Abstract
Pulmonary arterial hypertension (PAH) is a rare but fatal cardiovascular disorder with high morbidity and mortality. Diagnosis and treatment of this disease at an early stage would greatly improve outcomes. The molecular indicators of PAH are mostly nonspecific, and diagnostic and prognostic biomarkers are urgently needed. A more comprehensive understanding of the molecular mechanisms underlying this complex disease is crucial for the development of new and more effective therapeutics to improve patient outcomes. In this article, we review published literature on proteomic biomarkers and underlying molecular mechanisms in PAH and their value for disease management, aiming to deepen our understanding of the disease and, ultimately, pave the way for clinical application.
Introduction
Pulmonary arterial hypertension (PAH) is a rare but fatal cardiovascular disorder with high mortality despite therapeutic advances. PAH is characterized by a sustained increase in mean pulmonary arterial pressure (mPAP > 20 mmHg), normal pulmonary capillary wedge pressure (PCWP ≤ 15 mmHg) and increased pulmonary vascular resistance (PVR > 3 Wood units). 1 In PAH, the pulmonary vasculature is dynamically obstructed by vasoconstriction, structurally obstructed by adverse vascular remodeling, and pathologically noncompliant as a result of vascular fibrosis and stiffening. 2 According to the new classification, PAH is classified as follows: idiopathic (IPAH); heritable (HPAH); drug- and toxin-induced; associated with related factors, including connective tissue disease, HIV infection, portal hypertension, congenital heart disease (CHD) and schistosomiasis; PAH in long-term responders to calcium channel blockers; PAH with overt features of venous/capillary (PVOD/PCH) involvement; and persistent pulmonary hypertension ( PH of the newborn syndrome. 1 The incidence of PAH ranges from 2.0 to 7.6 cases per million adults per year, and its prevalence varies from 11 to 26 cases per million adults. 3 Early clinical symptoms of PAH are nonspecific and overlap with those of other cardiorespiratory diseases, leading to misdiagnosis or delayed diagnosis. The current treatment strategies for PAH are insufficient, and the prognosis is poor. There is undoubtedly an unmet need for the detection of dependable biomarkers and the development of novel therapeutic strategies.
Multiple advances have been made in the molecular analysis of the mechanisms underlying PAH. In particular, several genetic alterations have been identified, such as heterozygous germline mutations of bone morphogenetic protein receptor type-2 (BMPR2), a member of the transforming growth factor beta (TGF-β) family. 4 Mutations in this gene account for 70%–80% of HPAH cases, as well as for 15%–25% of IPAH cases; 2 multiple other genes are also related to this disease. 5 However, complex processes occur during gene expression, such as transcriptional regulation, alternative splicing and RNA editing, and proteins execute such functions. Thus, the measurement of protein levels is better for examining the clinical phenotype than raw transcriptome data. 6 Indeed, with the development of high-throughput mass spectrometry (MS) and analysis software, proteomics has become a vital complement to genomics approaches, providing valuable information about unresolved aspects of PAH.
Protein microarrays designed based on known proteins or peptides first enabled an increase in the throughput capacity to discover protein biomarkers. Then, gel-based and MS-based techniques, including two-dimensional gel electrophoresis (2-DE) and two-dimensional difference gel electrophoresis (2D-DIGE), were implemented as high-throughput technologies for proteomics. 7 However, the main problems with these methods are their poor reproducibility, small protein load capacity, and inability to detect proteins with low abundance and strong hydrophobicity. 8 The most commonly utilized method for analyzing proteins by MS is a bottom-up approach to identify and quantify enzymatically digested peptides. The two traditional methods for protein identification by MS are matrix-assisted laser desorption/ionization-time of flight (MALDI-TOF) and liquid chromatography (LC) with electrospray ionization (ESI). Nano ESI coupled with nano high-performance LC (nHPLC) has become the standard LC-MS approach in the majority of proteomics labs. 9 In addition, surface-enhanced laser desorption/ionization (SELDI), another strategy for the MS analysis of macromolecules, has been introduced, simplifying sample extraction and facilitating effective on-probe investigations of biopolymers when compared to conventional MALDI approaches. 10 Quantitative proteomics can be separated into shotgun proteomics and targeted proteomics approaches. The former can be performed based on mass tags, including isobaric tags for relative and absolute quantification labels (iTRAQ), tandem mass tag (TMT), and stable isotope labeling using amino acids in cell culture (SILAC) or by calculating the spectral count, which is a label-free quantitative approach. 11 Labeling combined with tandem mass spectrometry (MS/MS) strategies allow the quantification and identification of many different pooled samples in one experiment, with high sensitivity and reproducibility. 9 Moreover, label-free quantification overcomes the limitation of sampling conditions in label-based methods and has the capacity to evaluate large samples, although demands for data and information analysis are high. The methods mentioned above are frequently used, and new proteomic technologies are still evolving.
A wide variety of clinical specimens, including blood (plasma/serum), heart/lung tissues (fresh frozen and formalin-fixed paraffin-embedded tissue blocks) and different cell types from patients or murine models, such as monocrotaline (MCT)- and pulmonary artery binding (PAB)-induced PAH rats, are commonly used in PAH proteomic studies. Conversely, other body fluids, such as urine and bronchoalveolar lavage fluid, have not been used in PAH studies to date. The classical workflow for PAH proteomics studies using various biospecimens is shown in Figure 1. In this review, we summarize the current proteomic studies and outline those that are proposed to be valuable in the field of PAH. The main findings were summarized in Figure 2.
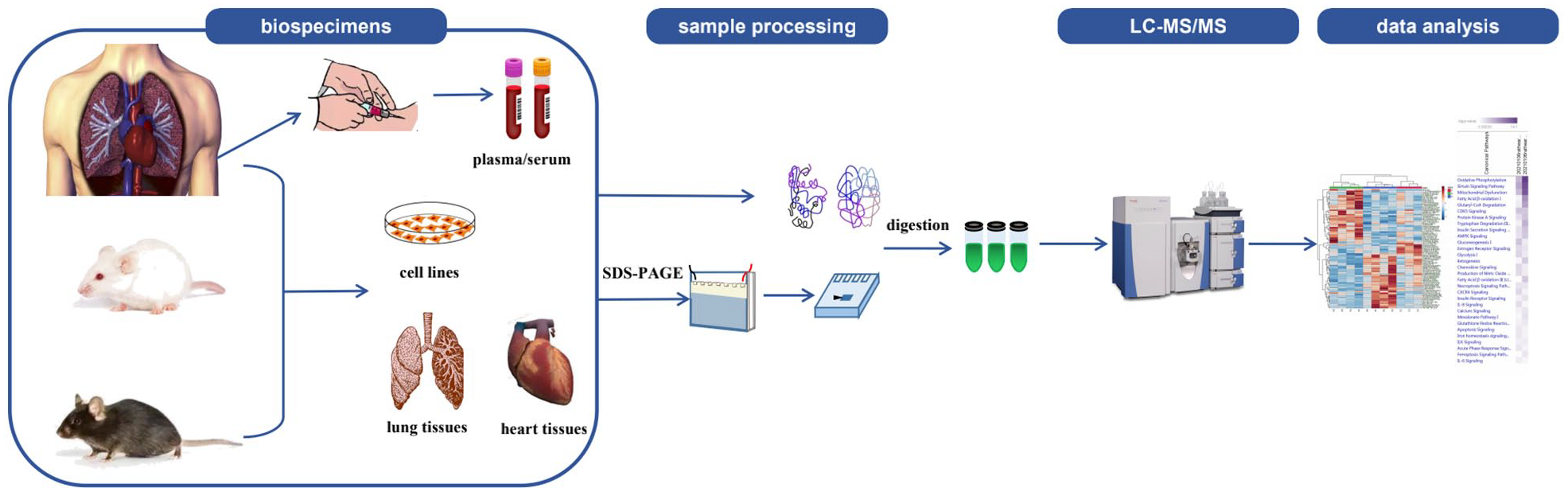
An overview of proteomics workflow and commonly used biospecimens for studying PAH. Commonly used biospecimens in proteomic studies of PAH include blood (plasma/serum), heart/lung tissues and different cell types from patients and murine models. Samples are extracted via gel-based or gel-free approaches and then digested enzymatically. Peptides are then fractionated using liquid chromatography (LC) and then subjected to mass spectrometry (MS). Proteins are identified in databases, and functional analyses are performed using bioinformatic methods.

The main findings from proteomic analysis of PAH.
Proteomic studies to identify PAH biomarkers for potential clinical use
Although the list of biomarkers is constantly growing,12–14 and includes TNF-like weak inducer of apoptosis (TWEAK), pro-atrial natriuretic peptide (proANP), growth differentiation factor-15 (GDF-15), soluble fms-like tyrosine kinase 1, placental growth factor, and galectin-3, guidelines recommend only a small number of them, such as brain natriuretic peptide (BNP), N-terminal proBNP (NT-proBNP), and cardiac troponin (cTn). Two major biomarkers of PAH were recently proposed in the 2019 European Respiratory Society (ERS) International Congress.
15
One study reported that the levels of endostatin (ES), a potent angiostatic factor, are elevated in PAH patients and distinguishes them from healthy controls (HCs) at a serum threshold of 66 ng/ml, with an area under the receiver operator characteristic (ROC) curve (AUC) of 0.879.
16
The ES level in PAH patients was found to correlate positively with the mPAP, right atrial pressure (RAP), 6-min walking distance (6MWD), and cardiac output (CO).16,17 ES could also identify nonsurvivors, with an AUC of 0.675 (
A comprehensive list of proteomic studies focusing on blood biomarkers of PAH.

ADIPO, adiponectin; ADMA, asymmetric dimethylarginin; ANPEP, alanyl membrane aminopeptidase; ApoE, apolipoproteins E; AUC, area under the ROC curve; DBH, dopamine β-hydroxylase; 2-DE, two-dimensional gel electrophoresis; Epo, erythropoietin; ELISA, enzyme-linked immunosorbent assay; FSTL3, Follistatin-like 3; GNAS, guanine nucleotide-binding protein Gs; GP1BA, glycoprotein Ib platelet-subunit; HC, healthy control; HPAH, heritable pulmonary arterial hypertension; IGFBP1, insulin-like growth factor binding protein-1; IL-1R1, interleukin-1 receptor-like 1; IPAH, idiopathic pulmonary arterial hypertension; iTRAQ, isobaric tags for relative and absolute quantification labels; LC-MS, liquid chromatography-mass spectrometry; LRG, leucine-rich a-2-glycoprotein; MALDI-TOF, matrix-assisted laser desorption/ionization-time of flight; MDK, Midkine; MS, mass spectrometry; 6MWD, 6-minute walking distance; NYHA, New York Heart Association; SSc, systemic sclerosis; TIMP, tissue inhibitors of metalloproteinases; TRF1, transferrin receptor 1; UHPLC, ultra-high-performance liquid chromatography; VSD, ventricular septal defect; PAH, Pulmonary arterial hypertension; WHO, World Health Organization.
Diagnostic biomarkers for PAH
Currently, the diagnosis of PAH is mainly based on symptoms, physical examination and several procedures, including right heart catheterization (RHC).
26
Several molecular biomarkers, such as pentraxin 3 (PTX3), caveolin-1 (Cav1), selenoprotein P and interleukin (IL)-32, have potential diagnostic value, but large-scale verification is lacking.27–30 Proteomic research can identify whole protein profiles and constitutes a potent method for novel biomarker discovery. Abdul-Salam
Prognostic/predictive biomarkers for PAH
Several recent registries have identified some effective predictors of survival outcomes, such as age, sex and factors reflective of right heart function. 31 Most prognostic markers recommended by the European Society of Cardiology (ESC) and ERS guidelines are related to right ventricular function and CO, including various indices of exercise tolerance, mean RAP, and cardiac index (CI). 26 At present, the most popular tools for PAH risk stratification and prognostication are the risk scores published by the US Registry to Evaluate Early and Long-Term PAH Disease Management (REVEAL), the 2015 ESC/ERS PH guidelines risk table and a multiparametric risk stratification approach raised in the sixth World Symposium on Pulmonary Hypertension (WSPH), in which BNP or NT-proBNP is the only laboratory biomarker, which is absolutely not specific in PAH.32–34 More recently, investigators have derived additional risk assessment strategies, but molecular biomarkers are still lacking.32,35–37
Using LC-MS/MS, Sandqvist
There is a growing consensus that the sensitivity and specificity of biomarker panels, multiple markers used in combination, are increased compared to those of proteins utilized individually.
9
Yuan
In addition, complex changes in cytokines, cellular immunity and autoantibodies suggest that PAH is, in part, an autoimmune, inflammatory disease.
2
A multiplex immunoassay performed by Sweatt
Nevertheless, blood samples do have some limitations. The presence of a few highly abundant proteins, such as albumin and immunoglobulin, can mask proteins of lower abundance that may be biologically important; ready-made albumin and IgG removal kits are available. Another limitation of serum samples is that a small series of protein changes might occur after plasma is allowed to clot, which may prevent the detection of some disease-related proteins. 19 However, the establishment of blood biomarkers that can be noninvasive is still an ideal choice for PAH patients. In addition, serum/plasma can be a true indicator of disease progression and can help in longitudinal studies monitoring minute changes in the proteome. 38
Proteomics studies to uncover the mechanisms underlying PAH
Progress has been made in understanding the pathophysiological mechanisms of PAH and approving drug therapies. For example, a cancer-like increase in cell proliferation and resistance to apoptosis reflects acquired abnormalities of mitochondrial metabolism and dynamics. Epigenetic dysregulation of DNA methylation, histone acetylation, and microRNAs also contributes to disease pathogenesis. 2 In addition to identifying protein biomarkers, proteomics has been useful for uncovering the mechanism of PAH, especially with the development of bioinformatic technology and analysis software. Comprehensive lists of proteomic studies focusing on PAH occurrence and progression and on PAH-related gene mutations as well as their main findings are presented in Tables 2 and 3, respectively.
A comprehensive list of proteomic studies focusing on PAH occurrence and progression.

ATP5F1A, ATP synthase subunit alpha; C3, complement 3; CFHR2, complement factor H-related protein 2; CHCHD3, MICOS complex subunit; CHD, congenital heart disease; CLIC4, chloride intracellular channel 4; CPS1, carbamoyl-phosphate synthetase I; 2D-DIGE, two-dimensional fluorescence difference gel electrophoresis; 2-DE, two-dimensional gel electrophoresis; DEPs, differentially expressed proteins; EIF5B, eukaryotic translation initiation factor 5B; FABP4, fatty acid binding protein-4; HADHA, trifunctional enzyme subunit alpha; HADHB, trifunctional enzyme subunit beta; HC, healthy control; HPLC-ESI, high-performance liquid chromatography-electrospray ionization; HSP-27, heat shock protein-27; IMMT, MICOS complex subunit MIC60; IPAH, idiopathic pulmonary arterial hypertension; iTRAQ, isobaric tags for relative and absolute quantification labels; LC-MS, liquid chromatography-mass spectrometry; MALDI-TOF, matrix-assisted laser desorption/ionization-time of flight; MCT, monocrotaline; MRPL34, mitochondrial ribosomal protein L34; MS, mass spectrometry; NDUFS1, NADH-ubiquinone oxidoreductase 75 kDa subunit; PAB, pulmonary arterial banding; PABPN1, polyadenylate-binding nuclear protein 1; PAEC, pulmonary artery endothelial cell; PAH, pulmonary arterial hypertension; PDE5A, phosphodiesterase type 5 A; PH, pulmonary hypertension; PKA, protein kinase; RV, right ventricle; SLC25A3, phosphate carrier protein; SUCLG2, succinate-CoA ligase subunit beta; SuHx, Sugen5416 with hypoxia; TGOLN2, trans-Golgi network integral membrane protein 2; TMT, tandem mass tag; UQCRC2, Cytochrome b-c1 complex subunit 2; ZC3 H4, zinc finger CCCH domain-containing protein 4.
A comprehensive list of proteomic studies focusing on PAH-related gene mutations.
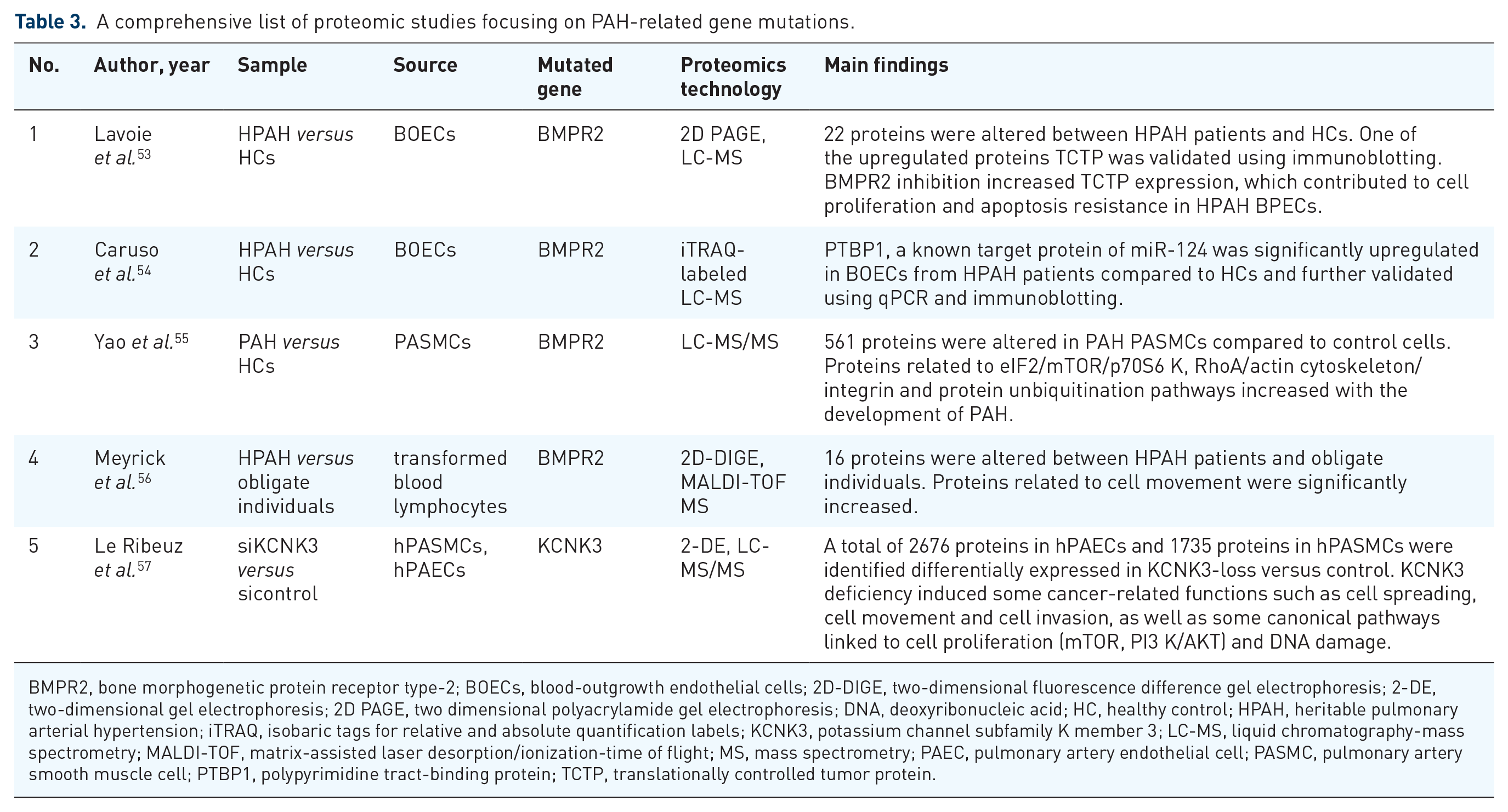
BMPR2, bone morphogenetic protein receptor type-2; BOECs, blood-outgrowth endothelial cells; 2D-DIGE, two-dimensional fluorescence difference gel electrophoresis; 2-DE, two-dimensional gel electrophoresis; 2D PAGE, two dimensional polyacrylamide gel electrophoresis; DNA, deoxyribonucleic acid; HC, healthy control; HPAH, heritable pulmonary arterial hypertension; iTRAQ, isobaric tags for relative and absolute quantification labels; KCNK3, potassium channel subfamily K member 3; LC-MS, liquid chromatography-mass spectrometry; MALDI-TOF, matrix-assisted laser desorption/ionization-time of flight; MS, mass spectrometry; PAEC, pulmonary artery endothelial cell; PASMC, pulmonary artery smooth muscle cell; PTBP1, polypyrimidine tract-binding protein; TCTP, translationally controlled tumor protein.
Disease occurrence and progression
Using a gel-based approach, Régent
By examining surgically acquired lung samples from IPAH/HPAH patients undergoing lung transplantation (
In addition to the commonly used samples, samples such as platelets and exosomes have received recent attention in the proteomic field. As platelets from IPAH patients exhibited typical metabolic shifts and activation defects, Aulak
Pathogenic alterations underlying PAH-related gene mutations
BMPR2 was the first mutated gene to be associated with PAH, although the mechanism by which it acts was not entirely clear. One of the notably upregulated proteins in the blood-outgrowth endothelial cells (BOECs) of HPAH patients with BMPR2 mutations was identified by proteomic analysis as translationally controlled tumor protein (TCTP), a protein promoting tumor cell growth and survival; TCTP was further validated in the Sugen5416 injection combined with hypoxia (SU5416) rat model of severe PAH. This result indicated that increased TCTP expression was associated with intimal lesions and proliferating ECs and may be an important mediator of vessel remodeling.
53
Using an iTRAQ-labeled proteomic approach, Caruso
Using an LC-MS/MS-based proteomics approach, Le Ribeuz
Response to treatment
PAH is characterized by elevated PAP and is managed by vasodilator therapies. Despite great advances in the treatment of PAH, it is still a devastating disease with high mortality. Novel targets and identification of suitable candidates for PAH therapy are urgently needed. Nogueira-Ferreira
Huang
Different alterations between RV and LV
In most cases, the study of PAH is limited to the right heart and pulmonary circulation system. Studies focused on different alterations between the left ventricle (LV) and RV have also been conducted. It is well established that MCT has toxic effects, which can be directly observed in the myocardium; thus, proteomic analysis might be biased by this fact. In the MCT model, both ventricles are affected by enhanced neuroendocrine stimulation, but only the RV exhibits excessive pressure load.
67
Schott
Proteomic studies to characterize deep phenotypes of PAH
Except for HPAH, PAH subtypes are mainly classified by clinical features, which are not relevant to molecular biology pathogenesis. Although they all belong to PAH based on the WSPH classification, different subtypes of PAH have both common and specific mechanisms. According to the abovementioned proteomic studies, we summarized the differences and similarities of the altered proteins and pathways among IPAH, HPAH, CHD-PAH, and SSc-PAH (Supplementary Table 1). Funded by the National Heart, Lung, and Blood Institute (NHLBI), the pulmonary vascular disease phenomics program (PVDOMICS, NCT02980887) aims to perform reclassification and deep phenotyping of PH based on molecular and cellular information using multiomics approaches (genomics, transcriptomics, proteomics, metabolomics, coagulomics, cell biomics), which could assist with personalized diagnoses, prevention and treatment in the long term. In the future, we expect more innovative findings from currently undergoing programs in this area will be published in article form.
Conclusion
To conclude, several studies have examined the proteome of PAH and successfully discovered potential biomarkers for disease occurrence, diagnosis, prognosis and response to treatment. Most of these biomarkers are related to previously accepted mechanisms but provide details about more concrete molecular changes, such as energy metabolism, endothelial function, regulation of cell growth, inflammation and the immune system. Moreover, some newly proposed biomarkers provide further insight into the disease. Although many candidate protein markers and latent mechanisms have been reported, few have been deeply investigated and implemented in the clinic. Whether the proteins and mechanistic pathways acquired from proteomic studies aid clinical decisions needs more validation in prospective clinical cohorts and deeper mechanistic mining grounded in basic research. Integration of multiomics approaches is imperative for the elucidation of molecular alteration at multiple levels and for providing more targeted directions for further in-depth research. We should pay continuous attention to the innovative outcomes from the large-scale project, such as the PVDOMICS, the U.K. PAH Cohort Study Consortium, and the U.S. PAH Biobank Consortium. While progress has been made regarding elucidation of the underlying mechanisms and approval of drug therapies, proteomics, and posttranslational modifications (PTMs) will continue to broaden our understanding of PAH pathogenesis and promote the development of proteomics-driven precision medicine (PDPM) for PAH. Ultimately, integrating the data from all platforms will deepen our understanding of the disease and pave the way for better outcomes for patients with PAH.
Supplemental Material
sj-docx-1-taj-10.1177_20406223211047304 – Supplemental material for Proteomic analysis of pulmonary arterial hypertension
Supplemental material, sj-docx-1-taj-10.1177_20406223211047304 for Proteomic analysis of pulmonary arterial hypertension by Xiaohan Qin, Tianhao Li, Wei Sun, Xiaoxiao Guo and Quan Fang in Therapeutic Advances in Chronic Disease
Footnotes
Author contributions
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Chinese Academy of Medical Sciences Innovation Fund for Medical Sciences (grant number 2018-I2M-1-001), the National Natural Science Foundation of China (grant number 81500306), and Xinxin-Merck Cardiovascular Research Fund (grant number 2017-CCA-xinxin merck fund-005).
Conflict of interest statement
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
