Abstract
This review summarizes the salient points of the symposium ‘Red Cell Genotyping 2015: Precision Medicine’ held on 10 September 2015 in the Masur Auditorium of the National Institutes of Health. The specific aims of this 6th annual symposium were to: (1) discuss how advances in molecular immunohematology are changing patient care; (2) exemplify patient care strategies by case reports (clinical vignettes); (3) review the basic molecular studies and their current implications in clinical practice; (4) identify red cell genotyping strategies to prevent alloimmunization; and (5) compare and contrast future options of red cell genotyping in precision transfusion medicine. This symposium summary captured the state of the art of red cell genotyping and its contribution to the practice of precision medicine.
Introduction
Molecular immunohematology is gaining clinical application in transfusion medicine. To focus this field of immunohematology, a conference was initiated in 2011 entitled Red Cell Genotyping: At the Tipping Point. After that conference, BloodCenter of Wisconsin and the Department of Transfusion Medicine, NIH Clinical Center have co-sponsored annual red cell genotyping symposia since 2012. Several molecular immunohematology applications are now firmly established.
Personalized medicine is the use of diagnostic tests to determine which medical treatments will work best for an individual patient [Personalized Medicine Coalition, 2017]. As personalized medicine is sometimes misinterpreted as implying that unique treatments can be designed for each individual, precision medicine refers to the tailoring of standard medical treatments to the patient, taking individual variability into account [Collins et al. 2015]. It is not intended to mean the development of drugs or treatments for a specific patient, but rather applying the concepts of human variability obtained from information in biologic databases, such as genomics, and to using them to guide clinical practice. The president initiated a Precision Medicine Initiative [The White House, 2016] and the National Institutes of Health has been supporting the concept through research on prevention and treatment strategies [Collins et al. 2015]. Hence, the Red Cell Genotyping 2015 symposium adopted the theme to exemplify how molecular immunohematology has been implementing precision medicine in clinical care to enhance patient safety.
The 2015 symposium reviewed applications in molecular immunohematology to benefit the patient. Clinical vignette presentations showcased transfusion recipient or obstetrical cases aimed to underscore challenging situations that were solved by applying molecular immunohematology. Mass-scale genotyping of blood donors, the perspective on weak D testing and quality assurance initiatives in the US and Europe all provided a perspective on state-of-the-art testing and current developments. A summary of an international perspective on molecular immunohematology provided a timely conclusion to the program. This review summarizes the symposium proceedings, documenting the state of the art and progress in the field of red cell genotyping.
Molecular immunohematology in the era of precision medicine: overview and role of AABB (Lynne Uhl, Beth Israel Deaconess Medical Center and Harvard Medical School)
The concept of personalized medicine is not new; following President Obama’s call out in his State of the Union address in 2015 supporting the scientific endeavors targeted toward improved health care delivery using a precision medicine approach [Collins et al. 2015], clinicians, researchers and patients are re-energized in their efforts to further the application of novel technologies and therapeutics. Those of us practicing in the area of transfusion medicine recognize clearly the benefits of a more personalized approach to managing our patients. It goes beyond the ‘simple’ provision of ABO and D antigen compatible blood products for transfusion, with blood typing acknowledged by Collins and Varmus as one of the first applications of precision medicine [The White House, 2016].
The application of molecular technology has opened the doors to a new opportunity in transfusion medicine: red cell genomics for clinical application and improved healthcare and patient safety. The scope of application is large and varied; it ranges from molecular testing to better manage presumed Rh-incompatibility in the prenatal setting [Sandler et al. 2015], to more specific red cell component selection for patients with hemoglobinopathies [Noizat-Pirenne et al. 2002], and to explain the basis of complex serologic assessment of many patients.
As the professional organization of transfusion medicine whose mission is to advance the practice and standards of transfusion medicine and cellular therapies to optimize patient and donor care and safety, AABB actively supports red cell genomics research, development and application in a number of ways. AABB spearheaded the first set of standards for molecular testing laboratories, initially published in 2008 with the third edition now available [AABB, 2016]. In addition, the organization has sponsored/co-sponsored workshops that brought together key thought-leaders and stakeholders, affording an opportunity for robust, thoughtful discussion on the applications of this increasingly sophisticated technology. As important, AABB has tirelessly advocated for avenues for reimbursement of molecular testing related to transfusion management [AABB, 2014]. AABB will continue to be a voice in this arena, endeavoring to smooth the way for new innovation through its regulatory advocacy efforts; provide financial support for research endeavors through foundation support and industry relationships; and disseminate cutting-edge findings through its educational initiatives and publications.
Clinical vignette: alloantibody formation to a high-frequency antigen in the Rh blood group in the setting of hematopoietic progenitor cell transplantation for sickle cell disease (SCD) (Celina Montemayor, National Institutes of Health)
A clinical case was discussed involving a 19-year-old male with SCD (HbS/HbS) treated by a 10/10 HLA-matched sibling hematopoietic progenitor cell (HPC) transplantation. Comprehensive serological and molecular data were presented demonstrating that both patient and donor carried a genotype comprising the Weak D type 4.2.2–ceAR and (C)ces type 1 haplotypes. One day after transplantation, the patient developed an unexpected alloantibody to a high-prevalence Rh antigen. Evidence was presented that transfusion of a crossmatch-compatible blood unit with the (C)ces/(C)ces genotype resulted in an acute hemolysis of the entire unit. Transfusion of a blood unit from the HPC donor was successful.
This report was the second case that we observed [Fasano et al. 2010], implying the need for genotype-identical, not just haplotype-compatible, transfusion in patients carrying an alloantibody to a high-prevalence Rh antigen (Figure 1). Interestingly, both reported cases involved a (C)ces type 1 haplotype [Flegel et al. 2014], an association of unknown significance at this time. In a large study of 226 patients with SCD, the two alleles comprising the (C)ces type 1 haplotype, the RHD allele DIIIa-CE(4-7)-D and the RHCE allele ces, were identified 21 (5%) and 26 times (6%), respectively [Chou et al. 2013].
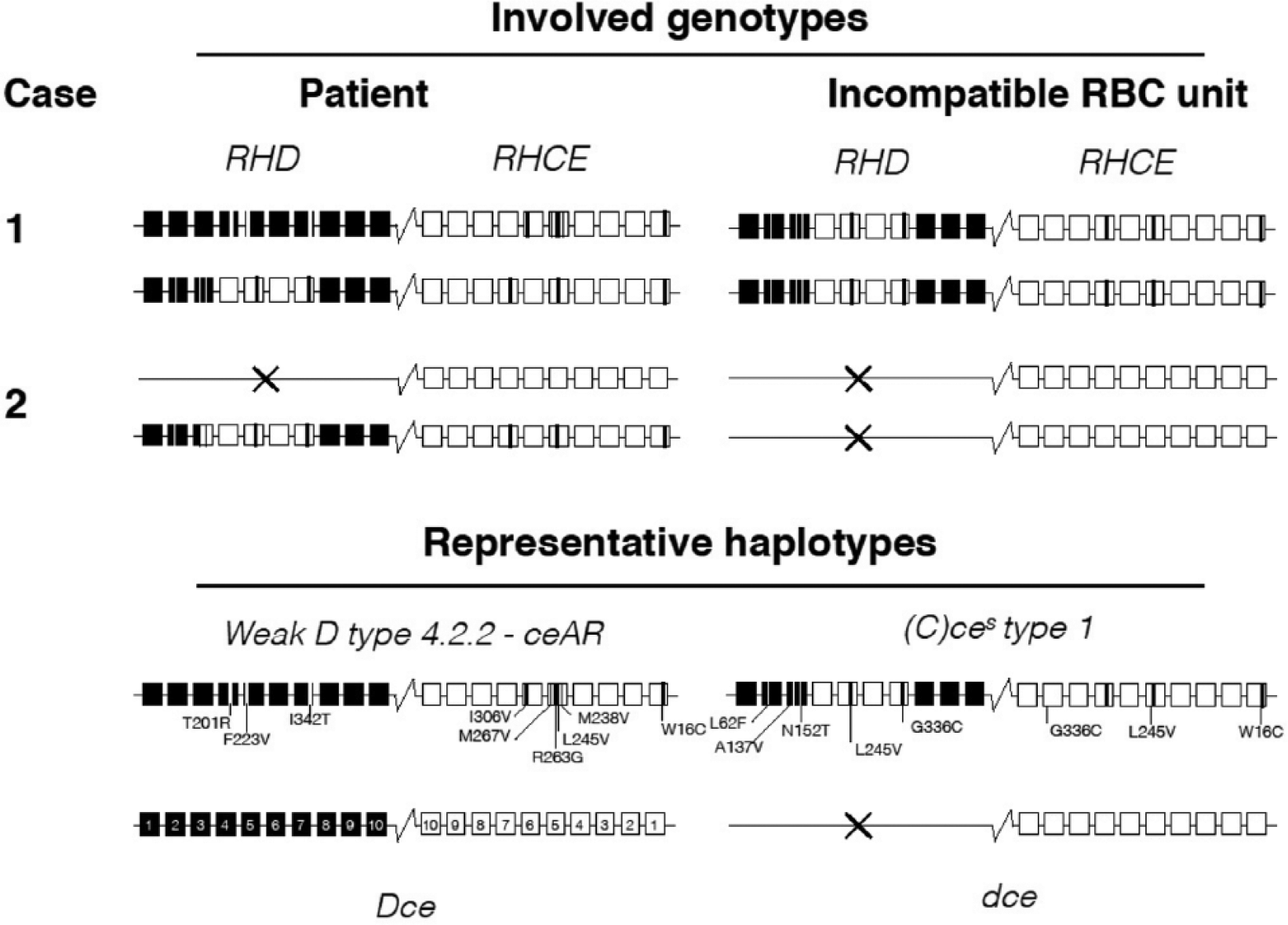
Schematic comparison of two patients’ RH genotypes with the genotype of the respective incompatible red blood cell units. Case 1 represents the patient reported at the conference (upper panel); case 2 has been published previously by Fasano et al. [2010]. Four representative haplotypes carrying distinct amino acid substitutions in the RHD and RHCE exons are shown for comparison (lower panel). A conventional Dce haplotype is depicted for reference; each square corresponds to an exon that is numbered in the Dce haplotype. The variant amino acid substitutions and their locations are indicated by bars (black or white lines in the squares of the representative haplotypes).
The major conclusion of this clinical vignette was that the molecular mechanism and antigenic determinants underlying the need for genotype matching in a subset of patients with certain alloantibodies to high-prevalence Rh antigens is intriguing and is not fully understood by current routine technologies. Further research is warranted to explore the benefit of a potential precision medicine application.
Clinical vignette: genotype dry matching (Razvan C Lapadat, BloodCenter of Wisconsin)
Increasing numbers of RHCE alleles have been identified. Many were detected because of weak or variable antigen expression, typically due to a decreased RhCE antigen density on the red blood cell surface. Some patients can be mistyped by routine hemagglutination as negative for the c or e antigen. Individuals of African ancestry have a higher incidence of these RH variants, many encoding partial antigens leading to Rh alloimmunization due to transfusion of normal high-prevalence Rh antigens. RH matching patients with SCD and selected African-American donors can improve transfusion therapy. We describe an unusual RHCE*ce48C allele found in a three-year-old African male with SCD [Lapadat et al. 2015].
Although phenotyped as D+c+e+, he developed an anti-e after being transfused. Red cell genotyping revealed he was homozygous for the RHCE*ce48C allele. Anemia and evidence of hemolysis were observed (hemoglobin concentration averaged 7.85 g/dl and total bilirubin 2.55 mg/dl). Besides the underlying SCD, these findings may be attributed to the anti-e driving hemolysis, because the transfused blood units were not matched for RHCE*ce48C [Vidler et al. 2015]. Subsequently, blood units were transfused based on genotyping results, a process being termed dry matching [Denomme et al. 2008], which is a type of precision medicine. Careful clinical follow up in similar patients is needed to document the extent of the possible clinical impact.
The RHCE*ce48C allele, frequently associated with an R0 phenotype (Dce), encodes a cysteine at amino acid position 16, but is not known to induce the presence or absence of a blood group antigen. Suitable antisera to detect the RHCE*ce48C allele are lacking, leaving red cell genotyping as the only resource to identify patients at risk and donors with matching blood.
Clinical vignette: Crawford variants (Waseem Q. Anani, BloodCenter of Wisconsin)
Choosing antibody reagents for a blood bank laboratory can be a difficult task. By understanding the limitations associated with specific monoclonal antibodies, one can be aware of potential testing pitfalls. Discrepant serologic testing results are often difficult to interpret. The advent of red cell genotyping resolved many of these interpretation difficulties. This case will illustrate the benefit of red cell genotyping in the context of a D typing discrepancy.
A 29-year-old African-American female (G2P1) arrived for her first obstetric visit at an outside hospital. Her pertinent history included no prior transfusions and no past RhIG administration. A blood sample was sent for a type and screen as a part of her routine prenatal care. Her forward and reverse type demonstrated that she was O D-positive, but her serum showed a clear anti-D. The outside blood bank suspected a D variant and applied a panel of 12 monoclonal anti-D (research use only) to detect D variants (Advanced Partial RhD Typing Kit, Alba Bioscience, Edinburgh, UK), and no agglutination occurred. By using four monoclonal anti-D reagents (FDA-licensed for clinical diagnostic use), agglutination was observed with two reagents (Gamma-clone: ImmucorGamma, Norcross, GA and Alba-clone: Alba Bioscience Limited, Edinburgh, UK). Recognizing a potentially rare phenotype, the outside blood bank referred the specimen to BloodCenter of Wisconsin Immunohematology Reference Laboratory for red cell genotyping. A partial RHD assay via gel electrophoresis, RH variant panel via polymerase chain reaction with sequence specific primers, and Sanger sequencing were performed. The most notable findings included the absence of an RHD allele and RHCE*ce697G and RHCE*ce733G single nucleotide substitutions.
Although the red blood cells were predicted to lack D antigen expression, agglutination was clearly observed with two monoclonal anti-D reagents. The incongruent results were first observed in 1980 when a reagent polyclonal anti-D was found to contain antibodies specific for the Crawford phenotype [Cobb, 1980]. With molecular methodologies, it was later demonstrated that the Crawford phenotype was composed of a triad of variants in the RHCE gene: 48G>C, 697C>G and 733C>G [Flegel et al. 2006]. The result is a D-like epitope in the RHCE gene that is recognized by some FDA-approved monoclonal typing reagents [Flegel et al. 2006; Roback et al. 2011]. The Gamma-clone has classically been observed to agglutinate at only immediate spin, because the IgM monoclonal antibody recognizes the D-like epitope. However, no agglutination is observed in the IAT or AHG phases, because the monoclonal IgG component of the reagent recognizes a different epitope.
Two clinical implications become apparent in the D-negative patient. Although the Crawford phenotype creates a D-like epitope, these patients are at risk of becoming alloimmunized when exposed to D-positive blood [Flegel et al. 2006]. Only D-negative blood should be transfused to Crawford individuals that lack an RHD (D-negative individuals), and RhIG administration in pregnant women is recommended. Second, the use of the Gamma-clone, while FDA approved, can incorrectly type patients with the D-like epitope present in Crawford patients [Flegel et al. 2006]. The rarity of the phenotype has limited epidemiologic studies, but this phenotype is estimated to occur in fewer than 0.1% of African-Americans [Moulds et al. 2003]. Although uncommon, any blood bank that transfuses a significant number of African-American patients or performs prenatal type and screens should consider discontinuing the Gamma-clone for D typing in these settings.
The molecular methodologies resolved a result that had been previously discrepant by serology alone and guided the management of the patient through the use of D-negative products, if necessary, and RhIG administration to reduce the risk of anti-D alloimmunization; another example of an inexpensive precision medicine application.
Weak D: a historical perspective (Neil D. Avent, University of Plymouth)
We are lucky to work in transfusion medicine at an age where the molecular characterization of most blood group active genes has been completed, and the major clinically significant antigens have been described at the DNA level. However, the path to this achievement sometimes included controversy and friendly competition between a number of research groups in Europe and North America. In some European healthcare systems, it is routine to be able to prenatally define fetal RHD status using maternal plasma [Damkjaer et al. 2012]. DNA analysis of D variants is routine and the next phase, including complete gene sequencing, is underway using the new techniques of next-generation sequencing [Flegel et al. 2016a].
Having worked on the purification of Rh proteins and their eventual cloning in the late 1980s [Avent et al. 1988, 1990], the next phase of work inevitably turned to the characterization of D variants, including the mysterious and legendary partial D phenotype DVI and, of course, the strange phenomena of the weak D phenotype. Key contributions to the body of knowledge come from the groups of Jean-Pierre Cartron [Mouro et al. 1994; Rouillac et al. 1996], Neil Avent [Avent et al. 1996; 1997a], Bill Flegel [Wagner et al. 1998, 1999], Ellen van der Schoot [Maaskant-Van Wijk et al. 1998], Cathy Hyland [Cowley et al. 2000], Greg Denomme [Denomme et al. 2005], Connie Westhoff and Marion Reid [Murdock et al. 2008], and several others.
The first partial Ds to be deciphered at the molecular level were the D category VI variants, which were found to be formed by gene conversions between RHCE and RHD, causing the expression of hybrid RhD–RhCcEe proteins [Mouro et al. 1994; Avent et al. 1997b]. Initial analysis of the weak D phenotype had incorrectly suggested that quantitative variation in RHD mRNA levels were responsible [Rouillac et al. 1996], which could not be corroborated [Beckers et al. 1997]. It was somewhat of a surprise to learn at the 1998 ISBT congress that an unknown group from the German Red Cross from Ulm in Germany had methodically analyzed vast numbers of weak D phenotypes. The work was published in 1999 [Wagner et al. 1999] and is indeed a landmark in molecular biology of blood groups. Single-nucleotide polymorphisms in the RHD gene were known prior to the 1999 paper to occur in several partial D phenotypes [Avent et al. 1996; Legler et al. 1998], but the scale of mutations in RHD were laid bare by this landmark publication. We have over 150 defined weak D type alleles [Wagner et al. 2014] and we are still gaining detailed knowledge of their clinical significance [Sandler et al. 2015]; some are capable of eliciting anti-D when individuals are exposed to D-positive blood [Wagner et al. 2000; Sandler et al. 2015].
During the mid-2000s the definition of the crystal structure of AmtB allowed the prediction of the structure of the Rh core complex trimer, comprising two Rh-associated glycoproteins (RhAG) and one Rh protein monomer. This information gave critical insight into the approach of the affected amino acids in partial and weak D phenotypes. Unsurprisingly, those amino acid exchanges causing partial D phenotypes occur on exofacial loops, although the extent by which these loops penetrate the membrane bilayer was not appreciated until these structural models appeared [Conroy et al. 2005]. The concept of the D antigenic vestibule is now fully appreciated. However, weak D mutations appear to affect the assembly of the Rh core complex trimer, and some indeed occur within G-XXX-G motifs, critical for inter-membrane protein interactions, namely that of RhD with RhAG [Avent, 2007].
The focus on Rh molecular biology has been for the past decade predominantly at the level of the gene. The implementation of RH genotyping has seen rapid uptake in implementation and, in particular within the US, this information is invaluable in the management of highly transfused patients – for example, those with SCD. The next decade of Rh research will be an interesting phase, and will certainly embrace molecular diagnostics even more strongly contributing to precision medicine procedures. The unknown function and protein chemistry of the RhD and RhCE proteins have been somewhat neglected, and the speaker looks forward to a new generation of Rh researchers picking up the mantle of the pioneering work done over the past 25 years.
Integration of red cell genotyping into the blood supply chain (Gregory A. Denomme, BloodCenter of Wisconsin)
One important role of a blood center is to provide antigen-negative blood units for the alloimmunized transfusion recipient [Denomme, 2013]. Presently, the identification of such blood units can be prohibitively time-consuming and expensive by phenotyping with serologic reagents. The molecular basis of blood group antigen expression has been confirmed for many more antigens than can be phenotyped using existing phenotyping techniques. Mass-scale red cell genotyping of blood donors provides the opportunity to create and maintain large databases that can serve the transfusion recipient with specific antigen-negative requirements beyond ABO and D. In 2010, a database of over 25,000 blood donors was created within six months at the BloodCenter of Wisconsin [Flegel et al. 2015a]. The database included all blood donors who self-declared their ethnicity as African-American and Hispanic, regardless of their frequency of donation to capture rare blood types found among these donor populations. The largest set of donors had a history of repeat donation following a 3:3:1:1 rule; three donations within the last 3 years of which one donation was within the last year.
Mass-scale genotyping was performed on 32 nucleotide polymorphism linked to the expression for 42 blood group antigens. Additional blood units were identified for 14 blood group antigens using phenotype techniques with unlicensed antisera. Within 3 years, 43,066 donors donated a total of 202,275 blood units that had a red cell genotype on file. Antigen-negative requests were filled for 99.8% of 5672 patient encounters, with an ‘encounter’ defined as one order for blood units regardless of the number of units requested. The database was comprised of 1.67 million genotypes that were assembled over 4 years. Contrasting that with 320,000 phenotypes accumulated over the past 30 years, red cell genotyping was able to assemble five times the serologic data in one-seventh of the time. Red cell genotyping met the demand for antigen-negative blood for 94.1% of all requests, with a total of 333 requests filled with phenotype data alone. It is important to note that individual donor records contained no fewer than 28 genotypes, whereas 95% of the phenotype data comprised no more than 15 antigens. In addition, the cumulative mass-scale data resulted in 30% of blood units having a genotype on any given day in the final year of the program. The database could be maintained annually by genotyping 4000 repeat donors each year, which reflected the year-over-year repeat donor loss-rate.
The construction of a large central database with nearly one-third of the blood units having historical red cell genotype information affords the opportunity to provide the information throughout the supply chain [Flegel et al. 2015b]. The BloodCenter of Wisconsin gave seven hospitals access to historical genotype and phenotype via a web-based antigen query portal. The hospital would scan the ISBT128 code available on blood units in their inventory into a query form, which is submitted electronically to the blood center along with the number of units and the antigen-negative attributes desired among the common blood group antigens: C, c E, e, M, N, S, s, K, k, Fy(a/b) and Jk(a/b). The result returned to the hospital identified the oldest blood units that meet the requirements. If the query is not successful, additional blood units could be scanned or the order can be ‘pushed’ to the blood center to be filled by the reference laboratory. This online approach helped hospitals up to 200 miles away from the blood center to locate antigen-negative blood units in their inventories. The 8-month pilot program identified 71 blood units from 52 antigen-negative requests. By the end of 2016, a total of 14 hospitals were using the portal to access the available antigen-negative information within their inventories. The program is poised to use two historical phenotypes or genotypes to enhance the ability to provide antigen-negative blood for patients who need such blood units, and enables a wide cost-effective implementation of precision medicine in the blood supply chain (Figure 2).
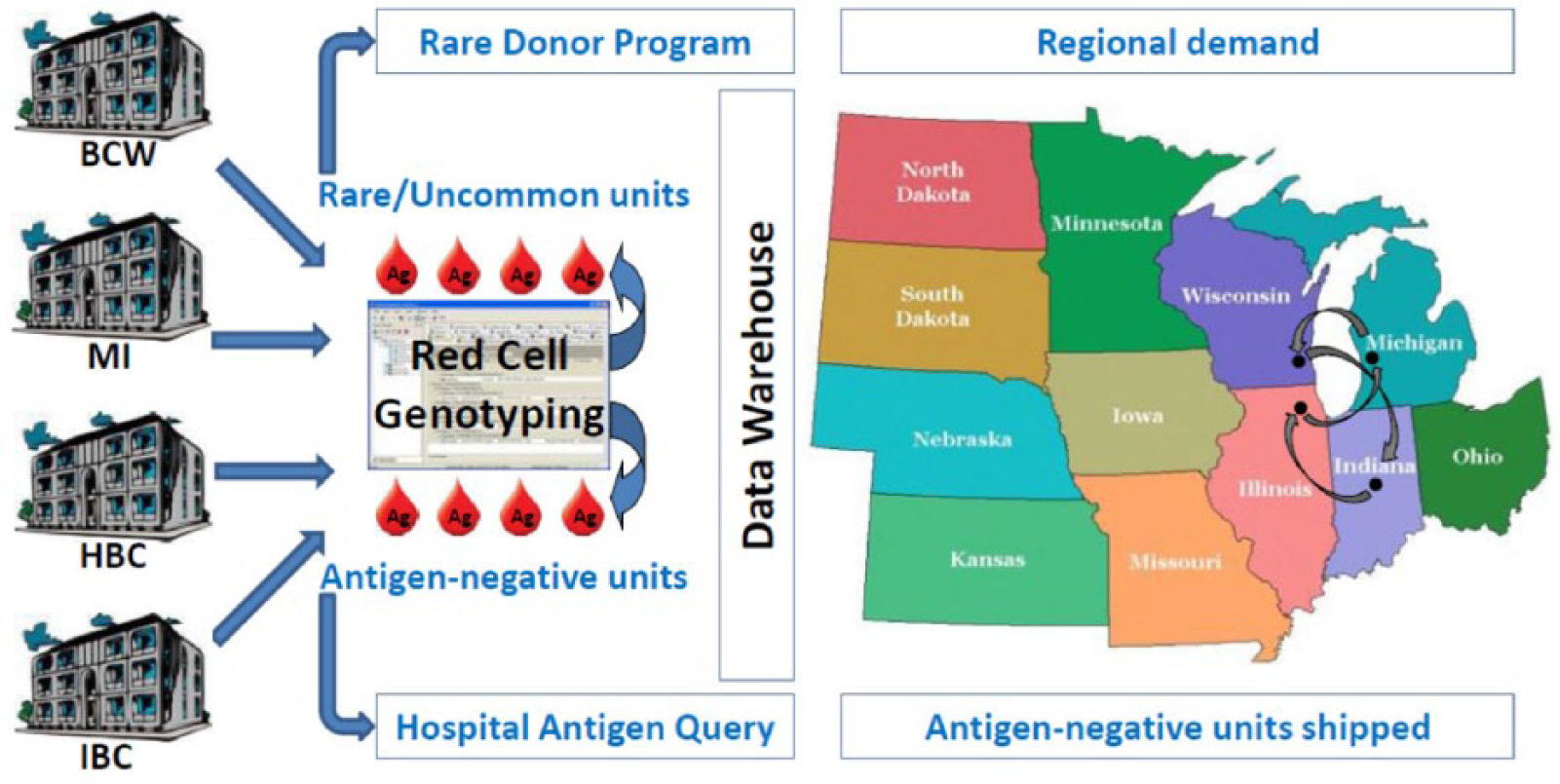
Red cell genotyping across multiple blood centers. All Versiti-affiliated blood centers participate in mass-scale red cell genotyping of donor units (left side) [Flegel et al. 2015a]. Rare units are identified and diverted to rare donor programs worldwide. Antigen-negative information is shared throughout the supply chain via a web-based antigen query portal in the cloud [Flegel et al. 2015b]. A data warehouse is accessible online and offers real-time web-based red cell genotype information on blood units. A blood center queries the data warehouse to fill orders that cannot be met from their current inventory. Units can be shipped between blood centers in a large region and across state lines to meet antigen-negative demands (right side). Versiti is composed of the BloodCenter of Wisconsin (BCW), Michigan Blood (MI), Heartland Blood Centers (HBC) and Indiana Blood Center (IBC).
Finding a needle in a haystack: technology enabling visibility of rare and uncommon blood units (Lynne B. Briggs, BloodCenter of Wisconsin/Versiti)
Information technology (IT) requirements to support a clinical need often move from the innovation pipeline to an industry-standard practice. The drivers of such innovation are a composite of ‘what is possible?’ and ‘what is desirable?’, from a customer perspective, to ‘what will be viable?’ in the marketplace. The development of an IT solution for a need in blood banking and transfusion medicine should have the goal of improved patient outcomes. For red cell genotyping data, the goal is to provide a fast and cost-effective means of identifying donors and their associated red cell genotypes that match the needs of a patient, while at the same time reducing or, where possible, eliminating data handling. The first step to translating mass-scale red cell genotype data in a meaningful and sustainable way to the benefit of the patient was to identify the current gap in handling such large amounts of data. The BloodCenter of Wisconsin recognized the gaps to be data transmission, the link between a genotype and phenotype, storage of historical results and the labeling of donor units.
For transmission, the output of data from an instrument is handled on the basis of the file type protocol and the format. Transfer of data is not new to blood centers, given the volume of tests and the need to add new tests, often on short notice, as deemed by regulatory entities. To address the link between a phenotype and a genotype, the BloodCenter of Wisconsin took the approach of maintaining the source of the results, be it a genotype or a phenotype, and to create a meaningful link between the two using a ‘Rosetta Stone’ governed by a rules engine [Bhagat et al. 2011]. From the linked phenotype and genotype data, exception files were designed to report phenotype–genotype discordant results or when duplicate data were entered. Those solutions were built keeping in mind the eventual need to comply with ICCBBA ISBT Standards for the labeling of blood units with historical results [ICCBBA, 2016].
Red cell genotyping technology, along with the ability to genotype thousands of donors, makes finding rare blood units a true ‘needle in a haystack’. IT and informatics changes are required to fully operationalize the use of red cell genotyping data throughout the supply chain, which includes the inventory in hospital transfusion services. The BloodCenter of Wisconsin included the development of an IT solution that makes available red cell genotype and phenotype data to hospitals for blood units in their inventory (Figure 3). In this way, antigen-negative blood units can be identified throughout the supply chain to the benefit of the patient [Flegel et al. 2015b].

The web-based solution for red cell genotype and phenotype data. Hospitals can access such data in the cloud for all blood units in their inventory. The hospital enters patient information (top portion), the blood type and antigen-negative attributes (middle portion) and number of units required (lower left). Blood unit ISBT 128 identification numbers are scanned (not shown) and then the electronic query is sent to the blood center (lower center-left). The query returns the ISBT identification number of the oldest unit(s) with the required attributes (not shown). If the required units are not found, the process can be started again (lower center) by scanning additional inventory, or the hospital can place the order with the blood center (lower center-right).
PCR-free red cell genotyping using a nanobiosensor (Maryse St-Louis, Héma-Québec)
For decades serology was the sole technique used in immunohematology. It depends on reliable sera, is labor-intensive at times and could present major limitations. In the molecular world, actual single-nucleotide polymorphism (SNP) detection depends on different techniques, most using enzymatic amplification, such as PCR-RFLP, PCR-SSP, modular and multiplex PCR. However, enzymatic-based amplification presents many drawbacks: it is prone to contamination and enzyme inhibition, time-consuming, costly and the economy of scale is met only for batches. We explored a PCR-free red cell genotyping technique using a nanobiosensor [Brouard et al. 2015].
The goal of this study was to amplify the signal generated by the recognition of hybridization events between oligonucleotide probes and perfectly complementary targets rather than amplifying the number of targets by PCR. The challenge was to maintain good sensitivity and specificity. Nanomaterials were selected based on their multiple properties: color changes, fluorescent signal or luminescent signal. Nanoscopic properties are nothing new. Several examples can be found throughout history. The Lycurgus cup held at the British Museum contains nanoparticles made of silver and gold, causing light transmission and scattering. The Damascus blade was made of carbon nanotube steel. Nanoparticles can be described as antennae because of the metal-enhanced fluorescence (MEF). When a fluorophore is in close proximity to a metal, after excitation, the electron in the metal creates an electron cloud called surface plasmon, increasing the fluorescent signal. The distance between the fluorophore and the metal has to be 5–50 nm for this to happen.
The strategy was to make silver nanoparticles (d = 50 nm) with a silica shell containing eosin (d = 10 nm) to which were grafted DO*01 and DO*02 single-stranded probes. These nanoparticles tend to aggregate in the presence of a cationic polymer. Once the genomic DNA sample is added, the polymer changes its conformation and emits light. The MEF amplifies the light signal enough to be detected by cytometry.
DNA hybridization was performed at 58.5°C, although the optimal temperature for the DO*02 probe was higher. Nine samples were assayed, three of each genotype: DO*01/DO*01, DO*01/DO*02, DO*02/DO*02. Results were compared to classic PCR and were concordant. The turnaround time for the nanobiosensor scheme was less than 30 min once the DNA was extracted. The hybridization time could be further reduced.
Results are reliable, even for homozygous and heterozygous fluorescent signals that are sometimes difficult to differentiate with other techniques. These nanobiosensors present advantages over PCR techniques by being fast, low-cost and specific, but are limited to known SNPs.
Red cell genotyping assays: FDA regulations and development of reference reagents (Maria Rios, Food and Drug Administration)
Molecular testing of donors has proven useful in identifying blood units that are more likely to match alloimmunized patients, and these genotyping assays have been used to assist patient care. Due to the common chemistry involved, genotyping may be carried out in high-throughput platforms as multiplex assays. However, the use of red cell genotyping in clinical settings presents some challenges.
Regulatory considerations for devices
Red cell genotyping platforms are classified as high-risk Class III devices, for which the pathway to market requires an investigational device exemption (IDE) to perform clinical studies and a premarket approval (PMA) submission to demonstrate reasonable assurance of safety and effectiveness based on scientific evidence. A PMA has a 180-day review, which may be extended up to 320 days if the submission is sent to an advisory committee.
Well-characterized specimens are needed as raw material to prepare reference reagents for use in the development, validation and manufacturing of genotyping tests, and for the optimal use of Class III devices in routine practice. Currently, there is a paucity of well-characterized material for use as reference reagents for extensive red cell genotyping. In order to demonstrate efficacy of assay performance and to fulfill regulatory requirements, OBRR is developing DNA reference reagents covering genetic variants for antigens from the 17 blood group systems recommended by AABB. These reference reagents consist of well-characterized genomic DNA from human cells containing several red cell genetic variants that may not be readily available during clinical trials. This material will be made available to facilitate analytical evaluation during assay development, for proficiency testing and for QC.
Development of reference reagents
OBRR has supported a project for the development of comprehensive DNA reference panels for red cell genotyping (Table 1). This project was initiated in 2012, led by Dr. Jason Liu with intramural funding by the CBER/FDA Office of Blood Research and Review, with the overall goal of producing reference reagents to support development and validation of new assays and to fulfill regulatory requirements for the application submission and approval process.
Reference reagent development project at CBER since 2012: specific aims.

The specimen acquisition and processing has been achieved through recruitment of donors with the desired historical phenotype and the genotype characteristics of the homozygous and heterozygous genotypes for each allele. An IRB-approved project was initiated that provided de-identified donor samples under contract with the BloodCenter of Wisconsin. Whole-blood units have been collected from 53 specially recruited donors with desired genetic variants. The whole-blood units were shipped to CBER for processing and preparation of frozen red blood cells, plasma and buffy coat components. Genomic DNA from each donor was isolated from leukocytes and characterized with molecular typing assays developed in the laboratory to determine/confirm blood genotypes associated with 56 blood group antigens. Laboratory methods include TaqMan allelic discrimination assays, PCR-SSP and PCR-RFLP assays, Sanger bi-directional sequencing and next-generation sequencing. The samples have been characterized for 39 alleles associated with 56 antigens from 17 blood group systems by TaqMan assays, and the genotypes confirmed by Sanger sequencing. Immortalized cell lines were generated as renewable sources of DNA using PBMC obtained from buffy coats by transformation with EBV. To date, 51 cell lines have been produced and stored in liquid nitrogen tanks. The verification of the integrity of target genes after transformation is ongoing by testing the DNA from the cell lines by PCR and TaqMan. We have selected 19 of these 51 established cell lines that cover all genetic variants collected to date. Cell line expansion and bulk genomic DNA has been extracted for 14 of these 19 samples. The effects of EBV transformation on allelic discrimination assay results were also assessed. The most appropriate formulation for freeze-drying has been chosen after pilot lyophilization runs, and the resultant materials were assessed for reproducibility of previously performed assays by testing target genes by PCR and/or TaqMan to verify performance of reagents after reconstitution. To date, of the 123 allele combinations proposed, blood samples covering 80 antigens have been acquired and cell lines established (Table 2) [Vokkova et al. 2016].
EBV-transformed cell lines for renewable DNA for reference purposes.

Office of Blood Research and Review (CBER/FDA) has identified 41 target alleles across 18 blood group systems among 53 individuals. A reference panel of DNA from 17 cell lines, ensuring a renewable DNA supply, will be made available to manufacturers for developing and calibrating red cell genotyping kits and to laboratories for validating and monitoring the performance of such kits [Vokkova et al. 2016].
The next steps will be formulation of the panel and initiation of stability and accelerated degradation studies. In parallel, external evaluations will be performed for final characterization, which will include identifying appropriate laboratories to participate in the study, setting up agreements with collaborating laboratories, shipping panel material, developing testing protocols and reporting worksheets and establishing deadlines for participation. At the time of this writing, we are collecting and analyzing the results to formally publish this collaborative study [Vokkova et al. 2016].
Application of external quality assessment in the clinical immunohematology laboratory (Gregor Bein, Justus-Liebig-University)
Genotyping for red blood cell, platelet (PLT) and granulocyte antigens is a new tool in the diagnostic evaluation of immunohematological disorders and typing of blood donors and recipients. Proficiency in laboratory tests can be established by external quality assessments (EQAs), which are required for clinical application in many health care systems and to maintain accreditation. There are few EQAs for molecular immunohematology [Delaney, 2013]. One of the first workshops on molecular immunohematology was originally supported by a grant from the German Federal Ministry of Health in 1998 [Kroll et al. 2001]. This funding jumpstarted an EQA program organized by the German Society for Transfusion Medicine and Immunohematology, which was transitioned to a nonprofit provider in 2006 (INSTAND, Dusseldorf, Germany) [Flegel et al. 2013]. Other international EQA programs include workshops that are organized by the College of American Pathologists (CAP) [Allen et al. 2011], the International Society of Blood Transfusion (ISBT) [Daniels et al. 2011], and the Consortium for Blood Group Genes [Denomme et al. 2010]. The INSTAND EQA offers proficiency sets twice per year. Each set is always composed of four specimens of genomic DNA. Participants are requested to test all four specimens, even if only a single blood group system and a single proficiency certificate are of interest. A list of predefined mandatory alleles per blood group system guarantees maximum transparency [Flegel et al. 2013]. The number of participants is steadily increasing and now encompasses 96 participants from 18 countries, mainly from central Europe (October 2015). Almost all institutions participate in red blood cells, 60% in PLTs and 25% in granulocyte proficiencies. A recent summary of the workshop results revealed that the overall pass rates per system exceed 93% ± 4% (mean ± SD). However, low pass rates were obtained for RHD (82% ± 10%) and ABO (93% ± 4%). Frequent errors include failures of zygosity testing and identification of rare alleles (e.g. weak D types). High pass rates are obtained for blood group systems that are characterized by single nucleotide polymorphisms (e.g. JK, HPA) that can be genotyped without determination of extended haplotypes and/or haplotype phasing. The passing rates did not correlate with the experience of the participants with this EQA.
In summary, ABO and RHD systems seem to pose challenges for genotyping, possibly due to the large number of alleles caused by recombination. We conclude that intrinsically more robust technologies, such as sequencing platforms that allow for long-distance haplotype phasing, may be needed for reliable genotyping for diagnostic purposes, at least in some rare allele combinations. Future perspectives include the establishment of larger numbers of B-lymphoblastoid cell lines as a source of reference DNA, the inclusion of additional blood group systems (e.g. LU, DI, YT, HPA-2), and EQA for the non-invasive prenatal diagnosis of fetal blood groups [Kroll et al. 2001; Vokkova et al. 2016].
Molecular immunohematology and precision medicine: international perspective (Silvano Wendel, Hospital Sirio Libanês)
Molecular biology has considerably developed in the immunohematology field, not only confirming previous serological data, but also elucidating unsolved puzzles. The aim of this presentation was to compare what can be readily applicable for molecular immunohematology tests (MIT), taking into account the previous experience already acquired with molecular methods developed for screening of transfusion-transmitted diseases (TTID), which have now over a decade of international use.
General issues
Most TTID molecular tests adopted for blood donor screening are used as universal testing (HIV, HCV and HBV). Selective testing is applied mainly for West Nile virus (WNV), according to seasonal periods. This has a tremendous impact as far as the annual number of tests, given that more than 10 million donations are tested just in the US. Thus, one can expect a great interest from pharmaceutical industries not only to develop, validate and register commercial tests, but also to pursue continuous improvement in the test’s performance and related hardware.
On the other hand, MITs are targeted at genetic markers (red blood cells, leukocytes and platelets) from healthy donors or their recipients. Since the genetic code is permanent, one does not need to apply the same MIT to the same donor every time there is a blood donation; actually it is very likely that whenever a given donor is tested once (or twice to be certain), results will be considered as permanent in the donor’s file. Whatever the adopted strategy, blood centers usually face a restricted budget for this activity, making the final number of MIT intended to guarantee a stable widely tested donor panel much lower than those annually tested for infectious diseases.
The situation concerning recipients must be viewed from a different perspective. Although most recipients will not have their genetic code changed throughout their lifetime, the increasing number of transplants (bone marrow, cord blood and solid organs) poses a challenge in the follow up of patients. In this respect, there are clinical situations in which MITs could be quite useful (e.g. chronic transfusion recipients, pregnancies) [Legler et al. 1999; Rujirojindakul et al. 2014].
Finally, one cannot neglect the power that public perception or medical agencies exert over authorities to facilitate the introduction of new tests in order to improve blood safety. For example, the World Health Organization (WHO) Aide Memoire for Ministries of Health [World Health Organization, 2016a] defines in its section on clinical transfusion in patient management that there should be ‘Systems, processes and procedures for compatibility testing and issue of blood; the rational use of blood and blood products; safe transfusion practice at the bedside and patient monitoring and follow-up’ without mentioning which kind of testing should be performed. In addition, another important WHO document, the 2015 Global Reference List of 100 Core Health Indicators [World Health Organization, 2016b], when dealing with service utilization is quite vague; there is only one mention of health service access, hospital bed density, and availability of essential medicines and commodities. Thus, it is important that blood centers advocate for the benefits of MITs in order to raise public awareness. Nevertheless, despite the relative imbalance between infectious diseases and MITs, one can consider that the use of MIT will become a current practice [Flegel et al. 2015b]. An expansion in the use of MITs is expected in this decade.
MIT in the global economic crisis
Even when applied at a large scale, MITs are not a cheap procedure at this time, involving not only complex procedures but also expensive equipment and trained professionals. Thus, its immediate application requires a substantial place in the budgets of blood centers, which are affected at least partially by the economic situation of a given country. Naturally, the higher health expenditure per capita or the relative percentage of the gross domestic product provided by a country, the higher the chance that this problem could be overcome by blood services authorities. Detailed differences concerning health expenditures of all countries (low, medium and high income) can be found in the annual World Bank reports [The World Bank, 2016a, 2016b].
Current applications of MITs
Despite the numerous papers available in the literature dealing with highly specific antigens, large-scale testing of regular blood donors is a reality [Denomme, 2013; Haer-Wigman et al. 2013; Latini et al. 2014; St-Louis, 2014; Tanaka et al. 2014; Flegel et al. 2015a; Manfroi et al. 2015]. The Ebola outbreak of 2014/2015 has also raised some unique proposals for the use of molecular methods whenever Ebola-infected patients require a transfusion [Flegel et al. 2016b].
The dramatic development of molecular biology has facilitated its current application in a number of routine blood services procedures, enhancing blood safety worldwide. Technology is now available and might be used for several applications. However, due to its high costs, MIT will be used selectively for most immunohematological cases [Delaney, 2015], as opposed to universal adoption for some TTIDs. On the other hand, recent economic reports from several countries show progressive recovery, which will facilitate its adoption in the foreseeable future. Despite the lack of public awareness and support, its use cannot be limited and requires an active role be taken by professionals engaged in the field in order to show the public the benefits. Naturally, every new blood policy must be implemented based on judicious criteria, with balanced views, public acceptance and good sense being key points.
Footnotes
Acknowledgements
The authors thank Susan T. Johnson and Phyllis Kirchner, BloodCenter of Wisconsin, and Karen M. Bryne, Avis B. Brown-Williams and her team at the DTM Administrative office of the NIH Clinical Center for their contribution to the coordination of the symposium. The Red Cell Genotyping 2015: Precision Medicine conference received continuing medical education credit from the American College of Graduate Medical Education. We appreciate the scientific input provided by Jerome L. Gottschall and Jay S. Epstein as moderators of two sessions during the meeting. The authors acknowledge Harvey G. Klein for his comments on the symposium and this review.
Statement of disclaimer
The views expressed do not necessarily represent the view of the National Institutes of Health (NIH), the Department of Health and Human Services (DHHS), or the US Federal Government.
Authorship contribution
GAD and WAF organized the scientific content of the symposium and this review. The speakers provided summaries of their presentations. GAD compiled the report and WAF provided the edits.
Funding
Red Cell Genotyping 2015: Precision Medicine was made possible by funding from the National Institutes of Health, BloodCenter of Wisconsin, Héma-Québec and OneBlood. This work was supported by the Intramural Research Program (project IDZ99 CL999999) of the NIH Clinical Center and the Commonwealth Transfusion Foundation (BCW 01/2016).
Conflict of interest statement
GAD is on the authorship and speaker’s bureau of Grifols. NDA is a consultant for Grifols. GB is an inventor of a device patent and has contract research with BAG Health Care. WAF is inventor of patents owned by the German Red Cross Blood Service Baden-Württemberg-Hessen and holds intellectual property rights for RHD genotyping applications. The remaining authors declared no conflict of interests.
