, with the global prevalence reaching 25%. 4 Usually, NAFLD arises in the context of obesity and its related metabolic diseases. 5 The prevalence of NAFLD in population with diabetes goes up to 55–70%, which is much higher than that in population without diabetes. 6 An approximately 60–70% increase in the risk of fatal and nonfatal cardiovascular disease was also observed in patients with NAFLD. 7 Our previous work including 211 subjects with liver biopsy-proven NAFLD showed that the proportion of NASH and advanced fibrosis in NAFLD patients with type 2 diabetes (T2DM) was 96.1% and 56.5%, respectively. 8 Thus, there is a close association of NAFLD with T2DM, obesity, and cardiovascular disease.9–12 Recently, an international panel of experts has suggested using metabolic (dysfunction) associated fatty liver disease (MAFLD) to replace the nomenclature of NAFLD,13,14 triggering intense academic debate. Meta-analysis reveals that the global prevalence of MAFLD is 38.77%, which is much higher than that of NAFLD. 15 With the more accurate reflection of the underlying pathogenesis and the superior use of the MAFLD definition for patient awareness and management, global multi-stakeholders from over 134 countries endorsed MAFLD definition, 16 which will be a landmark of MAFLD renaming process.
The classic two-hit theory was first proposed to explain the pathogenesis of MAFLD in 1998, 17 and the hepatic accumulation of lipids secondary to metabolic disorders acts as the first hit to the liver. As time goes on, plenty of evidence indicates that multiple factors contribute to the initiation and progression of fatty liver disease, including but not limited to insulin resistance, adipocyte dysfunction, nutritional factors, gut microbiota, and genetic and epigenetic factors, bile acid and nuclear receptor signaling, and inflammatory and immune signaling.18,19 With the progress of high-throughput sequencing technology, the application of genome-wide association study (GWAS), chromatin immunoprecipitation sequencing (ChIP-seq), RNA sequencing (RNA-seq), and other methods has uncovered the features of genetics and epigenetics in MAFLD to a large extent, which brings us a new comprehension on the effect of genetics and epigenetics on the initiation and progression of MAFLD. In this review, we summarized the latest progress in genetic and epigenetic factors of MAFLD, especially focusing on epigenetic factor that regulates MAFLD.
Genetics in MAFLD
The heritability of NAFLD was first reported in 2000. 20 Over the past decade, the application of GWAS in the NAFLD research has helped us accelerate the discovery of single-nucleotide polymorphisms (SNPs) associated with NAFLD. More than 10 SNPs linked to risk of NAFLD in patients with metabolic dysfunction have been found at present (e.g. PNPLA3, TM6SF2, MBOAT7) which has been summarized in Table 1. Some classic SNPs (e.g. PNPLA3 rs738409 C > G, TM6SF2 rs58542926 C > T, MBOAT7 rs641738 C > T, GCKR rs780094 A > G) have been replicated in multiple independent studies, the effect of which has been well explored. These SNPs also contributed to the development of liver steatosis in patients with metabolic dysfunction 21 (Figure 1). Some new SNPs linked to risk of MAFLD have been discovered in recent years. Although genetic risk of MAFLD in individuals have gained considerable advance, the effect and application of these SNPs has not yet been explored adequately, which need a future investigation.
The effects and applications of single-nucleotide polymorphism (SNP) associated with MAFLD.
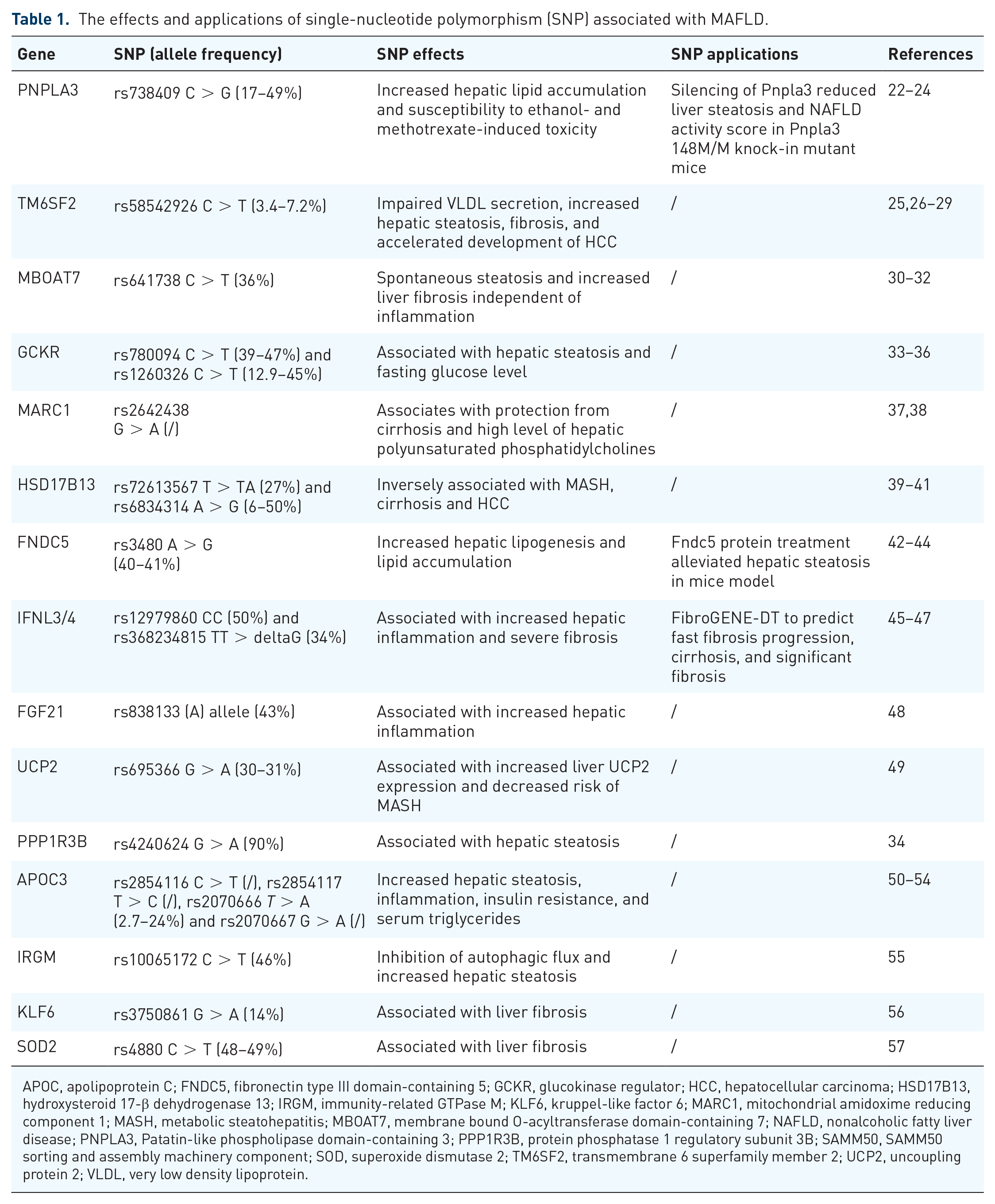
APOC, apolipoprotein C; FNDC5, fibronectin type III domain-containing 5; GCKR, glucokinase regulator; HCC, hepatocellular carcinoma; HSD17B13, hydroxysteroid 17-β dehydrogenase 13; IRGM, immunity-related GTPase M; KLF6, kruppel-like factor 6; MARC1, mitochondrial amidoxime reducing component 1; MASH, metabolic steatohepatitis; MBOAT7, membrane bound O-acyltransferase domain-containing 7; NAFLD, nonalcoholic fatty liver disease; PNPLA3, Patatin-like phospholipase domain-containing 3; PPP1R3B, protein phosphatase 1 regulatory subunit 3B; SAMM50, SAMM50 sorting and assembly machinery component; SOD, superoxide dismutase 2; TM6SF2, transmembrane 6 superfamily member 2; UCP2, uncoupling protein 2; VLDL, very low density lipoprotein.

The genetics and epigenetics of MAFLD. The genetics and epigenetics of MAFLD include gene polymorphisms, DNA methylation, histone modifications, non-coding RNAs regulations, and RNA methylation.
PNPLA3
The patatin-like phospholipase domain-containing 3 (PNPLA3) is located in the hepatic lipid droplets, which interferes with adipose triglyceride lipase activity by interacting with its cofactor, comparative gene identification-58 (CGI-58).
58
Since the identification of PNPLA3 as a modifier of NAFLD pathogenesis in the first reported NAFLD GWAS,
22
the PNPLA3 rs738409 C > G variant has been fully demonstrated to be the most robust genetic variant associated with NAFLD in populations from all ethnicities.59,60 This variant is also associated with metabolic steatohepatitis (MASH) and severe fibrosis in subjects with lean MAFLD.
61
Recent progress on the mechanism underlying the association between the PNPLA3 variant and fatty liver disease indicated that the accumulation of PNPLA3 variant (I148M) sequesters CGI-58 on hepatic lipid droplets, thus restricting its access to adipose triglyceride lipase and causing liver fat accumulation.
20
Human metabolomic study also found that the hepatic triglyceride export from the liver
TM6SF2
The transmembrane 6 superfamily member 2 (TM6SF2) is a membrane protein that located in the cellular endoplasmic reticulum membrane (ER) and the ER-Golgi intermediate compartment. 65 The TM6SF2 rs58542926 C > T variant was associated with hepatic steatosis and lower plasma cholesterol and triglycerides (TG) according to the exome-wide association study (EWAS) analysis in the multiethnic population from the Dallas Heart Study. 25 The TM6SF2 rs58542926 (T) allele was also associated with lean MAFLD, 66 reduced hepatitis C viral load, and increased hepatitis B viral load. 67 Knockdown of TM6SF2 in the mouse liver inhibited the lipidation of apolipoprotein B-containing lipoproteins, thus decreased hepatic VLDL-TG secretion and led to liver steatosis. 68 The TM6SF2 rs58542926 C > T variant was associated with increased risk of MAFLD, MASH, and advanced liver fibrosis, but protected against cardiovascular disease. 69 Consistent with the PNPLA3 variant, TM6SF2 variant promoted the development of MAFLD but not the fatty liver disease without metabolic dysfunction. 21
MBOAT7
The membrane-bound O-acyltransferase domain-containing 7 (MBOAT7), also known as lysophosphatidylinositol acyltransferase 1 (LPIAT1), is a protein with six transmembrane domains located in the endoplasmic reticulum membrane.
70
MBOAT7 has lysophospholipid acyltransferase activity that incorporates arachidonic acid into phosphatidylinositol (PI). The association between MBOAT7 rs641738 C > T variant and liver cirrhosis was first identified in GWAS of alcohol abusers.
71
Further analysis in individuals from the Dallas Heart Study and of European descent showed that the MBOAT7 variant carriers were associated with increased hepatic fat content, more severe liver damage, and increased risk of liver fibrosis compared with subjects without the variant.
30
Recent meta-analysis confirmed the association of MBOAT7 rs641738 C > T variant with hepatic fat content and MAFLD in Caucasian adults.
72
The MBOAT7 rs641738 variant was also associated with severe hepatic inflammation and increased risk of fibrosis in subjects with chronic hepatitis B or C.73,74 Liver-specific knockout of
MARC1
The mitochondrial amidoxime reducing component 1 (MARC1) is a protein located in the outer mitochondrial membrane. 78 Its function is to catalyze the reduction of N-oxygenated molecules in metabolism. 79 Recent study including 12,361 all-cause cirrhosis cases and 790,095 controls showed that a missense MARC1 variant (rs2642438 A > G) was associated with low liver fat content, low blood liver enzymes, and protection from all-cause cirrhosis. 37 Subsequent study found that MARC1 rs2642438 variant was associated with increased hepatic phosphatidylcholines and decreased steatosis, activity, and fibrosis (SAF) score in 369 subjects with liver biopsy. 38 The similar association between MARC1 rs2642438 variant and low risk of alcohol-related cirrhosis was also found in European cohorts. 80 However, the causal relationship between MARC1 variant and MAFLD is still unknown. Transgenic animal models are probably applied to further illustrate the function of MARC1 in the development and progression of MAFLD.
HSD17B13
Hydroxysteroid 17-β dehydrogenase 13 (HSD17B13) is expressed predominantly in the liver and is a liver-specific lipid droplet-associated protein.
81
Recently, studies have showed that a splice-site SNP in HSD17B13 (rs72613567 T > TA) and another SNP, rs62305723 (encoding a P260S mutation, considered loss of function variant), were significantly associated with decreased inflammation and ballooning in liver histology and low risk of MASH.39,82 This variant was also associated with low risk of alcohol-related hepatocellular carcinoma (HCC).83,84 It seems that there is a specific function of HSD17B13 in the development and progression of MAFLD. To explore the function of HSD17B13 in liver,
FNDC5
The membrane protein fibronectin type III domain-containing 5 (FNDC5) is a precursor of myokine, which is proteolytically cleaved in skeletal muscle to form irisin and secrete into blood.
86
Serum irisin level is negatively associated with liver triglyceride content and liver enzymes in obese subjects.
86
Irisin treatment induces browning and UCP1 expression of white adipose tissue, leading to increased energy expenditure.86,87 Studies found that the FNDC5 rs3480 A > G variant was associated with protection from significant fibrosis in patients with liver biopsy proved MAFLD.
88
Recent study including 987 Caucasian patients with MAFLD showed that the FNDC5 rs3480 variant was associated with advanced steatosis, but not inflammation or fibrosis,
42
and serum irisin was inversely associated with hepatic steatosis.
42
For the function of FNDC5,
IFNL3/4
The interferon lambda type 3 and 4 (IFNL3/4) is a cytokine that can be induced by viral infection and has an association with immune and inflammatory response. Studies revealed that IFNL4 rs12979860 and rs368234815 variants were associated with increased liver inflammation and severe fibrosis in Europeans with MAFLD.45,46 Based on the association with IFNL genotype and hepatic fibrosis, a prediction model (FibroGENE-DT) of fast fibrosis progression, cirrhosis, and significant fibrosis was developed and performed well in MAFLD subjects.
47
For the mechanism, the expression of downstream of IFNL4, including interferon regulatory factor 9 (IRF9), tyrosine kinase non receptor 1 (TNK1), signal transducer and activator of transcription 2 (STAT2), interleukin 10 receptor subunit beta (IL10RB), and tyrosine kinase 2 (TYK2), was reduced in subjects with IFNL4 rs368234815 TT/TT and rs12979860 CC, indicating that IFNL4 variant may affect hepatic inflammation and fibrosis
FGF21
Fibroblast growth factor 21 (FGF21) is a hepatokine that regulates metabolic homeostasis. However, whether FGF21 has beneficial effect on metabolism is controversial. 90 Recent study revealed that FGF21 rs838133 variant was associated with elevated serum FGF21 level and increased hepatic inflammation in subjects with MAFLD. 48 For the mechanism, FGF21 rs838133 minor allele increased FGF21 translation and protein stability, leading to elevated serum FGF21 level and subsequent liver inflammation. 48
CNV
The role of copy number variation (CNV) in MAFLD should not be ignored. CNV is a common form of genetic alterations ranging from 1 kb and up to the megabase scale. CNV is quite different from SNPs in size and mutation rate. An increasing body of evidence reveals that CNV can affect gene expression and is a significant factor of human diseases. However, current studies about CNV and MAFLD are limited. The first study about CNV and MAFLD was reported in 2014, showing that four rare and novel CNVs (12q24.33, 13q12.11, 21p11.1–11.2, and 12q13.2) were associated with MAFLD in 49 patients with NAFLD and 49 controls.
91
Subsequent study enrolled 646 Caucasian subjects with biopsy-proven MAFLD and 170 healthy controls confirmed that the exportin 4 (XPO4) CNV was associated with hepatic fibrosis. Functional experiments showed that XPO4 CNV reduced liver expression of XPO4, resulting in hepatic stellate cells (HSC) activation and fibrosis
Epigenetics in MAFLD
Genetics can only partially explain the epidemiological characteristics of MAFLD, suggesting that other hereditary factors may also play a role. Epigenetics is a study that explores the mechanism in which phenotypes are changed by non-DNA sequence variation, including DNA methylation, histone modifications, non-coding RNAs regulations, and, in recent years, RNA methylation. Over the last decade, epigenetics has become an important field to study physiology and pathophysiology of fatty liver disease. The association between epigenetics and MAFLD in humans has been well established.19,93 The interaction between epigenetics and environment, diet, aging, gut microbiota, and so on indicates an intriguing and complex role of epigenetic modification in MAFLD. The characteristics of epigenetics in MAFLD help us understand the development and progression of fatty liver disease from a different aspect and give us an opportunity to predict or even diagnose metabolic associated fatty liver (MAFL)/MASH. It also brings a new way to develop treatment for MAFLD. In this part, we focused on recent advance in the epigenetics of MAFLD including the characteristics and function of epigenetic modification in MAFLD and the application of epigenetics in prediction and treatment of MAFLD (Figure 1).
DNA methylation
The association between DNA methylation and MAFLD has been well established.94,95 Study including 56 subjects showed that a total of 69,247 differentially methylated CpG sites in the advanced liver fibrosis group compared with that in the mild liver fibrosis group (histologic fibrosis: F0–1 mild and F3–4 advanced). 94 The advanced MAFLD group displayed hypomethylated and overexpressed genes relative to tissue repair genes (e.g. transmembrane protein 204 TMEM204, fibroblast growth factor receptor 2 FGFR2) and increased expression of hypomethylated genes compared with the mild liver fibrosis group. 94 Another study found that nine differentially methylated genes (e.g. putative polypeptide N-acetylgalactosaminyltransferase-like protein 4 GALNTL4, ATP citrate lyase ACLY, glutamate receptor delta-1 GRID1, insulin-like growth factor binding protein 2 IGFBP2, phospholipase C-gamma-1 PLCG1, protein kinase C epsilon PRKCE, insulin-like growth factor 1 IGF1, inositol hexaphosphate kinase 3 IP6K3, pyruvate carboxylase PC) in the normal control, healthy obese, simple steatosis, and steatohepatitis groups. The methylation at the loci of homeobox B1 (HOXB1), protein kinase C zeta (PRKCZ), solute carrier family 38 member 10 (SLC38A10), and secreted and transmembrane 1 (SECTM1) could be reversible after bariatric surgery. 96 The association between hepatic DPP4 methylation and fatty liver disease was also reported. 97
Because liver biopsy is an invasive operation, it is hard to obtain the liver tissue in clinical practice. Therefore, a method of plasma cell-free DNA methylation is developed. It is reported that circulating DNA methylation could be detected in human plasma and hypermethylation at the PPARγ promoter of plasma DNA was associated with the methylation in hepatocellular DNA, suggesting that plasma cell-free DNA methylation measurement could be a potential non-invasive biomarker to distinguish liver fibrosis severity.95,98 But the DNA methylation model still needs to be verified in other large cohorts. Subsequent study using peripheral blood DNA in Europeans found that hypomethylation of cg08309687 (LINC00649) was associated with MAFLD and risk for new-onset type 2 diabetes mellitus (T2DM). 99 However, DNA methylation is dynamic and can be affected by environment, diet, aging, gut microbiota, and so on. It is hard to keep a stable DNA methylation condition at present. We are not sure whether there might be some stable DNA methylation loci that are only affected by environmental factors. Thus, it is still a long way to go to use DNA methylation for distinguish MAFLD.
DNA methylation has been proposed as a measure of human chronological and biological age, also called the ‘Horvath clock’, in recent years.100–102 The principle of DNA methylation age is an unbiased assessment of aging by calculating a predicted aging rate based on methylation patterns. If a DNA methylation age exceeds the chronologic age, it would suggest age acceleration. The deviation between DNA methylation age and chronologic age is usually associated with health issues, which is helpful to see health status. Recent study showed that DNA methylation age acceleration was existed in MASH subjects compared with controls and was associated with hepatic collagen content. 103
Histone modification
Histone is one of the basic structures of nucleosome, which assembles with DNA base pairs. There are four kinds of histones including H1, H3, H2, and H4 according to their molecular weight. For the structure of histone, an N-terminal amino acid tail on histone can be post-translational modificated including methylation, acetylation, phosphorylation, ribosylation, sumoylation, ubiquitination, and so on. The post-translational modification of histone affects multiple physiological processes (e.g. gene transcriptional regulation, alternative splicing, DNA repair, chromosomal condensation). However, inappropriate histone modification causes organism dysfunction and disease. The association between histone modification with MAFLD is established and the cause effect of histone modification in MAFLD is also explored.
Histone acetylation is the most reported histone modification in MAFLD at present. Cluster of differentiation 36 (CD36) is a fatty acid translocase that mediates fatty acid uptake in hepatocytes.
104
Recent study has demonstrated that nuclear receptor subfamily 2, group F, member 6 (NR2F6) directly bound to the CD36 promoter, recruited nuclear receptor coactivator 1 (SRC-1), and promoted histone acetylation at CD36 promoter in hepatocytes, leading to increased hepatic fat accumulation.
105
Another nuclear receptor, liver X receptor α (LXRα), is also involved in the pathophysiology of MAFLD. Impaired Ser196 (S196A) LXRα phosphorylation induced hepatic steatosis, but decreased hepatic inflammation and fibrosis in mice on a high cholesterol diet by histone acetylation (H3K27) and transcriptional regulation in pro-fibrotic and pro-inflammatory genes.
106
Histone acetylation also mediated the transcriptional regulation of fatty acid synthase (FASN) by carbohydrate responsive element binding protein (ChREBP)
The histone methylation is also involved in the pathophysiology of MAFLD. The histone demethylase, plant homeodomain finger 2 (Phf2), was co-recruited with ChREBP on stearoyl-CoA desaturase 1 (SCD1) promoter and promoted H3K9me2 demethylation at the SCD1 promoter, resulting in increased
However, little is known about the effect of histone phosphorylation, ribosylation, sumoylation, and ubiquitination in the development and progression of MAFLD at present, which should be further investigated.
Non-coding RNA
The non-coding RNAs (ncRNAs) are transcribed from the genome and perform its biological functions at the RNA level without being translated into proteins. Several ncRNAs have been reported including transfer RNA, ribosomal RNA, microRNA (miRNA), small nucleolar RNA, long ncRNA (lncRNAs), and recent circular RNA (circRNA). Evidence has suggested that ncRNA plays an important role in mediating the development and progression of MAFLD.
MicroRNA
miRNA is a class of ncRNA encoded by endogenous genes with a length of about 22 nucleotides. miRNAs that are associated with MAFLD have been founded. miRNA-223 global knockout mice on a HFD displayed increased hepatic steatosis, inflammation, fibrosis, and easier to develop into HCC.
111
Meanwhile, myeloid-specific IL-6 knockout in mice downregulated miRNA-223 expression in macrophages and promoted profibrotic gene expression in hepatocytes through exosomal transfer, worsening liver fibrosis.
112
miRNA-192-5p from lipotoxic hepatocyte-derived exosomes induced M1 polarization of macrophages and liver inflammation, resulting in the progression of MAFLD.
113
In addition of miRNA from exosomes, miRNA from hepatocyte-derived extracellular vesicles (EVs) also regulates MAFLD progression. miRNA-9-5p from hepatocyte-derived EVs induced M1 polarization of macrophages and inflammation.
114
Knockdown of miRNA-9-5p in mice reduced hepatic steatosis and inflammation.
114
Endoplasmic reticulum (ER) stress is one of the mechanisms that promotes MAFLD. Study showed that liver-specific deficiency of microRNA-26a in mice promoted HFD-induced ER stress and hepatic steatosis.
115
Mechanistically, miR-26a exerted its effect by directly targeting the eukaryotic initiation factor 2α (eIF2α).
115
miRNA-378 is also involved in the development of steatosis. Knockdown of miR-378
Recently, circulating microRNAs have been applied as biomarkers to diagnose and prognose MAFLD. Four serum miRNAs (miR-21-5p, miR-151a-3p, miR-192-5p, and miR-4449) were used to distinguish MASH from MAFL. 130 The area under the receiver operating characteristic curve (AUROC) value of four miRNAs for MASH diagnosis was 0.875, which had been confirmed in another 37 subjects with MAFLD. 130 However, the subject number is relatively small in this study, the result of which should be validated in a large population. Another study using a composite biomarker of miR-192, miR-21, miR-505, and alanine aminotransferase (ALT) to predict MASH in mice and the AUROC could be up to 0.913. 131 A study including 441 subjects with liver biopsy confirmed MAFLD showed that serum miR-122 levels was a significant risk factor of mortality, which could predict mortality of subject with MAFLD. 132
Long non-coding RNA
lncRNA is a non-coding RNA with a length greater than 200 nucleotides, which plays a significant role in epigenetic regulation, cell cycle, and cell differentiation. The association between lncRNA and MAFLD has been established.133–136 lncRNA (RP11-484N16.1), screened from sequencing of human liver, was associated with hepatic steatosis and MASH. Knockdown of IncRNA (RP11-484N16.1) inhibited cell growth and viability in hepatocyte cell lines. However, whether IncRNA (RP11-484N16.1) mediated liver steatosis, inflammation, and fibrosis or not is still unknown.
135
Studies showed that lncRNA Blnc1 mediated hepatic lipogenesis and the development of MASH. Liver-specific Blnc1 knockout mice on a HFD displayed reduced hepatic steatosis and insulin resistance and were resistant to MASH by inhibiting hepatic lipogenesis.
137
Recent study has reported that lncRNA Platr4 had a beneficial effect on liver inflammation. Platr4 knockdown reduced liver inflammasome activity and was much easier to develop MASH in mice on a methionine-choline-deficient diet, whereas overexpression of Platr4 reversed it.
138
lncRNA HULC was reported to regulate MAFLD-associated fibrosis. HULC siRNA injection
In addition, studies
Circular RNA
Study about circRNA and MAFLD is rare. circRNA expression profiles are changed in MAFLD, indicating that circular RNA may participate in the development and progression of liver steatosis.148,149 Knockdown of circScd1 in liver cell line AML12 promote hepatic fat content, while overexpression of circScd1 attenuated steatosis, suggesting that circScd1 mediated hepatic steatosis.
150
circRNA_0046366 increased hepatocellular FAO
Conclusion and future perspective
There is no doubt that genetics and epigenetics play a critical role in development and progression of MAFLD. The progress of technology helps us further uncover the detail of genetics and epigenetics on MAFLD. In addition to the classic SNPs in MAFLD, some new SNPs (e.g. MARC1, HSD17B13, FNDC5) have been reported, but their function on the development and progression of MAFLD requires further investigated. DNA methylation is the epigenetic modification mostly studied, whereas studies about m6A modifications or circRNA in MAFLD are still rare. As to the histone modification, the effect of phosphorylation, ribosylation, sumoylation, and ubiquitination should be further explored. Recently, a new concept of the shared genetic basis between MAFLD and other metabolic diseases has been proposed based on mounting evidence.90,153 In this situation, collaboration and partnerships between stakeholders need to be further strengthened to provide optimal care for patients with MAFLD and associated metabolic diseases 153 (Figure 2).

The perspectives of MAFLD based on the findings of genetics and epigenetics.
The potential translational impact of genetic and epigenetic findings on drug development and non-invasive biomarkers is also of great concern. Although the pathophysiology of MAFLD has been widely explored, no efficient drug for MAFLD is available at present. However, the genetics and epigenetics of MAFLD have a potential to identify new drug targets or opportunities for drug repurposing, elucidate pharmacological mechanism, and predict adverse events and a therapeutic window for new therapies 154 (Figure 2). In addition, the dynamic change of epigenetics makes it as a potential biomarker for disease diagnosis, monitoring, and treatment response 155 (Figure 2). Although the translational application of genetic and epigenetic findings is still in its infancy, this area will bring us the hope to conquer MAFLD in the future.
