Abstract
Many species of the order
Keywords
Introduction
The order of the
Materials and Reagents
BUNV-mCherry elite stock 3 (Prof. Xiaohong Shi at the University of St. Andrews, Scotland, UK)
Human lung epithelial cells (A549, ATCC®, catalog number: CCL-185™)
Filter tips 10 µl, 100 µl, 1250 µl (Sapphire, Greiner-Bio, catalog number: 771261, 737261, 750261)
Serological pipets 5 mL, 10 mL, 25 mL (Sarstedt, catalog number: 86.1253, 86.1254, 86.1685)
2 mL aspirating pipette (Corning Life Sciences, Falcon®, catalog number: 357558)
T-75 culture flasks (TPP®, catalog number: 90076)
96-well plate (Falcon, catalog number 353072)
Black 96-well plate with clear bottom (Greiner Bio-one, catalog number 655090)
DMEM (Thermo Fisher Scientific, Gibco™, catalog number: 41965039)
Fetal calf serum (FCS) (Thermo Fisher Scientific, Gibco™, catalog number: 10270106)
Sodium bicarbonate (Thermo Fisher Scientific, TMatalog number: 25080060)
Sodium pyruvate (Thermo Fisher Scientific, Gibco™, catalog number: 11360039)
Penicillin/streptomycin (Thermo Fisher Scientific, Gibco™, catalog number: 15140148)
Trypsin 0.25% (Thermo Fisher Scientific, Gibco™, catalog number: 25200056)
Colorless Minimal Essential Medium (MEM) (Thermo Fisher Scientific, GibcoTM, catalog number: 51200046)
Hoechst 33342 (Thermo Fisher Scientific, Invitrogen, catalog number: H1399)
Cell counter slides (Logos Biosystems catalog number: L12001)
Reagent reservoirs (VWR catalog number: 613-1184)
Cryotubes (Thermo Fisher Scientific, Thermo Scientific™, catalog number: 377224)
A549 culture medium (see below)
A549 assay medium (see below)
Barcodes (optional)
Cell media
Equipment
Overall
Laboratory suited for biosafety level (BSL)-2 work
Protective gear for working at BSL-2 laboratory (gloves, lab coat, shoe covers, safety glasses)
B. BSL-2 laboratory
BioVanguard Greenline Biosafety cabinet from Telstar Incubator (37°C, 5% CO2, humidified) Pipette set (P10, P100, P1000) Pipet controller Multichannel pipette Electric aspirator −80°C freezer Vortex Inverted light microscope Cell counter Arrayscan XTI from Thermofisher® Cellomics software of ‘HCS Studio’
Procedure
Cell culture and virus stocks
A549 cells are maintained in culture medium (see Materials and Reagents) in T-75 culture flasks at 37°(C) in a 5% CO2 humidified atmosphere. Cells are split 1:10 once per week. Cells used in assays are between passage 15 and 45. For this protocol, we selected A549 cells, since when infected with BUNV-mCherry these cells showed minimal CPE at the desired moment of readout compared to Vero E6 and BHK cells.
To create additional virus stocks cell cultures were infected with 0.001 MOI out of the elite stock and incubated for 4 days. The supernatant was collected after four freeze thaw cycles and then stored at −80°C. The viral titer of these stocks was determined by end-point titration.
Day -1
Assay plate preparation
Take a T-75 culture flask with A549 cells grown to ∼90% confluency.
Remove the culture medium and incubate for ∼5 minutes with trypsin to detach the cells.
Add 5–10 mL assay medium (see Materials and Reagents). Pipet up and down and against the side or bottom surface of the flask to make sure all cells are in suspension and unattached.
Count the cells and make the appropriate cell suspension of 50 000 cells/mL using assay medium.
Dispense 100 µL of the cell suspension in each of black ‘96 well’ plates with clear bottom. While the outer wells are not used in the assay, loading them with the cell suspension will help prevent volume changes due to evaporation during incubation.
Close the plates and incubate them overnight at 37°C in a 5% CO2 humidified atmosphere.
b. Compound preparation
Dissolve the test compounds in DMSO to make 10 mM solutions. Store them at room temperature, 4°C or −20°C (depending on the compound).
Prepare an overview of which compounds will be tested in which plate using barcode numbers. There are six compounds tested per plate (Figure 1).
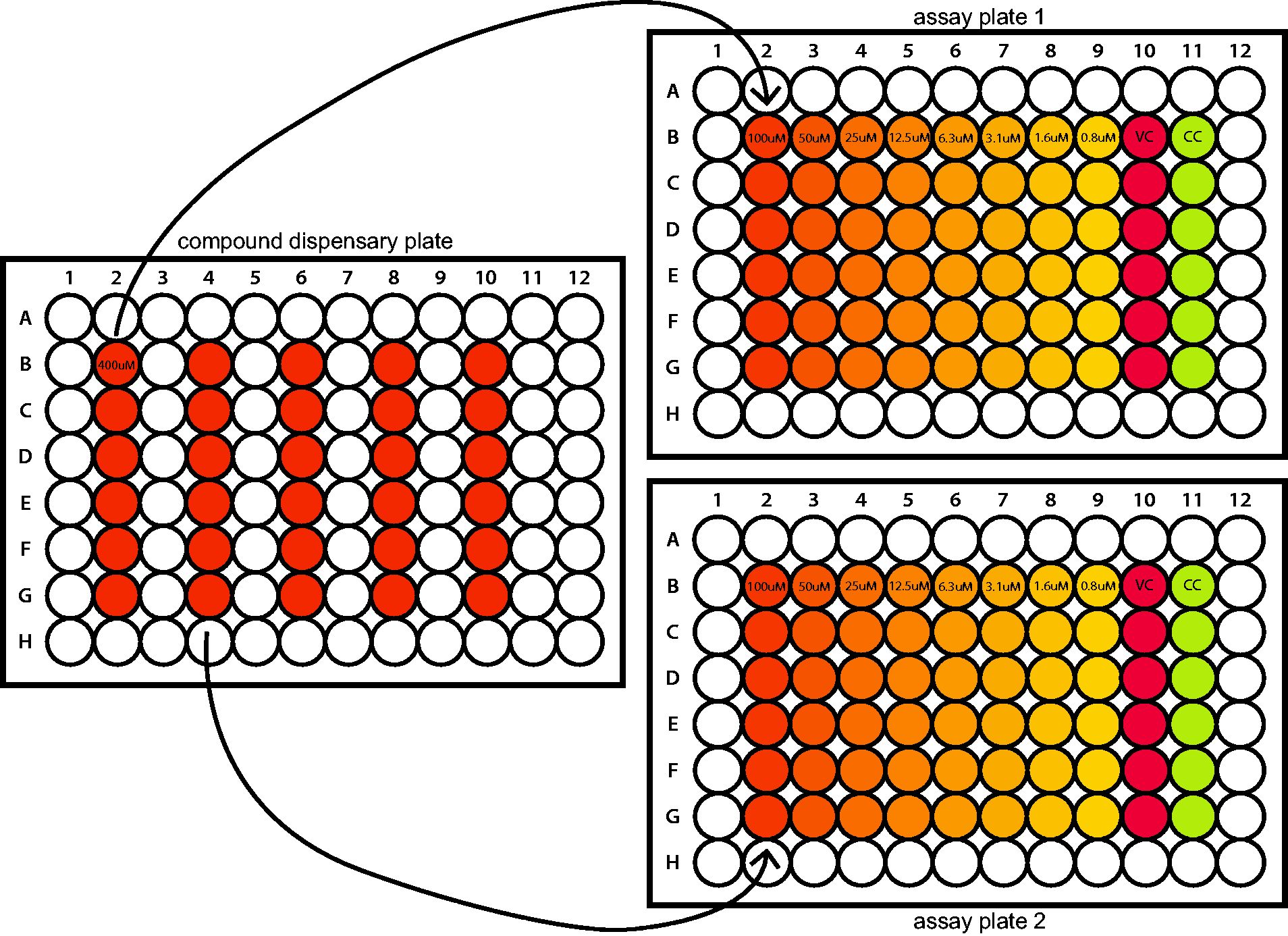
Schematic representation of the pipetting proceedings between the compound dispensary plate and the assay plates. The concentrations mentioned in the compound dispensary plate is the concentration of compound in the well just before transferring it to the assay plates. The concentration mentioned in the assay plates are the final concentrations in the well after finishing the step of infecting the assay plates with BUNV-mCherry. M: molecule/compound; VC: virus control; CC: cell control.
Day 0
Plate inspection
Inspect the assay plates using the inverted light microscope. Check whether the cells look healthy and are equally distributed throughout the plate. Label the plates using barcodes (optional).
Preparing compound dispensary plate
Take an empty 96-well plate and dispense 125 µL assay medium in columns of six wells over the plate. This will be the ‘compound dispensary plate’. The amount of columns depends on the amounts of assay plates that are used. Each column will correspond to one assay plate/barcode (Figure 1). Thaw and vortex your compounds and add 5.21 µL of the compound stock solution to the assay medium. Put the compound dispensary plate on a plate shaker for at least 5 minutes.
Compound dilution series
Use a multichannel pipette to pipet 100 µL from a column of the compound dispensary plate to the middle six wells (B to G) of column 2 of the corresponding assay plate (Figure 1). Pipet up and down three times and transfer 100 µL into the next column. Repeat until column 9 and discard the 100 µL after this last column. Repeat this for all assay plates. Add 100 µL fresh assay medium to the middle six wells (B to G) of lane column 11. This is the cell control (CC).
Infection
Place a container with decontaminant in the biosafety cabinet ( Prepare a BUNV-mCherry virus suspension with approximately a 0.1 MOI in assay medium (requires a total volume of 6 mL per assay plate). Vortex well. Pour the virus suspension in a reagent reservoir. Dispense 100 µL of the virus suspension over the middle six wells (B to G) of columns 2 to 10 (this includes the VC column) using a multichannel pipette. Close the assay plates and incubate them for 96 hours (4 days) at 37°C in a 5% CO2 humidified atmosphere.
4 Days post infection
Plate inspection
Inspect the assay plates using the inverted light microscope. Check if both the virus- and cell controls contain a (near) confluent monolayer of cells. CPE in the virus control should be minimal.
Nuclei staining using Hoechst
Place a container with decontaminant in the biosafety cabinet (i.e. Virkon-S). Use this to put in all waste generated from this point onward. Prepare a Hoechst solution of 2.5 µM in colorless MEM. You will need 6 mL per assay plate. Take the assay plates and aspirate the culture medium from all wells using a vacuum system that connects to a container with decontaminant. If desired, the supernatants could be collected and stored for virus yield assays, instead of being aspirated. Make sure not to disturb the monolayer. Dispense 100 µL of the Hoechst suspension over the middle 60 wells (rows B to G, columns 2 to 11) using a multichannel pipette. Close the assay plates and incubate them for 15 minutes at 37°C in a 5% CO2 humidified atmosphere.
High content imaging
Place an assay plate in the drawer of the High content imager (HCI).
Open HCS Studio software and create a new ‘AcquireOnly’ protocol and name it ‘HCI_BUNV-mCherry image acquisition protocol’. If done before, skip to step 3.
HCI_BUNV-mCherry image acquisition protocol
Image acquisition is performed on an Arrayscan XTI, Thermofisher, equipped with an X1 CCD camera with a 14-bit dynamic range and a maximal pixel array of 2208 × 2208 with a pixel size of 4.54 micron. A 10X objective is used to image 9 fields per well of the 96-well plate, allowing a sufficient amount of cells to be observed for robust statistical analysis, while maintaining a fast scanning speed. To further maintain scanning speed set the resolution to 1104 × 1104 with 2 × 2 binning and reduce autofocus search coarse planes count to ‘6’. The excitation in the primary channel is set on 386–23 nm with a dichroic mirror and emission filter on BGRFRN_BGRFRN. Apply this channel as autofocusing channel throughout the experiment. Optimal exposure time was set to 10 ms (based on 50% of the dynamic range of the 14-bit camera). The secondary channel is fixed on an excitation 560–25 nm with the dichroic mirror and emission filter on BGRFRN_BGRFRN with an exposure time of 0.059 seconds (Figure 2).
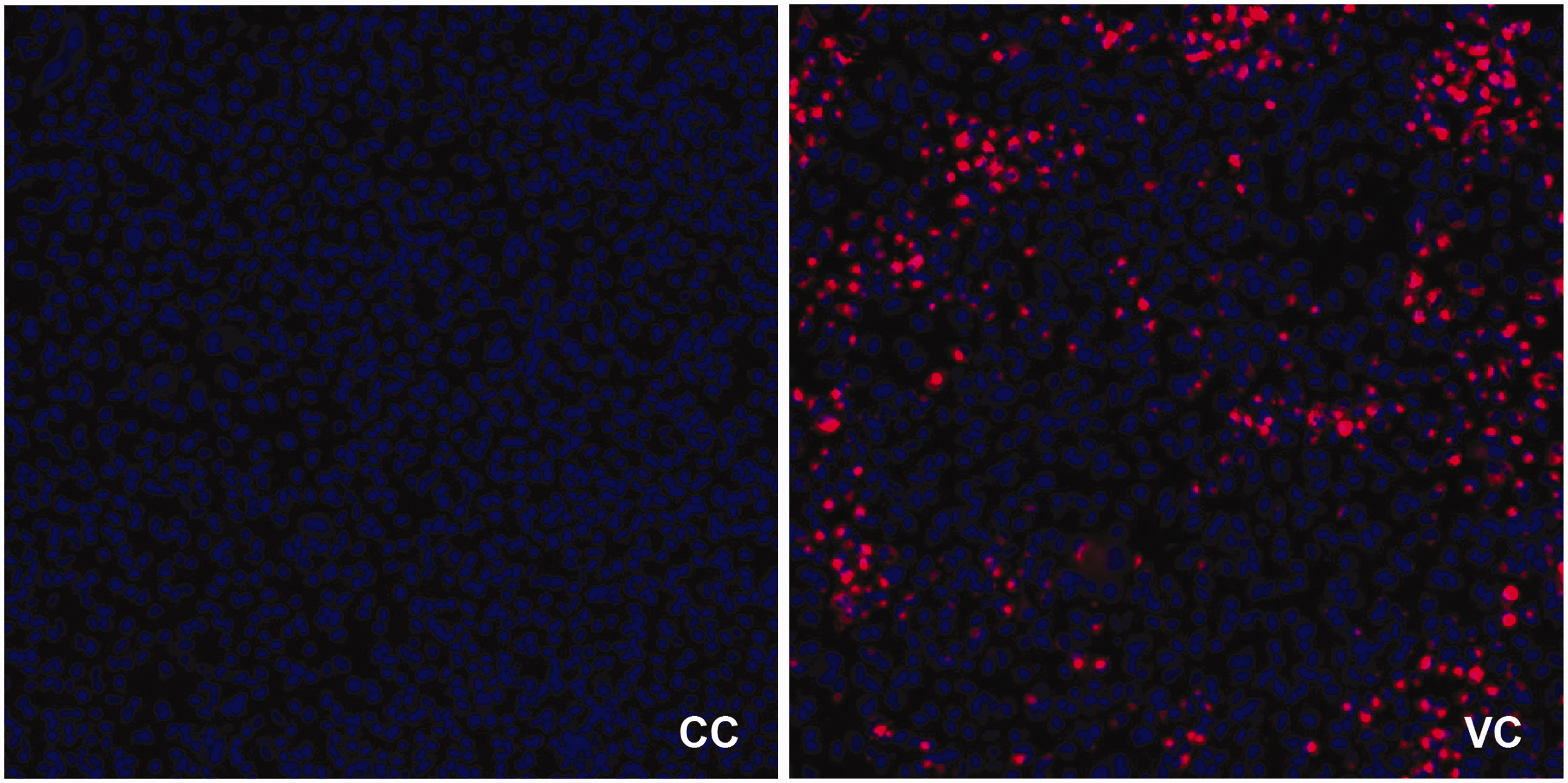
Images of single fields from a cell control (CC) and virus control (VC) with the cell nuclei stained with Hoechst 33342 (blue) and BUNV-mCherry infected cells expressing the mCherry fluorophore (red).
3. Select correct form factor.
4. Good practice but not necessary: check control wells:
i. Select control well.
ii. Autofocus channel 1 and check pixel intensity histogram.
iii. Acquire image from channel 2 and check pixel intensity histogram.
iv. If exposure times need to be adjusted, save protocol with a different name.
5. Select ‘Instrument Based’ in the Scan Plate-tab.
6. Select the middle 60 wells as ‘Scan Area’ in the Scan Settings button.
7. Give the plate a Plate Name or the barcode number as name.
8. Start. The HCI will now begin imaging.
9. When completed, inspect the images using the HCS Studio software. When satisfied with the image quality, proceed with the subsequent assay plates.
10. After imaging, decontaminate and discard the assay plates.
Data processing and analysis
a. Image scanning
1. Open the ‘Scanning’ tool in HCS Studio (Cellomics).
2. Create a new protocol by selecting the ‘SpotDetector.V4’ bio-application using the following settings in the ‘Configure Assay Parameters’ tab and using images acquired as described above. If this protocol already exists, skip to step 4.
BUNV-mCherry SpotDetector protocol
For HCI data processing the Cellomics software of ‘HCS Studio’ is used. Analysis is done by extracting and quantifying specific cell features from the acquired images. First, process the images by applying background reduction. (Setting the correct value for this parameter is crucial, since a background reduction set too low gives increased noise and unspecific signal, while a background reduction set too high filters out the desired fluorescent signal.) The background reduction in the primary channel was set on ‘30’ using the 3D_SurfaceFitting method. Preprocessing of the secondary channel is done by setting the LowPassFilter to ‘255’. This LowPassFilter method is more aggressive on background reduction and works specifically well in images with strong fluorescent signal and high contrast edges. Next, primary object identification (
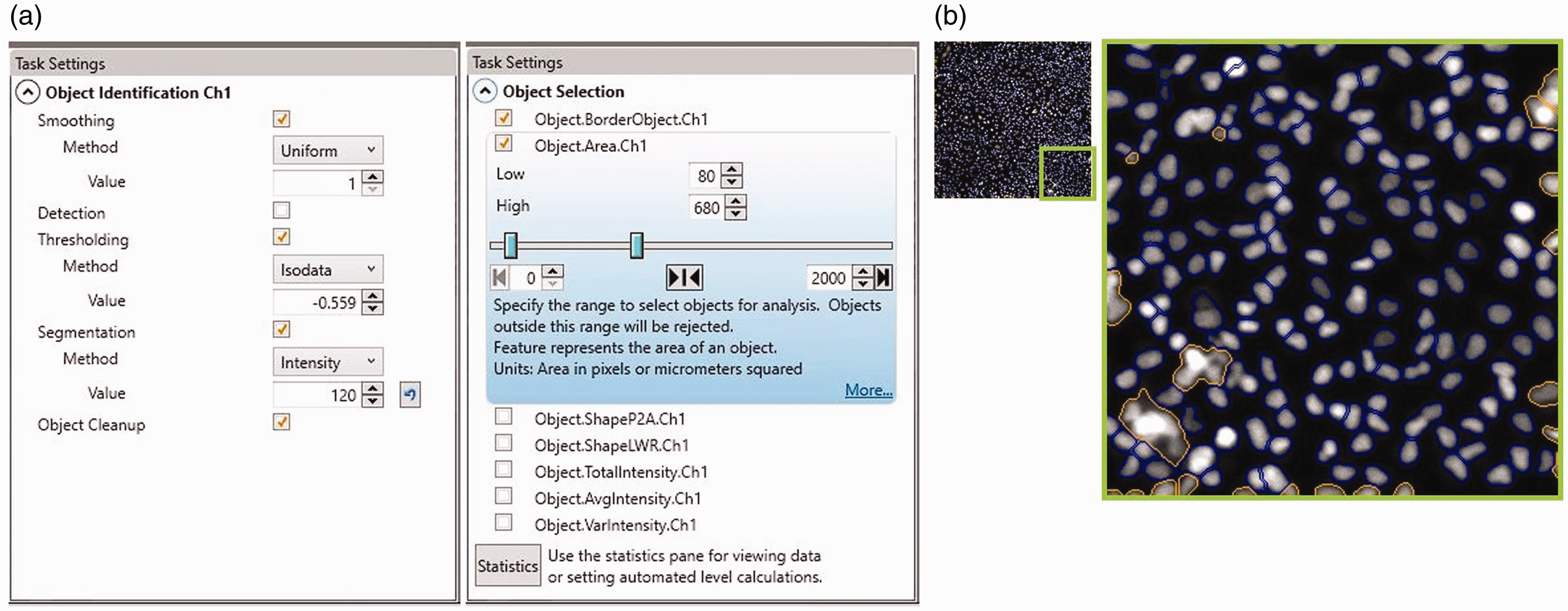
(a) Screenshots from the task settings panel from the HCS Studio software displaying the specific settings for the ‘Object identification’ (left panel) and ‘Object selection’ (right panel) tools showing the defined parameters for this protocol. (b) Original image (left) and enhanced outtake (right) of the cell control showing included objects (encircled in blue) and excluded objects (encircled in brown). The excluded objects are those objects that are partially outside the border of the field or outside the defined size parameter.
Analysis on the secondary channel is done by using the Spot Identification tool. The region of interest was set to the ‘Circ Mask’-type with an expansion of 3 pixels on the primary object mask (Figure 4). This allows the detection of mCherry signal inside a cell (object), and thus viral infection. The signal on channel 2 (mCherry) is detected by setting a fixed threshold value of ‘200’, which is based on the observation of minimal non-background signal on multiple plates across several days. The signal strength is highly dependent on the quality of the assay and the threshold should be optimized accordingly. To score the infection rate use the feature ‘%HIGH_ObjectSpotTotalCountCh2’ and configure ‘Object.SpotCount.Ch2’ by setting both low and high levels on ‘0’. This will score every cell (object) with at least one viral spot (mCherry) as infected cell. Select the following well features for data analysis: ‘ValidObjectCount’ and ‘SpotCountCh2’. Save the optimized protocol and it is ready for use.
(a) Screenshot from the task settings panel from the HCS Studio software displaying the specific settings for the ‘Spot identification’ tool showing the defined parameters for this protocol. (b) Outtakes of images of cell control (CC) and virus control (VC) image fields post-processing showing the included objects (encircled in green) identified in the primary channel and included viral spots (red) identified in the secondary channel. A single red spot within an included object will count as one infected cell by the image analysis algorithm.
3. Save the protocol as ‘BUNV-mCherry_SpotDetector_protocol’.
4. Load protocol by selecting Open → SpotDetector.V4 → ‘BUNV-mCherry_SpotDetector_protocol’.
5. Select ‘Disk Based’ in the Scan Plate-tab.
6. Select the middle 60 wells as ‘Scan Area’ in the Scan Settings button.
7. Give the plate a Plate Name or the barcode number as name.
8. Start.
b. Data collection
1. Open the ‘View’ tool in ‘HCS Studio’ (Cellomics).
2. Open the files you wish to analyze and make sure to include the ‘validObjectCount’, and ‘SpotCountCh2’. The software will now present the number of counted cells per well and the number of infected cells per well respectively. These can be exported into a desired file format, or copied into Excel or GraphPad Prism for further analyses.
c. Analysis of antiviral data
1. Use the data obtained from the ‘SpotCountCh2’ analyses.
2. Average the values of the six cell controls on the plate (avgCC). This number should be (close to) zero, since no virus signal should be obtainable. Signal that is picked up is background noise and will be accounted for in step 4.
3. Average the values of the six virus controls on the plate (avgVC). This will be the 100% value to which to compare the compound treated wells.
4. Calculate the fluorescent signal (% of control): 100 - ((spot count of well - avgVC)/(avgVC-avgCC)*100)
5. Plot the results in a bar graph as compound concentration versus fluorescent signal (% of control) (Figure 5). From here, the EC50 (the compound concentration that reduces the number of mCherry signal spots by half, compared to the virus control) can be determined by logarithmic interpolation.
Example graph of the effect of compound X on BUNV-mCherry replication in A549 cells. Quantified number of infected cells (as percentage of the virus control) are depicted as black bars and cell viability as solid circles.
d. Analysis of cell viability data
1. Use the data obtained from the ‘validObjectCount’ analyses.
2. Average the values of the six virus controls on the plate (avgOC). The virus control is used here, since the difference between the virus controls and the analyzed well is the added compound. Therefore, this gives a more accurate comparison than when comparing to the cell control.
3. Calculate the cell viability (% of control):(object count of well/avgOC)*100
4. Add the results to the graph in a line with markers as compound concentration versus cell viability (% of control) (Figure 5). From here, the CC50 (the compound concentration that reduces the number of viable cells by half, compared to the virus control) can be determined by logarithmic interpolation.
Conclusions
The assay described here allows the researcher to study the antiviral effect of small molecules in a cell-based assay, in a dose dependent manner, against BUNV-mCherry. We estimate that using this exact protocol will allow a single dedicated researcher can prepare well over 100 assay plates per day, assessing over 600 molecules in eight concentrations. One of the major benefits of using an assay not based on CPE scoring, but rather fluorescence as a marker of infection, is the fact that in addition to antiviral effect of the tested compounds also the cell viability can be determined within the same well. This is an important advantage when performing screening of large compound libraries. To further increase the throughput, the current assay could be adapted by using a 384-well plate format, assessing fewer concentrations per molecule and introducing liquid handlers and robotics. This assay methodology could be further applied using other non-CPE causing (fluorescent reporter) viruses within the
Footnotes
Acknowledgments
We would like to express our gratitude to Prof. Xiaohong Shi at the University of St. Andrews, Scotland, UK for providing us with the BUNV-mCherry virus.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
