Abstract
Gingival and periodontal ligament fibroblasts are functionally distinct cell types within the dento-gingival unit that participate in host immune response. Their microenvironment influences the behavior and immune response to microbial challenge. We developed three-dimensional gingival and periodontal connective tissue equivalents (CTEs) using human fibrin-based matrix. The CTEs were characterized, and the heterogeneity in their innate immune response was investigated. The CTEs demonstrated no to minimal response to planktonic
Keywords
Introduction
The dento-gingival unit that forms a collar around the teeth comprises of epithelial (gingival, sulcular, and junctional) and connective tissue (gingival and periodontal) components. 1 In health, the oral commensals and the host defense mechanism offered by the dento-gingival tissues exist in a state of homeostasis. Dysbiosis in the microflora with the colonization of periodontal pathogens leads to a disruption in the homeostasis, local tissue destruction, and periodontitis. 2 Along with the epithelium, the gingival and periodontal connective tissues actively participate in innate immune responses against the bacteria in health and diseased states.3 –6 Despite being covered by gingival epithelium, the gingival, and periodontal connective tissues are not completely sterile even in healthy states. Instead, they are constantly exposed to microbes and/or their products, 7 which can drive the selective expansion of regional fibroblast sub-population with differential immune responses. 8 Hence, it becomes imperative to understand fibroblast heterogeneity within the dento-gingival unit and their immune response to early microbial colonization.
Gingival fibroblasts (GFs) and periodontal ligament fibroblasts (PDLFs) are the primary cell types within the connective tissues of superficial gingival and deeper periodontal tissues, respectively.
8
Although the GFs and PDLFs are spatially located close to each other and have a similar spindle-shaped morphology in vitro, they have distinct functional characteristics.8
–10 GFs and PDLFs are known to produce a variety of pro-inflammatory cytokines, including interleukin (IL)-1β, IL-6, IL-8, stromal-derived factor (SDF)-1, and tumor necrosis factor-alpha (TNF-α).11
–14 Interestingly, stimulation of monolayer cultures of GFs and PDLFs with periodontopathogens or their virulence factors like lipopolysaccharide (LPS) demonstrate heterogeneity in their pro-inflammatory cytokine response.4,11,14
–16 However, much of the literature is based on studies on monolayer cultures, and the findings are conflicting. GFs exposed to LPS from
There has been growing interest in the application of in vitro three-dimensional (3D) organotypic cultures of reconstructed epithelium,17 –20 connective tissue,21 –23 and full-thickness6,24 –30 equivalents to recapitulate native tissue microenvironment. The 3D culture models have recently enhanced our understanding of the immune response of the gingival epithelium against commensal,28,31 intermediate, 25 and late17,18,20,28 colonizing pathogenic microbiome. However, the isolated response of connective tissues to early microbial colonizers is poorly understood. Secondly, the use of 3D cultures to capture the topological heterogeneity of superficial gingival and deeper periodontal tissues and their immune responses to the oral biofilm colonizers has not been explored.
Therefore, the present study aimed to reconstruct and characterize connective tissue equivalents (CTEs) representative of gingival and periodontal tissues; and investigate their immune response to early microbial colonization represented by oral commensals
Methods
Cell culture and fabrication of gingival and periodontal CTEs
Primary PDLFs and GFs were isolated from non-carious human impacted third molars extracted with informed consent from healthy donors following Institutional Review Board approval (No. 2018/00256). The PDLFs were isolated from the tissue fragments scrapped from the middle one-third of the roots. The GFs were isolated from the gingival tissues attached to the peri-coronal area of the molar tooth. The tissue fragments were cultured as explants in Dulbecco’s modified Eagle’s medium (DMEM)-low glucose (Gibco) supplemented with 10% fetal bovine serum (Biowest), 1% penicillin-streptomycin (Gibco), and 0.50 µg/mL amphotericin-B (Gibco). The expanded fibroblasts were cultured under the same media conditions without amphotericin-B, and passage 4–7 were used for the fabrication of the CTEs.
Gingival and periodontal CTEs were fabricated using a fibrin-based hydrogel matrix as previously described. 30 Briefly, human fibrinogen (40 mg/mL, Merck Millipore) and polyethylene glycol, succinimidyl glutarate-terminated (PEG, 10 mg/mL, Sigma-Aldrich, 10 kDa) were mixed in a volume ratio of 4:1. After incubation at 37°C for 30 min, the PEG-fibrinogen solution was mixed with fibroblast cell suspension containing 25,000 GFs or PDLFs. The gelation of this mixture was initiated by adding an equal volume of human thrombin (6.25 U/mL, Sigma-Aldrich) in calcium chloride solution (40 mM, Sigma-Aldrich). The constructs (50 µL in volume) were cultured for 9 days within a low-serum media containing ascorbic acid, hydrocortisone, basic fibroblast growth factor, aprotinin, and 1% penicillin-streptomycin. 30 The antibiotics were eliminated from the culture media 3 days before challenge with oral microbes.
Bacterial culture
Exposure of CTEs to pattern recognition receptor agonists
CTEs were exposed with TLR-2 agonist (Pam3CSK4, Invivogen), and TLR-4 agonist (ultrapure
Bacterial challenge
For the bacterial challenge in planktonic state, the gingival and periodontal CTEs were challenged with the respective microbes (106 CFU/cm2) for 4 and 24 h. To recapitulate post-colonization events,
Whole-mount visualization using confocal reflectance microscopy
To visualize the collagen and other extracellular matrix (ECM) fibers, whole-mounts of the formalin-fixed CTEs were imaged using laser scanning confocal microscopy (Olympus FluoView™ FV1000) under reflectance mode as previously described. 34 Briefly, the samples were illuminated at 488 nm, and the reflected light was detected with photomultiplier tube detectors. Z-stack images were processed using Fiji/Image J (NIH, USA) and Imaris software (Oxford Instruments).
Histology, immunostaining, and fluorescence in situ hybridization
The CTEs were fixed in 10% neutral-buffered formalin (Sigma-Aldrich), processed, and embedded in paraffin. Tissue sections (5 µm thick) of formalin-fixed, paraffin-embedded (FFPE) tissues were stained with hematoxylin-eosin (H-E) for histological evaluation.
For immunostaining, the deparaffinized sections were subjected to heat-induced epitope recovery at 121°C in a pressure vessel (Retriever™ 2100, Aptum Biologics) and 0.01M (pH6) citrate buffer (Sigma-Aldrich). Following blockage of nonspecific staining, the sections were incubated with respective primary antibodies overnight at 4°C. The primary antibodies include rabbit polyclonal anti-collagen-I (Abcam, ab34710, dilution 1:600) and mouse monoclonal anti-vimentin (Novocastra, NCL-VIM-V9, dilution 1:500, Clone V9). Sections were washed, labeled with appropriate secondary antibodies (goat anti-mouse alexa fluor 488 and goat anti-rabbit alexa fluor 594, Molecular Probes) for 45 min, counterstained with DAPI, and mounted with anti-fade fluorescent mounting medium (Abcam). To visualize the bacteria on the CTEs, fluorescence in situ hybridization (FISH) was performed as per manufacturer’s specifications (Biovisible BV). Briefly, the deparaffinized and dehydrated tissue sections were hybridized using EUB 338 probe (5′-GCTGCCTCCCGTAGGAGT-3′) at 60°C in a humidified chamber for 16 h. The slides were washed and mounted for microscopy.
H-E stained slides were visualized using a brightfield microscope (Nikon Eclipse E600 equipped with NIS-Elements software). Immunostained slides were visualized under a fluorescence microscope (Leica DMi8, Leica Microsystems equipped with Leica Application Suite X software) and FISH slides were visualized using laser scanning confocal microscopy (Olympus FluoView™ FV1000).
Enzyme Linked Immunosorbent Assay (ELISA) for secretome analysis
In accordance with manufacturer’s instructions, ELISAs for cytokines IL-1β, IL-6, IL-8, and TNF-α (all from Biolegend) were performed using the culture supernatants. The absolute cytokine values were normalized to the total protein content using BCA assay (Thermo Fisher Scientific).
Statistical analysis
All experiments were performed as biological triplicates, and results are presented as mean ± SD. Statistical significance between groups was determined using ANOVA with Bonferroni corrections and independent t-test using Stata 16 statistical software. Differences were considered significant if
Results
Fabrication and characterization of gingival and periodontal CTEs
Donor-derived GFs and PDLFs were characterized for the expression of mesenchymal (CD26, CD44, CD73, CD90, CD105), neural/neural-crest (CD56, CD271) and major histocompatibility complex (HLA-ABC and HLA-DR) markers (Supplemental Figure S1 and Table T1). Results showed that there was no significant difference in the surface marker expression among the two fibroblasts. The GFs and PDLFs were embedded within a human fibrin-based matrix to construct 3D gingival and periodontal CTEs respectively (Figure 1). The fibrin-based matrix was stable, without any visible contraction or degradation over the 9-day culture period. H-E and immunostained sections of the CTEs revealed the presence of vimentin-positive, spindle-shaped fibroblasts that are uniformly distributed within the fibrin matrix (Figure 2(a) and (b)). The fibrin-based matrix provided a natural provisional matrix for the vimentin-positive fibroblasts to produce lamina propria-like ECM. This is evident by the strong expression of cell-derived fibrillar collagen-I in the gingival and periodontal CTEs (Figure 2(a)). Further, whole-mount label-free imaging using confocal reflectance microscopy demonstrated the presence of the fibroblasts within a 3D network of thick cell-derived collagen fibers interspersed with fine collagen fibrils (Figure 2(c) and (d), and Supplemental Videos SV1 and SV2). In most places, the fibroblasts and their cellular processes were embedded within the dense cell-derived collagen network (Figure 2(b)). Amorphous material representative of native fibrin matrix was also seen interspersed between the newly formed collagen fibers. This data correlated well with the immunostaining images wherein a stronger signal of collagen-I was present in the proximity and periphery of the fibroblasts of both gingival and periodontal CTEs. Taken together, these findings suggest matrix remodeling and de novo deposition of collagen-containing ECM by the respective fibroblasts.

Schematic depicting the workflow for fabrication of gingival and periodontal CTEs and subsequent microbial challenge.

Characterization of connective tissue equivalents (CTEs): (a) H-E and immunostaining of gingival and periodontal CTEs with uniformly distributed, vimentin-positive fibroblasts and cell-derived collagen-I rich extracellular matrix (
CTEs exhibit differential responses to TLR-2 and TLR-4 agonists
Before experiments with bacteria, we investigated the response of gingival and periodontal CTEs to surrogate challenge with TLR-2 (Pam3CSK4) and TLR-4 (

Secretome analysis of connective tissue equivalents (CTEs) exposed to TLR agonists. Graphs showing IL-6 and IL-8 production by gingival (a) and periodontal CTEs (b) after dose- and time-dependent challenge with TLR-2 agonist (Pam3CSK4). Graphs showing IL-6 and IL-8 production by gingival (d) and periodontal CTEs (e) after time-dependent challenge with TLR-4 agonist (
CTEs elicit a heterogenous innate immune response to planktonic bacteria
We next investigated the innate immune response of gingival and periodontal CTEs to live bacteria (early and intermediate colonizers) exposed in a planktonic state. To recapitulate the initial colonization events, the CTEs were challenged with commensals or primary colonizers
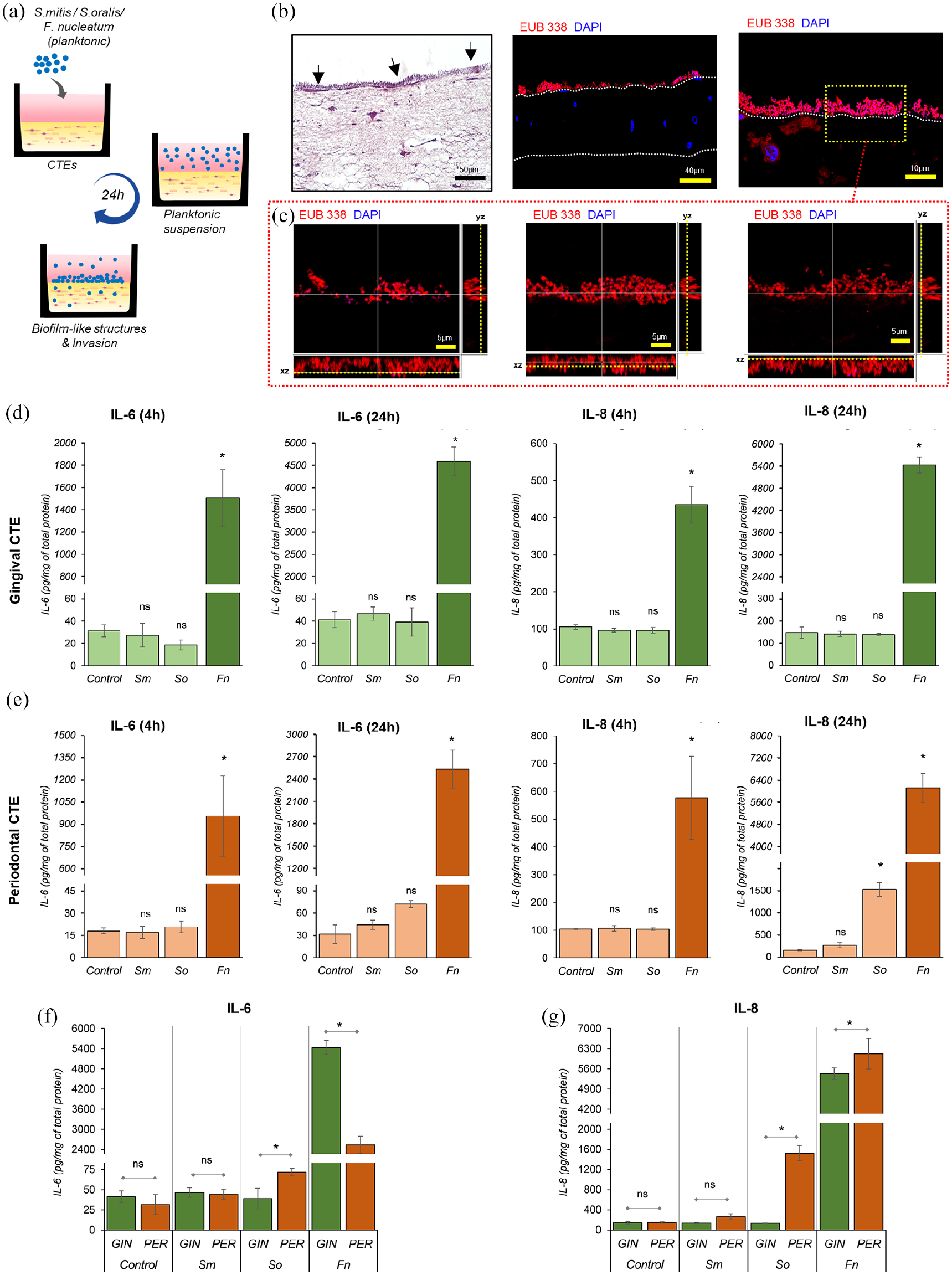
Innate immune response by CTEs after co-culture with bacteria in planktonic state: (a) Schematic representing the exposure of CTEs with primary and intermediate colonizers in planktonic state and formation of biofilm-like clusters on tissue equivalents after 24 h of co-culture, (b) H-E micrographs showing bacterial clusters (black arrows) on the CTEs (
The innate immune response of the CTEs was assessed by quantifying the production of IL-1β, IL-6, IL-8, and TNF-α in the culture supernatants. There was no significant difference in the IL-6 and IL-8 production by gingival CTEs co-cultured with
In contrast to the primary colonizers,
Overall, these results show the contrasting cytokine response of gingival and periodontal CTEs to primary and intermediate colonizers in the planktonic state.
Gingival and periodontal CTEs elicit differential immune responses to bacterial biofilms
We next investigated the immune response of the two CTEs to bacterial biofilms post-colonization, wherein the cultures were exposed to 24 h-old biofilms (Figure 5(a)). The total biomass of the microbial biofilms computed based on the 3D reconstruction of confocal z-stack images showed similar biomass across all bacterial species (Figure 5(b) and (c)). Viability assessment of bacterial biofilms of

Response by CTEs after exposure with commensal biofilms and the impact of tissue heterogeneity: (a) Schematic showing method employed to culture
In contrast to the planktonic state, the CTEs exposed to primary colonizers in the biofilm elicited a moderate immune response demonstrated by significant secretion of IL-6 and IL-8 (Figure 5(f)). However, the IL-6 and IL-8 levels were significantly lower than that elicited by
Discussion
This study presents a method to fabricate 3D gingival and periodontal CTEs using a human fibrin-based matrix and investigate the heterogeneity in their innate immune response to early microbial colonization. To understand the host-microbiome interaction of the CTEs, an in vitro infection model employing a range of bacterial challenges that include surrogate TLR-2 and TLR-4 agonists, commensal bacteria (

Schematic shows that tissue topography influences immune responses to early colonizers wherein the superficial connective tissue (gingival CTE) responds through higher pro-inflammatory cytokine IL-6 secretion and the deeper connective tissue (periodontal CTE) through higher IL-8 production.
Exposure of GFs and PDLFs cultured as monolayers are commonly used for host-microbiome studies, till date. They are the simplest models, easy to setup, and effective for mechanistic studies. However, the results from monolayer culture models may not represent the features of complex 3D cell-cell and cell-matrix interactions 35 and their impact on the tissue response to bacterial challenge. Therefore, in this study, we developed a 3D culture model to represent the gingival and periodontal connective tissue elements of the dento-gingival unit, respectively. ECM is an integral component of connective tissue and the host cellular microenvironment. Collagen-based matrices (typically from rat-tail and bovine origin) are commonly used for the fabrication of 3D cultures.22,23,36 However, it has technical and translational limitations owing to its contractile nature and xenogeneic origin. 36 Contraction of the collagen gels due to polymerization and fibroblast-mediated process raises technical concerns on reproducibility and long-term cultures. 36 Polymeric scaffolds are thus commonly used to counteract the contraction and handling issues with collagen matrices. 21 Unlike commonly used collagen matrices,22,36 the human fibrin-based matrix used in this study was stable without any visible contraction or degradation issues, thus enabling culture over a long period. Non-invasive and label-free imaging using confocal reflectance microscopy supplemented with immunostaining for collagen-1 revealed that the GFs and PDLFs deposited de novo cell-derived collagen and ECM fibers.
The ECM plays a critical role in providing substrate for microbial adhesion and penetration in the host. There is growing evidence that various ECM components and their degradation products also contribute to tissue inflammation and innate immune response to microbial pathogens.37,38 ECM components and degradation products of collagen, glycoproteins (e.g. fibrinogen, fibronectin, tenascin-C), proteoglycans (e.g. aggrecan, biglycan, decorin, versican), and glycosaminoglycans (e.g. hyaluronan, heparan sulfate) which can be potentially released from damaged tissues have been identified as capable of acting as danger-associated molecular patterns. 37 Hence, the choice of ECM components used to fabricate 3D tissue equivalents can potentially impact the innate immune response against microbial pathogens. For instance, low molecular weight hyaluronic acid has been shown to activate TLR-2 signaling, while its high molecular counterpart inhibits TLR-2 signaling. 39 Similarly, fibrin and its precursor fibrinogen have been demonstrated to display differential regulation of macrophages upon stimulation with LPS and interferon-γ. 40 Macrophages cultured on fibrin gels displayed an anti-inflammatory phenotype (increased production on IL-10 and decreased levels of tumor necrosis factor alpha), while its precursor fibrinogen stimulated inflammatory activation. Future studies on the impact of various ECM components and their degradation products (in response to host or bacterial proteases) on the innate immune response against periodontopathogens can provide insights on developing ECM-based immunomodulatory strategies.41,42
The in vitro infection of CTE models used in this study, enabled the potential to recapitulate spatiotemporal early microbial colonization by planktonic oral commensals. The planktonic oral commensals formed well-defined biofilm-like structures over the tissues after 24 h of co-culture. Alternatively, as demonstrated in this study and others,27 –29,43 the 3D organotypic tissues could be challenged with preformed bacterial biofilms. The 3D architecture of organotypic cultures also offers the advantage of visualizing microbial adhesion, biofilm formation, and microbial invasion.28,29 The bacterial challenge in the form of both live planktonic and biofilm states used in this study and studies using organotypic gingival epithelium,27 –29,43 results in the recapitulation of the host-microbiome interface of native gingival and periodontal tissues (albeit without the overlying gingival epithelium). In contrast, host-microbiome studies on monolayer cultures are typically based on exposure of microbiome in planktonic states using multiplicity of infection or CFU/mL.4,16,44 However, exposure on 3D culture models offer the potential for exposure based on surface area and as microbial biofilms. This precludes the ability for direct comparison of the cellular responses between monolayer and 3D culture systems.
Previous studies using monolayer cultures of GFs and PDLFs and gene expression or microarray analysis have demonstrated the heterogeneity in constitutive expression of various genes, including inflammatory cytokines among the two cell types.8,11
–14,45 Further, this translates to contrasting cytokine responses of the GFs and PDLFs to periodontopathogens like
Previous studies have demonstrated the involvement of TLR and nucleotide-binding oligomerization domain (NOD)-containing protein-like receptors (NLR) signaling pathways to mediate the innate immune response through the production of cytokines like IL-6 and IL-8.5,43,48
–52 TLRs are a family of membrane-associated PRRs that lead to the activation of nuclear factor kappa B (NF-κB), interferon regulatory factors (IRF), and mitogen-activated protein kinases (MAPK) signaling pathways.43,52,53 On the other hand, NLRs are located in the cytoplasm, and are predominantly activated through bacterial invasion into the cells. Activation of intracellular NOD1 and(or) NOD2 in GFs and PDLFs could mediate the release of IL-6 and IL-8 through the activation of NF-κB and MAPK signaling pathways.
49
In the present study, exposure to TLR-2 agonist triggered a significantly higher IL-6 and IL-8 among gingival CTEs than its periodontal counterpart. On the contrary, we did not observe any immune response by both CTEs exposed to TLR-4 agonist (ultrapure
In this study, the exposure of gingival and periodontal CTEs to oral commensals in the planktonic state triggered a none to mild innate immune response. In contrast to
Future studies on exposure of the CTEs to pathogenic biofilms, particularly from diseased sites, could help provide a deeper understanding of the priming ability of commensals and immune evasion strategy of pathogens. Further, to mimic the microphysiology, it would be beneficial to investigate the response of the CTEs primed by commensal biofilms before exposure to pathogenic biofilms. However, such a setup would require a prolonged culture period, which may not be feasible in the current 3D culture system due to bacterial overgrowth and nutrient depletion. Next-generation tools like microfluidic organ-on-chip systems provide the opportunities to continuously perfuse the tissues for better nutrient delivery and drainage of metabolic wastes. 59 Further, the microfluidic channels and compartmentalization can enable the potential to introduce different bacterial challenges at desired time points and over the long-term culture period.60,61 Microfluidic gut-on-chip models have demonstrated the ability to model and study long-term host-microbiome interactions between intestinal bacteria and gut mucosa. 60 One of the limitations of our model is the lack of immune cells that help to amplify and translate the innate immune response of the connective tissues. Microfluidic systems that incorporate or present the immune cells at defined time points can also biomimic the recruitment and migration of immune cells into the tissue.62,63 Future studies combining the 3D culture and microfluidic systems can be transformative and provide novel insights into host-microbial interactions and opportunities to develop novel therapeutic strategies.
Supplemental Material
sj-docx-1-tej-10.1177_20417314221111650 – Supplemental material for Differential immune responses of 3D gingival and periodontal connective tissue equivalents to microbial colonization
Supplemental material, sj-docx-1-tej-10.1177_20417314221111650 for Differential immune responses of 3D gingival and periodontal connective tissue equivalents to microbial colonization by Hardik Makkar, Srividya Atkuru, Yi Ling Tang, Tanya Sethi, Chwee Teck Lim, Kai Soo Tan and Gopu Sriram in Journal of Tissue Engineering
Supplemental Material
sj-docx-2-tej-10.1177_20417314221111650 – Supplemental material for Differential immune responses of 3D gingival and periodontal connective tissue equivalents to microbial colonization
Supplemental material, sj-docx-2-tej-10.1177_20417314221111650 for Differential immune responses of 3D gingival and periodontal connective tissue equivalents to microbial colonization by Hardik Makkar, Srividya Atkuru, Yi Ling Tang, Tanya Sethi, Chwee Teck Lim, Kai Soo Tan and Gopu Sriram in Journal of Tissue Engineering
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from iHealthtech Microbiome Seed Grant (A-0002947-00-00, A-0002947-01-00), and Singapore Ministry of Education Academic Research Fund Tier 1 (A-0002090-00-00; A-0002084-00-00). Hardik Makkar is supported by President Graduate Fellowship, and Srividya Atkuru is supported by NUS Research Scholarship.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
