Abstract
Nucleoside and nucleotide analogs have played significant roles in antiviral therapies and are valued for their impressive potency and high barrier to resistance. They have been approved for treatment of herpes simplex virus-1, HIV, HBV, HCV, and influenza, and new drugs are being developed for the treatment of RSV, Ebola, coronavirus MERS, and other emerging viruses. However, this class of compounds has also experienced a high attrition rate in clinical trials due to toxicity. In this review, we discuss the utility of different biochemical and cell-based assays and provide recommendations for assessing toxicity liability before entering animal toxicity studies.
Nucleoside analogs were first studied as antimetabolites in the 1950s by George Hitchings and Gertrude Elion. 1 Their work led to the discovery of treatments for leukemia, gout, hyperuricemia, and parasitic protozoan infection, as well as an immunosuppressant for organ transplant. These efforts paved the way for the discovery of a series of nucleoside analogs as antitumor agents in the 1970s, some of which, such as ara-C and gemcitabine (Figure 1), are still in use today.2,3 Many of these antitumor nucleoside analogs showed antiviral activity, but none of them possessed an adequate clinical safety profile for antiviral use until Acyclovir (Figure 1) was discovered for the treatment of herpes simplex virus (HSV) infection, a breakthrough that started a new era in antiviral therapy. 1 Acyclovir’s remarkable safety profile stemmed from two properties: (1) activation of the compound to its monophosphate (MP) only occurs in virus-infected cells, since this process relies on a viral thymidine kinase; and (2) acyclovir 5’-triphosphate (TP) selectively inhibits viral DNA polymerase over host DNA polymerase α. 4 Follow on generations of HSV nucleoside antivirals (penciclovir and ganciclovir, Figure 1) and prodrugs were developed to improve potency and resistance profiles. 5
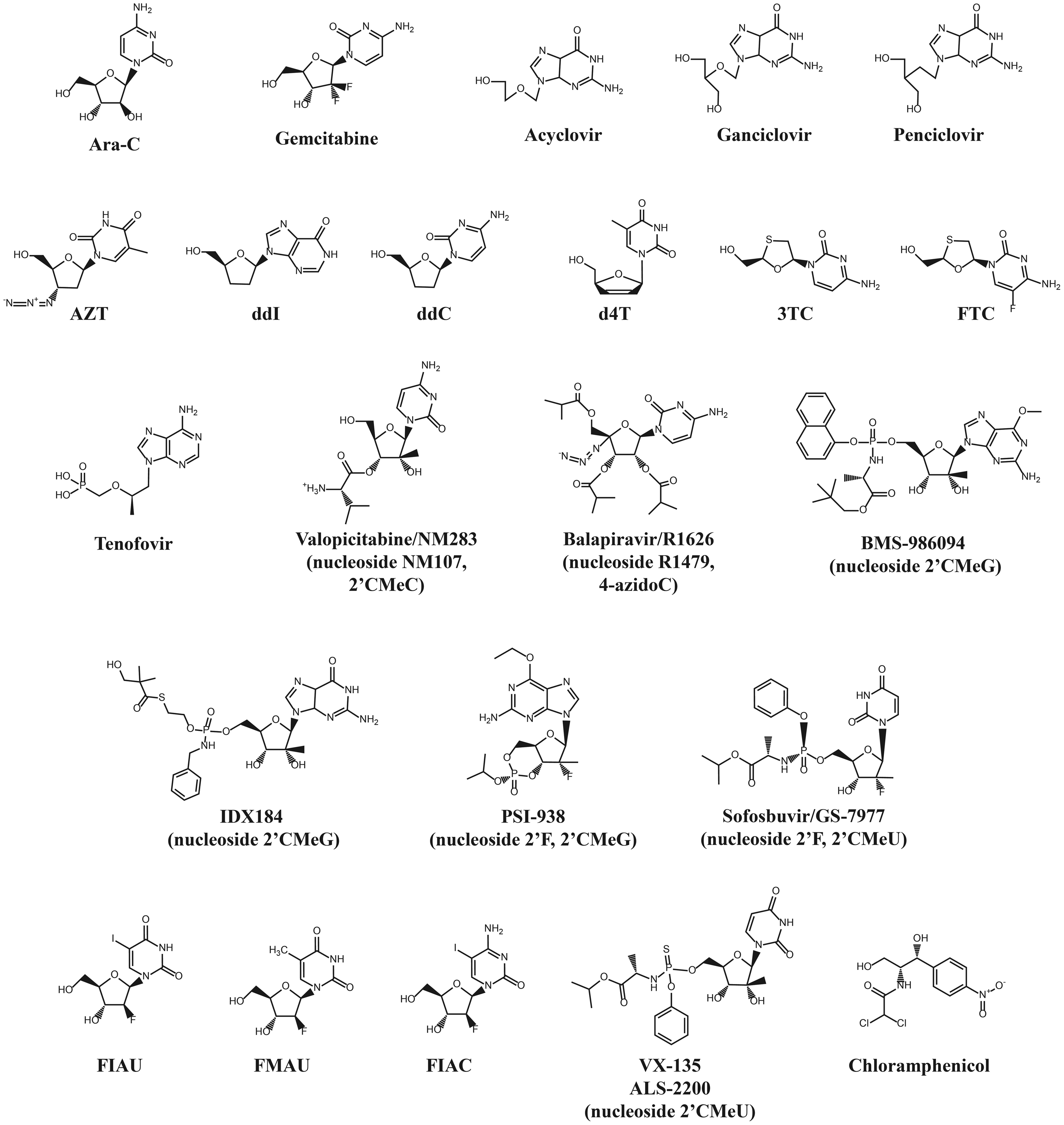
Chemical structures of compounds discussed in this review.
The HIV/AIDS epidemic in the 1980s presented unprecedented demand for effective antivirals. In 1986, azidothymidine (AZT, zidovudine, Figure 1), a chain-terminating inhibitor of HIV-1 reverse transcriptase (RT), was approved, signifying the first step towards control of this devastating disease. The first generation of nucleoside analogs for HIV was efficacious, but these compounds also showed considerable toxicity, including cardiomyopathy (AZT, ddI, and ddC, Figure 1), peripheral neuropathy (d4T, ddI, and ddC), sensorineural deafness (ddC), lactic acidosis (AZT, ddI, and d4T), diabetes (ddI), and cytopenia (AZT).6,7 The toxicity of ddC and AZT was associated with inhibition of mitochondria DNA synthesis in cell culture8–10 and in patient tissue samples. 11 Later on, detailed kinetic studies of HIV nucleosides using recombinant human mitochondrial DNA polymerase (Pol γ) demonstrated the importance of selective inhibition of on-target HIV-1 RT over that of the off-target human Pol γ.7,12 Unfortunately, this knowledge came 10 years too late to inform the 1993 Phase II clinical trial of FIAU in patients infected with HBV, where 7 out of 15 patients developed severe liver toxicity and lactic acidosis, and 5 of these patients died. 13 Follow-up biochemical, cellular, and in vivo studies showed that FIAU was readily incorporated into mitochondrial DNA, leading to a decrease in mtDNA abundance and thus rendering the mitochondria dysfunctional.14,15 The prevalence of mitochondrial toxicity associated with 2′-deoxy nucleoside analogs led to the FDA’s recommendation to test drug candidates in a set of mitochondrial toxicity assays including Pol γ inhibition, lactic acid formation, mitochondrial DNA content, and glucose utilization. 16 With the use of these tests, the second-generation antiviral nucleoside/tide analogs such as 3TC, FTC, and tenofovir were shown to be poor substrates of Pol γ and demonstrated significantly improved clinical safety profiles.
Despite our collective knowledge of nucleoside analog-associated mitochondrial toxicity, safety concerns continued in the discovery of HCV antivirals. A number of HCV nucleoside/tide analogs failed in Phase II due to toxicity or association with a toxic analog. These include valopicitabine (NM283, gastrointestinal toxicity, Figure 1), 17 balapiravir (R1626, hematologic toxicity, Figure 1), 17 BMS-986094 (cardiac and kidney toxicity, Figure 1), 18 IDX184 (clinical hold due to association with BMS-986094, Figure 1), 19 and PSI-938 (liver toxicity, Figure 1). 17 With the exception of PSI-938, the TP forms of these compounds are substrates of mitochondrial RNA polymerase (POLRMT). Balapiravir is a specific mitochondria toxin, inhibits POLRMT, reduces mitochondrial protein synthesis, and eventually decreases mitochondrial respiration.20,21 Our data showed that 2′CMeG-TP is incorporated into RNA by POLRMT, decreases mitochondria protein synthesis, and decreases mitochondrial cellular respiration.20,22 Furthermore, Jin et al. demonstrated that the toxicity of BMS-986094 is directly correlated to its 2′CMeGMP nucleotide moiety rather than the naphthalene-phosphorylamino propanoate prodrug moiety. 21 Interestingly, when compared to selective mitochondria inhibitors such as ddC, 4-azidoC, and chloramphenicol (Figure 1), the inhibition profile of BMS-986094 is distinctly different for mitochondrial protein synthesis and respiration, indicating it may hit other off-targets such as RNA polymerase I. 20 As of today, the only approved nucleotide prodrug for the treatment of HCV is sofosbuvir (Figure 1), a 2′F, 2′CMe uridine prodrug that is well tolerated in patients. 23 The TP form of sofosbuvir showed no inhibition of any of the human polymerases tested and is a poor substrate of POLRMT. 20 The relevance of POLRMT inhibition as a marker for nucleotide toxicity has been further supported by works from multiple groups.20,21,22,24 A clear outlier is PSI-938, a 2′F, 2′CMe guanosine prodrug, which showed liver toxicity in clinical trials but is a poor substrate of POLMRT and showed no toxicity in any of the in vitro studies conducted so far. 20
Host polymerases have been generally regarded as the primary off-targets for this class of compounds. However, the observed toxicities tend to be highly unpredictable.7,25 These may be attributed to complex uptake, distribution, and accumulation of different compound in different organs. For example, three closely related analogs, FIAU, FMAU, and FIAC (Figure 1), showed distinctly different toxicities in animals and human. FIAU showed no toxicity in non-clinical animal species including mice, rats, dogs, and monkeys, but conflicting toxicity findings from two separate studies in woodchucks.15,26 In contrast, FIAU-induced hepatotoxicity was observed in clinical trials in week 13 and beyond under potentially much lower exposures than the animal models.15,26 In addition, FMAU, a metabolite of FIAU, was tested in a Phase I clinical trial for the treatment of murine leukemia. It caused severe neurologic toxicity and two deaths among eight patients after 8–10 days of treatment. Toxicity, primarily hematopoietic, was observed with FIAC, the cytidine analog and known metabolic precursor of FIAU, when used to treat HSV and vesicular stomatitis virus infection. 26 Interestingly, while the more recent HCV nucleotide antivirals, e.g. BMS-986094, PSI-938, and VX-135 (also known as ALS-2200, Figure 1) were all liver-targeting nucleotide prodrugs and the administration of PSI-938 and VX-135 resulted in liver toxicity,27,28 BMS-986094 showed cardiovascular and renal toxicity. 29
It is worthwhile mentioning that antiviral nucleosides could also exert toxicity through disruption of natural NTP pools. AZT only showed modest inhibition of mitochondrial DNA polymerase 7 but is a competitive inhibitor of thymidine kinase 2-catalized phosphorylation of thymidine to TMP. 30 Ribavirin is a poor substrate for mitochondrial RNA polymerase, 22 but it reduces the size of the guanosine triphosphate (GTP) pool via inhibition of inosine MP dehydrogenase. 31
As one might expect, there are many possible ways that a nucleoside/tide analog can elicit toxicity that are still beyond our knowledge. In this review, we focus on the host polymerase mechanism-based toxicity and summarize different biochemical and cellular assays to evaluate potential toxicity liabilities, and the strength and limitations of each method.
Biochemical evaluations
As most nucleoside/tides analogs need to be transformed into 5’-TP to exert on-target and off-target effects, their TP forms are necessary for elucidating these mechanisms (Table 1). The typical initial enzymatic assay for polymerases is a radioactivity-based polymerization assay, in which the inhibition by nucleotides is measured as an IC50 value. The inhibition IC50 assays can be easily adapted to a 96-well filter binding assay, and multiple compounds can be accessed simultaneously. However, based on inhibition studies alone, one cannot establish whether the nucleotide was actually incorporated. Therefore, a single nucleotide incorporation (SNI) assay can be particularly useful to determine if a NTP analog actually served as a substrate for the polymerases. Under a high concentration of NTP analogs (e.g. 500 µM ribose nucleotide TP (rNTP) or 50–100 µM 2′-deoxyribose nucleotide TP (dNTP)), the relative rate of incorporation of an analog can be compared to that of its natural NTP counterpart. For further mechanistic insight, a pre-steady state kinetic analysis of the NTP incorporation offers detailed kinetic parameters such as pre-steady state polymerization rate constant kpol and NTP dissociation constant Kd. In these studies, incorporation rates are studied using multiple time points and nucleotide concentrations, often using quench-flow techniques, making these studies reagent- and time-consuming. Finally, the mechanism of a nucleotide analog-derived enzyme inhibition can be revealed by an elongation assay, in which a stable elongation complex is formed, and the incorporation of the next few natural NTPs is measured. Normally, an NTP analog can be characterized into one of the three groups: chain-terminators, delayed chain-terminators, or stably incorporated analogs. Stably incorporated NTP analogs are difficult to study for toxicity as they could lead to little or no quantitative changes in DNA, RNA, or protein expression. 36
Biochemical and cellular assays to evaluate general and mitochondria toxicities of nucleoside/tide analogs.
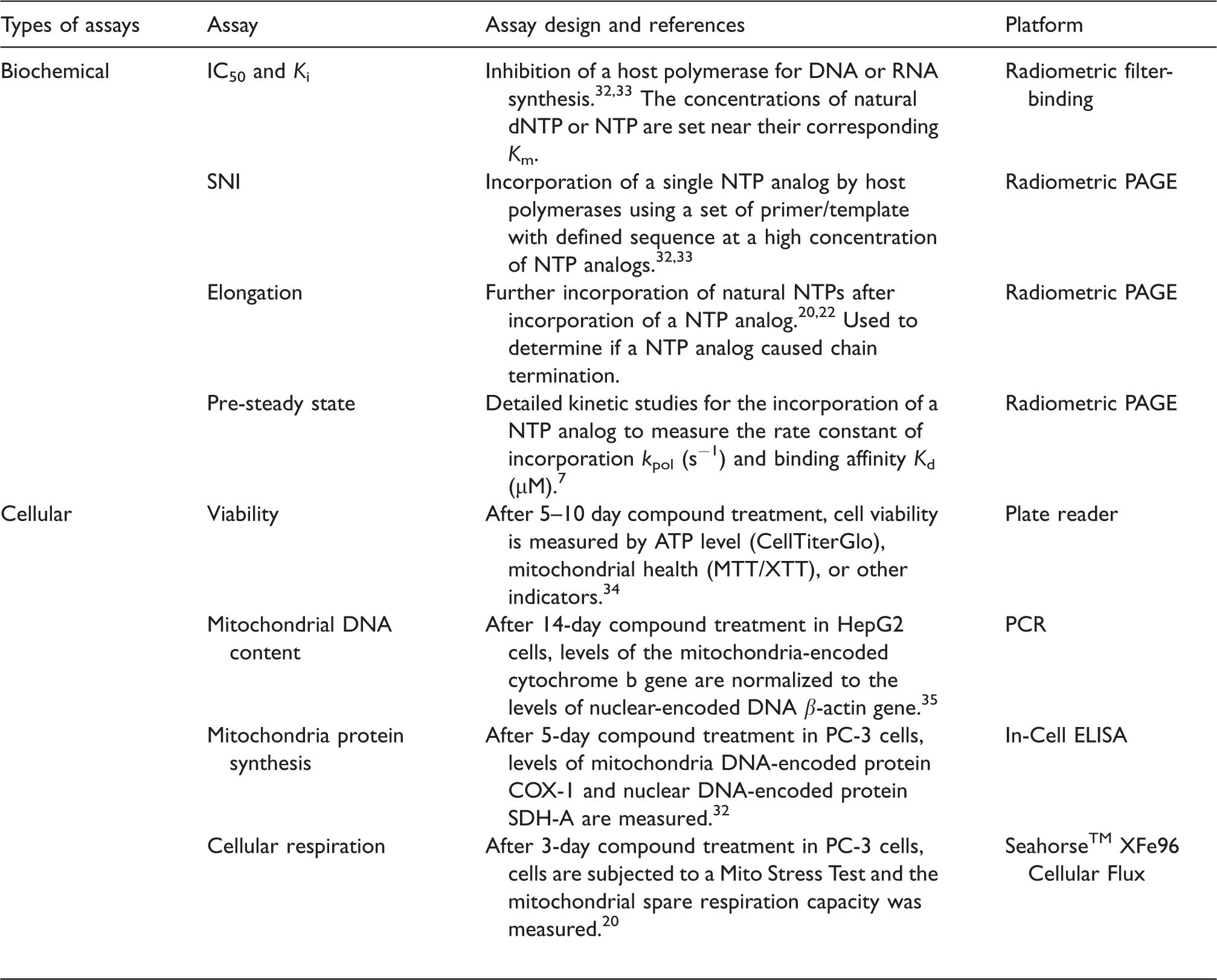
SNI: single nucleotide incorporation; NTP: nucleotide triphosphate; PAGE: polyacrylamide gel electrophoresis; PCR: polymerase chain reaction.
The quality of the above-mentioned biochemical assays relies heavily on the availability of high quality TP active forms, which can be time- and resource-consuming to synthesize, and the TP salts are subject to degradation overtime. At times, certain inorganic contaminants could lead to apparent inhibition of polymerases.
Cellular evaluations
Toxicity is cell line-dependent
Due to their mechanism of action, nucleoside analogs are generally evaluated for 5–8 days in both replicating laboratory-adapted cell lines and non-replicating primary cells. 18 A standard readout is the effect of compound on the ATP level within the cells. We found that among 20+ laboratory-adapted cells we tested, prostate cancer-derived PC-3 cells and T-cell leukemia-derived MT4 cells are generally the most sensitive to toxins, and are significantly more sensitive than HepG2 cells, a widely used cell line for in vitro toxicity studies. In addition, we found that laboratory-adapted cells are as useful for detecting potential toxicity liabilities as primary cells.
In certain cases, testing in a specific cell assay is warranted. For example, a number of the first-generation HIV nucleoside analogs showed myelosuppressive effects. As a result, the FDA recommended evaluating this toxicity on the growth of human bone marrow progenitor cells in colony formation assays. 37
The importance of measuring cellular TP levels
A nucleoside/tide analog’s cytotoxicity is often directly related to its cellular level of the active TP metabolite. BMS-986094 and IDX184 are both 2′CMe guanosine nucleotide prodrugs, but the former formed >37-fold higher TP, and this correlated with its significantly higher toxicity. 18 Some nucleoside analogs are poorly phosphorylated in cells, such as balapiravir. Even though it is a specific inhibitor of POLRMT, balapiravir showed little or no toxicity in a large panel of cell lines and primary cells tested, but development of this compound was stopped at Phase II due to toxicity.
Evaluation of mitochondrial toxicity
As shown in Table 1, we employed three mitochondria-focused cell-based assays to evaluate potential mitochondrial toxicity. The mitochondrial DNA content assay has been widely used to study 2′-deoxy nucleoside analogs, but it does not assess the effect of ribose nucleoside/tide analogs on mitochrondrial RNA synthesis.20,29 ddC serves as a good positive control and HepG2 cells seem to be an ideal cell line for this assay.
The mitochondrial protein synthesis assay has been useful to evaluate ribonucleoside/tide analogs. In this assay, the levels of two proteins are measured simultaneously: the mitochondrial DNA-encoded protein cytochrome c oxidase 1 (COX-1) and a nuclear DNA-encoded protein succinate dehydrogenase A (SDH-A). 32 For data analysis, we found that comparing the individual levels of the COX-1 and SDH-A proteins to the untreated DMSO control yielded more reproducible data than using the ratio of COX-1:SDH-A. Chloramphenicol, a specific inhibitor of mitochondrial protein synthesis, is used as the positive control.
Since the 1950s, mitochondrial respiration has been regarded as the “gold standard” for the measurement of mitochondrial function. 38 It has been used extensively to study mitochondrial toxins that directly impair the electron transport chain but has not been used to study the nucleoside/tide analogs until recently.39–42 This assay provides a sensitive functional readout and does not require knowledge of the mechanism of action. 20 A complete loss of mitochondrial respiration capacity can be detected before any change in the cellular ATP level, and this phenomenon has been observed for inhibitors with diverse molecular targets including ddC (inhibitor of mitochondrial DNA synthesis), 4′-azido cytidine (inhibitor of mitochondrial RNA synthesis), and chloramphenicol (inhibitor of mitochondrial protein synthesis). For compounds that show both mitochondrial and general toxicity (such as BMS-986094), this method makes it possible to identify whether mitochondria toxicity occurs at a lower drug concentration. 20 For both mitochondrial protein and respiration assays, we found that PC-3 cells offer higher sensitivity and reproducibility than HepG2 cells. 20
An assay with limited predictive value: The glucose–galactose CC50 assay
When faced with mitochondria damage, cells are known to switch to glycolysis to generate ATP; therefore the glucose–galactose CC50 assay has been widely used to detect mitochondria toxicity. 43 In the absence of glucose, the galactose-adapted cells are forced to rely heavily on mitochondrial OXPHOS and thus would show a ≥3-fold decrease in CC50. 44 However, the predictivity of this assay was challenged by Hyne’s 2013 report showing that this assay only detected 2–5% of the 200 potential mitochondrial toxins tested.44,45 Even though no nucleoside/tide analogs were included in Hyne’s study, it is consistent with our finding from testing >30 clinically relevant nucleoside/tide analogs. None of the known mitochondrial toxins such as ddC, FIAU, or 4′-azido C showed enhanced cytotoxicity in galactose-adapted cells.
Evaluation of toxicity in animal studies
Research and development of a significant number of nucleoside/tide analogs were terminated in the pre-clinical stage due to toxicity in the animal models; though only a few were reported24,46,47 and mitochondrial toxicity was indicated or suspected. To date, due to a number of factors outlined above, mitochondrial toxicity has been difficult to detect in animal species and correlation to humans is at best tenuous. 48 Potentially, animal models with genetic mitochondrial abnormalities or humanized animals may be more sensitive to nucleoside/tide toxicity. For example, mice models with genetic mitochondrial abnormalities such as heterozygous superoxide dismutase 2 knockout (Sod+/−) mice were more susceptible to mitochondria toxins than the wild-type animals. 49 Recently, Xu et al. reported FIAU-induced liver toxicity in chimeric TK-NOG mice grafted with humanized livers. 50 Animal models like these may improve our ability to detect nucleoside toxicity.
A comment on therapeutic window
As viral diseases vary greatly in duration and severity, the safety threshold of an antiviral nucleoside/tide needs to be considered in the context of its efficacy and treatment duration. For example, compounds to treat acute and deadly viral infections such as Ebola would likely have different safety criteria than drugs used to control chronic infections such as HIV-1 and HBV. Existing nucleoside/tides are routinely screened against emerging viruses, and it is possible that a compound that failed to achieve an adequate therapeutic window against one virus may be suitable for treating other viral infections.
Summary
Anti-viral nucleoside/tide analogs target the viral DNA and RNA replication machinery, and this mechanism makes them potential inhibitors of host DNA and RNA synthesis as well, especially mitochondrial DNA and RNA synthesis. Mitochondrial toxicity manifests in multiple forms in vivo and underlies many clinical-stage failures and has been difficult to detect in animal studies. Over the past 30 years, a battery of in vitro tests for mitochondrial toxicity has evolved and continues to expand, allowing for improved early detection of mitochondrial liability. As for any drug molecule, the actual toxicity liability cannot be assessed fully until tested in animal toxicity studies and eventually in human clinical trials. Nevertheless, we believe that employing the proper biochemical and cellular assays early on will have a meaningful impact on reducing pre-clinical and clinical toxicity liability for this class of compounds.
Footnotes
Acknowledgement
We thank Dr. Brian Schultz, Dr. Roman Sakowicz, and Dr. Chin Tay for careful review of this manuscript and insightful comments.
Declaration of conflicting interests
The author is an employee and shareholder of Gilead Sciences, Inc.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
