Abstract
Intervertebral disc degeneration (IDD) is a common condition and a leading cause of chronic low back pain, affecting millions of individuals worldwide. Human Umbilical Cord Mesenchymal Stromal Cell (hUCMSC)-derived extracellular vesicles (EVs) are emerging as a promising therapeutic strategy for IDD. However, the limited production yield and unclear mechanisms by which EV contents mediate their therapeutic effects have hindered the clinical application of EVs. In this study, using transcriptomic data and single-cell RNA sequencing, we identify PANoptosis as a key mechanism driving the progression of IDD. Furthermore, parathyroid hormone (PTH) enhances the secretion of hUCMSC-derived EVs and alters their cargo composition, which may contribute to their improved therapeutic effects. Mechanistically, PTH-preconditioned EVs, enriched with ZDHHC5, ameliorate PANoptosis by modulating ZBP1 transcription through competitive inhibition of YBX1 phosphorylation via palmitoylation. Our findings provide strong support for a cell-free therapeutic strategy utilizing EVs from PTH-preconditioned MSCs for IDD treatment and propose the ZDHHC5/YBX1/ZBP1 axis as a novel molecular target for inhibiting PANoptosis, thus paving the way for clinical translation and broader healthcare applications.
Introduction
Intervertebral disc degeneration (IDD) is a prevalent condition and a leading cause of chronic low back pain, affecting millions of individuals worldwide. The global impact of chronic low back pain is substantial, ranking as a top cause of years lived with disability (YLDs), with the prevalence projected to rise significantly as populations age.1,2 In recent years, the prevalence of back pain has also been rising among adolescents, further highlighting IDD as a significant public health concern with both physical and socioeconomic implications. 3 The nucleus pulposus (NP) cells, located at the center of the intervertebral disc, are essential for maintaining the disc’s biomechanical properties by producing an extracellular matrix (ECM) rich in proteoglycans and collagen. During intervertebral disc degeneration (IDD), NP cells lose the self-repair capacity and undergo regulated cell death (RCD), resulting in ECM degradation and disc collapse.4,5 Inflammatory responses, driven by cytokines such as IL-1β and TNF-α, further accelerate this process, creating a cycle of cell death and matrix breakdown.6,7 This continuous degradation reduces disc hydration and elasticity, impairing its shock-absorbing function and leading to pain and reduced mobility. 8 Thus, a combined approach addressing multiple RCD pathways may offer a more effective strategy for treating IDD.
PANoptosis is a newly recognized form of regulated cell death that integrates apoptosis, necroptosis, and pyroptosis, all of which play pivotal roles in IDD.9,10 Apoptosis, a programmed form of cell death, contributes to reducing the cell population responsible for homeostasis and triggering catabolic signaling cascades, which leads to ECM breakdown, and reduced disc integrity, thus exacerbating degeneration despite its non-inflammatory nature. 11 In contrast, necroptosis, an inflammatory cell death pathway regulated by RIPK1 and RIPK3, leads to cell membrane rupture and amplifies inflammation, accelerating ECM degradation and disc destruction. 12 Pyroptosis, driven by caspase-1 and gasdermin D, releases pro-inflammatory cytokines, further sustaining the chronic inflammatory environment and contributing to IDD progression. 13 The convergence of these pathways in PANoptosis creates a vicious cycle of inflammation and cell death, which accelerates degeneration. Given the limited success of therapies targeting individual pathways, jointly regulating these mechanisms may offer a more effective approach for protecting nucleus pulposus cells, slowing IDD progression, and enhancing therapeutic outcomes.14,15
Many studies have highlighted the promising potential of stem cell-derived extracellular vesicle-based therapies in the treatment of musculoskeletal disorders.16–20 Mesenchymal stromal cell (MSC)-extracellular vesicles (EVs) have been demonstrated to provide similar reparative effects in IDD as MSCs, while offering additional advantages such as low immunogenicity, enhanced product stability, and greater feasibility for clinical applications. 21 Although these advantages of EVs have garnered significant research interest in the academic community, their efficacy in disc repair remains lower than that of MSCs, highlighting the need for further improvement. 22 Previous studies have shown that when parent MSCs are exposed to various environmental stimuli or drug treatments, the bioactive content of their extracellular vesicles, including mRNAs, non-coding RNAs, proteins, and lipids, undergoes changes.23,24 Recent research has reported that engineered EV delivery systems target the subchondral bone for osteoarthritis treatment and modulate inflammation to support disc repair.25,26 These alterations enhance the EVs’ biological properties, enabling them to achieve more potent therapeutic effects.
Parathyroid hormone (PTH), produced by the chief cells of the parathyroid glands, is a key regulator of calcium and phosphate metabolism, playing a crucial role in maintaining calcium balance through its effects on bones and kidneys.27,28 Intermittent PTH injections have been shown to enhance intervertebral disc height and mitigate degeneration by activating TGF-β pathway, thereby promoting ECM integrity. 29 Additionally, studies have demonstrated that PTH improves nucleus pulposus cell function, enhances ECM synthesis, and reduces inflammation, oxidative stress, and apoptosis through activation of Sonic Hedgehog (SHH) signaling pathway. 30 Notably, PTH-preconditioned bone marrow multipotent mesenchymal stromal cell-derived exosomes have shown superior effects on cell proliferation and ECM formation in osteoarthritis chondrocyte repair. 31 Based on this, we hypothesize that PTH-preconditioned human umbilical cord mesenchymal stromal cells (hUCMSC)-derived extracellular vesicles (PTH pre-EVs) may have distinct protein profiles and offer enhanced therapeutic efficacy compared to unconditioned hUCMSC-derived EVs (un-EVs) in treating IDD.
In this study, we reported that PANoptosis contributed to the pathogenesis of IDD. We found that both PTH pre-EVs and un-EVs inhibit PANoptosis in NP cells, while PTH pre-EVs exhibited stronger anti-PANoptosis properties compared to untreated hUCMSC-derived EVs. Moreover, using LC/MS, we discovered that ZDHHC5, which was highly concentrated in PTH pre-EVs, could be transported to NP cells via EVs, effectively suppressing PANoptosis both in vitro and in vivo. Mechanistically, ZDHHC5 palmitoylated YBX1, competitively inhibiting its phosphorylation and subsequently preventing its nuclear translocation. This, in turn, downregulates ZBP1 expression at the transcriptional level, effectively inhibiting PANoptosis in NP cells. The findings from our study provided evidence that PTH enhanced the therapeutic effect of hUCMSC-derived EVs by inhibiting PANoptosis in NP cells. This research aimed to identify therapeutic targets and elucidate the underlying signaling pathways, presenting opportunities for clinical translation and potential applications in clinical practice.
Materials and methods
hUCMSCs culture and treatment
The hUCMSCs, generously donated by Dr. Ya-nan Pu and sourced from Cyagen Biotechnology Inc. (Catalog No. HUXUC-01001, Cyagen Biotechnology Inc., Guangzhou, China), underwent identification procedures as outlined in the Supplemental Methods and Results, following the MISEV2014 guidelines. 32 The hUCMSCs were cultured according to the manufacturer’s instructions. Briefly, hUCMSCs were expanded in OriCell® Medium, which contains OriCell® Basal Medium For Cell Culture (BLDM-03011) and OriCell® Fetal Bovine Serum Without EXO + Culture Supplement (For Human Umbilical Cord Mesenchymal Stem Cells; HUXUC-05002). The cells were maintained at 37°C in a humidified atmosphere of 5% CO₂. Seeding density for hUCMSC expansion was 2.5 × 10⁴ cells/cm². Culture media were changed every 2–3 days, and the cells were passaged upon reaching approximately 80%–90% confluence using 0.25% trypsin-EDTA. hUCMSCs were pretreated with PTH (10 nM) for 6 h, then the medium was replaced, and the medium was collected after 48 h. The differentiation potential of hUCMSCs was evaluated by inducing differentiation into osteogenic, adipogenic, and chondrogenic lineages. The characterization of hUCMSCs were analyzed by flow cytometry for the expression of CD73, CD90, and the absence of CD14, CD19, CD34, CD45, and HLA-DR, according to the guidelines. 32
hUCMSC-EVs isolation and identification
The hUCMSC-EVs were prepared through culturing in serum-free medium for 24 h at 37°C and 5% CO2. Subsequently, the supernatants were centrifuged at 300 × g for 10 min to remove cells. The supernatants were centrifuged at 2000 × g for 20 min and subsequently filtered through a 0.22 μm filter to eliminate cell debris. The filtered supernatant was then ultracentrifuged at 100,000 × g and 4°C for 90 min using a SW32Ti Beckman rotor. The pelleted EVs were re-suspended in phosphate-buffered saline (PBS) and ultracentrifuged again at 100,000 × g for 90 min (Beckman Coulter Optima L-100 XP ultracentrifuge). The final pellet was re-suspended in PBS, divided into aliquots to avoid repeated freeze-thaw cycles, and stored at -80°C for subsequent use. In accordance with the instrument’s guidelines, the morphology and size distribution of hUCMSC-EVs were determined by transmission electron microscopy (TEM), while particle size distribution was measured using the Zetasizer Nano ZS. Nanoparticle tracking analysis of hUCMSC-EVs was conducted using the NanoSight NS300. The protein markers of EVs (CD63, CD73, and CD90), as well as the hUCMSC marker (calreticulin), were analyzed by Western blot.
Extracellular vesicles uptake assay
The hUCMSC-EVs were labeled with CM-DiI (red fluorescent dye, 1 μM) for 30 min. Subsequently, the excess dye was removed by ultracentrifugation at 100,000 × g at 4°C for 90 min using a SW32Ti Beckman rotor, followed by two washes (Beckman Coulter Optima L-100 XP ultracentrifuge). NP cells were then incubated with CM-DiI-labeled EVs for 2 h, followed by fixation with 4% paraformaldehyde (PFA) at room temperature for 10 min. The cells were then washed three times with PBS. NP cell nuclei were stained with DAPI (4′,6-diamidino-2-phenylindole) for 5 min at room temperature. All reagents were obtained from Invitrogen (Carlsbad, CA, USA). Images depicting EV uptake were obtained using a Nikon Eclipse Ti confocal laser scanning microscope.
Human NP tissue collection and cell culture
The study strictly followed the guidelines outlined in the Declaration of Helsinki and was approved by the Institutional Review Board of the First Affiliated Hospital of Nanjing Medical University (2024-SR-170). For the purposes of histological staining, RNA, and protein extraction, NP tissue samples were obtained from patients diagnosed with lumbar disc herniation who underwent procedures including endoscopic discectomy, decompressive laminectomy, and lumbar interbody fusion. For the purposes of cell culture, NP tissues obtained from patients with MRI T2-weighted images rated as Pfirrmann grades I or II were specifically selected. A comparison of sociodemographic and clinical characteristics between Control and IDD groups was provided in Supplemental Tables S1 and S2. The procedures for human NP cell isolation were conducted following the ORS Spine Section consensus guidelines. 33 When reaching 80%–90% confluence, the cells were trypsinized, seeded into various culture plates, and maintained in a 37°C, 5% CO2 incubator. Human NP cells within passages 2–4 were used in this study.
Animal model of IDD
Approval for animal use was obtained from the Institutional Animal Care and Use Committee of Nanjing Medical University (IACUC-2402011). Male C57BL/6 mice, 8 weeks old and weighing between 23 and 27 g, were maintained under a 12-h light/12-h dark cycle and housed at the Experimental Animal Center of Nanjing Medical University. During the study period, food and water were provided ad libitum. The mice were anesthetized using 3% isoflurane for induction and 1.5% isoflurane for maintenance under aseptic conditions. The Co6/7 and Co7/8 intervertebral discs (IVDs) were located by palpation of the coccygeal vertebrae and confirmed through trial radiography. Once located, the corresponding IVDs (Co6/7 and Co7/8) were marked. A 26G needle was then used to completely penetrate the IVD under a stereo microscope, followed by a 180° rotation and held in place for 10 s. A 26 G needle was used to completely penetrate the intervertebral disc of the Co6-7 and Co7-8 tail, followed by a 180° rotation, and held in place for 10 s. Mice without any intervention were designated as the Control group. Animals subjected to needle puncture were randomly assigned to the following groups: the Needle Puncture group (treated with PBS, 1 μL), the EVs group (treated with un-EVs (1 μL, 5 μg/μL per segment)), the PTH pre-EVs group (treated with PTH pre-EVs (1 μL, 5 μg/μL per segment)), the PTH pre-EVs-si-ZDHHC5 group (treated with EVs derived from hUCMSCs transfected with ZDHHC5 siRNA (1 μL, 5 μg/μL per segment)), and the PTH pre-EVs and AAV9-shZBP1 group (treated with PTH pre-EVs (1 μL, 5 μg/μL per segment) and AAV9-shZBP1 (1.5 μL, 3 × 10¹² vg/mL)). Subsequent weekly EV injections were administered at the marked site of the caudal vertebrae and confirmed by palpation. The mice were euthanized by intraperitoneal injection of 1% sodium pentobarbital at 150 mg/kg after 4 weeks, and intact Co6-7 and Co7-8 tail samples were collected for further study.
X-ray and MRI analysis
Before radiology evaluation, mice were euthanized by inhaling an excessive amount of isoflurane. Radiographs were taken (exposure time, 5 s; distance, 520 cm; current, 180 μA; voltage, 40 kV) using the Panther system (X-viewer, PINGSENG Healthcare). Prone-position MRI was performed using a 3.0 T system (Biospec 7T/20 USR) to obtain T2-weighted images (TR, 3000 ms; TE, 45 ms; FA, 90°; field of view, 35 mm × 30 mm). The Pfirrmann grading system was employed to evaluate the degree of disc degeneration by MRI scan. The disc height index (DHI) was calculated by averaging values from the posterior, middle, and anterior parts of the disc and dividing by the average height of the adjacent vertebral body. 34
Cell counting kit-8 (CCK8) assay
Cell viability was assessed using Cell Counting Kit-8 (CK04, DOJINDO, Shanghai, China). Ninety-six-well plates were seeded with human NP cells. The cells were then incubated in complete culture medium with various reagents for 24, 48, and 72 h following cell attachment. CCK8 assays were conducted to measure optical density (OD) values at 0-, 24-, 48-, and 72-h time points.
Flow cytometric analysis of apoptosis
Apoptosis was analyzed using the Annexin V-FITC Apoptosis Detection Kit (C1062L, Beyotime, Shanghai, China) according to the manufacturer’s instructions. Briefly, after the human NP cells were treated for 24 h, they were stained with FITC-conjugated Annexin V and propidium iodide (PI), followed by flow cytometric analysis. The percentages of early and late apoptotic cells were quantified based on Annexin V and PI staining patterns. Data were analyzed using flow cytometry software.
ELISA for IL-1β and IL-18 detection
The concentrations of IL-1β and IL-18 in culture supernatant were measured using ELISA kits (RK00001 and RK00176, ABclonal, Wuhan, China) following the manufacturer’s protocol. Briefly, after the human NP cells were treated for 24 h, 100 μL of standards and samples were added to a 96-well plate and incubated at 37°C for 2 h. After washing, biotin-conjugated secondary antibodies were added and incubated for 1 h, followed by another washing step. Horseradish peroxidase (HRP)-streptavidin solution was added and incubated for 30 min. After washing, 100 μL of TMB substrate was added, and the reaction was stopped with stop solution after 15 min. The absorbance measured at 450 nm was adjusted by subtracting the background absorbance recorded at 630 nm and concentrations were calculated using a standard curve.
Terminal deoxynucleotidyl transferase dUTP Nick-end labeling (TUNEL) assay for apoptosis detection
Cell apoptosis was detected using the One Step TUNEL Apoptosis Assay Kit (C1088, Beyotime, Shanghai, China) according to the manufacturer’s instructions. Briefly, after the human NP cells were treated for 24 h, they were fixed in 4% paraformaldehyde at room temperature for 30 min, followed by permeabilization with 0.3% Triton X-100 for 5 min. After washing with PBS, cells were incubated with the TUNEL reaction mixture at 37°C for 1 h in a humidified chamber. After incubation, the cells were washed with PBS and counterstained with DAPI (4′,6-diamidino-2-phenylindole) to visualize cell nuclei. TUNEL-positive cells, indicating DNA fragmentation, were observed using a fluorescence microscope.
Cell transfection and adeno-associated virus (AAV)-mediated gene downregulation
Si-RNAs, Si-NC, or plasmids were transfected into human NP cells or hUCMSCs using Lipofectamine 3000 (L3000001, Invitrogen, Shanghai, China) or X-treme GENE HP DNA Transfection Reagent (6366244001, Roche, Shanghai, China) in OPTI-MEM medium. After 6 h of transfection, the medium was replaced with complete medium, and the cells were cultured for an additional 24–48 h. AAV9-mediated ZBP1 knockdown vectors were provided by GeneChem (Shanghai, China). AAV9-shZBP1 was administered via intervertebral disc injection. The siRNA and shRNA sequences were shown in Supplemental Tables S3 and S4, respectively.
Bioinformatics analysis: Single-cell sequencing data and bulk transcriptome sequencing data processing
For the single-cell data processing of the “normal→degenerative” and “moderate→severe” trajectories, the analysis data sets from multiple public datasets were integrated. All collected samples were explicitly derived from intervertebral disc tissues (excluding non-disc tissues) to ensure tissue specificity. Among them, the “normal→degenerative” trajectory was derived from GSE167056, GSE205535, and GSE233666, and after integration, it included four normal intervertebral disc nucleus tissue samples and five degenerative intervertebral disc nucleus tissue samples. The “moderate→severe” trajectory was derived from GSE244889, which included three moderately degenerative intervertebral disc nucleus tissue samples and three severely degenerative intervertebral disc nucleus tissue samples. We used the Seurat package to perform a workflow analysis of the single-cell data: after filtering out low-quality cells, we used the sctransform function to normalize and standardize the count matrix. We then ran the RunPCA, FindNeighbors, FindCluster, and RunUMAP functions on the default parameters. We selected the first 20 principal components for further UMAP visualization. To address batch effects between different cohorts, we applied batch correction using the Harmony package with the number of principal components used set to 50. Following dimensionality reduction and clustering, cell annotation was performed with strict criteria: nucleus pulposus cells were defined by co-expression of COL2A1 and ACAN, based on their unambiguous tissue origin (nucleus pulposus rather than annulus fibrosus or endplate tissues).
For transcriptomics data, we also obtained high-confidence datasets from the GEO database for human transcriptomics sequencing data (GSE266883 and GSE245147). The differential analysis was performed by limma and DEseq2, and the enrichment analysis was performed by Metascapes. The visualization was completed by the ggplot2.
Extracellular vesicles proteomics and bioinformatics analysis
Proteins were denatured using TCEP buffer containing protease and phosphatase inhibitors. After high-temperature incubation and overnight trypsin digestion, peptides were extracted and concentrated via centrifugal evaporation. Dried peptides were reconstituted in 0.1% formic acid and desalted using a C18 column. Peptides were separated using an EASY-nLC 1200 system with nanoscale flow rate. The mobile phases were Phase A (0.1% formic acid in water) and Phase B (0.1% formic acid in 80% acetonitrile). Columns (2 cm C18 pre-column, ID 100 and 3 μm; 30 cm C18 analytical column, ID 150 and 1.9 μm) were equilibrated with 100% Phase A. Separation was carried out at 600 nL/min with a gradient: 6% Phase B for 8 min, increasing to 12% over 8 min, from 16 to 108 min increasing from 13% to 30% Phase B, followed by 45% Phase B for 10 min, then 100% Phase B for 1 min, holding at 100% Phase B for 9 min.
Analysis was performed using a Q Exactive HF-X mass spectrometer in positive ion mode, with m/z range 300–1400. MS scan resolution was 120,000 at 200 m/z, AGC target 3e6, and Maximum IT 30 ms. Dynamic exclusion was set to 12.0 s. For MS/MS analysis, 60 fragment spectra were collected per scan using HCD fragmentation at Normalized Collision Energy 27% and an isolation window of 1.6 m/z. MS/MS resolution was 7500 at 200 m/z.
Protein clustering was performed using the ComplexHeatmap R package with Euclidean distance and average linkage. GO annotation was done using Blast2GO, with additional InterProScan annotation. KEGG pathway annotation was performed using KAAS, and enrichment analysis was conducted using the hypergeometric distribution. Protein-protein interaction analysis was performed using the STRING database and visualized in CytoScape (version 3.2.1).
Liquid chromatography-tandem mass spectrometry (LC-MS/MS)
The prepared samples were mixed with 40 μL of Pepsin buffer for enzymolysis of peptide fragments, followed by chromatographic separation. The resulting products were separated by capillary high-performance liquid chromatography and then subjected to MS using a Q Exactive HF-X mass spectrometer (Thermo Fisher Scientific, USA). Next, software MaxQuant 1.5.5.1 was used to search corresponding databases (uniprot_Homo_sapiens), and the results of protein identification and quantitative analysis were obtained.
Protein extraction and western blot
Total proteins from human NP tissues, murine NP tissues and cultured human NP cells were extracted and quantified utilizing the BCA Protein Assay Kit (P0012, Beyotime, Shanghai, China). EV-derived proteins were extracted utilizing the Exosomes Protein Extraction Kit (41223ES10, Yeasen, Shanghai, China) and quantified utilizing the BCA Protein Assay Kit. Western blot was performed in accordance with a previously established protocol. 35 Protein samples (20 μg per lane) were resolved on 10% SDS-PAGE gels and subsequently transferred onto polyvinylidene difluoride membranes. After a blocking step with a 5% skim milk solution, membranes were incubated overnight at 4°C with specific primary antibodies. Thereafter, they were incubated with species-specific secondary antibodies. Enhanced chemiluminescence reagents and the ChemiDoc XRS system (Bio-Rad, Shanghai, China) were utilized to visualize immunoreactive bands. A comprehensive list of the primary and secondary antibodies in this study could be found in Supplemental Table S5.
Histopathology and immunohistochemistry (IHC)
Histopathology and IHC procedures were conducted following a previously outlined protocol. 35 The hematoxylin-eosin (H&E; Jiancheng, Nanjing, China), Safranin-O/fast green (SO&FG; Solarbio, Beijing, China), and Masson staining (G1340, Solarbio, Beijing, China) were performed according to the manufacturer’s instructions. For the IHC staining images, image analysis was performed using Image Pro Plus software. H&E staining was used to analyze pathological morphology as described in the previous study. 36 Histological scoring was performed by two independent raters in a blind manner. For a detailed list of primary and secondary antibodies used, please refer to Supplemental Table S5.
Co-immunoprecipitation (Co-IP) assay for protein-protein interaction detection
Protein-protein interactions were assessed using the Pierce Crosslink Magnetic IP/Co-IP Kit (88805, Thermo Fisher Scientific, USA) according to the manufacturer’s protocol. In brief, cell lysates were prepared and incubated with the specific primary antibody and protein A/G agarose beads provided in the kit overnight at 4°C. After washing, the bound proteins were eluted and subjected to SDS-PAGE, followed by Western blot to detect the proteins of interest. Negative controls with antibody against rabbit IgG were included to ensure the specificity of the interactions.
Immunoprecipitation-acyl-biotin exchange (IP-ABE) for protein palmitoylation detection
Protein palmitoylation was assessed using the IP-ABE Palmitoylation Kit for WB (AM10314, Aimsmass, Shanghai, China) according to the manufacturer’s instructions. Briefly, cells were lysed and the lysates were incubated with specific antibodies to immunoprecipitate the target proteins. Following immunoprecipitation, hydroxylamine (HAM) treatment was applied to remove palmitoyl groups, and proteins were subsequently labeled with biotin. The biotinylated proteins were detected using streptavidin-HRP and analyzed by Western blot to confirm palmitoylation status. Negative controls without HAM treatment were included to ensure specificity.
CUT&RUN
CUT&RUN was performed using Hyperactive pG-MNase CUT&RUN Assay Kit for PCR/qPCR (Vazyme, HD101). Briefly, 100,000 treated human NP cells were collected and counted. Then the cells were captured with ConA Beads Pro and incubated with primary antibodies against rabbit IgG or YBX1 overnight at 4°C. After washing, the complex was incubated with pG-MNase Enzyme premix for 1 h at 4°C with rotation. Following that, fragmentation was conducted with CaCl2 premix at 4°C for 2 h. After termination and release of fragmentation, DNA was extracted by column centrifugation and was analyzed by qRT-PCR. The primer sequences for CUT&RUN-qPCR were shown in Supplemental Table S6.
Luciferase activity assay
Human NP cells were seeded into 24-well plates and allowed to adhere overnight. Subsequently, ZBP1 wild-type or mutant luciferase reporter vectors, along with YBX1 overexpression plasmids, were co-transfected into NP cells. Luciferase activity was determined 36 h post-transfection using the Dual-Luciferase Reporter Gene Assay Kit (11402ES60, Yeasen Biotechnology, Shanghai, China).
Statistical analyses
All data were collected from a minimum of three independent experiments and are presented as the mean ± standard deviation (SD). Differences among treatment groups were assessed using either the student’s t-test or one-way analysis of variance (ANOVA). All statistical analyses were performed using GraphPad Prism (Version 9.0, La Jolla, CA, USA). Statistical significance was defined as p < 0.05, with *p < 0.05, **p < 0.01, and ns indicating no significant difference.
Results
PANoptosis plays a significant role in the progression of IDD
We used the transcriptomic intervertebral disc dataset that included 12 samples with transcriptomic data and 18 samples with single cell RNA sequencing (scRNA-seq) results to evaluate the PANoptosis features and explore the potential opportunity for the precise targeting of PANoptosis in IDD (Figure 1(a)). After taking the intersection, we found that no matter in any data set, the programmed cell death pathway was enriched, which was specifically manifested as the overall up-regulation of pyroptosis, apoptosis and necroptosis (Figure 1(b) and (c)). To explore the specific altered genes in these pathways, we used heat map formation to show the up-regulated genes in the aforementioned programmed cell death pathways in NP cells from IDD tissues. Subsequently, we found that proteins that play central roles in these pathways, such as Caspase-1, Caspase-3, Caspase-7, and Caspase-8 in pyroptosis; TRAF2, TRAF5, and RIPK3 in necroptosis; and SOD1 and SOD2 in apoptosis; were significantly upregulated in degenerative NP cells. Such results suggested an important role of PANoptosis in IDD (Figure 1(d)).

PANoptosis plays a significant role in the progression of IDD: (a) workflow of the analytical process performed in this study, (b) volcano plot of differentially expressed genes between IDD tissues and normal tissues in GSE266883, (c) dotplot of the results of enrichment analysis of upregulated genes in IDD tissues, (d) heatmap of up-regulated PANoptosis-related genes in IDD tissues, (e) featurePlot of PANoptosis-related genes in the single-cell cohort, (f) representative images of sagittal T2-weighted MRI scans of patients according to Pfirrmann grades, (g) representative histological staining (H&E staining and Masson staining; upper images, scale bar: 100 μm; lower images, scale bar: 200 μm), and IHC staining (scale bar:100 μm, lower images scale bar: 200 μm) images of disc specimens between the Control group and the IDD group, (h) immunoblot analysis of degeneration-associated and PANoptosis-associated markers of disc specimens in the Control and IDD groups. At least three independent experiments were performed. Data are presented as mean ± SD. *P < 0.05, **P < 0.01.
In order to further verify the important role of PANoptosis in IDD, we constructed two comprehensive cohorts based on the publicly available single cell RNA sequencing cohorts of IDD. The specific composition of the cohort was as described in Methods. The degeneration cohort consisted of four normal NP tissues from two datasets and five degenerated NP tissues from two different datasets. After quality control, feature screening, and dimension reduction annotation, we divided the cells into NP cells and non-nucleus pulposus cells for subsequent research. NP cells with normal secretion of collagen type Ⅱ and aggrecan were defined as non-degenerated NP cells, while NP cells with loss of collagen type Ⅱ expression, including activated expression of collagen type Ⅰ, were defined as degenerated nucleus pulposus cells. The core functional proteins related to PANoptosis, such as Caspase-3, Caspase-7, and GSDMD, were significantly upregulated in degenerative NP cells. The score of pan-apoptotic gene set based on ssGSEA principle also showed that PANoptosis gene set score was higher in NP cells of degenerated intervertebral disc tissue. Similarly, to decipher the role of PANoptosis in the progression of IDD, we analyzed data from a single cell sequencing cohort of moderately and severely degenerative intervertebral disc tissues. We found that whether Caspase-3, Caspase-7, or GSDMD was upregulated in NP cells originating from severely degenerative disc tissues (Figure 1(e)). Taken together, the results from a single cell sequencing cohort of real-world samples from multiple sources fully demonstrated that PANoptosis is activated in degenerative intervertebral disc tissues and is closely related to the progression of IDD.
As depicted in Figure 1(f), the Control group exhibited higher signal intensity on T2-weighted MRI, while lower T2-weighted signal intensity was observed in the IDD group. H&E staining revealed an irregular tissue structure, featuring loss of vacuolated nucleus pulposus in the IDD group. Masson staining illustrated a reduction in ECM and a disrupted arrangement of collagen fibers in the IDD group. IHC staining exposed that NP samples in the IDD group displayed lower Col-II expression levels compared to those in the Control group (Figure 1(g)).
Subsequently, the expression levels of ECM metabolism-, apoptosis-, pyroptosis-, and necroptosis-associated proteins in disc samples were analyzed by utilizing Western blot. The findings revealed notably decreased levels of Col-II and ACAN and an increased level of MMP3 in the IDD group compared to the Control group. Notably, active forms of Caspase3, Caspase7, and Caspase8 (apoptosis markers), active forms of Caspase1, GSDMD, and GSDME (pyroptosis markers), and phosphorylation of MLKL, RIPK1, and RIPK3 (necroptosis markers) were observed in the IDD group (Figure 1(h)). Therefore, the findings indicated that cell death characterized by activation of PANoptosis in NP tissues contributed to the pathogenesis of IDD.
PTH promotes hUCMSC-derived EV secretion quantitatively
To determine whether PTH could be utilized to alter the MSC-derived EVs production for hUCMSC-based therapy, hUCMSCs were pretreated with PTH (10 nM) for 6 h, then the medium was replaced, and the medium was collected after 48 h. Both hUCMSCs and PTH-preconditioned hUCMSCs exhibited a spindle-like cell morphology at 90% confluency (Figure 2(a)). Following induction for 18 days with osteogenic, adipogenic, and chondrogenic stimuli, Alizarin Red, Oil Red O, and Alcian Blue staining revealed distinct calcified nodules, small cytoplasmic lipid droplets, and cartilage formation, affirming the multilineage differentiation potential of both hUCMSCs and PTH-preconditioned hUCMSCs into osteocytes, adipocytes, and chondrocytes (Figure 2(b)).

PTH promotes hUCMSC-derived EV secretion quantitatively: (a) both hUCMSCs and PTH-preconditioned hUCMSCs exhibited a spindle-like cell morphology (scale bar: 100 μm), (b) representative images of Alizarin Red (scale bar: 200 μm), Oil Red O (scale bar: 50 μm), and Alcian Blue (scale bar: 100 μm) staining following osteogenic, adipogenic, and chondrogenic induction, (c) flow cytometric analysis of cell surface markers on both hUCMSCs and PTH-preconditioned hUCMSCs (CD14, CD19, CD34, CD45, CD73, CD90, and HLA-DR), (d) transmission electron microscopy revealed that hUCMSC-EVs and PTH pre-EVs (black arrows) exhibited typical round-shaped membranous vesicles (scale bar: 200 μm), (e) NTA assessed the size distribution of hUCMSC-EVs and PTH pre-EVs, (f) quantification of NTA experiments of EVs isolated from the same volume of cell culture supernatant from hUCMSCs and PTH-preconditioned hUCMSCs, (g) western blot analysis depicted the expression of protein markers (CD63, CD73, and CD90) in hUCMSCs, hUCMSC-EVs, and PTH pre-EVs, with calreticulin (an intracellular contaminant) absent in EVs and PTH pre-EVs, (h) quantification of total protein of hUCMSC-EVs and PTH pre-EVs was analyzed by BCA assay. At least three independent experiments were performed.
Additionally, flow cytometric analysis confirmed positive expression of CD90 and CD73, while CD34, CD19, CD14, HLA-DR, and CD45 were not expressed (Figure 2(c)), meeting the criteria established by the International Society for Cellular Therapy. Transmission electron microscopy (TEM) showed that extracellular vesicles derived from both hUCMSCs and PTH-preconditioned hUCMSCs exhibited typical round-shaped membranous vesicles (Figure 2(d)). Nanoparticle tracking analysis (NTA) revealed higher EV production in the PTH-preconditioned hUCMSCs than in the Control group, with an average diameter of 108 nm, with the original concentration at 3.5E + 10 particles/mL in EVs group, and an average diameter of 116.6 nm, with the original concentration at 8.2E + 10 particles/mL in PTH pre-EVs group (Figure 2(e) and (f)). Western blot analysis confirmed the enrichment of CD63, CD73, and CD9 in both hUCMSC-EVs and PTH pre-EVs compared to hUCMSCs, while the intracellular contaminant calreticulin was absent in EVs and PTH pre-EVs (Figure 2(g)). Importantly, EV secretion was remarkably enhanced in PTH-preconditioned hUCMSCs, as indicated by both measuring the total amount of EV proteins (Figure 2(h)). These findings collectively demonstrated the successful isolation and characterization of hUCMSCs, hUCMSC-EVs, and PTH pre-EVs and that PTH enhanced hUCMSC-derived EV secretion quantitatively.
EVs derived from PTH-preconditioned hUCMSCs exhibit enhanced effects against NP cell PANoptosis in vitro and in vivo
To determine whether TNF-α could induce NP cell PANoptosis and the optimal timing, NP cells were exposed to different concentrations of TNF-α (0, 5, 10, and 15 ng/mL) for 24, 48, and 72 h. As shown in Supplemental Figure S1, TNF-α significantly decreased the viability of NP cells in a time- and dose-dependent manner and activated the degradation of ECM in NP cells in a dose-dependent manner, which effectively induced PANoptosis of NP cells. Thus, we used 10 ng/mL of TNF-α as the working concentrations and 24 h as the stimulation duration for the following studies.
To further validate the effects of hUCMSC-EVs and PTH pre-EVs in vitro, NP cells were exposed to TNF-α (10 ng/mL) and treated with hUCMSC-EVs (100 μg/mL) or PTH pre-EVs (100 μg/mL) for 24, 48, and 72 h. As depicted in Figure 3(a), a decrease of cell viability was observed in TNF-α group at 24, 48, and 72 h, compared with the Control group. Both groups (TNF-α and EVs group and TNF-α and PTH pre-EVs group) could partially rescue suppression of cell viability at 24, 48, and 72 h while PTH pre-EVs presented a better rejuvenating effect than EVs. Flow cytometry analysis showed that TNF-α could induce NP cell apoptosis; both hUCMSC-EVs and PTH pre-EVs group could partially inhibit NP cell apoptosis while the apoptosis rate of cells with PTH pre-EVs incubation was lower than that of EVs group cells (Figure 3(b)). Besides, the cell apoptotic effect was evaluated by TUNEL staining assay. Clearly, compared with the Control group, obvious green fluorescence could be observed in TNF-α group. Following treatment with EVs and PTH pre-EVs, a decrease of apoptosis was observed, whereas EVs derived from PTH-preconditioned hUCMSCs exhibited a more pronounced effect than EVs (Figure 3(c)). Moreover, IL-1β and IL-18 release were observed in response to TNF-α treatment, with significant reductions in both the EVs and PTH pre-EVs groups (Figure 3(d)).

hUCMSC-EVs and PTH pre-EVs alleviate TNF-α-induced NP cell PANoptosis: (a) the OD values of NP cells exposed to TNF-α (10 ng/mL) and treated with hUCMSC-EVs (100 μg/mL) or PTH pre-EVs. (100 μg/mL) at 24-, 48-, and 72-h time points detected by CCK-8 assay (*p < 0.05, Control group vs. TNF-α group; #p < 0.05, TNF-α group vs. TNF-α and EVs group; &p < 0.05, TNF-α group vs. TNF-α and PTH pre-EVs group), (b) the apoptosis rate of NP cells analyzed by flow cytometry, (c) representative images of TUNEL staining (scale bar: 25 μm). (d) IL-1β and IL-18 levels in the cell culture supernatant, (e, f) immunoblot analysis of degeneration-associated and PANoptosis-associated markers. At least three independent experiments were performed.
Next, the expression levels of ECM metabolism- and PANoptosis-associated proteins in NP cells were analyzed. The findings revealed notably decreased levels of Col-II and ACAN in the TNF-α group compared to the Control group. Conversely, these levels were noticeably increased in the EVs group and PTH pre-EVs group, compared with the TNF-α group. The level of MMP3, cleavage of Caspase3, Caspase7, Caspase8, Caspase1, GSDMD, and GSDME, and phosphorylation of MLKL, RIPK1, and RIPK3 were increased in the TNF-α group. However, these levels were noticeably decreased in the EVs group and PTH pre-EVs group compared to the TNF-α group. Notably, PTH pre-EVs exhibited a more pronounced effect than EVs (Figure 3(e) and (f)).
To further validate the effect of hUCMSC-EVs and PTH pre-EVs in vivo, hUCMSC-EVs or PTH pre-EVs were injected to the NP tissue in needle puncture-induced IDD mice. We evaluated the PANoptosis and degeneration degree of intervertebral disc by using X-ray, MRI, histological staining, IHC staining, and Western blot. As depicted in Figure 4(a)–(c), in the Control group, X-ray analysis revealed a normal height of the intervertebral disc, and MRI exhibited high T2-weighted signal intensity. Following needle puncture surgery, the Needle Puncture group exhibited a reduction in disc height and lower T2-weighted signal intensity compared with the Control group. Administration of hUCMSC-EVs or PTH pre-EVs effectively postponed the decrease of disc height and maintained higher T2-weighted signal intensity. The histomorphology analysis using H&E, Safranin-O/fast green, and Masson staining revealed that fibrochondrocytes replaced the NP cells in the Needle Puncture group. Additionally, the volume of the NP decreased significantly and eventually vanished, while the annulus fibrosus structure suffered severe damage. Furthermore, the staining with Safranin-O/fast green suggested a decrease in the glycosaminoglycan and proteoglycan levels of NP following needle puncture. The histological scores of the Needle Puncture group were considerably greater than those of the Control group, as evidenced by the images of histological staining. The histological scores of the group that received needle puncture and EVs or PTH pre-EVs were considerably lower compared to the group that only received needle puncture (Figure 4(d) and (e)). IHC staining showed that the expression levels of Col-Ⅱ were decreased in the NP following needle puncture. Administration of hUCMSC-EVs or PTH pre-EVs effectively postponed the decrease caused by needle puncture. Notably, PTH pre-EVs exhibited a more pronounced effect than EVs (Figure 4(f)).

hUCMSC-EVs and PTH pre-EVs alleviate NP cell PANoptosis in needle puncture-induced IDD mice: (a) representative pictures of mouse IVD samples in Control group, needle puncture group, needle puncture group and needle puncture and PTH pre-EVs group detected by X-ray, MRI, (b) analysis of disc height index (DHI) based on X-ray images (n=6), (c) Pfirrmann grades based on MRI images (n=6), (d) representative pictures of mouse IVD samples in diverse groups detected by histological staining (H&E staining, Masson staining and Safranin-O/fast green staining; scale bar:100 μm), and IHC staining (scale bar: 100 μm), (e) analysis of histological scores based on the images of histological staining (n=6), (f) quantitative analysis of Col-II-positive areas (n=6), (g) immunoblot analysis of degeneration-associated and PANoptosis-associated markers (n=3). At least three independent experiments were performed.
Following the needle puncture and injection of hUCMSC-EVs or PTH pre-EVs, the expression levels of ECM metabolism- and PANoptosis-associated proteins in mice NP samples were analyzed. Consistent with the results in vitro, both hUCMSC-EVs and PTH pre-EVs alleviated NP cell PANoptosis, whereas PTH pre-EVs exhibited a more pronounced effect than EVs (Figure 4(g)). Collectively, EVs derived from PTH-preconditioned hUCMSCs present enhanced effects against NP cell PANoptosis in vitro and in vivo.
Alterations of cargo contents in EVs derived from PTH-preconditioned hUCMSCs
After establishing that EVs derived from hUCMSCs could ameliorate IDD by inhibiting PANoptosis, and that EVs derived from PTH-preconditioned hUCMSCs could amplify this therapeutic effect, we performed proteomic sequencing to clarify the mechanism by which EV contents mediated their therapeutic effects. After quality control of the data, we co-identified 1780 proteins that were more highly expressed in the PTH-treated group and 1203 proteins that had a higher abundance in the Control group (Figure 5(a)). By principal component analysis, we found significant differences in the distribution and content of protein species in EVs between PTH-treated and Control groups (Figure 5(b)). Differential analysis identified a total of 118 proteins that were significantly upregulated in PTH pre-EVs (Figure 5(c)). Through functional enrichment analysis based on GO, KEGG, and Reactome databases, we found that most of these proteins were enriched in lipid metabolism, cell connection, and other pathways, among which lipid metabolism attracted our attention (Figure 5(d)). PPI network analysis showed that the palmitoyl-acyltransferase ZDHHC5 played a potential key role in the inhibition of PANoptosis in IDD (Figure 5(e)). In our constructed single-cell cohorts, ZDHHC5 was downregulated in NP cells of degenerated intervertebral discs, but not further downregulated in severely degenerated intervertebral discs (Figure 5(f)). Similarly, the transcriptome cohort showed that ZDHHC5 was significantly downregulated in degenerated NP tissues (Figure 5(g)). Pan-palmitoyl-acyltransferase screening revealed that ZDHHC5 was one of the most abundant palmitoyl-acyltransferases in intervertebral disc tissues (Figure 5(h)).

Alterations of cargo contents in EVs derived from PTH-preconditioned hUCMSCs: (a) heatmap of differentially expressed proteins between PTH group and Control group, (b) principal component analysis plots of the PTH and Control groups, (c) volcano plot of differentially expressed proteins between PTH group and Control group, (d) dotplot of the results of enrichment analysis of up-regulated proteins in the PTH group, (e) PPI network based on the up-regulated proteins in PTH group, (f) violin plot of differential expression of ZDHHC5 in two single-cell cohorts, (g) bar graph of differential ZDHHC5 expression between IDD and Normal groups in GSE266883, (h) heatmap of the expression abundance of entire ZDHHC family members in GSE266883, (i) immunoblot analysis of ZDHHC5 in human NP tissues, mouse NP tissues and NP cells, (j) immunoblot analysis of ZDHHC5 in hUCMSCs or EVs treated with ZDHHC5 siRNA. At least three independent experiments were performed.
To validate whether ZDHHC5 could be the promising candidate for ameliorating IDD by inhibiting PANoptosis, the protein expression levels of ZDHHC5 were analyzed. The level of ZDHHC5 was decreased in human degenerated NP tissues, needle puncture-induced degenerated mouse NP tissues, and TNF-α induced-degenerated NP cells (Figure 5(i)). To verify the function of ZDHHC5 in EVs, we knocked down the expression of ZDHHC5 in hUCMSCs using siRNA to obtain the EVs with ZDHHC5 knockdown (Supplemental Figure S2). Western blot analysis revealed a significant increase in ZDHHC5 levels both in PTH-preconditioned hUCMSCs and EVs derived from PTH-preconditioned hUCMSCs. However, the level of ZDHHC5 was reduced in both hUCMSCs transfected with siRNA (hUCMSCs-Si-ZDHHC5) and in EVs derived from PTH-preconditioned hUCMSCs transfected with siRNA (PTH pre-EVs-Si-ZDHHC5; Figure 5(j)). Collectively, these data pointed to the possibility that ZDHHC5 in the EVs might play an important role in the pathogenesis of IDD.
PTH pre-EVs transport ZDHHC5 to inhibit PANoptosis in NP cells
To further investigate the exact effects of ZDHHC5 enriched in the PTH pre-EVs, the NP cells were stimulated with TNF-α (10 ng/mL) and various modified PTH pre-EVs for 24 h. The EVs secreted from the PTH-preconditioned hUCMSCs after transfection with ZDHHC5 siRNA were collected. Purified EVs or PTH pre-EVs could be internalized by NP cells (Supplemental Figure S3). Consistent with the results in Figure 3, a decrease of cell viability, a higher apoptosis rate of cells, obvious green fluorescence, increased IL-1β and IL-18 release, and notably increased levels of degeneration, apoptosis, pyroptosis, and necroptosis markers was observed in TNF-α group compared with the Control group, while PTH pre-EVs could partially rescue suppression of cell viability, enhanced apoptosis and increased release of IL-1β and IL-18. Notably, inhibition of ZDHHC5 in the PTH pre-EVs partially or almost completely diminished the therapeutic effect of PTH pre-EVs on TNF-α-induced NP cell PANoptosis (Figure 6). Collectively, these findings validated that ZDHHC5 had significant involvement in the therapeutic impact of PTH pre-EVs on TNF-α-induced NP cell PANoptosis.

PTH pre-EVs transport ZDHHC5 to inhibit PANoptosis in NP cells: (a) the OD values of NP cells exposed to TNF-α (10 ng/mL) and treated with PTH pre-EVs (100 μg/mL), PTH pre-EVs-si-NC (100 μg/mL) or PTH pre-EVs-si-ZDHHC5 (100 μg/mL) at 24-, 48-, and 72-h time points detected by CCK-8 assay (* p < 0.05, Control group vs. TNF-α group; #p < 0.05, TNF-α group vs. TNF-α and PTH pre-EVs group; &p < 0.05, TNF-α and PTH pre-EVs-si-NC group vs. TNF-α and PTH pre-EVs-si-ZDHHC5 group), (b) the apoptosis rate of NP cells analyzed by flow cytometry, (c) representative images of TUNEL staining (scale bar: 25 μm), (d) IL-1β and IL-18 levels in the cell culture supernatant, (e) immunoblot analysis of degeneration-associated and PANoptosis-associated markers. At least three independent experiments were performed.
ZDHHC5 regulates ZBP1 transcription by competitively inhibiting YBX1 phosphorylation via palmitoylation
To confirm the potential target proteins of ZDHHC5, we used Co-IP-based LC-MS/MS to identify the ZDHHC5-interacting proteins. Among them, YBX1 attracted our attention. YBX1 is a member of the cold shock protein superfamily. As a transcription factor, it binds specifically to the promoters of target genes and regulates their expression (Figure 7(a)). We used AlphaFold3 to predict the binding probability of ZDHHC5 and YBX1 and found that there were regional peptides with high binding probability between ZDHHC5 and YBX1, thus supporting the results of mass spectrometry (Figure 7(b)). Moreover, the endogenous and exogenous Co-IP assays confirmed that ZDHHC5 could interact with YBX1 in NP cells and 293T cells (Figure 7(c) and (d)). Given that ZDHHC5 is a palmitoyl-acyltransferase, we hypothesized that ZDHHC5 might hinder NP cell PANoptosis by manipulating palmitoylation of YBX1. As expected, a noticeable increase in YBX1 palmitoylation was detected in NP cells overexpressing ZDHHC5 (Figure 7(e)). Given YBX1 is required to be phosphorylated to function as a transcriptional regulator, 37 we detected the phosphorylation level of YBX1. Notably, a significant decrease in YBX1 phosphorylation was detected in NP cells overexpressing ZDHHC5 (Figure 7(f)). Moreover, treatment with the palmitoylation inhibitor 2-BP effectively reversed the enhanced palmitoylation of YBX1 and decreased phosphorylation of YBX1 induced by ZDHHC5 overexpression (Figure 7(g) and (h)). It is well known that PANoptosis sensors, including ZBP1, AIM2, RIPK1, and NLRP12, play a vital role in PANoptosis. 14 Hence, to validate how YBX1 regulated PANoptosis, the mRNA expression of above PANoptosis sensors was detected in NP cells overexpressing YBX1. We found only ZBP1 mRNA expression was increased significantly in NP cells overexpressing YBX1, and there was no statistical significance for the remaining sensors (Supplemental Figure S4). To confirm whether YBX1 can regulate ZBP1 expression, we simultaneously predicted the binding ability of YBX1 to the promoter region of ZBP1 by two methods. On the one hand, we used JASPAR to predict the promoter sequence binding score of YBX1 and ZBP1 and found that the score was high and statistically significant. On the other hand, alphafold3 was used to predict the binding ability of YBX1 to the DNA in the ZBP1 promoter region, and we found that YBX1 had a high binding probability to the ZBP1 promoter region (Figure 7(i)). These results indicated that the transcription factor YBX1 might act directly as an upstream modulator of ZBP1.

ZDHHC5 regulates ZBP1 transcription by competitively inhibiting YBX1 phosphorylation via palmitoylation: (a) Venn plot of the mass spectrum showing the intersection between proteins bound to ZDHHC5 and proteins upregulated in IDD tissues, (b) schematic diagram of the docking of ZDHHC5 with YBX1 based on Alphafold3, (c) Co-IP analysis of the interaction between endogenous ZDHHC5 and endogenous YBX1 in NP cells, (d) Co-IP analysis of the interaction between exogenous ZDHHC5 and exogenous YBX1 in 293T cells, (e) the alterations of YBX1 palmitoylation in NP cells with ZDHHC5 overexpression, (f) the alterations of YBX1 phosphorylation in NP cells with ZDHHC5 overexpression, (g) the alterations of YBX1 palmitoylation in NP cells treated with ZDHHC5 plasmid or the palmitoylation inhibitor (2-BP), (h) the alterations of YBX1 phosphorylation in NP cells treated with ZDHHC5 plasmid or the palmitoylation inhibitor (2-BP), (i) schematic representation of AlphaFolD3-based binding of YBX1 (protein) to the ZBP1 promoter sequence (DNA), (j) the alterations of ZBP1 promoters in NP cells treated with or without ZDHHC5 plasmid or the palmitoylation inhibitor (2-BP) by using antibodies against YBX1, (k) results of relative luciferase activities, (l) the alterations of ZBP1 in NP cells treated with ZDHHC5 plasmid or the palmitoylation inhibitor (2-BP). (m) FeaturePlot of YBX1 and ZBP1 expression in the single-cell cohort. At least three independent experiments were performed.
To further verify that YBX1 modulated ZBP1 transcription, we conducted CUT&RUN and luciferase reporter assays. The enrichment of YBX1 on the ZBP1 promoter in NP cells significantly increased after ZDHHC5 overexpression, and 2-BP effectively abolished the increased enrichment of YBX1 on the ZBP1 promoter (Figure 7(j)). Luciferase reporter assays further verified that the reporter expression levels were significantly increased in 293T cells overexpressing YBX1, and the mutation of the binding site effectively abolished the YBX1 overexpression-mediated increase in luciferase activity (Figure 7(k)). Moreover, ZBP1 protein expression was decreased in NP cells overexpressing ZDHHC5, and 2-BP effectively reversed the decreased protein levels of ZBP1 (Figure 7(l)). Moreover, we confirmed that YBX1 and ZBP1 were upregulated in degenerative NP cells and associated with IDD based on single cell sequencing data from public cohorts (Figure 7(m)). Overall, ZDHHC5 inhibited YBX1 phosphorylation to suppress ZBP1 transcription, thereby hindering PANoptosis by promoting YBX1 palmitoylation.
PTH pre-EVs ameliorate NP cell PANoptosis by inhibiting ZDHHC5/YBX1/ZBP1 axis in needle puncture-induced IDD mice
To unveil the function of ZDHHC5 and ZBP1 in the pathogenesis of IDD in vivo, PTH pre-EVs, either modified or unmodified, or AAV9-shZBP1, were injected to the NP tissue in needle puncture-induced IDD mice. We evaluated the apoptosis, pyroptosis, necroptosis, and degeneration degree of intervertebral disc by using X-ray, MRI, histological staining, IHC staining, and Western blot. Consistent with the results in Figure 4, administration of PTH pre-EVs effectively postponed the decrease of disc height and maintained higher T2-weighted signal intensity. Notably, the PTH pre-EVs-Si-ZDHHC5 group demonstrated a reduction in disc height and lower T2-weighted signal intensity compared to the PTH pre-EVs group. However, the combination of PTH pre-EVs-Si-ZDHHC5 and AAV9-shZBP1 partially restored disc height and increased T2-weighted signal intensity compared to the administration of PTH pre-EVs-Si-ZDHHC5 alone (Figure 8(a) – (c)). The histomorphology analysis using H&E, Safranin-O/fast green, and Masson staining revealed that fibrochondrocytes replaced the NP cells in the Needle Puncture group. The histological scores of the group that received needle puncture and PTH pre-EVs were considerably lower compared to the group that only received needle puncture. The PTH pre-EVs-Si-ZDHHC5 group showed lower histological scores compared to the PTH pre-EVs group. However, the combination of PTH pre-EVs-Si-ZDHHC5 and AAV9-shZBP1 partially restored histological scores to higher levels compared to the administration of PTH pre-EVs-Si-ZDHHC5 alone (Figures 8(d)–(f) and 9).

PTH pre-EVs ameliorate NP cell PANoptosis by inhibiting ZDHHC5/YBX1/ZBP1 axis in needle puncture-induced IDD mice: (a) representative pictures of mouse IVD samples in Control group, needle puncture group, needle puncture and PTH pre-EVs group, needle puncture and PTH pre-EVs-si-ZDHHC5 and needle puncture and PTH pre-EVs-si-ZDHHC5 and AAV9-shZBP1 detected by X-ray and MRI, (b) analysis of DHI based on X-ray images (n=6), (c) Pfirrmann grades based on MRI images (n=6), (d) representative pictures of mouse IVD samples in diverse groups detected by histological staining (H&E staining, Masson staining, and Safranin-O/fast green staining; scale bar:100 μm), and immunohistochemistry staining (scale bar: 100 μm), (e) analysis of histological scores based on the images of histological staining (n=6), (f) quantitative analysis of Col-II-positive areas (n=6), (g) immunoblot analysis of degeneration-associated and PANoptosis-associated markers (n=3). At least three independent experiments were performed.
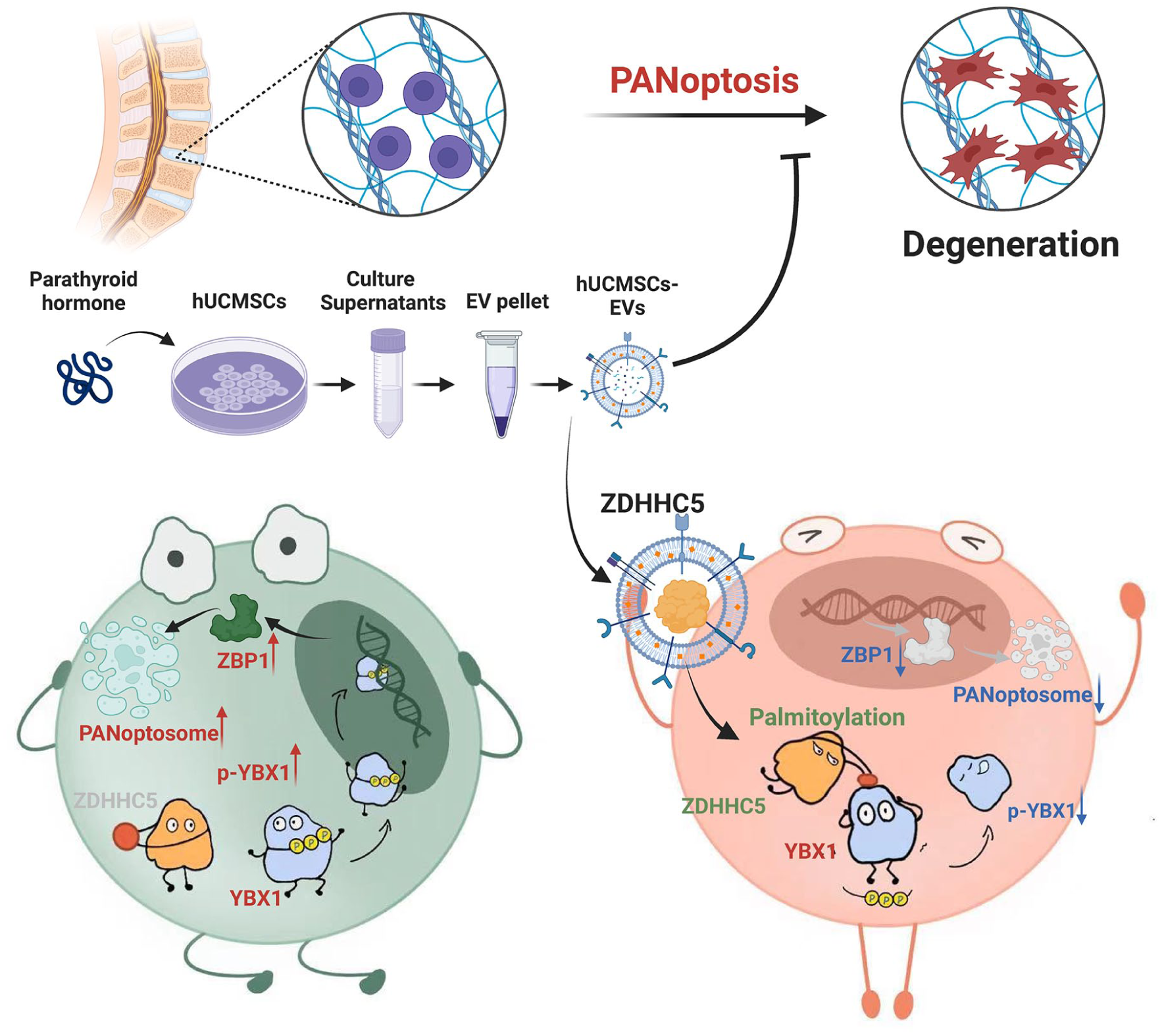
Schematic diagram demonstrating the potential mechanism by which hUCMSCs-EVs exert their therapeutic efficacy in the development of IDD. PTH enhances the secretion of hUCMSC-derived EVs and alters their cargo composition, which may contribute to their improved therapeutic effects. Mechanistically, PTH-preconditioned EVs, enriched with ZDHHC5, ameliorate PANoptosis by modulating ZBP1 transcription through competitive inhibition of YBX1 phosphorylation via palmitoylation.
Following the needle puncture and injection of PTH pre-EVs, PTH pre-EVs-Si-ZDHHC5, or AAV9-shZBP1, the expression levels of ECM metabolism- and PANoptosis-associated proteins in mouse NP samples were analyzed. Consistent with the results in vitro, inhibition of ZDHHC5 in the PTH pre-EVs partially or almost completely diminished the therapeutic effect of PTH pre-EVs on needle puncture-induced IDD via regulating PANoptosis. However, the combination of PTH pre-EVs-Si-ZDHHC5 and AAV9-shZBP1 partially reduced the levels of degeneration, apoptosis, pyroptosis, and necroptosis markers compared to the administration of PTH pre-EVs-Si-ZDHHC5 alone, suggesting the involvement of ZDHHC5 and ZBP1 in hUCMSC-EVs-based therapy for IDD (Figure 8(g)). Collectively, the findings suggested that PTH pre-EVs hindered PANoptosis and degeneration in intervertebral discs by inhibiting ZBP1 through the regulatory effect of ZDHHC5 in needle puncture-induced IDD mice.
Discussion
EVs, particularly stem cell-derived EVs, demonstrate significant therapeutic potential for intervertebral disc degeneration by reducing inflammation, inhibiting apoptosis, promoting tissue regeneration, and enhancing intercellular communication. However, current research also indicates that MSC-derived EVs may not be the optimal candidate for IDD therapy. 38 Although some studies have reported that engineered exosomes are more effective than natural exosomes for treating disc degeneration, further exploration is needed to enhance their therapeutic effects. Moreover, characterizing EV components is crucial for understanding their potential in cell-free therapies for IDD. In our study, scRNA-seq, RNA-seq data analysis, and validation with human disc samples showed that the biomarkers promoting PANoptosis are notably upregulated in the IDD group, while the biomarkers suppressing PANoptosis are downregulated, suggesting PANoptosis as a potential therapeutic target in the pathogenesis of IDD. Using an TNF-α-stimulated PANoptosis cell model, we demonstrated that both hUCMSC-EVs and PTH pre-EVs effectively mitigate PANoptosis in NP cells. Given EVs’ potential as delivery vehicles, we employed scRNA-seq and RNA-seq to identify bioactive molecules. Our bioinformatic analysis yielded significant findings, identifying ZDHHC5 as a crucial downstream target in hUCMSC-EVs-based therapy for IDD development. Subsequently, we demonstrated that PTH pre-EVs containing high levels of ZDHHC5 ameliorate PANoptosis through internalization by NP cells, both in vitro and in vivo. Moreover, through loss- and gain-of-function experiments, we found that ZDHHC5 regulates ZBP1 transcription by competitively inhibiting YBX1 phosphorylation via palmitoylation. To our knowledge, this study is among the first to reveal the protein composition of EVs derived from PTH-preconditioned hUCMSCs and to assess the efficacy of PTH pre-EVs-based therapy in treating IDD by inhibiting PANoptosis.
MSCs, with their innate immunomodulatory capacity, are a promising therapeutic approach that could help mitigate the inflammatory environment of the IVD upon transplantation. However, their characteristics still require further investigation. While MSCs have been shown to survive and differentiate in the IVD across various animal models, their ability to fully express NP cell phenotypical characteristics has not yet been documented. 39 Studies indicate that MSC transplantation, especially when combined with biomaterials such as hydrogels or atelocollagen, significantly enhances proteoglycan synthesis, preserves disc height, and restores disc structure in animal models.40,41 Recent studies suggest that the therapeutic benefits of MSCs are largely due to paracrine signaling, particularly through the release of exosomes and other bioactive molecules that affect the biological activities of surrounding cells.42–44 MSC-derived exosomes offer distinct advantages over MSC therapies. They are safer, as they avoid risks such as immune rejection and tumor formation. 45 Exosomes are also easier to store and standardize, providing a more stable therapeutic option. 46 They retain MSCs’ regenerative benefits by delivering bioactive molecules, such as cytokines and miRNAs, to support tissue repair and immune modulation without live cell transplantation. 47 Ambrosio et al. demonstrated that NPC-derived EVs have a more significant therapeutic effect compared to MSC-derived EVs. 38 Despite the lack of FDA-approved EV therapeutics for IDD, many preclinical studies support the potential of EVs as a promising cell-free strategy for treating IVDD and promoting endogenous repair. 17 MSCs have been shown to respond to a variety of biophysical and biochemical stimuli, which can significantly enhance the secretion and immunomodulatory properties of MSC-derived EVs.48–50 Shao et al. reported that parathyroid hormone enhances the therapeutic efficacy of bone marrow multipotent mesenchymal stromal cell-derived exosomes by suppressing proinflammatory cytokine expression, thereby promoting chondrocyte repair in osteoarthritis (OA). 31 Simultaneously, our previous research has reported that exogenous PTH alleviates intervertebral disc degeneration through sonic hedgehog signaling pathway mediated by CREB. 30 In our study, we used PTH as a pretreatment agent for hUCMSCs and found that the administration of PTH pre-EVs significantly enhanced therapeutic effects in ameliorating IDD by inhibiting PANoptosis. However, EVs secreted by unstimulated hUCMSCs also alleviated IDD, though they were less effective than those secreted by PTH-preconditioned hUCMSCs. These findings underscore that appropriate pretreatment of hUCMSCs can enhance the therapeutic efficacy of hUCMSC-derived EV-based therapy for IDD. However, we should acknowledge the short-lived effects of EV therapy, as retention within the disc is a known challenge. Leakage and clearance from the injection site could limit long-term efficacy.
PANoptosis is a unique form of programmed cell death that integrates three major pathways: pyroptosis, apoptosis, and necroptosis, orchestrated by a multi-protein complex known as the PANoptosome.9,11,51 Unlike traditional cell death mechanisms that operate independently, PANoptosis activates elements from all three pathways simultaneously, particularly in response to immune challenges or infections.14,52 This coordinated inflammatory cell death ensures the effective clearance of infected or damaged cells, especially when pathogens attempt to block other death pathways. Dysregulation of PANoptosis has been implicated in various diseases, including cancer, neurodegenerative disorders, inflammatory diseases, and cytokine storms, marking it as a critical factor in immune response modulation.14,52,53 Consequently, researchers are investigating therapeutic approaches to modulate PANoptosis by targeting specific components of the PANoptosome, either to enhance or inhibit the process depending on the disease context, presenting new opportunities for innovative treatments.14,54 Although direct studies on PANoptosis in intervertebral disc degeneration are limited, related cell death mechanisms, such as apoptosis and pyroptosis, are well-documented in IDD pathology.9,14,15 Apoptosis contributes to the depletion of nucleus pulposus cells and the degradation of the extracellular matrix, both of which are critical drivers of disc degeneration.10,11 Similarly, pyroptosis, a highly inflammatory type of cell death, aggravates IDD by activating inflammasomes and triggering inflammatory responses that further deteriorate disc structure.13,55 The convergence of these cell death processes in PANoptosis suggests that this combined mechanism may play a role in IDD, presenting new avenues for targeting and managing cell death and inflammation in degenerative disc diseases. In our study, we verified the elevation of pro-PANoptosis genes and proteins and the reduction of anti-PANoptosis genes and proteins through RNA-seq data analysis and validation using human degenerative disc samples. Additionally, we observed signs of extracellular matrix degradation. Thus, further research is essential to comprehensively understand PANoptosis mechanisms in IDD and develop effective therapeutic interventions.
Our study showed that both EVs and PTH pre-EVs could alleviate PANoptosis in NP cells, whereas EVs derived from PTH-preconditioned hUCMSCs exhibited enhanced effects against PANoptosis of NP cells in vitro and in vivo. In view of the enhanced anti-PANoptosis effect of PTH pre-EVs on the modulation of NP cells compared to that of EVs, we suggest that PTH precondition may change the protein profiles of exosomes, some remarkably expressed protein in PTH pre-EVs account for their superior anti-PANoptosis properties. Therefore, the protein expression profiles of EVs and PTH pre-EVs were compared through proteomic sequencing and bioinformatics analysis. Through differentially expressed proteins and their relative functional enrichment analysis based on GO, KEGG, and Reactome databases and PPI network analysis, we speculated that the palmitoyl acyltransferase ZDHHC5 might play a key role in the inhibition of PANoptosis in IDD. However, the role of ZDHHC5 in IDD has not been elucidated. In the present study, we verified the key role of ZDHHC5 in inhibiting PANoptosis by knocking down the level of ZDHHC5 in hUCMSCs or PTH pre-EVs. We found that PTH pre-EVs with downregulated ZDHHC5 expression weakened the enhanced anti-PANoptosis effect of PTH pre-EVs in IDD. Thus, the key effect of ZDHHC5 in inhibiting PANoptosis was revealed.
In the present study, we confirmed that ZDHHC5 could downregulate the downstream molecule ZBP1. ZBP1 is a crucial sensor in the innate immune system, known for detecting foreign nucleic acids, particularly in Z-DNA or Z-RNA configurations that are often present in viral genomes.13,56 Well known as a PANoptosis sensor, ZBP1 interacts with key proteins such as RIPK1, RIPK3, and caspase-8, forming the PANoptosome complex, which facilitates inflammatory cell death pathways necessary to eliminate infected or damaged cells.57,58 This pathway, known as PANoptosis, is integral to immune defenses against pathogens and involves multifaceted cell death mechanisms that combine pyroptosis, apoptosis, and necroptosis. 59 Moreover, ZBP1 activation through viral RNA detection has been shown to promote cytokine expression and inflammatory responses, underscoring its critical regulatory role in immune signaling. 60 Our data confirmed that ZBP1 was upregulated in degenerative NP cells and associated with intervertebral disc degeneration based on the single cell sequencing data from public cohorts.
Palmitoylation has emerged as a critical posttranslational modification that regulates protein–membrane associations and protein–protein interactions.61,62 Although nearly 3000 proteins are known to undergo palmitoylation, its role in modulating inflammation and innate immune responses remains unclear. 63 Given that ZDHHC5 functions as a palmitoyl acyltransferase, we hypothesized that ZDHHC5 might inhibit PANoptosis in NP cells by influencing palmitoylation processes. Co-IP-based LC-MS/MS was used to identify the ZDHHC5-interacting proteins. Our findings confirmed ZDHHC5 regulates YBX1 phosphorylation via palmitoylation. YBX1, as a transcription factor, binds specifically to the promoters of target genes, regulating transcription via phosphorylation. Here, ZDHHC5 inhibited YBX1 phosphorylation to suppress ZBP1 transcription, thereby hindering PANoptosis by promoting YBX1 palmitoylation.
This study has several limitations. First, we examined the effects of EVs in a mouse IDD model induced by needle puncture. Although this IDD model supported our findings, more advanced models, such as gene-edited mice or primate models, are necessary to further assess the therapeutic application of EVs. Second, only male mice were used in this study, despite documented sex-based differences in IDD progression and treatment response. Future studies should include both male and female subjects to improve the generalizability of our findings. Additionally, pain and functional disability were not assessed, which limits the translational relevance of our study. Third, additional factors in PTH-preconditioned EVs, including differentially expressed miRNAs, proteins, and non-coding RNAs, may also contribute to anti-PANoptosis effects. Furthermore, ZDHHC5 may have other molecular targets in NP cells, warranting further investigation. We used human-derived EVs to treat mice in a cross-species approach; while cross-species EV therapy has been explored previously, further studies are needed to confirm the biosafety and immunoreactivity of EVs. Finally, the mechanisms underlying PTH-induced EV secretion discussed in this study require additional exploration.
Conclusions
Collectively, the data from our study demonstrate that EVs secreted by PTH-stimulated hUCMSCs exhibit enhanced effects against PANoptosis in NP cells compared to unstimulated EVs. Furthermore, PTH promotes the quantitative secretion of hUCMSC-derived EVs and alters the cargo composition within EVs derived from PTH-preconditioned hUCMSCs. Additional analyses revealed that PTH-preconditioned EVs, containing elevated levels of ZDHHC5, ameliorate PANoptosis by regulating ZBP1 transcription through competitive inhibition of YBX1 phosphorylation via palmitoylation. Our findings strongly support a cell-free therapeutic strategy using EVs derived from PTH-preconditioned MSCs for the treatment of IDD and propose the ZDHHC5/YBX1/ZBP1 axis as a novel molecular target for inhibiting PANoptosis.
Supplemental Material
sj-docx-1-tej-10.1177_20417314251351011 – Supplemental material for Amelioration of intervertebral disc degeneration using engineered extracellular vesicle-delivered ZDHHC5 via inhibiting PANoptosis
Supplemental material, sj-docx-1-tej-10.1177_20417314251351011 for Amelioration of intervertebral disc degeneration using engineered extracellular vesicle-delivered ZDHHC5 via inhibiting PANoptosis by Tong Chen, Helong Zhang, Songtao Lv, Yue Xu, Wei Ren, Yifan Wei, Tianyou Zhang, Yunru Ge, You Lv, Ruya Li, Hui Che, Kunxin Xie, Yongxin Ren and Cheng Ma in Journal of Tissue Engineering
Footnotes
Acknowledgements
Thanks to the team of Research Center of Bone and Stem Cells of Nanjing medical university for their help.
Author contributions
Tong Chen, Helong Zhang, and Songtao Lv performed most of the experiments and wrote the manuscript. Yue Xu and Wei Ren contributed to the experiments. Yunru Ge and Yifan Wei, You Lv, Ruya Li, and Tianyou Zhang analyzed the data. Kunxin Xie contributed to the discussion. Hui Che reviewed the manuscript. Yongxin Ren and Cheng Ma designed the study and reviewed the manuscript. All the authors read and approved the submitted manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study received financial support from the National Natural Science Foundation of China (82372483), the Sino-German Mobility Program (M-0332) granted by the Sino-German Center (SGC) and funded directly by the NSFC and the German Research Foundation (DFG), the China Postdoctoral Science Foundation (Certificate Number: 2023M741464), the Jiangsu Funding Program for Excellent Postdoctoral Talent (2023ZB018), and the Postgraduate Research & Practice Innovation Program of Jiangsu Province (JX10214175).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
