Abstract
Objective
Circadian rhythms in cartilage homeostasis are hypothesized to temporally segregate and synchronize the activities of chondrocytes to different times of the day, and thus may provide an efficient mechanism by which articular cartilage can recover following physical activity. While the circadian clock is clearly involved in chondrocyte homeostasis in health and disease, it is unclear as to what roles it may play during early chondrogenesis.
Design
The purpose of this study was to determine whether the rhythmic expression of the core circadian clock was detectable at the earliest stages of chondrocyte differentiation, and if so, whether a synchronized expression pattern of chondrogenic transcription factors and developing cartilage matrix constituents was present during cartilage formation.
Results
Following serum shock, embryonic limb bud–derived chondrifying micromass cultures exhibited synchronized temporal expression patterns of core clock genes involved in the molecular circadian clock. We also observed that chondrogenic marker genes followed a circadian oscillatory pattern. Clock synchronization significantly enhanced cartilage matrix production and elevated SOX9, ACAN, and COL2A1 gene expression. The observed chondrogenesis-promoting effect of the serum shock was likely attributable to its synchronizing effect on the molecular clockwork, as co-application of small molecule modulators (longdaysin and KL001) abolished the stimulating effects on extracellular matrix production and chondrogenic marker gene expression.
Conclusions
Results from this study suggest that a functional molecular clockwork plays a positive role in tissue homeostasis and histogenesis during early chondrogenesis.
Introduction
Circadian (~24-hour) rhythms in mammals are orchestrated by a hierarchical system of self-sustained tissue clocks. The central, light-synchronized circadian pacemaker is the suprachiasmatic nucleus (SCN) situated in the hypothalamus, which drives circadian timing of behavioral, neuroendocrine, and metabolic rhythms in multicellular organisms. 1 These SCN rhythms in turn transmit timing information to synchronize molecular clocks in peripheral organs, such as the lung, liver, adrenal gland, or pancreas. 2 Self-sustained circadian rhythms are an inherent property of individual cells, 3 which can persist throughout the body in vivo, and also in isolated tissues/cells for many days in vitro. 4 The molecular circadian clocks have emerged as key regulators of cellular activity and regulate cell type–specific gene expression patterns over ~24-hour time frames. The proportion and identity of genes that are rhythmically transcribed differ between tissues and can range from 5% to 20% of the cell-specific transcriptome, 5 and include genes that play key roles in tissue homeostasis. 6
The molecular mechanisms that underlie the molecular clockwork are well characterized.6,7 The core oscillator is driven by a transcriptional/translational feedback loop (TTFL), which consists of interconnected positive and negative limbs. In the positive limb, a heterodimer of BMAL1 (aryl hydrocarbon receptor nuclear translocator-like ARNTL/BMAL1) and CLOCK (circadian locomotor output cycle kaput) drives rhythmic transcription of downstream genes by binding to E-box elements in their promotor regions. Among these are other clock genes, including period (PER1 and 2), cryptochrome (CRY1 and 2), and REV-ERB (nuclear receptor subfamily 1 group D member 2 [NR1D2/REV-ERB]), as well as a wide array of clock-controlled genes (CCGs) that influence physiology in a tissue-specific manner. 8 PER and CRY genes constitute the negative limb of the TTFL, and the heterodimers of their protein products interact with the CLOCK:BMAL1 complex, thereby inhibiting gene transcription. The positive and negative limbs are also interconnected by retinoic acid–related orphan nuclear receptors (i.e., REV-ERBs), which form part of a “stabilizing loop”; REV-ERBs repress the transcription of BMAL1. These autoregulatory loops constitute the circadian molecular clock, which takes approximately 24 hours to complete.2,6 Despite advances in understanding the rhythmic outputs of the circadian clock, less is known about how these highly conserved molecular clocks adapt to the local microenvironment within individual cells of peripheral tissues.
Diurnal rhythms in cartilage metabolism have been known for some time. 9 These cycles are hypothesized to temporally segregate and synchronize the activities of chondrocytes to different (optimal) times of the day. This may provide an efficient mechanism by which cartilage extracellular matrix (ECM) can recover following physical activity. 10 It has been established that the most rapid phase of chondrocyte proliferation in the epiphyseal growth plate is early in the morning, which induces growth plate expansion and increase in ECM synthesis that peaks at noon. 11 Since then, the expression of clock proteins in knee articular cartilage has been demonstrated.12,13 The intrinsic nature of cartilage circadian rhythms has been revealed through observations of cell-autonomous rhythms in gene expression in a human chondrocyte–derived cell line in vitro.14,15
Perturbations of the peripheral clocks has been linked to a number of pathologies, including diabetes, tumorigenesis, and neurodegeneration, which result from abnormal cell activity and metabolism.16-18 Indeed, the chondrocyte clock was found to be disturbed in osteoarthritis (OA), suggesting that circadian rhythms may be a critical step that is disrupted in OA development. 19 Knockdown of BMAL1 in healthy chondrocytes has resulted in upregulation of ECM-degrading enzymes in vitro, and conversely, restoring BMAL1 expression in OA chondrocytes has led to the downregulation of the same set of genes.20,21 More recently, overexpression of PER2 has been observed to upregulate ECM-degrading enzymes ADAMTS5 and MMP13 and downregulated the key chondrogenic transcription factor SOX9 in the H5 chondrocyte cell line in vitro. 22
At present, we lack information on the development of circadian rhythms in early chondrogenesis. This question is particularly relevant given the association between perturbed circadian rhythms in chondrocytes and OA, but it is unclear as to what roles the molecular clock may play during early chondrogenesis. Moreover, given the relevance of the production of cartilage ECM constituents in these early developmental stages, new information in this field will be critical for stem cell-based cartilage regeneration techniques. Here we set out to study whether the rhythmic expression of the core circadian clock was detectable at the earliest stages of chondrocyte differentiation, and if so, whether a circadian expression pattern of chondrogenic transcription factors and developing cartilage ECM constituents was present during cartilage formation.
Methods
Primary High-Density Chondrifying Cell Cultures
Chondrifying high-density (micromass) cell cultures were established from chicken embryos (Ross hybrid) of Hamburger-Hamilton stages 22 to 24 as previously described.
23
In this model, chondroprogenitor cells differentiate to matrix-producing chondrocytes on culturing days 2 to 3, and form hyaline cartilage by the end of the 6-day-long culturing period. Work on cells derived from early stage (4-day-old) chick embryos does not require an approval from the Ethics Committee of the University of Debrecen. Distal parts of forelimbs and hindlimbs of embryos were isolated, pooled, and dissociated in 0.25% trypsin-ethylenediaminetetraacetic acid (Sigma-Aldrich, St. Louis, MO, USA; pH 7.4) at 37°C for 1 hour. Dissociation was terminated by adding equal volumes of fetal bovine serum (FBS; Gibco, Gaithersburg, MD, USA). The digested limb buds were then filtered through a 20-μm pore size plastic filter (Millipore, Billerica, MA, USA) to generate a single-cell suspension of chondrogenic mesenchymal cells. Cells were then centrifuged (800 × g for 10 minutes at room temperature) and resuspended in Ham’s F12 culture medium (Euroclone, Pero, Italy) supplemented with 10% FBS at a concentration of 1.5 × 107 cells/mL. One hundred microliter droplets were inoculated into plastic 6-well plates (Eppendorf, Hamburg, Germany). After allowing the cells to adhere to the surface for 2 hours in a CO2 incubator (5% CO2 and 90% humidity), 2 mL of Ham’s F12 supplemented with 10% FBS, 0.5 mM
Validation of Cartilage Formation in Micromass Cultures by Metachromatic Staining
Micromass cultures were inoculated onto the surface of 30-mm round cover slips (Menzel-Gläser, Menzel GmbH, Braunschweig, Germany) placed into 6-well culture plates. For qualitative and semiquantitative evaluation of cartilage matrix production, on each day from day 1 until day 6, cultures were stained with dimethyl methylene blue (DMMB; pH 1.8; Sigma-Aldrich) or toluidine blue (TB; pH 2; Reanal, Budapest, Hungary) metachromatic dyes as previously described. 23 DMMB and TB metachromatic staining procedures were carried out on separate colonies; DMMB-stained specimens are visual representations of TB assays. Optical densities of extracted TB samples were determined at 620 nm using a microplate reader (Chameleon, Hidex Ltd., Turku, Finland) from 3 cultures on each day in 3 independent, biological replicate experiments.
Synchronizing the Circadian Clock in Chondrifying Cultures
In order to synchronize the molecular circadian clocks between cells in micromass cultures prior to de facto cartilage formation (during the earliest stages of chondrogenesis), chondrogenic cultures were serum-shocked using the established method of replacing the culture media with F12 supplemented with 50% FBS for 2 hours as described earlier. 24 Serum-shock synchronization was also applied on mature (6-day-old) colonies containing mature chondrocytes. After 2 hours, cultures were rinsed with phosphate buffered saline (PBS) and fresh F12 medium (containing 10% FBS and 1% P/S) was added. Serum shock was considered as time point 0. After an initial washout of the immediate effects of serum shock, samples were collected every 8 hours between 24 and 72 hours postsynchronization. Control, nonsynchronized cultures from the same experiments received a medium change with fresh F12 containing 10% FBS at time point 0 and harvested at the same time points as the serum-shocked colonies. Serum-shocked cultures (both early and late stage) were also collected 2 and 4 days postsynchronization (on culturing days 3 and 6) to monitor long-term effects on cartilage-specific gene expression patterns compared with nonsynchronized colonies. Harvested micromass cultures were immediately frozen in liquid nitrogen and stored at −80°C until RNA isolation.
Influencing the Function of the Molecular Clock by Small Molecule Compounds
Longdaysin (abbreviated as LDS; Cat. No.: SML0127; Sigma-Aldrich) is a clock modifier that inhibits casein kinase I (CKI). CKI phosphorylates the period (PER) protein and promotes its degradation through a proteasomal pathway. 25 KL001 (Cat. No.: SML1032; Sigma-Aldrich) is a small molecule that specifically interacts with the cryptochrome (CRY) protein and prevents its ubiquitin-dependent degradation. 26 Both compounds were dissolved in dimethyl sulfoxide (DMSO) at 5 mM and then applied separately to micromass cultures at the final concentration of 5 µM for 24 hours on culturing day 1, with or without serum shock. For these experiments, control cultures were treated with equal amount of the vehicle (DMSO).
Mitochondrial Activity (MTT) Assay
For mitochondrial activity (cell viability) assays, cells cultured in 24-well plates were used. Tel microliters of MTT reagent (3-[4,5-dimethylthiazolyl-2]-2,5-diphenyltetrazolium bromide; 5 mg MTT/1 mL PBS) were pipetted into each well on culturing day 3. Cells were incubated for 2 hours at 37°C, and following the addition of 500 μL MTT solubilizing solution, optical density was measured at 570 nm (Chameleon, Hidex). Optical density readings of the experimental groups have been normalized to those of the vehicle controls and shown as percentage changes.
RNA Isolation and Reverse Transcription
Total RNA was isolated from micromass cultures using the RNeasy kit (Qiagen, Hilden, Germany) according to the instructions of the manufacturer. Genomic DNA was eliminated from samples using gDNA spin columns. RNA concentration and purity were determined by a Nanodrop 1000 UV-Vis spectrophotometer (Thermo Fisher Scientific). For gene expression analyses, 1 μg of RNA was reverse transcribed into complementary cDNA using the High-Capacity cDNA Reverse Transcription Kit (Thermo Fisher Scientific), as per the protocol supplied by the manufacturer. cDNA was stored at −20°C.
Quantitative Real-Time Polymerase Chain Reaction (RT-qPCR) Analyses
The expression patterns of clock genes, osteo/chondrogenic transcription factors, and genes coding for cartilage ECM structural proteins were determined using RT-qPCR by absolute quantification using the standard curve method. Primer pairs were ordered from Integrated DNA Technologies (Coralville, IA, USA). For sequences of primer pairs please see Table 1 . First, standard curves had been generated by conventional PCR using the Promega GoTaq Flexi DNA Polymerase kit (Promega, Madison, WI, USA) by adding the following components (per 50 μL reaction): 1.25 U GoTaq polymerase; 3 mM MgCl2; 0.2 mM dNTP; 200 nM primers; and 50 ng cDNA (pooled from 3- and 6-day samples). Amplification was performed in a programmable thermal cycler (MultiGene 96-well Gradient Thermal Cycler; Labnet International, Edison, NJ, USA) using the following thermal profile: initial denaturation at 95°C for 5 minutes, followed by 40 cycles of denaturation at 95°C for 15 seconds, annealing at 58°C for 20 secconds, extension at 74°C for 20 seconds; followed by final extension at 74°C for 5 minutes. PCR products were loaded on 1.5% agarose gels and were isolated using a Roche High Pure PCR Product Purification Kit (Roche, Basel, Switzerland) as per the instructions of the manufacturer. The original gel images are shown in the Supporting Information (Supplemental Figure S1). DNA concentration of purified PCR products was determined using a Nanodrop 1000 UV-Vis spectrophotometer (Thermo Fisher Scientific). Standard curves were prepared by a serial (10-fold) dilution of the samples starting from 1 ng/μL.
Forward and Reverse Primer Sequences, Gene Accession Numbers, and Amplicon Length for the Primer Pairs Employed in This Study.
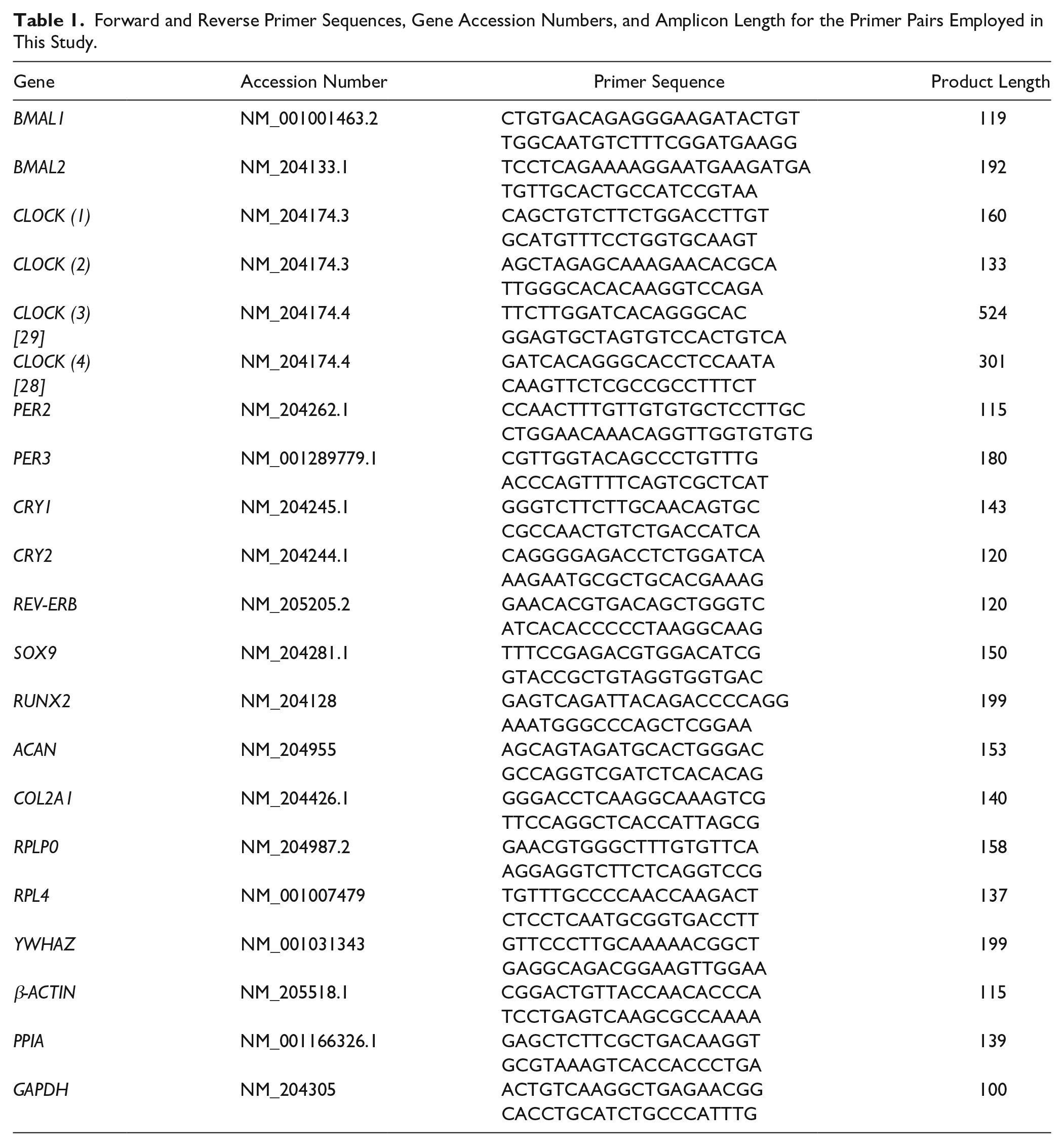
SYBR Green–based RT-qPCR reactions were set up using the GoTaq qPCR Master Mix (Promega) and 20 ng input cDNA per each 10-μL reaction. Reactions were run in a QuantStudio 3 Real-Time PCR System (Thermo Fisher Scientific) using the following standard thermal profile: activation and initial denaturation at 95°C for 2 minutes, followed by 40 cycles of denaturation at 95°C for 3 seconds, annealing and extension at 60°C for 30 seconds, and then final extension at 72°C for 20 seconds. Data were collected during the extension step. Amplification was followed by a melt curve stage consisting of 3 steps: denaturation at 95°C for 15 seconds, annealing at 55°C for 1 minute, followed by a dissociation step at 0.15°C/s increments between 55°C and 95°C. Data collection was enabled at each increment of the dissociation step. Amplification data were analyzed using the QuantStudio Design and Analysis Software (version 1.4.3) and exported data were processed using Microsoft Excel (version 15.14).
Based on previous literature, six reference genes were analyzed for stability during in vitro chondrogenesis at each time point as follows: glyceraldehyde-3-phosphate dehydrogenase (GAPDH), β-actin, peptidylprolyl isomerase A (PPIA), tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein zeta (YWHAZ), 60S acidic ribosomal protein P0 (RPLP0), and ribosomal protein L4 (RPL4). NormFinder software 27 was employed to determine the optimal normalization gene between the chosen reference genes based on their expression stability. RT-qPCR data for each gene of interest were normalized to the most stable reference gene levels in the same sample.
Interestingly, we were unable to detect transcripts for CLOCK in chondrogenic cultures, a key component of the positive limb of the molecular clock, with our custom-designed primers, despite amplifying the correct sequence in positive control samples. We were unable to amplify CLOCK in this model using published primer sequences either28,29 (see Supplemental Figures S2 and S3 in the supporting information).
Gene Expression Data Analysis and Cosine Fits
Expression data were detrended based on a 3-sample window size, to account for any noncircadian variation in expression that may occur, such as associated with proliferation and differentiation. A standard cosine function 30 was then fitted in Prism (version 7.04, GraphPad software, San Diego, CA, USA), with Period, Acrophase, Amplitude, and Mesor as free, unconstrained variables. As the only rhythmic positive transcription factor, all fits for BMAL1 were accepted. Cosine fits for transcription factors in the negative limb were accepted if the period of a fit was within 1 standard deviation of the mean. For phase angle calculations, we used the acrophase in the second (middle) cycle.
Statistical Analyses
All experiments were performed 3 times (3 biological replicates). For RT-qPCR reactions, one representative data set is shown out of 3 parallel experiments, each showing identical trends of changes. SPSS and Microsoft Excel were employed to analyze all data. For the assessment of cartilage ECM production, absorbance values are shown as means ± standard deviation (SD); statistical differences were determined using paired Student’s t test. For RT-qPCR data analysis, a Shapiro-Wilk test, skewness and kurtosis Z-values, and a visual inspection of histograms were used to determine the normality of all continuous samples. Normal probability plots (Q-Q plots) were also generated in SPSS and analyzed accordingly. Parametric tests were used for normally distributed data, while nonnormally distributed data were treated as nonparametric.
With regards to gene expression analyses, means with standard errors (SEs) and medians with interquartile ranges (IQRs) of normally and nonnormally distributed quantified genes ratios, respectively, were calculated and plotted for each time point, using Microsoft Excel. Standard errors for the ratio were calculated using the Taylor expansion of random variables equation:
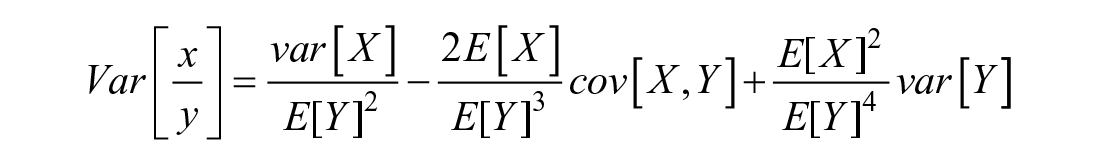
For independent variables, the covariance = 0 →
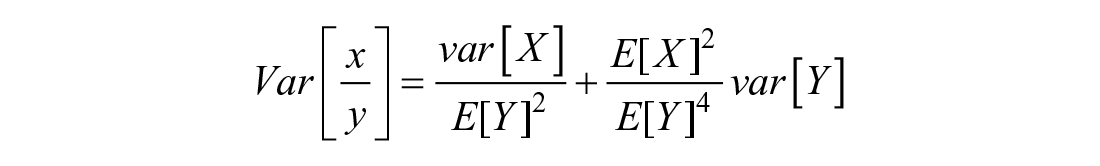
For calculating the statistical significance between time intervals, in the assumption that our paired data (quantified from the same micromass cultures) are parametric and in view of the fact that our predictor variables (time points) and dependent variables (quantified genes) of different cells are categorical and numerical continuous, respectively, a 1-way repeated-measures (within subject) analysis of variance test followed by least significant difference multiple comparisons post hoc test were used to identify specific significant differences between the means. These were considered statistically significant if P < 0.05. Statistical significance between each time point and the subsequent one is indicated by asterisks as follows: P < 0.05 = *; P < 0.01 = **; P < 0.001 = ***. The results of the statistical tests are shown in Supplemental Table S1 in the supporting information.
Results
Metachromatic Cartilage Matrix Production in Micromass Cultures Follows a Temporal Pattern
To determine the best period for studying the operation of the molecular clockwork during this dynamic system modelling in vitro chondrogenesis, we first assessed the temporal sequence of cartilage-specific ECM production in micromass cultures. The first metachromatic matrix territories around precartilage nodules produced by differentiating chondroblasts and chondrocytes were detected on day 3 as visualized by DMMB staining ( Fig. 1A ), indicating that the earliest matrix-producing cells (chondroblasts) appear in the cultures on day 3. The background-corrected absorbance of matrix-bound TB started to increase from as early as day 2 (from 0.012 ± 0.006 on day 1 to 0.071 ± 0.003 on day 2; P = 0.00001), with a further steep increase on days 5 (0.108 ± 0.006; P = 0.004) and 6 (0.141 ± 0.002; P = 0.00004) ( Fig. 1B ). Since we aimed to study the contribution of the circadian clock to early differentiating chondroblasts during the most intense phase of ECM synthesis, we concluded that the ideal time for synchronizing the molecular clock was culturing day 1, and the first time point for sample collection was at 24 hours postsynchronization (2-day-old cultures). To look at the role of the molecular clock at later stages of chondrogenic differentiation, serum shock was applied on 6-day-old cultures and the first time point for sample collection at 24 hours postsynchronization was on culturing day 7. For both early and late stages of chondrogenesis, control (nonsynchronized) cultures were exposed to medium change at the same time points as the serum-synchronized colonies.
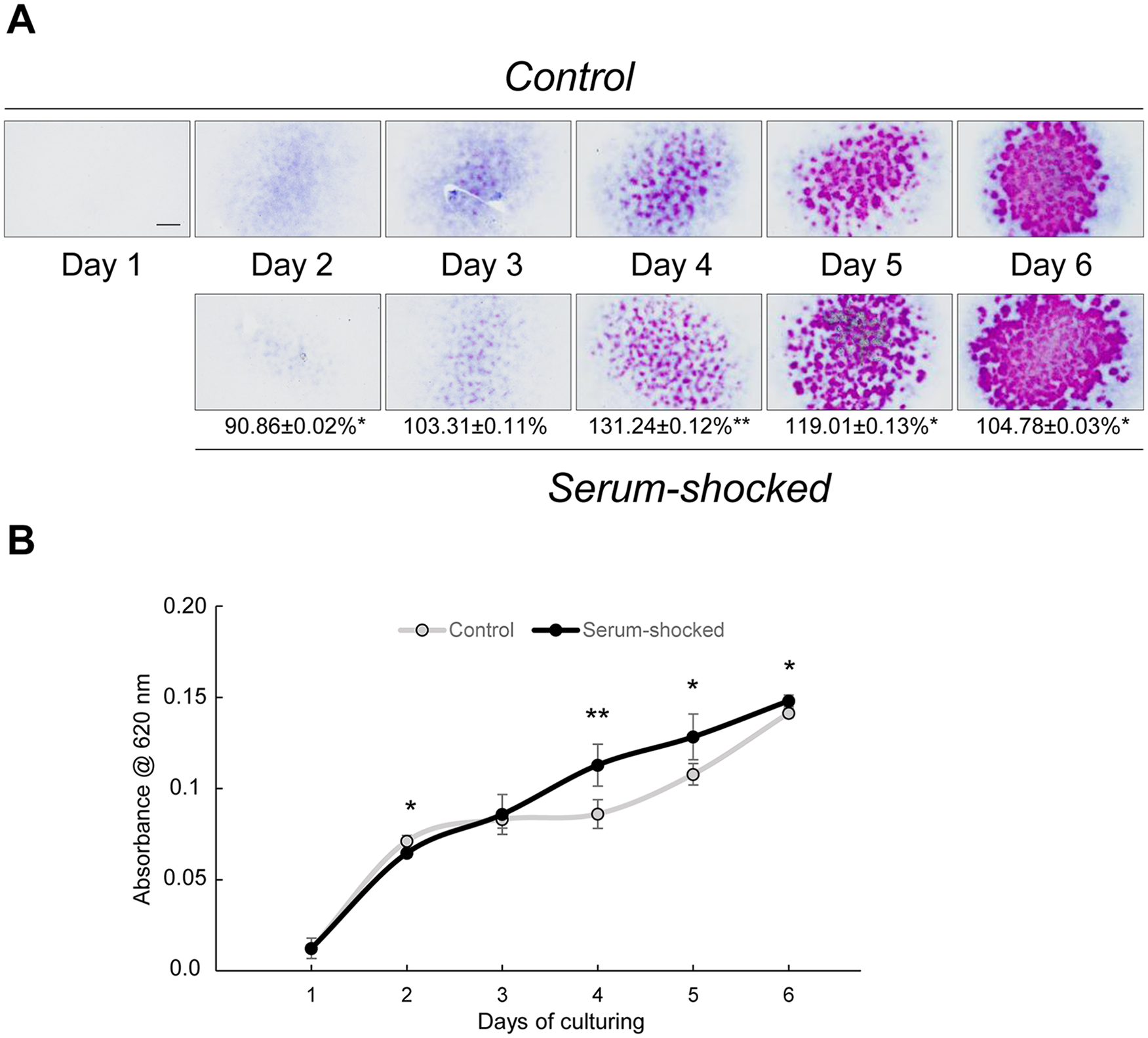
Serum shock (culturing the cells in F12 medium containing 50% fetal bovine serum [FBS] for 2 hours) on day 1 augments chondrogenesis and metachromatic matrix production in micromass cultures. (
Choosing the Optimal Reference Genes during Chondrogenesis of Serum-Synchronized Cultures
YWHAZ and RPLP0 were shown to be the most stable internal control genes during in vitro chondrogenesis on days 1 and 6, respectively, which all other genes were normalized to. For chondrogenic marker gene expression analysis on days 3 and 6, PPIA was showing the lowest stability value; therefore, SOX9, COL2A1, and ACAN expression values were normalized to PPIA expression. Supplemental Table S2 in the supporting information shows the expression stability values for the reference genes (GAPDH, β-actin, PPIA, YWHAZ, RPLP0, and RPL4) as determined using NormFinder.
Serum Shock Promotes Chondrogenesis in Micromass Cultures
Two-hour-long treatment with 50% FBS significantly augmented the amount of metachromatic cartilage ECM compared with untreated cultures starting from day 3. The largest, 1.3-fold elevation was observed on culturing day 4 (control, 0.086 ± 0.008 versus serum-shocked, 0.113 ± 0.011; P = 0.026) ( Fig. 1A and B ), when already differentiated chondroblasts and chondrocytes deposit cartilage ECM at high rates. These results indicate a strong chondrogenesis-promoting effect of the synchronizing serum-shock in chondrifying micromass cultures.
In order to confirm whether the observed increase in metachromatic matrix production was indeed attributable to a synchronized molecular clock, small molecule inhibitors of the clockwork (LDS and KL001, both applied at 5 µM for 24 hours) were added to early differentiating (day 1) colonies. Both compounds prevented the chondrogenesis-promoting effect of serum shock on metachromatic ECM production as detected on day 6 ( Fig. 2A ). Moreover, 24-hour treatment with 5 µM LDS significantly reduced the amount of cartilage ECM production (to 76.3% ± 0.28%; P = 0.04). Both compounds caused a significant decrease in mitochondrial activity by day 3, reflecting on a reduced proliferation rate in early micromass cultures on clock disruption ( Fig. 2B ).
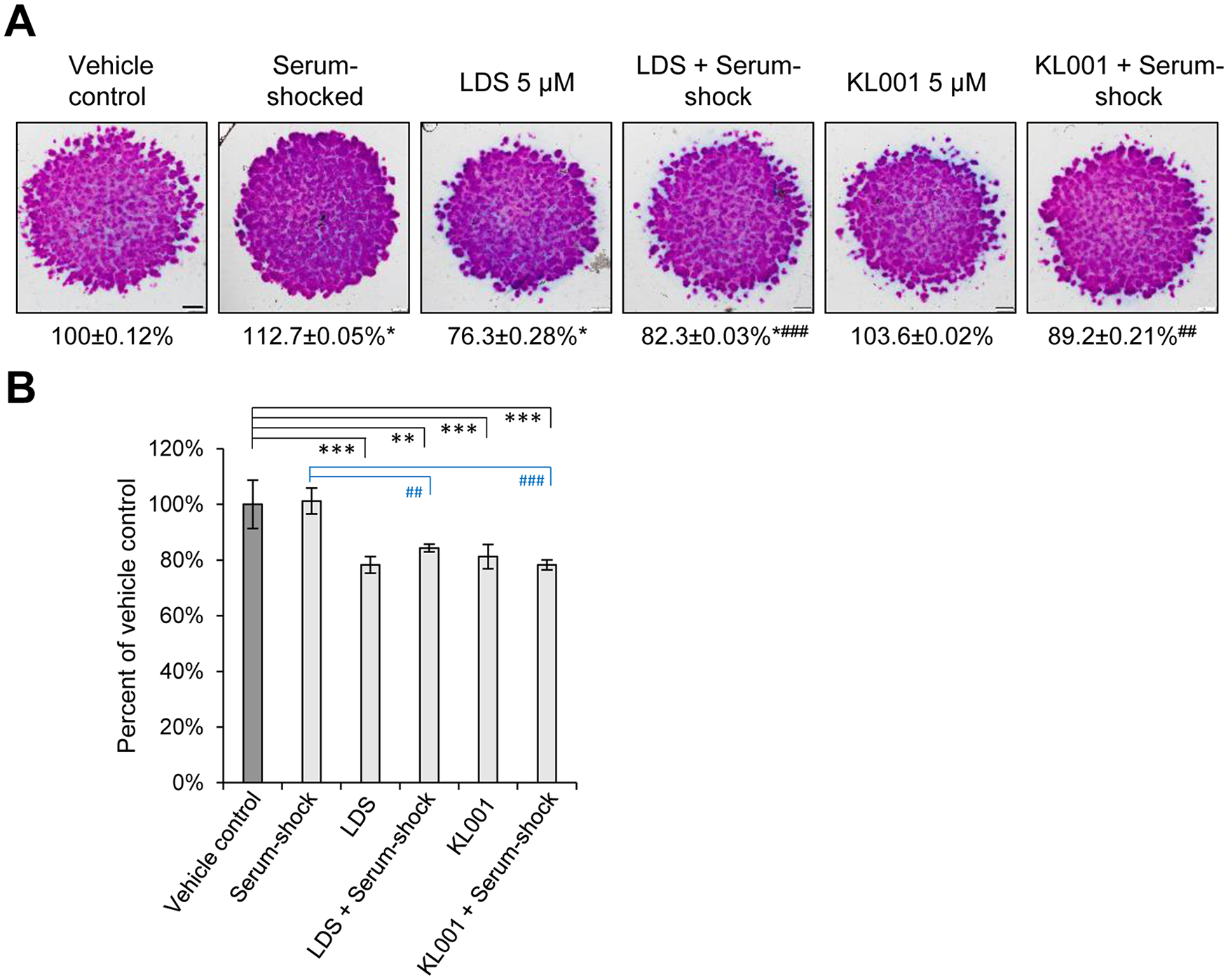
Effects of small molecule circadian clock modulators (longdaysin, LDS and KL001; both at 5 µM for 24 hours on day 1) with or without serum shock on cartilage matrix production in micromass cultures. (
To establish whether the observed changes in cartilage ECM production were detectable at the molecular level, the gene expression patterns of key chondrogenic marker genes, including SOX9, COL2A1, and aggrecan (ACAN), were tested using RT-qPCR. Two and 4 days postsynchronization (on days 3 and 6) all chondrogenic marker genes studied showed a significantly increased mRNA expression in serum-shocked cultures ( Fig. 3 ), which is in line with the augmented cartilage ECM production observed. Simultaneous treatment with either LDS and KL001 together with serum shock resulted in significantly lower gene expression levels for the chondrogenic marker genes relative to serum shock alone, on both days 3 and 6 ( Fig. 3 ).
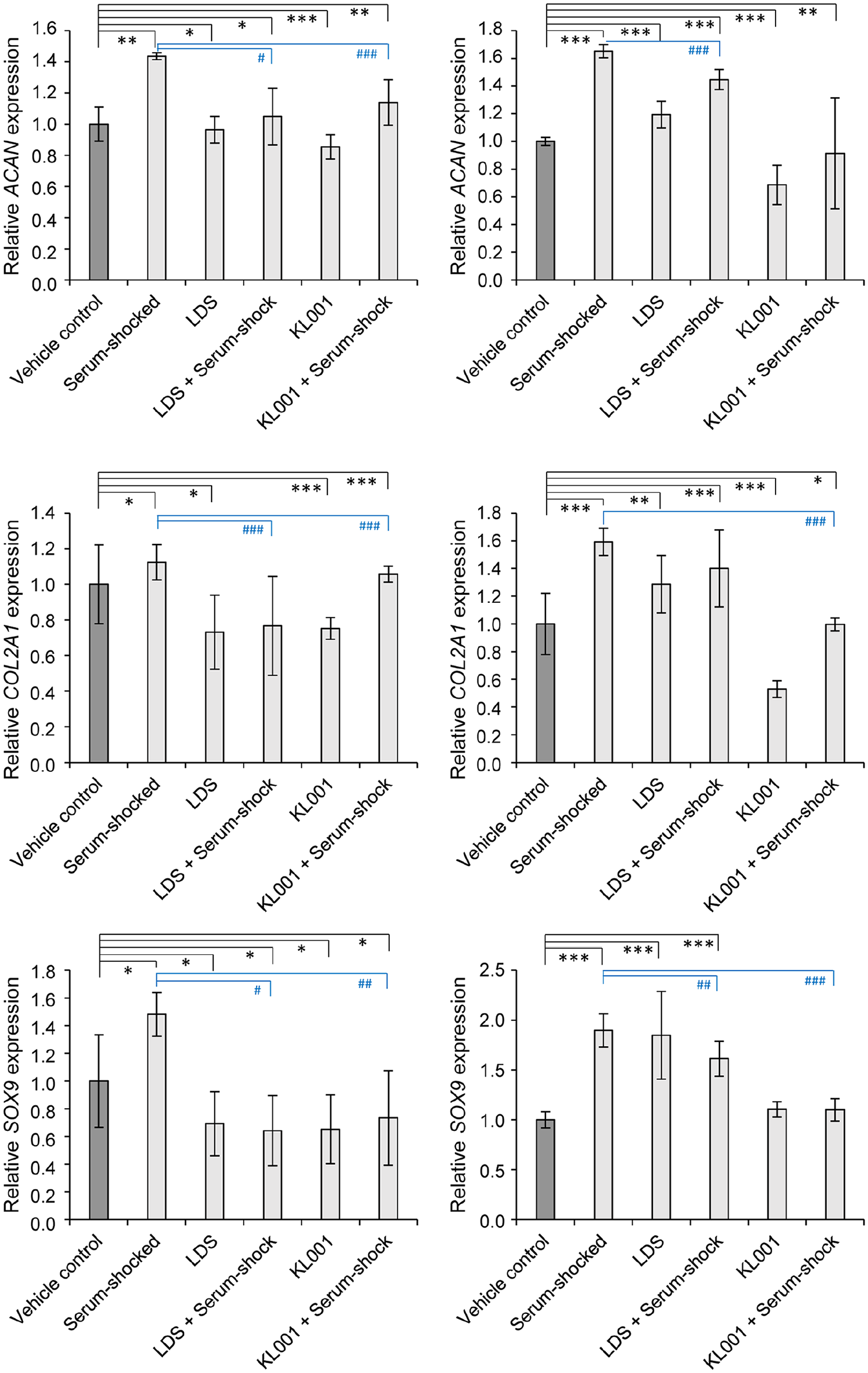
Cartilage-specific marker gene expression on days 3 and 6 following serum-synchronization on day 1, with or without treatment with the circadian clock regulators LDS or KL001. Data are expressed as the mean ± SD relative to the control and normalized against the reference gene PPIA. Statistical significance between gene expression levels is indicated by asterisks (relative to vehicle control) or hash signs (relative to serum-shocked cultures) as follows: P < 0.05 = */#; P < 0.01 = **/##; P < 0.001 = ***/###.
The Molecular Clock Genes Exhibit a Synchronized Oscillatory Pattern during Early Chondrogenesis
Transcripts for the circadian clock genes BMAL1, PER2/3, CRY1/CRY2, and REV-ERB were detected in chondrifying cells of micromass cultures, indicating that these genes were already expressed during early chondrogenesis. To establish whether the clock genes exhibited a rhythmic expression over time, 1-day-old chondrifying micromass cultures were clock-synchronized using 50% FBS for 2 hours, after which media was replaced with normal F12 medium. Samples for total RNA isolation were collected at 24 hours postsynchronization and then at every 8 hours for the period of 24 to 72 hours to identify oscillatory peaks in clock gene expression. The relative quantity (RQ) and relative SD (RSD) values for the core clock genes after being normalized to YWHAZ at each time point are shown in Supplemental Table S1 in the supporting information. As indicated, the normalized relative quantity for each gene at the 24-hour time point was set to 1.00.
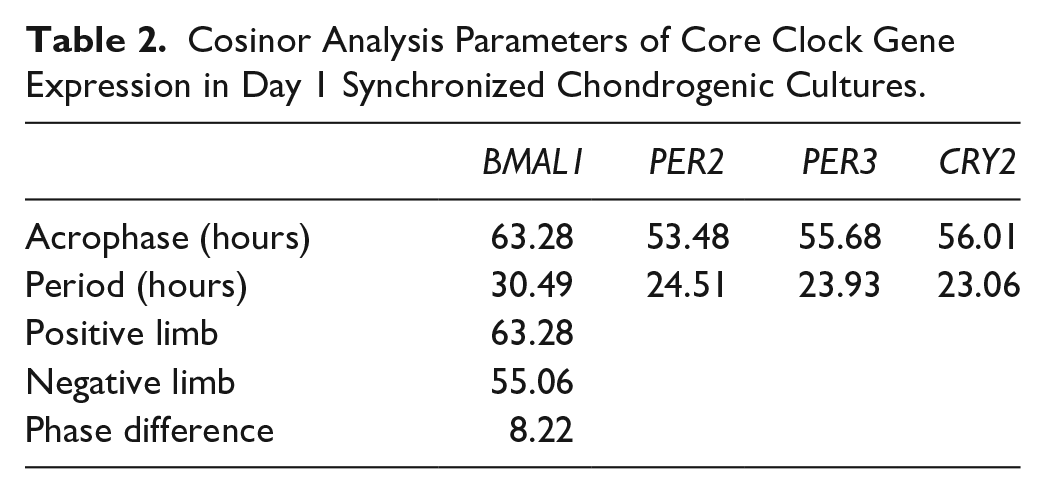
We detected a synchronized oscillatory expression pattern for some of the clock genes (BMAL1, PER2, PER3, and CRY2) over the investigated 72-hour period; the rhythmic patterns for the genes could be fitted with a nonlinear cosinor regression ( Fig. 4 ). There was an approximately 8-hour phase angle between the positive phase and negative phase of the clock (i.e., BMAL1 versus PER2/3 and CRY1). The parameters of the observed circadian expression patterns for day 1 synchronized chondrogenic cultures are shown in Table 2 . The expression patterns of the circadian clock genes after being normalized to the most stably expressed reference gene are shown in Supplemental Figure S4 in the supporting information.
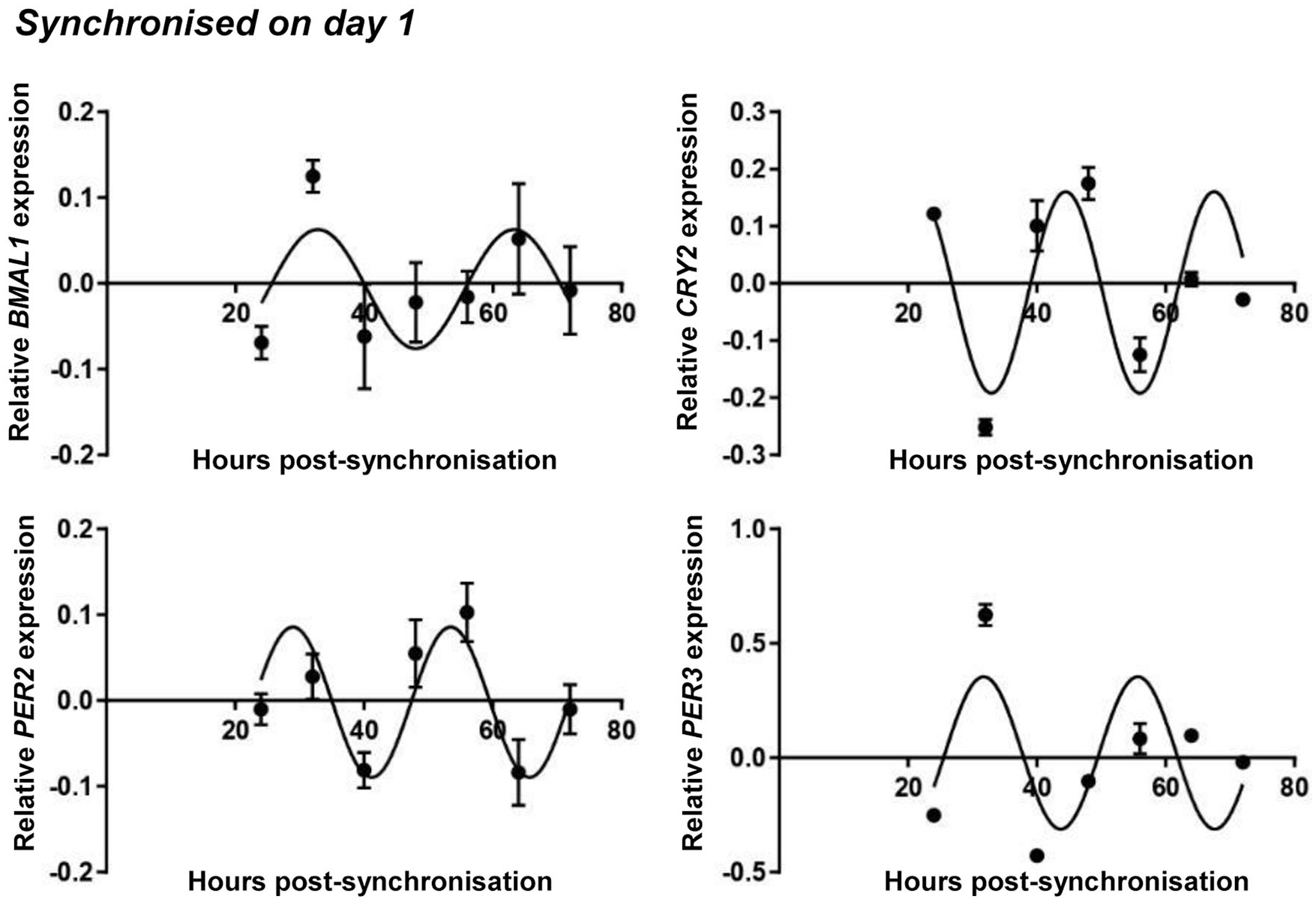
Circadian rhythm dynamics in clock gene expression in differentiating embryonic limb bud–derived chondroprogenitor cells of micromass cultures following synchronization with 50% fetal bovine serum (FBS) on culturing day 1. Quantitative RT-PCR analyses followed by cosine fits showing temporal expression profiles of the clock genes BMAL1, CRY2, PER2, and PER3 collected every 8 hours between 24 and 72 hours postsynchronization with serum shock. Data are expressed as the mean of transcript levels as determined by absolute quantification ± SD relative to the 24-hour time point and normalized against the housekeeping gene YWHAZ. Representative data are shown out of 3 independent experiments, each exhibiting similar patterns of gene expression profiles.
Cosinor Analysis Parameters of Core Clock Gene Expression in Day 1 Synchronized Chondrogenic Cultures.
Similar gene expression patterns were detected at a later stage of chondrogenesis ( Fig. 5 ). Serum shock was applied on day 6 of culturing, and samples were collected at similar time points as for day 1 cultures. The relative quantity (RQ) and relative SD (RSD) values for the core clock genes after being normalized to RPLP0 at each time point are shown in Supplemental Table S1. The normalized relative quantity for each gene at the 24-hour time point was set to 1.00. As seen in Figure 5 , not all 7 clock genes showed a circadian pattern over the investigated 72-hour period. BMAL1, CRY1, and PER2 were rhythmically expressed, and an almost anti-phasic expression (~8 hours phase angle) between BMAL1 and PER2/CRY1 was observed, indicating the operation of the molecular clockwork at the mRNA level. The cosinor parameters of the observed circadian expression patterns for day 6 synchronized chondrogenic cultures are summarized in Table 3 . The raw expressional data of the circadian clock genes after being normalized to the most stably expressed reference gene are shown in Supplemental Figure S5 in the supporting information. The normalized clock gene expression plots detected in 1- and 6-day-old nonsynchronized chondrogenic cultures are presented in Supplementary Figures S6 and S7 in the supporting information, respectively.
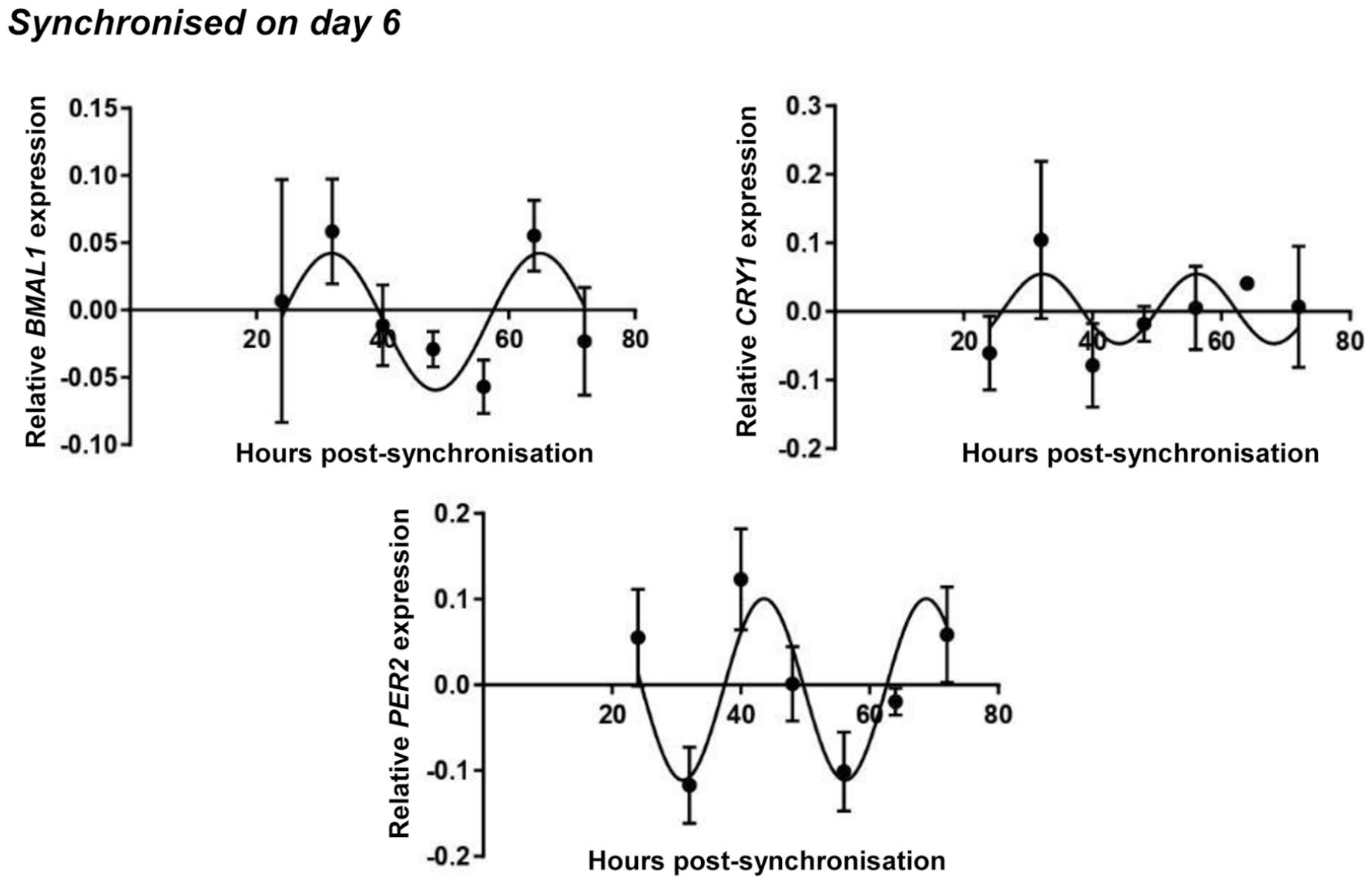
Circadian rhythm dynamics in clock gene expression in differentiating embryonic limb bud–derived chondroprogenitor cells of micromass cultures following synchronization with 50% fetal bovine serum (FBS) on culturing day 6. Quantitative RT-PCR analyses followed by cosine fits showing temporal expression profiles of the clock genes BMAL1, CRY1, and PER2 collected every 8 hours between 24 and 72 hours postsynchronization with serum shock. Data are expressed as the mean of transcript levels as determined by absolute quantification ± SD relative to the 24-hour time point and normalized against the housekeeping gene RPLP0. Representative data are shown out of 3 independent experiments, each exhibiting similar patterns of gene expression profiles.
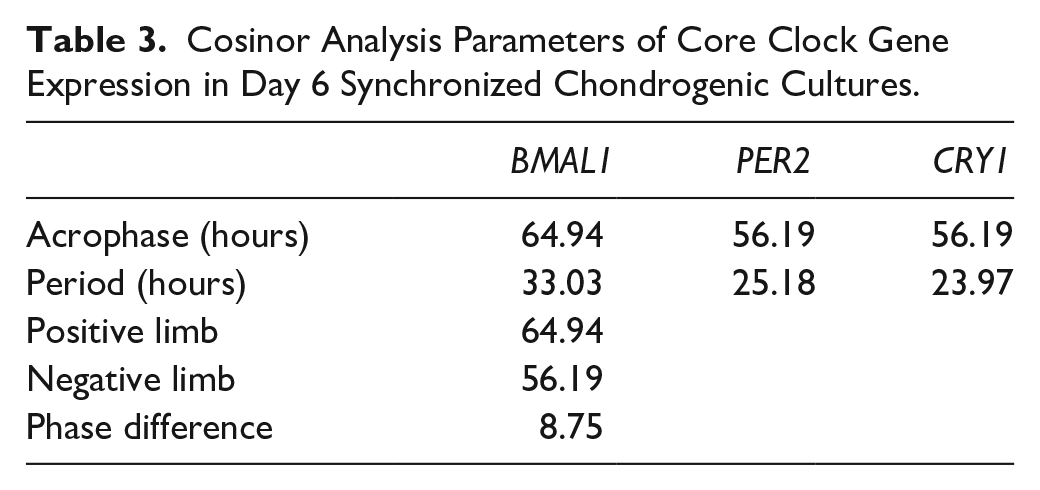
Cosinor Analysis Parameters of Core Clock Gene Expression in Day 6 Synchronized Chondrogenic Cultures.
The Chondrogenic Marker Genes Also Show a Circadian Expression Pattern During Chondrogenesis
Although embryonic limb bud–derived micromass cultures are widely accepted models of in vitro chondrogenesis, the progenitor cells represent a heterogeneous population, which also preferentially differentiate through the osteogenic lineage. This is highlighted by the concomitant expression of SOX9 and RUNX2, key chondro- and osteogenic transcription factors, respectively. 31 In order to establish whether their expression also showed an oscillatory pattern, SOX9 and RUNX2 transcript levels were analyzed by RT-qPCR at the previously indicated time points, and then the expression values were fitted with the cosine function to reveal circadian patterns. The cosinor parameters of the observed circadian expression patterns for the marker genes in chondrogenic cultures are summarized in Table 4.
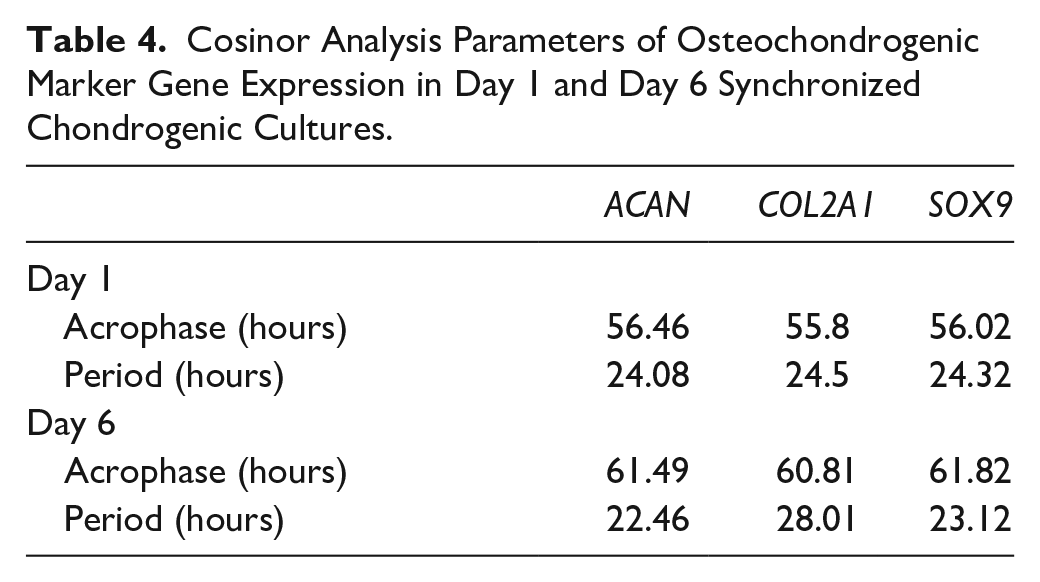
Cosinor Analysis Parameters of Osteochondrogenic Marker Gene Expression in Day 1 and Day 6 Synchronized Chondrogenic Cultures.
In day 1 synchronized micromass cultures, the acrophase for SOX9 was 56 hours, with a ~7-hour phase difference to BMAL1 ( Fig. 6 ). In contrast, no circadian pattern could be detected for RUNX2 (Supplemental Figure S8 in the supporting information). In day 6 synchronized cultures, SOX9 transcript levels were also showing a circadian pattern as revealed by the cosine fit and were largely phasic with those of BMAL1 ( Fig. 7 ). RUNX2 mRNA levels showed a downward trend, rather than an oscillatory pattern, in 6-day-old cultures (Supplemental Figure S9 in the supporting information). These findings indicate that the 2 transcription factors SOX9 and RUNX2 may be differentially influenced by the molecular clock on days 1 and 6.
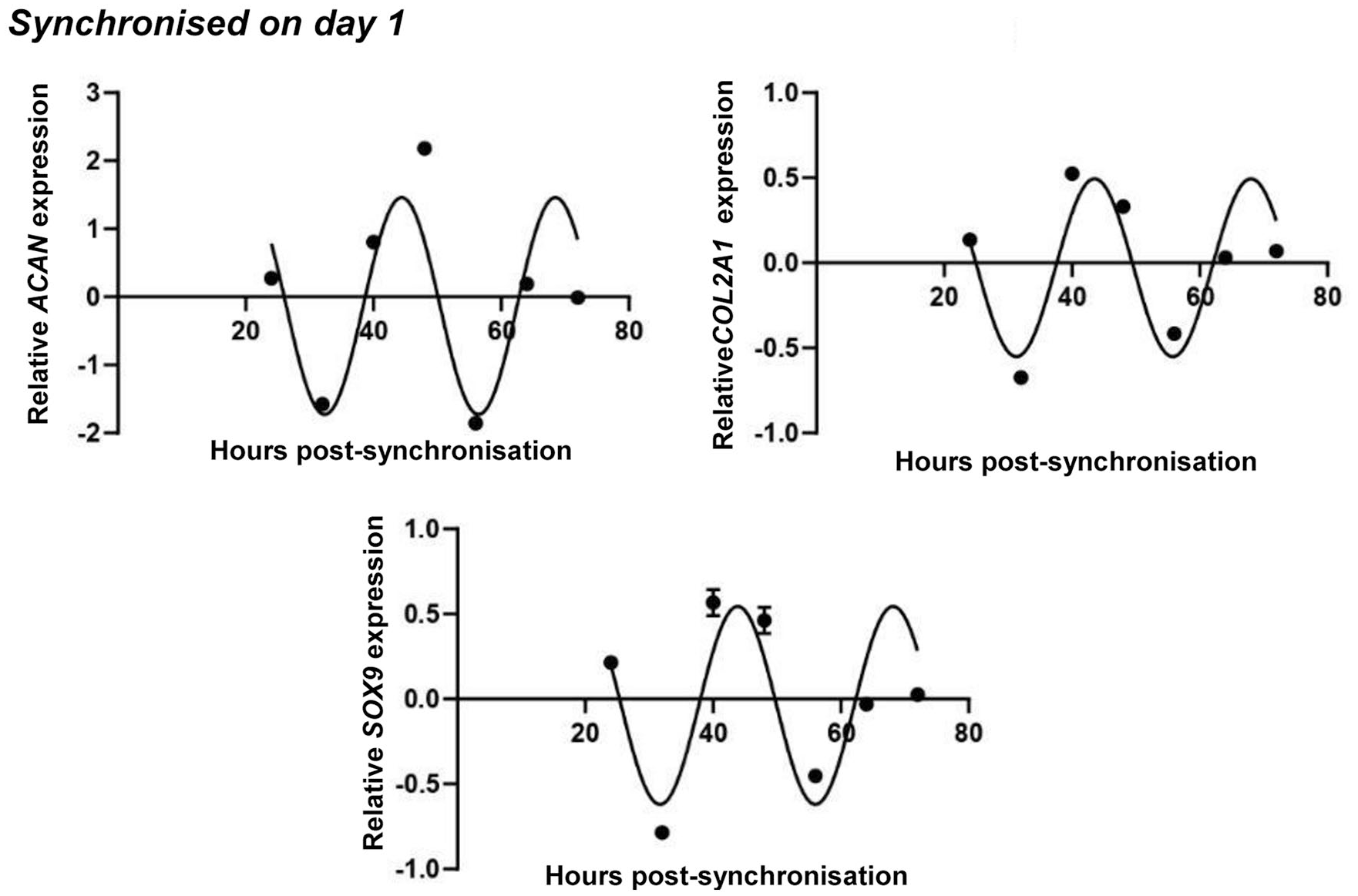
Circadian rhythm dynamics in SOX9 transcript levels and the expression of genes coding for cartilage extracellular matrix (ECM) constituents (ACAN and COL2A1) in differentiating embryonic limb bud–derived chondroprogenitor cells of micromass cultures following synchronization with 50% fetal bovine serum (FBS) on culturing day 1. Quantitative RT-PCR analyses followed by cosine fits showing temporal expression profiles of clock genes collected every 8 hours between 24 and 72 hours postsynchronization with serum shock. Data are expressed as the mean of transcript levels as determined by absolute quantification ± SD relative to the 24-hour time point and normalized against the housekeeping gene YWHAZ. Representative data are shown out of 3 independent experiments, each exhibiting similar patterns of gene expression profiles.
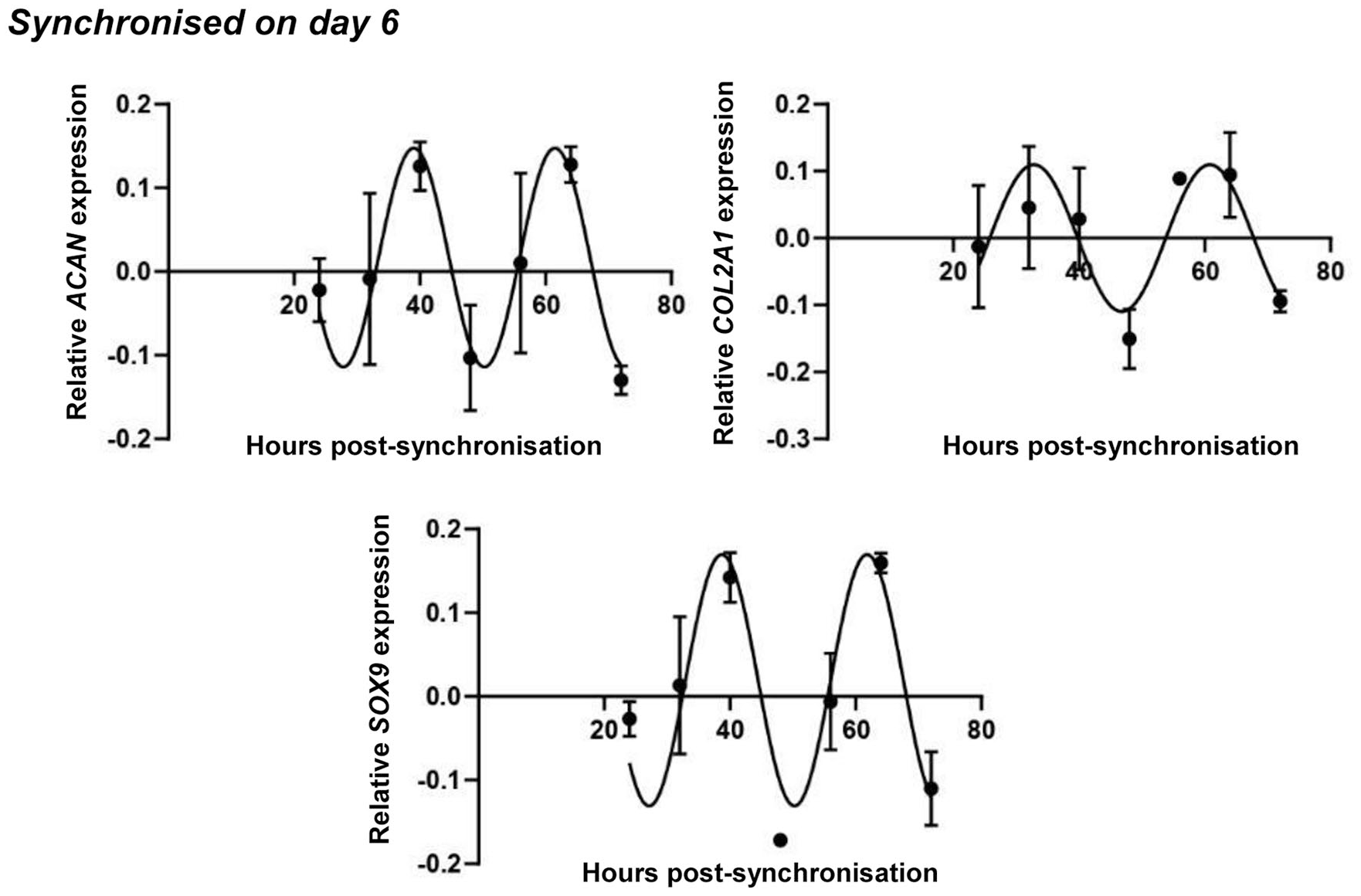
Circadian rhythm dynamics in SOX9 transcript levels and the expression of genes coding for cartilage extracellular matrix (ECM) constituents (ACAN and COL2A1) in differentiating embryonic limb bud–derived chondroprogenitor cells of micromass cultures following synchronization with 50% fetal bovine serum (FBS) on culturing day 6. Quantitative RT-PCR analyses followed by cosine fits showing temporal expression profiles of clock genes collected every 8 hours between 24 and 72 hours postsynchronization with serum shock. Data are expressed as the mean of transcript levels as determined by absolute quantification ± SD relative to the 24-hour time point and normalized against the housekeeping gene RPLP0. Representative data are shown out of 3 independent experiments, each exhibiting similar patterns of gene expression profiles.
The expression pattern of ACAN, which codes for the core protein of the proteoglycan aggrecan, followed a similar circadian rhythm to what has previously been seen for SOX9 in cultures synchronized on both days 1 and 6, respectively ( Figs. 6 and 7 ). COL2A1, coding for the alpha chain of collagen type 2, exhibited a circadian trend very similar to those of SOX9 and ACAN in both young and mature cultures ( Figs. 6 and 7 ). The raw expressional data of the osteo-chondrogenic marker genes after being normalized to the most stably expressed reference gene are shown in Supplemental Figures S8 and S9 in the supporting information, along with osteochondrogenic master gene expression plots detected in nonsynchronized chondrogenic cultures in 1- and 6-day-old colonies, respectively. The data suggest a circadian regulation of chondrogenic marker genes in micromass cultures during chondrogenesis.
Discussion
Given the role of circadian rhythms in maintaining normal cartilage tissue homeostasis and the fact that clock disruption in chondrocytes may lead to an osteoarthritic phenotype, 9 the purpose of this study was to identify circadian oscillations in the expression patterns of the core clock genes in a well-established chondrogenic model, and to determine whether a synchronized clock may contribute to augmented in vitro chondrogenesis. Whilst the presence of a cell-autonomous peripheral clock in mature chondrocytes has increasingly been recognized, to our knowledge, the current study is the first to demonstrate the synchronized operation of a core clock during the early stages of in vitro chondrogenesis. More important, this is the first study to demonstrate circadian expression patterns for major chondrogenic marker genes (SOX9, ACAN, and COL2A1) during the early and late stages of in vitro cartilage formation.
It is widely accepted that the molecular clockwork is maintained in cells cultured ex vivo. 32 However, since immortalized cell lines often have reduced tissue-specific functions and a disturbed phenotype, we have chosen to use primary micromass cultures established from cells isolated directly from chicken embryos, a well-established and reliable model for in vitro chondrogenesis. 33 Within the 6-day-long culturing period, the chondroprogenitor cells in micromass cultures differentiate into cartilage ECM-producing chondroblasts and start producing cartilage-specific ECM on culturing days 2 to 3. 34 In order to investigate whether a synchronized pattern in clock gene expression is detectable during early and late stages of in vitro chondrogenesis, we have chosen to synchronize the molecular clock with serum shock on culturing days 1 and 6.
The results obtained in cultures synchronized on day 1 indicate that chondroprogenitor mesenchymal cells differentiating to early chondroblasts in chicken micromass cultures readily express all core circadian clock genes during early chondrogenesis, and their expression followed a synchronized, rhythmic pattern. We detected an ~8-hour phase angle difference between the positive limb (BMAL1) and the negative limb (CRY2/PER2 and PER3) of the clock. Adult mesenchymal stem cells (MSC) such as bone marrow–derived or adipose-derived MSC, are also capable of circadian synchronization by hormonal and growth factor signals such as serum shock. 35 In contrast, there is a lack of a functional circadian clock in undifferentiated embryonic stem (ES) cells, despite the expression of core clock genes. 6 However, on differentiation into cardiomyocytes, ES cells in culture gained a molecular circadian rhythm. 6 Given that the primary cells used to establish micromass cultures were isolated from young (4-day-old) chicken embryos, it is plausible that the circadian molecular clock in those cells already resembles the developmental stage that characterizes MSCs and is already functional.
In more mature (6-day-old) serum-synchronized micromass cultures, a similar pattern was observed. While not all 7 clock genes showed a sinusoid pattern over the investigated 72-hour period, an expression pattern with an approximately 8-hour phase angle between BMAL1 and PER2/CRY1 was seen, indicating a functional molecular clockwork, at least at the mRNA level. These results show that more mature, ECM-producing chondroblasts and early chondrocytes are also characterized by a functional clock, which has been reported in mature chondrocytes cultured in vitro. 15 The periodicity of the circadian rhythm dynamics in micromass cultures was comparable to what has been reported in MSCs35,36 and in chondrocytes. 37
A key finding of our study is that a synchronized, rhythmic expression pattern was characteristic to the major chondrogenic transcription factor SOX9, as well as key genes coding for cartilage ECM constituent proteins and proteoglycans (COL2A1 and ACAN) following serum shock. To our knowledge, this is the first study to report such a rhythmic expression pattern for these gene products during early chondrogenesis. The pluripotency factor SOX2 has been shown to exhibit an oscillatory pattern in serum or dexamethasone-synchronized MSCs. 35
In addition, serum-synchronized micromass cultures appeared to produce more abundant cartilage ECM by day 6, and this was detectable at the mRNA expression level of the cartilage-specific marker genes SOX9, ACAN, and COL2A1. The observed chondrogenesis-promoting effect of the serum shock in early chondrogenic micromass cultures is likely attributable to its synchronizing effect on the molecular clockwork, as co-application of small molecule modulators (LDS and KL001) abolished the stimulating effects on ECM production and chondrogenic marker gene expression. Disruption of the molecular clock in early chondrogenic cultures likely has a negative effect on cell proliferation, as we detected significantly lower cellular viability following 24-hour-long treatment with LDS or KL001 using MTT assay. Although revealing the molecular mechanism of enhanced chondrogenesis followed by serum shock was beyond the scope of this study, it is plausible to hypothesize that the downward trend in transcript levels and the lack of a circadian expression profile for RUNX2 observed in day 1 and day 6 serum-shocked cultures could at least partially be accounted for it. Attenuated RUNX2 transcript levels may indicate preferential differentiation toward the chondrogenic, rather than the osteogenic lineage,31,38 a key observation which could be taken into account in future attempts at cell-based articular cartilage regeneration techniques.
Conclusions
Our results reveal that following serum shock, embryonic limb bud–derived chondrifying micromass cultures exhibit synchronized temporal expression patterns of core clock genes and stabilizing loop genes involved in the molecular circadian clock. We also observed that key chondrogenic marker genes followed a diurnal oscillatory pattern. Furthermore, cultures with a synchronized molecular clock produced more abundant cartilage ECM by day 6, which was abolished by clock disruption with the small molecule inhibitors LDS or KL001. Results from this study suggest that a functional molecular clockwork plays a positive role not only in the tissue homeostasis of mature articular cartilage but also during chondrogenesis. Since the circadian clock has extensively been shown to play a key regulatory role in various mature tissues, including those of the musculoskeletal system, 10 our findings on the role of the molecular clock in differentiating chondroprogenitor cells has key implications. What internal or external “zeitgeber” mechanisms exist in developing cartilage that would initiate and then entrain cell-autonomous clocks are still elusive. However, identifying the optimal synchronization methods relevant to chondroprogenitor cells or early chondroblasts may offer insightful ways by which the osteochondroprogenitor stem cells could be preferentially directed toward the chondrogenic lineage, thus generating high-quality cartilage for the desired tissue engineering applications.
Supplemental Material
revised-supplementary-material – Supplemental material for A Synchronized Circadian Clock Enhances Early Chondrogenesis
Supplemental material, revised-supplementary-material for A Synchronized Circadian Clock Enhances Early Chondrogenesis by M. Abdulhadi Alagha, Judit Vágó, Éva Katona, Roland Takács, Daan van der Veen, Róza Zákány and Csaba Matta in CARTILAGE
Footnotes
Supplemental Material
Acknowledgments and Funding
The authors are greatly indebted to Mrs Krisztina Biróné Barna for skillful technical assistance. The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: CM is supported by the Premium Postdoctoral Research Fellowship by the Hungarian Academy of Sciences, and a Bridging Fund from the Faculty of Medicine, University of Debrecen. The research was financed by the Thematic Excellence Programme of the Ministry for Innovation and Technology in Hungary, within the framework of the Space Sciences thematic program of the University of Debrecen (ED 18-1-2019-0028).
Author Contributions
MAA, JV, EK, RT, DvdV, RZ, and CM have made substantial intellectual contributions to the conception and design of the study, data acquisition, analysis and interpretation. MAA, VJ, EK, and CM carried out the bulk of the experimental work and contributed to data collection, interpretation and analysis. DvdV, RZ, and CM conceived the study design and coordinated the experimental work. MAA, VJ, EK, and CM were in charge of data acquisition and preliminary analysis. MAA and DvdV performed statistical data analyses. MAA, JV, EK, RT, DvdV, RZ, and CM contributed to data interpretation, and drafting the article or revising it critically for important intellectual content. MAA, JV, EK, RT, DvdV, RZ, and CM have approved the final version submitted.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Work on cells derived from early stage (4-day-old) chick embryos does not require an approval from the Ethics Committee of the University of Debrecen.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
