Abstract
The current model predicts that MDM2 is primarily overexpressed in cancers with wild-type (WT) p53 and contributes to oncogenesis by degrading p53. Following a correlated expression of MDM2 and NF-κB2 transcripts in human lung tumors, we have identified a novel transactivation function of MDM2. Here, we report that in human lung tumors, overexpression of MDM2 was found in approximately 30% of cases irrespective of their p53 status, and expression of MDM2 and NF-κB2 transcripts showed a highly significant statistical correlation in tumors with WT p53. We investigated the significance of this correlated expression in terms of mechanism and biological function. Increase in MDM2 expression from its own promoter in transgenic mice remarkably enhanced expression of NF-κB2 compared with its non-transgenic littermates. Knockdown or elimination of endogenous MDM2 expression in cultured non-transformed or lung tumor cells drastically reduced expression of NF-κB2 transcripts, suggesting a normal physiological role of MDM2 in regulating NF-κB2 transcription. MDM2 could up-regulate expression of NF-κB2 transcripts when its p53-interaction domain was blocked with Nutlin-3, indicating that the MDM2-p53 interaction is dispensable for up-regulation of NF-κB2 expression. Consistently, analysis of functional domains of MDM2 indicated that although the p53-interaction domain of MDM2 contributes to the up-regulation of the NFκB2 promoter, MDM2 does not require direct interactions with p53 for this function. Accordingly, MDM2 overexpression in non-transformed or lung cancer cells devoid of p53 also generated a significant increase in the expression of NF-κB2 transcript and its targets CXCL-1 and CXCL-10, whereas elimination of MDM2 expression had the opposite effects. MDM2-mediated increase in p100/NF-κB2 expression reduced cell death mediated by paclitaxel. Furthermore, knockdown of NF-κB2 expression retarded cell proliferation. Based on these data, we propose that MDM2-mediated NF-κB2 up-regulation is a combined effect of p53-dependent and independent mechanisms and that it confers a survival advantage to lung cancer cells.
Introduction
The human homologue of the mouse double minute-2 (mdm2) gene is frequently overexpressed in many human cancers and has been predicted to be a prognostic marker for breast cancer.1-3 Amplification or overexpression of the mdm2 gene enhances the tumorigenic potential of murine cells,4,5 suggesting MDM2 has a dominant oncogenic function. MDM2 recognizes the transactivation domain of p53 and inactivates p53-mediated transcriptional activation (reviewed in references 2, 3, 6-14). Work from our laboratory showed that the interaction of MDM2 with p53 is necessary for inhibition of p53-mediated transactivation.9,11 MDM2 degrades p53 by targeting it for ubiquitination (reviewed in references 2, 3, 6, 7). These properties of MDM2 predict that MDM2 contributes to oncogenesis by inactivating p53, and this possibility has been studied widely (reviewed in references 2, 15). MDM2 overexpression has also been reported in cancers with mutant p53, and several laboratories have reported p53-independent functions of MDM2. In recent years, a growing body of literature has documented p53-independent oncogenic function of MDM2 (reviewed in references 2, 3, 6, 7, 15-17).
Both WT and mutant p53 can modulate transcription, and MDM2 interferes with this activity.9,11,18 Therefore, we sought to determine altered gene expression that co-occurs with MDM2 overexpression in human lung cancer that may explain induction of oncogenesis and enable us to identify a biochemically linked group of markers that could be used as markers for treatment of cancer. The role of MDM2 in transcription has not been investigated in depth. Particularly, not investigated is its functional role using its transcriptional activity. Although not a general regulator of transcription, MDM2 has been shown to regulate transcription when recruited to a promoter. 19 MDM2 represses the telomerase RNA gene promoter and induces NF-κB p65 expression at the level of transcription through its interaction with the retinoblastoma gene product (Rb) and the transcription factor Sp1.20,21 MDM2 interacts and mono-ubiquitinates histone H2B, and this function of MDM2 has been related to its ability to inhibit gene expression. 22 MDM2 overexpression in cancer cells, therefore, could be associated with alteration of gene expression either because of its direct effect on transcription or through degradation of WT or mutant p53. Using this hypothesis, we investigated the levels of MDM2 expression, status of p53 mutation, and p53- or MDM2-regulated gene expression in human lung cancer tumors. Our data revealed a novel transcription regulatory function of MDM2 due to which MDM2 overexpression in human lung cancers up-regulates expression of the transcription factor p100/NF-κB2 and thus may enhance cell survival and proliferation.
Nuclear factor of κB (NF-κB) is a collection of dimeric transcription factors that control diverse biological processes. One of these members, NF-κB2/p52, is generated from the precursor proteins p100. In contrast to the canonical pathway that involves NF-κB1 (p50/p105), the protein precursor of NF-κB2 (p100) is activated by selective degradation at its C-terminus at the posttranslational level (reviewed by Xiao et al. 23 ). Mechanisms that regulate cross talk between p53 and NFκB family members, and how deregulation of these interactions contributes to oncogenesis, have been well-studied (reviewed by Schneider & Kramer 24 ).
Previous reports have suggested that MDM2 can up-regulate NF-κB/p65 expression. 20 We reported earlier that tumor-derived p53 mutants up-regulate expression of p100/NF-κB2. 25 Following a correlated expression of MDM2 and NF-κB2 transcripts in human lung tumors, here we report that MDM2 is capable of up-regulating expression of p100/NF-κB2 and its target gene CXCL-1 and 10. MDM2-mediated up-regulation of NF-κB2 expression contributes to its ability to induce cell proliferation and to resist cell death by the chemotherapeutic drug paclitaxel.
Results
MDM2 overexpression and p53 mutation are frequent events in human lung cancer
We identified single base-pair substitution mutation of p53 in 12 of 30 human lung cancer samples using a method described by Sjogren et al. 26 Approximately 30% of the samples showed 5-fold higher than normal levels of MDM2 irrespective of their p53 status (data not shown), indicating that incidence of MDM2 overexpression and p53 mutation in human lung cancer are not mutually exclusive.
MDM2 overexpression correlates with expression of p100/NF-κB2 transcript
Since mutant p53 can up-regulate p100/NF-κB2 25 and MDM2 can degrade p53, we investigated NF-κB2 expression in human tumor samples overexpressing MDM2. Relationships between the MDM2 (independent variable) and NF-κB2 p100 expression levels (dependent variables) were determined using separate linear regression models. The relationship between MDM2 and NF-κB2 in ungrouped cancer samples showed marginal statistical significance (P < 0.0895). Since mutant p53 up-regulates NFκB2 expression, 25 we reasoned that the tumor samples harboring mutant p53 would express higher levels of NFκB2 transcript and that this expression pattern may mask the correlation between MDM2 and NFκB2 expression. Indeed, samples harboring mutant p53 showed elevated levels of NF-κB2 transcripts compared with samples with WT p53 (Fig. 1A). However, the extent of increase varied with the type of p53 mutation, suggesting that the transcriptional activation property of some of the p53 mutants is stronger than others.
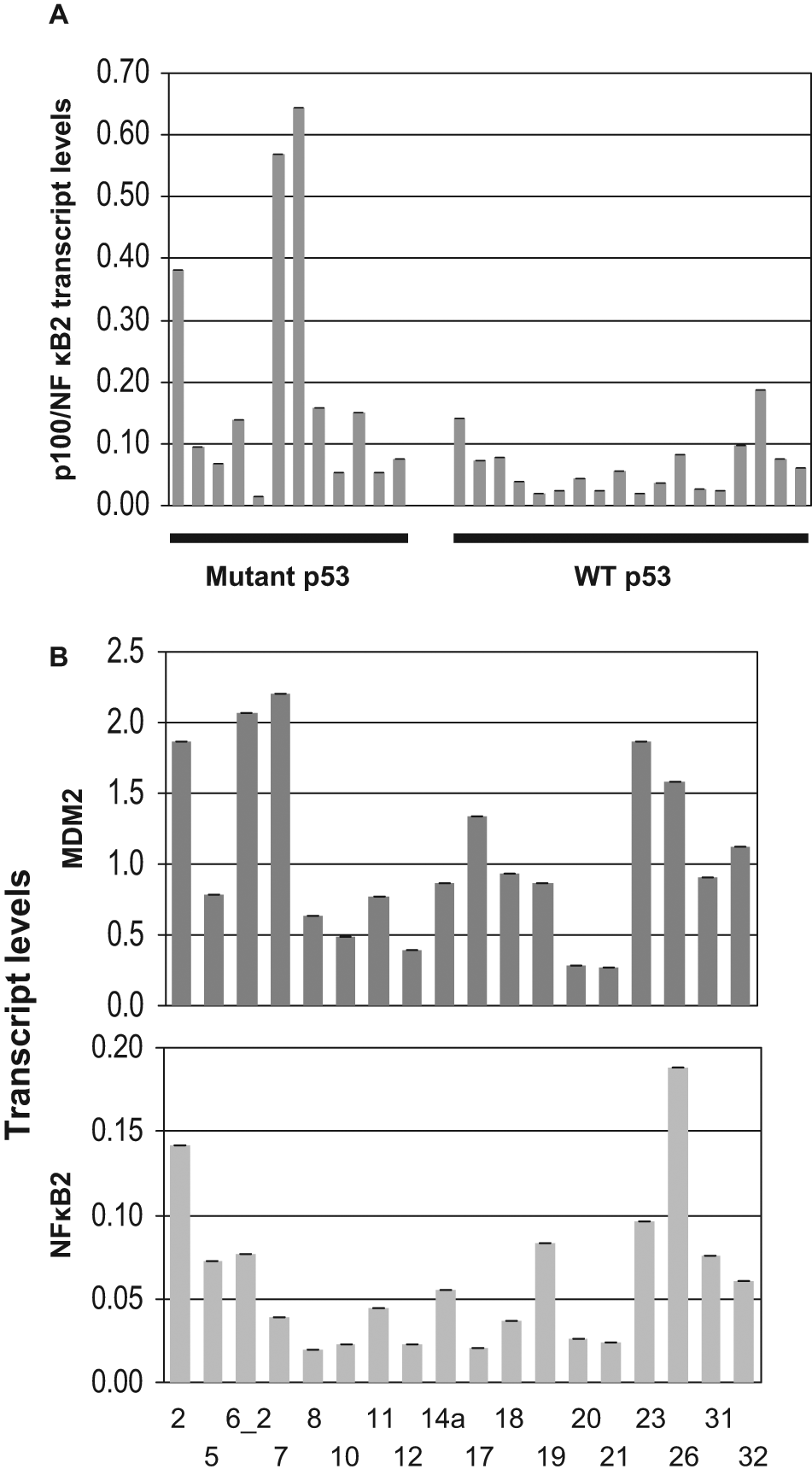
p100/NF-κ B2 expression in human lung tumors with WT and mutant p53 (A) and correlation of MDM2 and p100/NF-κB2 expression in human lung tumor samples without p53 mutation (B). Transcript levels were determined using QPCR of cDNA prepared from RNA from tumor samples and were normalized with GAPDH transcript levels. Methods used for generation of cDNA and QPCR and determination of p53 mutation are described in Materials and Methods. Expression of MDM2 and NF-κB2 in tumor samples without p53 mutations was found to be statistically significant (P < 0.001).
Therefore, we examined the relationship between MDM2 and NFκB2 by grouping the lung cancer samples into WT and mutant p53. Relationships between the MDM2 (independent variable) and NF-κB2 p100 expression levels (dependent variables) in WT or mutant p53 groups were determined using separate linear regression models. The relationship between MDM2 and NFκB2 expression in the WT p53 group was found to be highly significant (P < 0.001, Fig. 1B). This observation suggests that MDM2 may up-regulate NF-κB2 expression.
Enhanced expression of MDM2 in transgenic mice increases NF-κB2 expression
To determine whether MDM2 can elevate NF-κB2 protein expression, we generated MDM2 transgenic mice with p53+/+ and p53−/− background. QPCR analysis of genomic DNA isolated from these cells showed a 10-fold higher copy number of MDM2 gene than littermate non-transgenic mice. Lung fibroblast cells from these mice were isolated and cultured for a short period following a method described earlier. 27 The levels of MDM2 and NF-κB2 were determined by Western blot analysis of cell lysate. Densitometric analysis indicated that MDM2 overexpression elevates p100, and p52 expression approximately 12- and 6-fold respectively (Fig. 2A, B) for p53+/+ background.
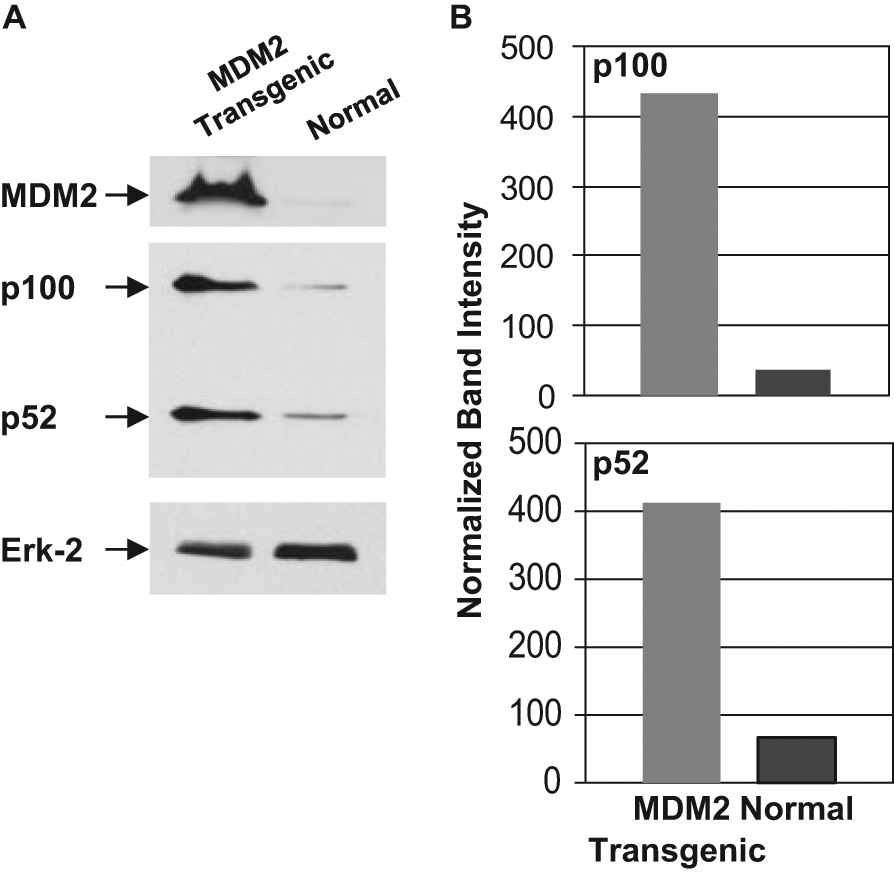
Enhanced expression of MDM2 in transgenic mice increases NF-κB2 expression. Western blot analysis (A) of extracts prepared from lung cells derived from normal and MDM2 transgenic mice (p53+/+ background) for expression of MDM2, NF-κB2 p100, p52, and a loading control Erk-2 were identified using respective antibodies. Migrations of the MDM2, p100, p52, and Erk2 bands are shown by arrows. Densitometric analyses (B) of p100 and p52 expression are also shown (B). Band intensities were normalized to levels of Erk-2.
Reduction of endogenous MDM2 expression with shRNA against MDM2 knocks down expression of p100/NF-κB2 transcript
Since enhanced expression of MDM2 transcripts correlates with an increased expression of p100/NF-κB2 transcripts in human lung tumor samples and NF-κB2 expression could be elevated by increasing copy number of MDM2 in transgenic mice (Fig. 2), we investigated whether knockdown of endogenous MDM2 in lung cancer H460 or lung fibroblast WI38 cells could down-regulate expression of p100/NF-κB2 transcripts. H460 and WI38 cells, both of which harbor WT p53, 28 were infected with lentiviral vectors expressing shRNA against MDM2 (shMDM2-1 and shMDM-2) or a non-endogenous luciferase gene (shLuc) as described earlier, 29 and the levels of MDM2 and p100/NF-κB2 transcripts were determined by QPCR. As shown in Fig. 3, MDM2 shRNA expression vectors reduced the expression of MDM2 and p100/NF-κB2 transcript in both H460 (Fig. 3A, 3B) and WI38 (Fig. 3C, 3D) cells in repeated experiments. These data suggest that endogenously expressed MDM2 up-regulates the expression of p100/NF-κB2. Since knockdown of MDM2 should stabilize p53, and both WI38 and H460 cells harbor WT p53, it is possible that the observed decrease in the NF-κB2 transcript level after knockdown of MDM2 in Fig. 3 is a result of accumulation of p53 and consequent p53-mediated repression of NF-κB2 expression. 25 To exclude this possibility, we investigated whether knockdown of both p53 and MDM2 would down-regulate expression of p100/NF-κB2 transcripts. Therefore, we infected H460 cells with 2 lentiviral vectors, one expressing shRNA against MDM2 and the other expressing shRNA against p53. 30 The control cells were infected with a non-endogenous luciferase gene (shLuc) as described earlier. 29 The cells were selected with appropriate antibiotics encoded by the viral vectors, and the levels of MDM2 and p53 transcripts in the pooled colonies were determined by QPCR to ensure knockdown of the two proteins (Fig. 3E and F). Western blot analysis of extracts prepared from the pooled colonies showed a drastic reduction in the levels of NF-κB2/p100 and p52 (Fig. 3G). These data show that endogenously expressed MDM2 up-regulates the expression of p100/NF-κB2.
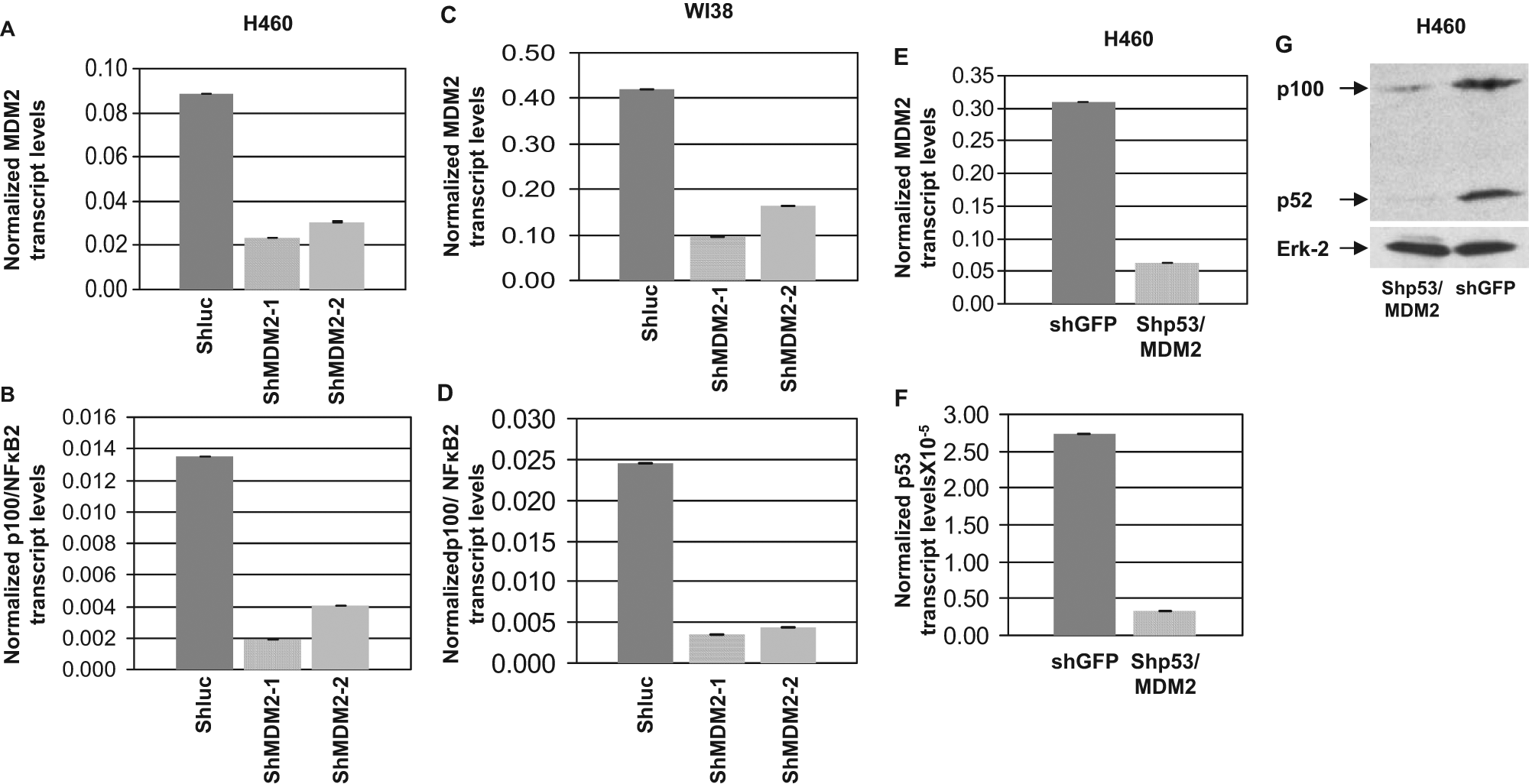
Reduction of MDM2 expression using lentiviral vectors expressing MDM2-specific shRNA lowers p100/NFκB2 expression. RNA and cDNA was prepared from H460 (A, B, E, F) and WI38 (C, D) cells infected with lentiviruses expressing shRNA against either the luciferase gene (shLuc) as negative control or two shRNAs directed against MDM2 (shMDM2-1 or -2), or against p53 (shp53) and MDM2 (shMDM2-2). MDM2 and NF-κB2 transcript levels were determined by QPCR. The bar graphs show the transcript levels normalized to endogenous GAPDH levels. The assays were performed in triplicates. The error bars are shown. Western blot analysis (G) of extracts prepared from pooled colonies of H460 cells stably infected with either shLuc or shp53 and shMDM2-2.
MDM2 elevates the expression of p100/NF-κB2 transcripts when its ability to interact with p53 is blocked
We further investigated whether the p53-MDM2 interaction is essential for MDM2 to up-regulate NF-κB2 expression. Therefore, we treated A549 cells, which harbor WT p53, 28 with a well-known small molecule inhibitor of MDM2, Nutlin-3, and examined the expression of NF-κB2 transcripts. It is known that Nutlin-3 specifically binds to the p53-binding region of MDM2 to prevent formation of p53-MDM2 complex. Although this event stabilizes WT p53, protecting it from MDM2-mediated degradation, it also releases the transactivation ability of WT p53 and thus up-regulates MDM2 expression. 31 Since Nutlin-3 specifically blocks the p53 interaction function of MDM2, treatment with Nutlin-3 enabled us to determine the transcription function of MDM2 independent of its ability to interact with and degrade p53. Thus, A549 lung cancer cells were treated with Nutlin-3 or vehicle (DMSO). The levels of MDM2 and p100/NF-κB2 transcripts were determined by QPCR of cDNA prepared from RNA isolated from the respective cells. The data obtained clearly demonstrated that Nutlin-3 treatment increased expression of MDM2 (Fig. 4A) and p100/NF-κB2 (Fig. 4B) transcripts, indicating that up-regulation of endogenous MDM2 elevates expression of p100/NF-κB2 transcripts even when its ability to interact with p53 is blocked by Nutlin-3. The data presented in Fig. 3 indicating that knockdown of MDM2, which is known to stabilize p53, drastically decreases expression of NF-κB2 transcripts exclude the possibility that the observed up-regulation of NF-κB2 transcript in Fig. 4 is a result of accumulation of WT p53. Thus, MDM2 is capable of up-regulating expression of p100/NFκB2 independent of p53-interaction despite the presence of higher levels of WT p53, which could repress this transcription. 25
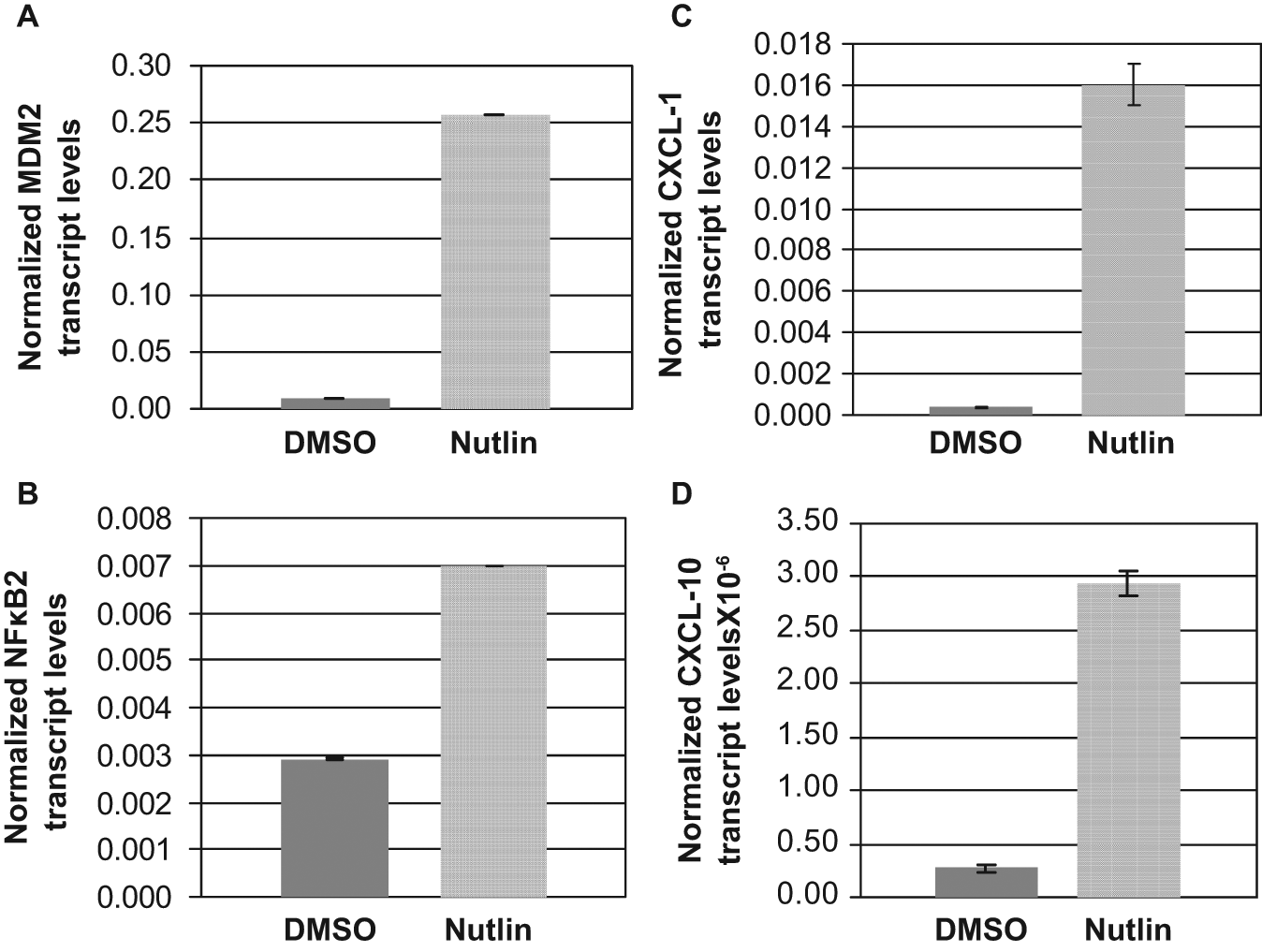
MDM2 elevates expression of p100/NF-κB2 transcripts when its ability to interact with p53 is blocked. Elevation of endogenous transcript levels of MDM2 (A), p100/NF-κB2 (B), and NF-κB2-target genes CXCL-1 (C) and CXCL-10 (D) in A549 lung cancer cells treated with DMSO (vehicle) or Nutlin (10 µM) for 24 hours); Transcript levels were determined by QPCR of cDNA prepared as described in the text. The bar graphs show the transcript levels normalized to endogenous GAPDH levels. The assays were performed in triplicates. The error bars are shown.
MDM2 elevates expression of NF-κB2 target gene CXCL-1 or 10
To determine the functional consequences of up-regulation of p100/NF-κB2 by MDM2, we investigated whether MDM2 enhances transcription of known NF-κB2 target gene CXCL-1 or CXCL-10. 32 For this purpose, we determined CXCl-1 and CXCL-10 transcript levels in A549 lung cancer cells treated with Nutlin-3 or vehicle (DMSO), by QPCR. The data (Fig. 4) clearly demonstrated that along with MDM2 and NF-κB2 transcripts, Nutlin-3 treatment increased expression of CXCL-1 (Fig. 4C) and CXCL-10 (Fig. 4D) transcripts, indicating that MDM2 up-regulates expression of CXCL-1 and 10 transcripts. This demonstrates that up-regulation of p100/NF-κB2 by MDM2 is functionally relevant.
MDM2 protein sequence elements required for up-regulation of p100/NF-κB2 promoter activity overlap with the domain of MDM2 needed to interact with the TATA-binding protein (TBP)
To gain an insight into the mechanism by which MDM2 may activate expression of p100/NF-κB2 transcripts, we determined the structural and functional domain requirement of MDM2 needed for up-regulation of p100/NF-κB2 promoter activity (Fig. 5). The functional domains of MDM2 are shown in Fig. 5A. We performed transient transcriptional analysis in A549 lung cancer cells by introducing a luciferase reporter plasmid containing the p100/NF-κB2 promoter and plasmids expressing full-length MDM2 or its deletion mutants11,33 following a method described earlier. 25 Our results indicated that MDM2 up-regulates the p100/NF-κB2 promoter activity (Fig. 5B). The deletion mutants (Del 491-154 and Del 491-110) encoding the p53-interaction region of MDM2 in the absence of its ubiquitin ligase domain up-regulated the p100/NF-κB2 promoter (Fig. 5B and 5C). Since the N-terminal 100 amino acid residues of MDM2 are responsible for its interaction with p53 11 (Fig. 5A) and the ubiquitin ligase domain of MDM2 is situated at its C-terminus 34 (Fig. 5A), this observation suggests that interaction of MDM2 with WT p53 contributes to MDM2-mediated up-regulation of the NF-κB2 promoter, whereas its ubiquitin ligase domain is dispensable for this function. Our data also showed that the N-terminal 120 amino acid residues of MDM2 (Del 1-120) were dispensable for up-regulating p100/NF-κB2 promoter activity, whereas deletion of the N-terminal 189 amino acid residues inactivated this function (Fig. 5B), implicating 120 to 189 amino acid residues for up-regulation of the p100/NF-κB2 promoter. Since the N-terminal 120 amino acid residues of MDM2 are essential for its interaction with p53, our analysis indicates that MDM2 can up-regulate NF-κB2 promoter by using two independent non-overlapping domains, the p53-interaction domain and the central acidic activation domain that overlaps with its TBP interaction site. 27 This observation, therefore, suggests that MDM2-mediated up-regulation of the NF-κB2 promoter could be mediated by a combination of two independent mechanisms, one through its interaction with p53 and the other independent of p53-interaction.
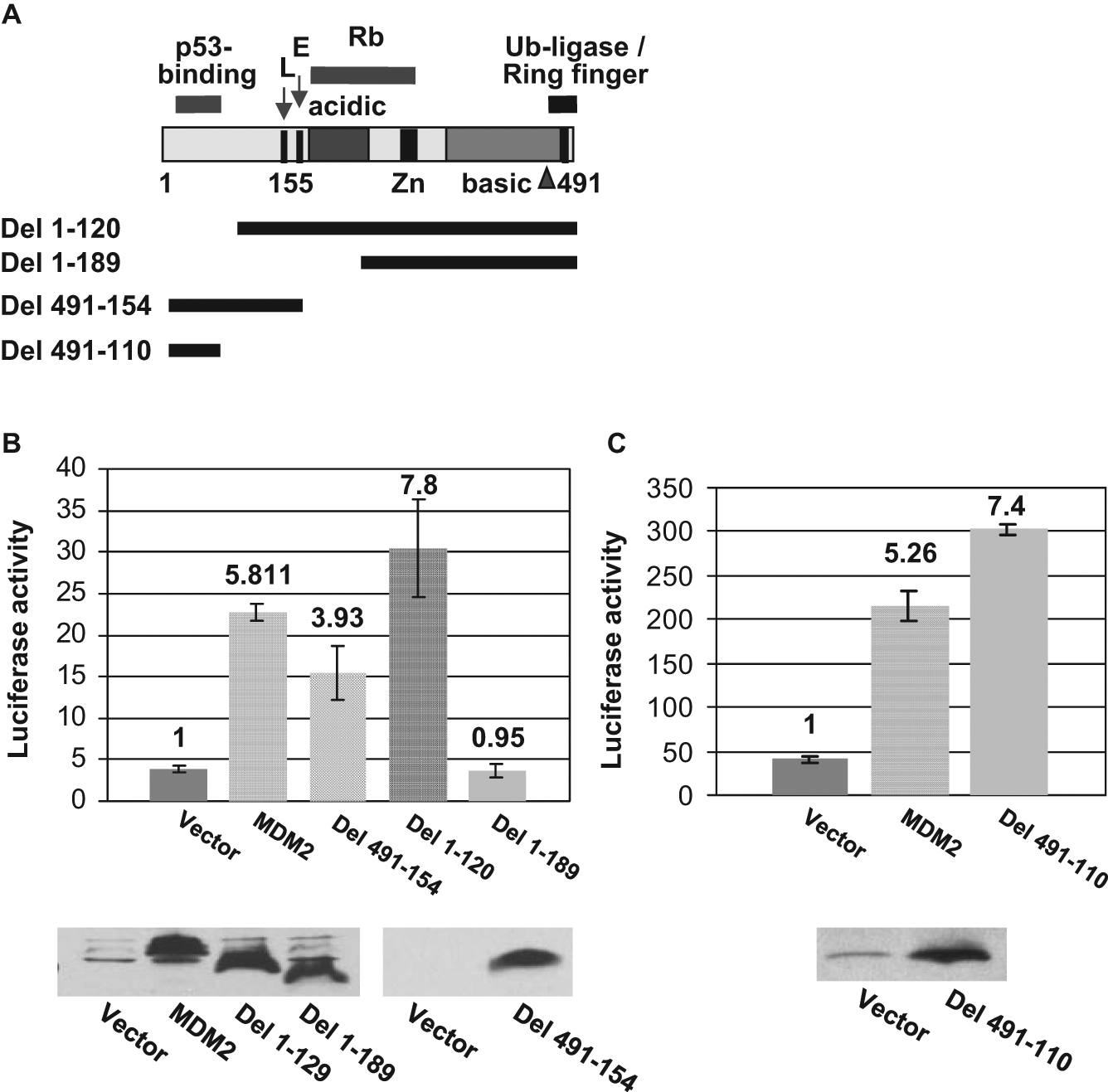
MDM2 requires its TBP interaction domain for up-regulation of p100/NF-κB2 promoter activity and up-regulates a minimal p100/NF-κB2 promoter activity. (A) Functional domains of MDM2. Scheme of MDM2 deletion mutants is shown at the bottom by solid lines. (B, C) The bar graph compares luciferase activities of extracts prepared from A549 cells nucleofected with a plasmid expressing luciferase reporter gene under the control of NF-κB2 promoter (NF-κB2-luc) and a plasmid expressing MDM2 or its deletion mutants (or empty expression vector). The assays were performed in triplicates. The error bars are indicated. Fold-increase is shown at the top of the bar graphs. The expression of the MDM2 deletion mutants was determined by Western blot analysis and is shown in the bottom panel. N-terminal deletion mutants were detected by the antibody 2A10, and the C-terminal deletion mutants were detected by N-20.
We reported earlier that MDM2 interacts with the conserved dimerization domain of TBP. This interaction also requires the 120 to 189 amino acid residues of MDM2. 33 This observation, therefore, suggests that interaction of MDM2 with TBP may be involved in up-regulation of p100/NF-κB2 promoter activity.
MDM2 up-regulates transcription from the P2 start site of p100/NF-κB2 in the presence or absence of p53
Since NF-κB2 promoter has two major start sites, P1 and P2,35,36 we investigated whether MDM2 preferentially increases transcription from the P1 or P2 promoter. For this purpose, H460 lung cancer cells were infected with lentiviral vectors expressing shRNA against MDM2 (shMDM2-1 and shMDM2-2) or luciferase gene (shLuc), and RNA and cDNA were prepared as described above and as described earlier. 29 Levels of MDM2 and p100/NF-κB2 transcripts were determined by QPCR. Exon 1a (transcribed from the P1 start sites) and exon 1b (transcribed from the P2 start sites) specific primers (Table 1) 25 were used for QPCR to determine whether knockdown of MDM2 would reduce expression of 1a or 1b transcript of p100/NF-κB2. As shown in Fig. 6, MDM2 knockdown reduced expression of MDM2 (Fig. 6A), p100/NF-κB2 (Fig. 6B), and Exon 1b transcript of p100/NF-κB2 (Fig. 6D) but did not significantly reduce the expression of the exon 1a transcript (Fig. 6C), indicating up-regulation of expression from the P2 start sites of p100/NF-κB2 promoter by MDM2.
Primer Sequences Used for QPCR
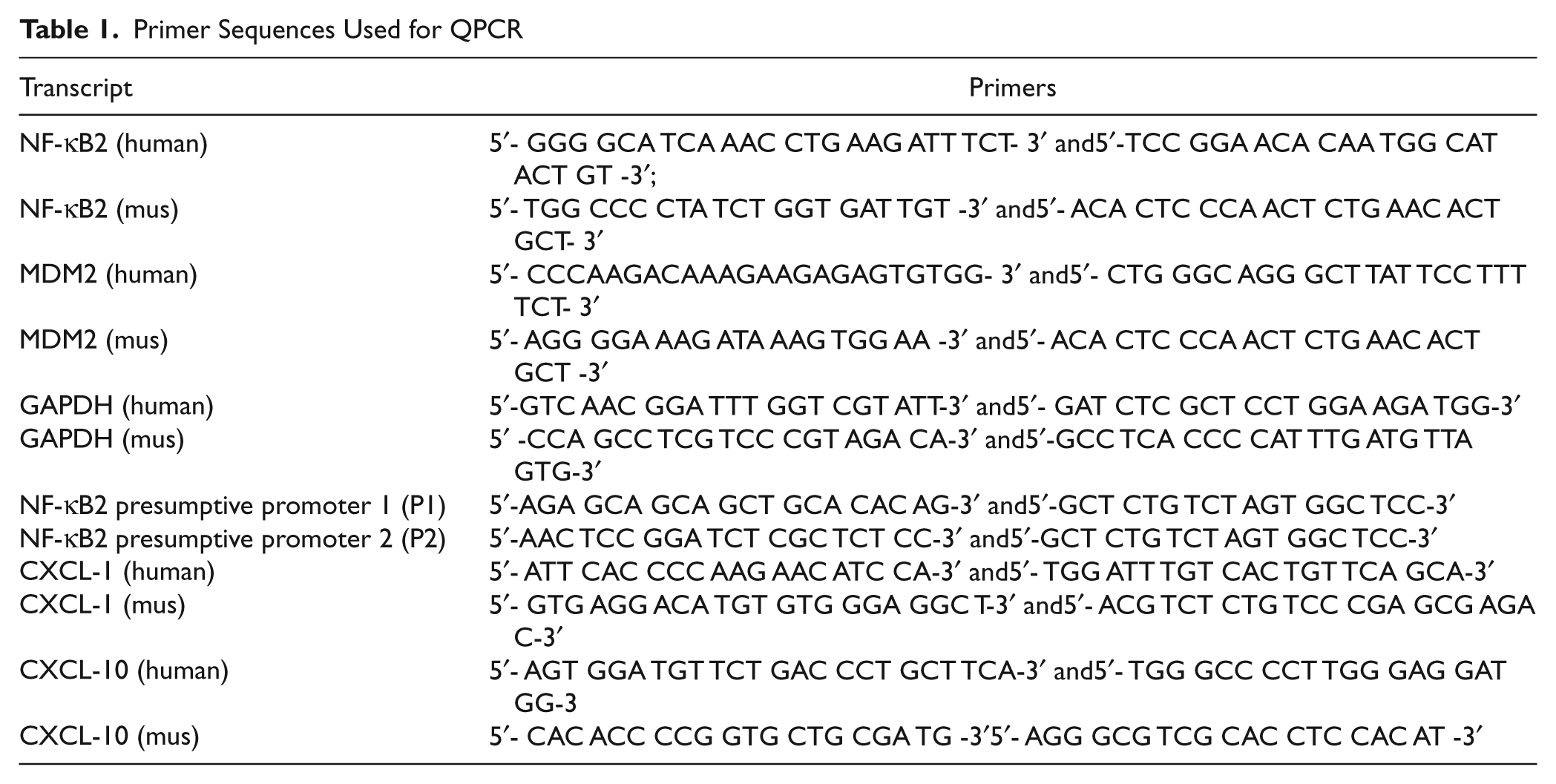
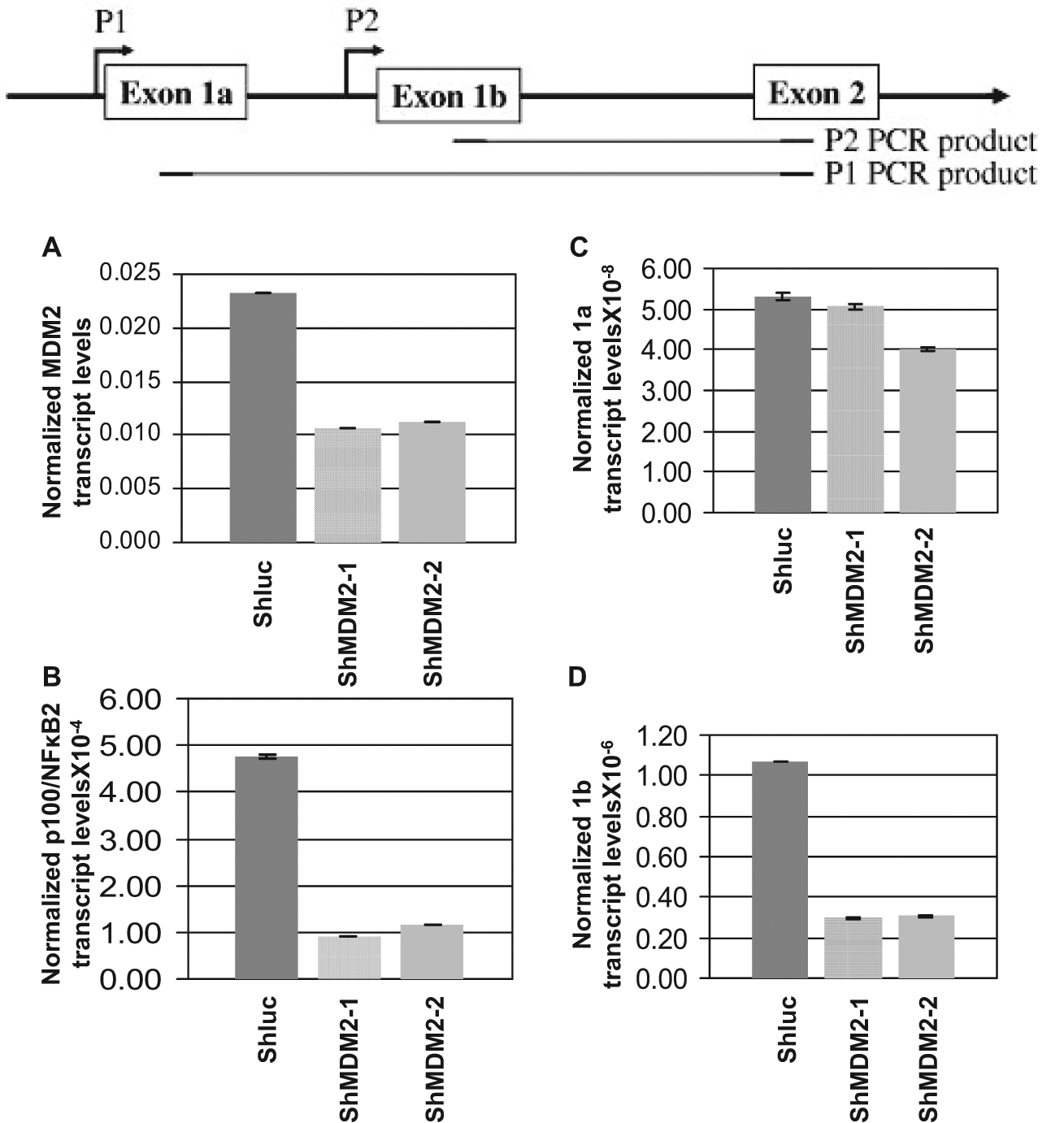
Reduction of MDM2 expression using lentiviral vectors expressing MDM2-specific shRNA lowers NF-κB2 transcription from P2 start site. Promoter specific QPCR was carried out to quantitate MDM2 and p100/NF-κB2 transcripts and transcripts initiated from P1 or P2 start sites of p100/NF-κB2 gene using RNA extracted from H460 cells infected with lentivirus expressing shRNA against the luciferase gene (shLuc) as negative control or 2 shRNAs directed against MDM2 (shMDM2-1 and-2). The bar graphs show the transcript levels normalized to endogenous GAPDH levels. The assays were performed in triplicates. The error bars are shown. Illustration at the top shows the P1 and P2 start sites and PCR products.
Since blocking (Fig. 4) or deleting the p53-interaction domain of MDM2 (Fig. 5B, Del1-120) retained its ability to up-regulate the expression of NF-κB2 transcription or the NF-κB2 promoter (Fig. 5B), we investigated whether MDM2 could up-regulate NF-κB2 expression in cells devoid of WT p53. Stable overexpression of MDM2 in H1299 cells, which do not express p53, 37 enhanced expression of the exon 1b from the P2 start site of the p100/NF-κB2 promoter but not 1a transcript (Fig. 7). These data indicate that MDM2 up-regulates transcription from the P2 start site of p100/NF-κB2 even in the absence of WT p53.
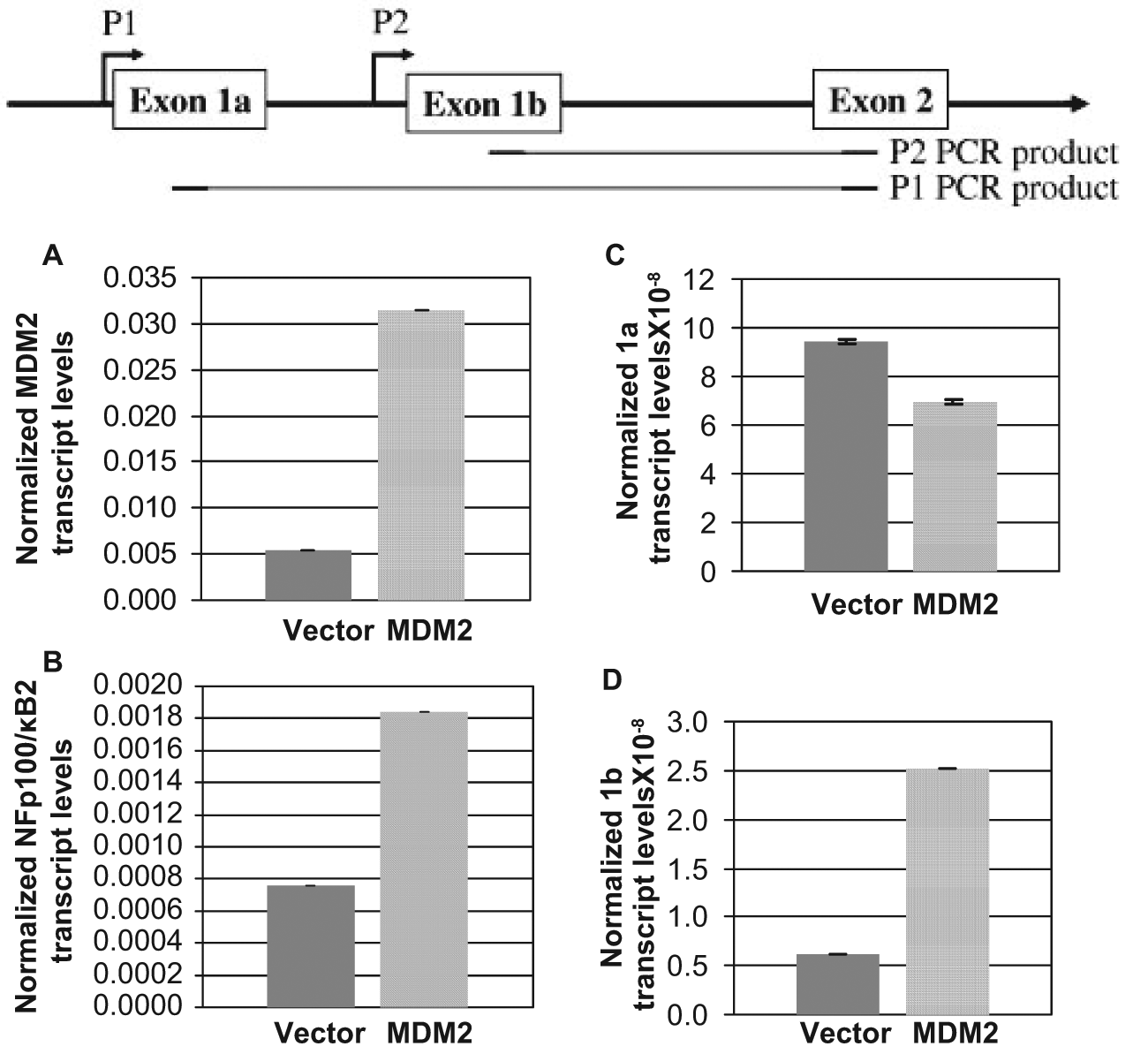
An increase in MDM2 expression in the absence of p53 up-regulates transcription from the P2 start sites of the NF-κB2 promoter. Promoter specific QPCR was carried out to quantitate MDM2 and NF-κB2 transcripts, and transcripts initiated from P1 or P2 start sites of the NF-κB2 gene using RNA were extracted from p53-null H1299 cells stably transfected with MDM2 expression plasmid (or vector alone). Relative RNA transcript levels in comparison to GAPDH levels are shown by bar graphs. The assays were performed in triplicates. The error bars are shown. Illustration at the top shows the P1 and P2 start sites and PCR products.
MDM2 elevates expression of p100/NF-κB2 transcripts and its target genes CXCL-1 and CXCL-10 in the absence of WT p53
We investigated whether endogenous MDM2 present in non-transformed cells in the presence or absence of WT p53 participates in the regulation of NF-κB2 expression. Therefore, we determined MDM2 and p100/NF-κB2 transcript levels in mouse embryo fibroblast (MEF) cells from p53−/− and p53−/−: MDM2−/− mice by generation of cDNA and QPCR. The data obtained clearly demonstrated a reduction in the levels of MDM2 (Fig. 8A) and p100/NF-κB2 transcripts (Fig. 8B) in p53−/−: MDM2−/− MEF cells compared with p53−/− MEF, indicating that MDM2 facilitates p100/NF-κB2 expression in non-transformed cells. Thus, MDM2 enhances p100/NF-κB2 transcription even in the absence of p53.
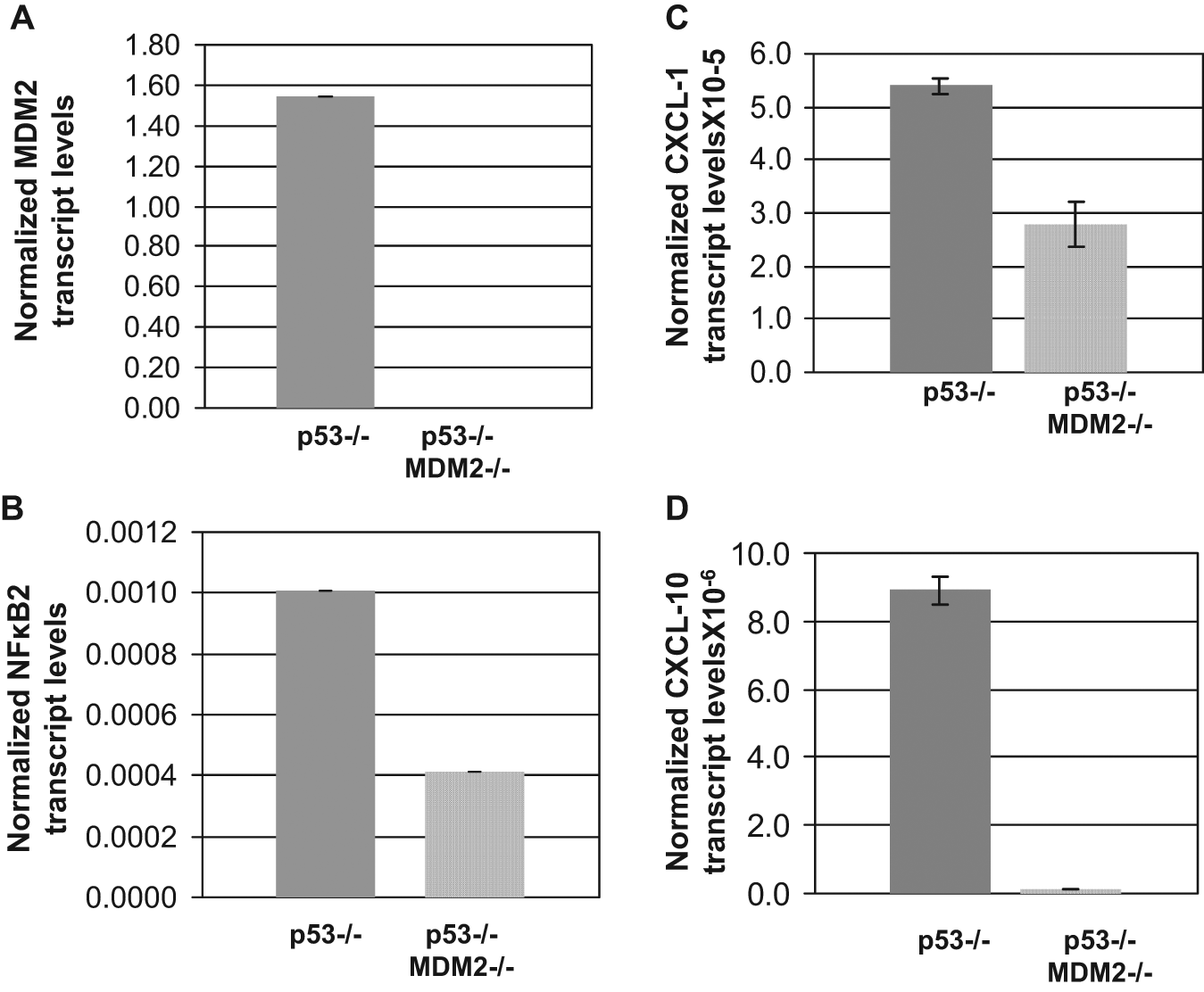
MDM2 up-regulates transcription of p100/NF-κB2 and NF-κB2 target genes CXCL-1 and CXCL-10 in the absence of p53. MDM2 (A), NF-κB2 (B), CXCL-1 (C), and CXCL-10 (D) transcript levels in MEF cells from p53−/− and p53−/−: MDM2−/− mice were determined by QPCR of respective cDNA. An endogenous GAPDH control was used to ensure equal mRNA levels in each sample. The normalized transcript levels are shown by a bar graph. The assays were performed in triplicates. The error bars are shown.
To determine whether up-regulation of p100/NF-κB2 by MDM2 is functionally relevant in p53−/− cells, we further investigated whether MDM2 up-regulates known NF-κB2 target genes CXCL-1 or CXCL-10 (32) in the absence of WT p53. For this purpose, we measured CXCl-1 and CXCL-10 transcript levels in p53−/− and p53−/−: MDM2−/− MEF cells by QPCR. The data (Fig. 8) clearly demonstrated that along with MDM2 and NF-κB2 transcripts, p53−/−: MDM2−/− MEF cells contained reduced levels of CXCL-1 (Fig. 8C) and CXCL-10 (Fig. 8D) transcripts compared with those of p53−/− MEF, indicating that elimination of MDM2 reduces CXCL-1 and 10 expression even in the absence of WT p53. This demonstrates that up-regulation of p100/NF-κB2 by MDM2 is functionally relevant, even in the absence of WT p53.
As evident from the data shown in Fig. 8C and 8D, the reduction of the CXCL-10 transcript levels was higher than that of CXCL-1 transcript levels in p53−/−: MDM2−/− MEF cells. Since p53 has been implicated in suppressing CXCL-1, 38 basal levels of CXCL-1 could be elevated in the absence of p53 in p53−/− MEF. CXCL-10 may not show this effect. However, our data show that MDM2 is involved in the regulation of CXCL-1 in both p53−/− MEF and A549 cells.
Up-regulation of MDM2 in lung cells of transgenic mice with a p53−/− background enhances expression of p100/NF-κB2
To determine whether MDM2 can elevate NF-κB2 protein expression in the absence of p53, we generated littermate p53−/− mice and MDM2 transgenic mice in a p53−/− background. QPCR analysis of genomic DNA isolated from these cells showed a 23-fold higher copy number of the MDM2 gene than littermate p53−/− mice. Lung fibroblast cells were isolated from these mice following a method described earlier. 27 The levels of MDM2 and NF-κB2 were determined by Western blot analysis of cell lysate. Densitometric analysis indicated that MDM2 overexpression elevated p100 and p52 expression approximately 2.9- and 1.7-fold, respectively (Fig. 9A, B).
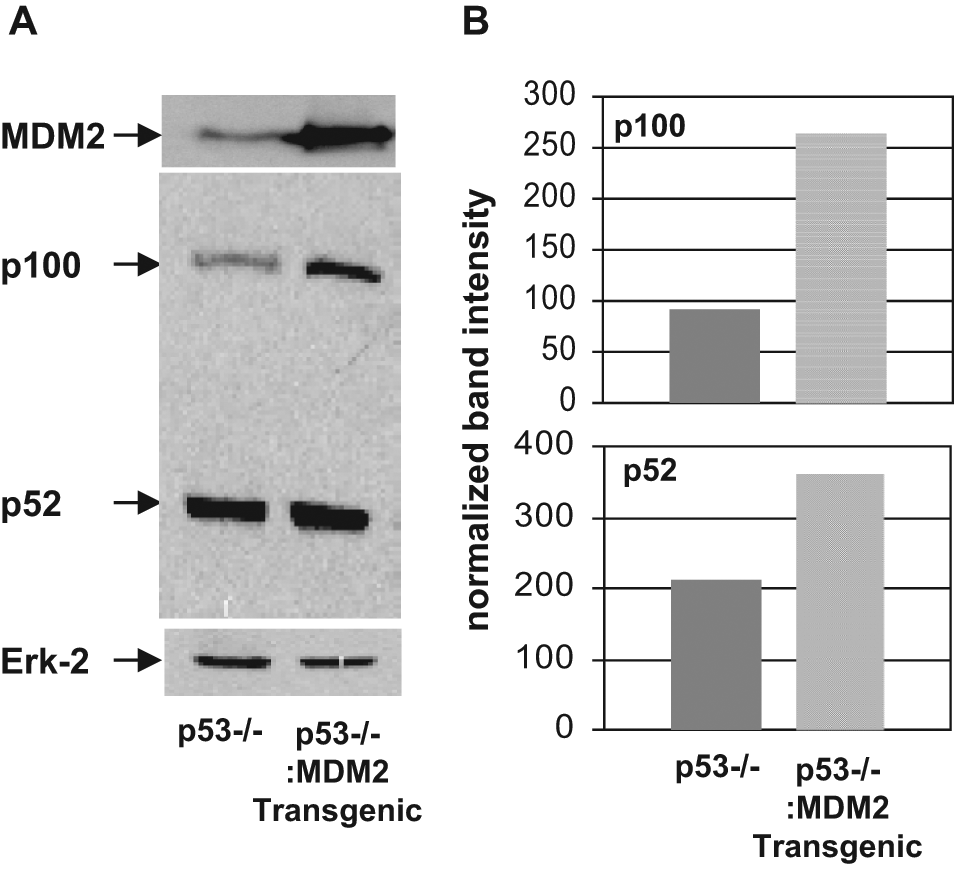
Enhanced expression of MDM2 in transgenic mice with a p53−/− background increases p100/NF-κB2 expression. Western blot analysis (A) of extracts prepared from lung cells derived from p53−/− and p53−/−: MDM2 transgenic mice for expression of MDM2, NF-κB2 p100, and p52. MDM2, NF-κB2, and a loading control Erk-2 were identified using respective antibodies, and their migrations are shown by arrows. Densitometric analyses (B) of p100 and p52 expression are also shown by bar graphs. Band intensities were normalized to the levels of Erk-2.
MDM2 requires NF-κB2 to prevent cell death by paclitaxel: NF-κB2 up-regulation has been related to chemoresistance of cancer cells.23,25,39 MDM2 overexpression has also been associated with reduced death of cancer cells in response to chemotherapeutic drugs.40,41 Therefore, we investigated whether the ability of MDM2 to up-regulate p100/NF-κB2 expression contributes to a lack of response of cancer cells to chemotherapeutic drugs. For this purpose, we used H1299 cells stably expressing MDM2 (or stably transfected with the expression vector alone) as described above (Fig. 10). A lentivirus expressing shRNA against p100/NF-κB2 (sh NF-κB2) was used to knockdown p100/NF-κB2 expression, whereas a lentivirus expressing shRNA against the luciferase gene (sh Luc) was used as a control. H1299 cells stably expressing MDM2 or the control cell line were infected with the lentiviruses, and subsequently an equal number of infected cells were treated with 25 nM paclitaxel (or DMSO) and subjected to colony formation assay as described in Materials and Methods. Our data (Fig. 10A) indicated increased colony forming ability of H1299 cells overexpressing MDM2 compared with the vector transfected cells. However, this difference is abrogated with knockdown of NF-κB2 expression. These data show that silencing of NF-κB2 expression sensitizes MDM2 overexpressing cells to paclitaxel. As shown in Fig. 10B, p100/NF-κB2 shRNA lentiviruses lowered the expression levels of NF-κB2 as expected. Since the cytoplasmic NF-κB2 protein levels in H1299 cells is very high, the differences in the NF-κB2 protein levels in MDM2 expressing whole cells extracts were masked by high levels of cytoplasmic protein. However, the difference in NF-κB2 protein levels in the vector transfected and MDM2-expressing cell lines can be detected in the respective nuclear extracts (data not shown). Since we intended to show overall knockdown of NF-κB2 protein expression, we used whole cell extract for this experiment.
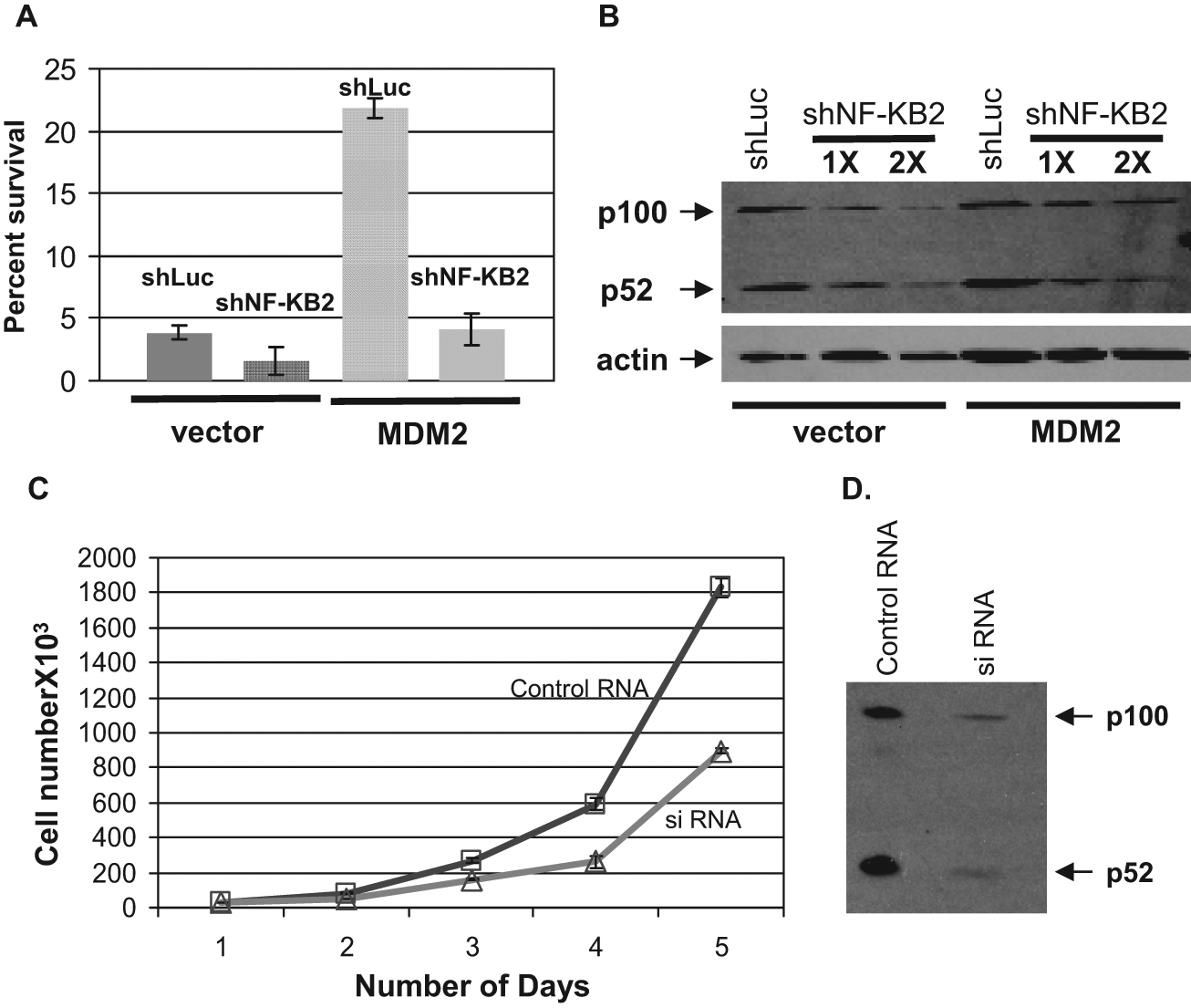
Silencing of NF-κB2 expression reverses MDM2-mediated paclitaxel insensitivity and decreases cell proliferation. The percentage survival of vector or MDM2 overexpressing H1299 cells after infection with the lentiviral vectors expressing shRNA against p100/NF-κB2 or luciferase followed by treatment with 25 nM paclitaxel was determined by colony formation assay (A). Number of colonies generated from DMSO (vehicle) treated cells in each case was considered to be 100%. Western blot analysis shows silencing of NF-κB2 expression by shNF-κB2 (B). 1X and 2X indicate the amount of virus used for silencing. 2X amount of the virus was used for the colony formation assay shown in (A). Lowering of the p100/NF-κB2 level was accompanied by a lowering of chemoresistance against paclitaxel. Equal numbers of H460 lung cancer cells were plated after transfection with NF-κB2 or control siRNA (C, D). The curve (C) shows the rate of cell growth generated by plotting cell number (average of triplicate experiments) counted at 24-hour intervals. The right panel (D) shows expression of NF-κB2 after transfection with the siRNA or control siRNA. All experiments were performed in triplicates. The error bars are indicated.
These data indicate that MDM2-mediated up-regulation of p100/NF-κB2 expression contributes to the reduced response of MDM2 overexpressing cancer cells to chemotherapeutic drugs.
Down-regulation of p100/NF-κB2 expression reduces cell proliferation
We next determined whether endogenously expressed NF-κB2 is involved in cell proliferation. H460 cells were transfected with siRNA targeted against p100/NF-κB2. After transfection, an equal number of cells were plated and counted at 24-hour intervals. The results of this experiment showed that introduction of p100/NF-κB2 siRNA down-regulates the rate of cell proliferation (Fig. 10C). Western blot analysis of the cells extracts showed that the siRNA against NF-κB2 down-regulates both p100 and p52 levels (Fig. 10D). This result suggests that endogenous levels of NF-κB2 determine the rate of cell proliferation, implying that up-regulation of p100/NF-κB2 expression by MDM2 contributes to an enhanced growth of human lung cancer cells.
Discussion
Cancer is a disease of multiple genetic abnormalities, and identification of abnormally expressed genes in human cancers has often been used as prognostic markers. Knowledge of a specific genetic lesion may not predict prognosis, chemotherapeutic resistance, or disease recurrence because downstream mutations may alter the expected consequence. It is therefore essential to identify functionally related groups of markers that can be used to predict chemotherapeutic drug response or other oncogenic parameters in human lung tumor.
We have identified a novel transactivation function of MDM2 by investigating a correlation of MDM2 and p100/NF-κB2 expression in human lung tumor samples. Our analysis revealed MDM2 overexpression in both WT and mutant p53 expressing tumors (data not shown) and did not find any significant co-occurrence of MDM2 overexpression with WT or mutant p53. This suggests that MDM2 overexpression may have a WT p53-independent role in oncogenesis. Our earlier studies demonstrated that p53 mutation induces expression of the transcription factor NF-κB2. 25 Here, we have presented data to show that expression of MDM2 and p100/NF-κB2 transcript levels is strongly related in human lung tumors with WT p53.
Correlated expression of MDM2 and NF-κB2 transcripts in human lung tumors prompted us to investigate whether MDM2 up-regulates p100/NF-κB2 expression. Data presented in this article show that down-regulation or elimination of MDM2 expression in apparently normal or lung cancer cells lowers p100/NF-κB2 transcript levels demonstrating that MDM2 regulates expression of p100/NF-κB2 transcripts (Fig. 2). This ability of MDM2 to up-regulate NF-κB2 expression is a combined effect of interaction of MDM2 with p53 25 and a p53-independent transcription regulatory function of MDM2. Consistently, the p53- interaction domain of MDM2 could up-regulate expression of NF-κB2 transcript, while blocking or deletion of the p53-interaction domain retains this function (Figs. 4 and 5). MDM2 could up-regulate p100/NF-κB2 expression in the absence of p53 (Figs. 7-9). These experimental data convincingly demonstrate the presence of a p53 independent mechanism in MDM2-mediated transactivation of p100/NF-κB2.
However, the levels of NF-κB2 up-regulation by MDM2 were consistently higher in the presence of WT p53 than in its absence, although MDM2 expression is also higher in cells with WT p53, because of p53’s ability to up-regulate MDM2. Therefore, the observed difference in MDM2- mediated up-regulation of NF-κB2 could be a consequence of p53-mediated up-regulation of MDM2. Our data also indicate that MDM2 is a facilitator rather than an essential factor required for p100/NF-κB2 expression, because, although at a lower level, p53−/−: MDM2−/− MEF cells express p100/NF-κB2 transcripts (Fig. 8D). These observations argue for a mechanism that involves interaction of MDM2 with a transcription factor needed for NF-κB2 expression.
Since MDM2 can inhibit the transactivation function of p73 even though it does not cause p73 degradation (e.g., Balint et al. 42 ), it is possible that MDM2 up-regulates NF-κB2 expression by inactivating p73. Therefore, we have investigated how p73 modulates NF-κB2 promoter activity. Our data showed that p73 did not significantly alter NF-κB2 promoter activity (data not shown). However, these experiments do not exclude the possibility that p73 may interfere with MDM2 mediated NF-κB2 up-regulation by blocking its p53-interaction site.
Our previous studies indicated interactions of MDM2 with TBP at the conserved domain of TBP encompassing amino acid residues 100 to 275, which harbors its dimerization domain. 33 MDM2 also interacts with TAFII250. 43 Overlapping domain requirements of MDM2 for its interaction with TBP 33 and up-regulation of p100/NF-κB2 expression argue for the possibility that the interaction of MDM2 with TBP enhances NF-κB2 expression. Since MDM2 is capable of activating p100/NF-κB2 transcription in the absence of p53 (Figs. 7-9), the p53-binding MDM2 deletion mutants (Fig. 5, Del 491-154) may release the endogenous MDM2 from the p53-MDM2 complex, making MDM2 available for p100/NF-κB2 transcription.
Our analysis of the P1 and P2 transcription start sites of p100/NF-κB2 revealed that MDM2 up-regulates transcription from the P2 site and accordingly generates more exon 1b transcripts than exon 1a (Figs.6 and 7).
We provide clear evidence that in the absence of p53, MDM2-induced paclitaxel insensitivity is dependent on the p100/NF-κB2 expression, indicating that MDM2-mediated p100/NF-κB2 up-regulation might contribute to chemotherapeutic resistance of MDM2 overexpressing cancer cells (Fig. 10A, 10B). We also present data suggesting that endogenous levels of NF-κB2 determine the rate of cell proliferation (Fig. 10C, 10D). Since endogenous levels of MDM2 influence p100/NF-κB2 levels, this implies that MDM2 overexpression in human lung cancer may enhance cell proliferation by up-regulating p100/NF-κB2.
The novel observations presented in this report suggest that MDM2-mediated up-regulation of p100/NF-κB2 expression may confer a selective survival advantage in human lung cancer. The strength and significance of the observation are compounded by the fact that MDM2-mediated up-regulation of p100/NF-κB2 expression observed in human lung cancer samples can be reproduced in cultured cells. These observations also predict a novel pathway responsible for MDM2-mediated oncogenesis.
Materials and Methods
Human lung cancer samples
The Tissue and Data Acquisition Core (TDAAC) laboratory at the Virginia Commonwealth University (VCU) acquires human residual lung tumor samples under a VCU IRB-approved protocol (IRB number 2471). Thirty human lung cancer specimens consisting of 21 adenocarcinomas and 9 squamous cell carcinomas were used in the present study. All tumors were classified according to standard histopathological criteria. 44 Histological evaluation of the frozen tumor tissues showed that on average, all specimens studied consisted of 68% ± 15% tumor cells. Adjacent non-neoplastic tissues were used as control tissues for the expression studies.
Plasmids
Construction of plasmids expressing full-length MDM2 and its deletion mutants has previously been described in detail.9,11 Construction of the NF-κB2 promoter was described earlier. 25 A plasmid containing Renilla luciferase under the control of thymidine kinase promoter (TK Renilla luciferase) was purchased from Promega (Madison, WI).
Generation of MDM2 transgenic mice with p53+/+ and p53−/− background
To determine whether MDM2 can elevate NF-κB2 protein expression, we generated MDM2 transgenic mice with p53+/+ and p53−/− background by crossbreeding p53+/−: MDM2 transgene+/− mice (gift from Stephen Jones). 45 The mice were genotyped by analyzing DNA from tail biopsies. 45
Cells, transfections, and generation of stable transfectants
A549, H460, and H1299 cells were purchased from American Type Culture Collection (Manassas, VA) and were maintained in media as suggested by the suppliers. Mouse embryo fibroblast (MEF) cells from normal or p53−/−: MDM2−/− mice and tumor cells from p53−/−: MDM2−/−; mice were gifts from Guillermina Lozano. 46 We also generated cultured lung cells from littermate MDM2 transgenic and non-transgenic mice with p53+/+ and p53−/− background.
For transfection of expression plasmids, cells were seeded 48 hours before transfection at a density of 3 × 106 cells per 10 cm dish and transfected (nucleofected) using the Nucleofector and kit reagents (Amaxa, Gaithersburg, MD) following the supplier’s protocol. Cells were harvested 24 hours after transfection for transient transfection or Western blot analysis unless otherwise indicated. To generate MDM2 stable transfectants, H1299 cells were transfected with MDM2 expression plasmids or the empty vector plasmid harboring a neomycin resistance gene and selected with appropriate antibiotic and pooled.
Lentivirus generation and infection
Plasmids (pLKO.1) expressing short hairpin (sh) RNA against MDM2 from the U6 promoter and harboring a puromycin resistance gene were purchased from Open Biosystems (Lafayette, CO) and tested for their ability to lower MDM2 expression. The control plasmid expresses shRNA against a non-endogenous luciferase gene. The lentiviral vectors to express shRNA against MDM2 (shMDM2-1 and -2), p53 (shp53), or luciferase (shLuc) were generated by transfecting the respective recombinant plasmid encoding shRNA into 293T cells (ATCC) by following the supplier’s protocol as described previously.29,30 For expression of shRNA, cells were infected with lentiviruses. Infected cells were incubated for 72 hours, after which cells were harvested for RNA preparation.
RNA extraction
RNA preparation from lung tumor samples was performed using methods described earlier.25,29 Total RNA was isolated from exponentially growing cultured cell lines using TRIzol reagent (Life Technologies, Invitrogen, Carlsbad, CA) following a protocol supplied by the manufacturer. Quality of RNA was checked by 1.2% agarose Tris-borate-EDTA gel electrophoresis.
Generation of cDNA and quantitative PCR (QPCR)
cDNA was synthesized using the Thermoscript RT-PCR system (Invitrogen) and amplification of the cDNA by PCR used sequence-specific primers. QPCR was performed using a LightCycler system (Roche, Branchburg, NJ) as described previously.25,29 Primers were designed using OLIGO 5 software (Molecular Biology Insights, Cascade, CO) and synthesized by Sigma Genosys. The QPCR primers used are shown in Table 1. p100/NF-κB2 promoter PCR was performed using the conditions described. 25 Reactions were performed in triplicate using SYBR green dye, which exhibits a higher fluorescence upon binding of double-stranded DNA. The data obtained (fluorescent units) were normalized with GAPDH, actin, or tubulin, and transcript levels per unit of GAPDH, actin, or tubulin transcripts were plotted in the bar graphs.
Transient transcription assays
MDM2-mediated up-regulation of the NF-κB2 promoter was determined by transient transcription assays as described earlier.18,25,47 Briefly, 3 × 106 H460 or A549 human lung cancer cells that harbor WT p53 were nucleofected with 800 ng of a reporter plasmid containing the p100/NF-κB2 promoter (1645 base pairs of sequences upstream from the ATG site of exon 2) inserted upstream of the luciferase gene and 2 µg of MDM2 expression plasmid (or vector plasmid). Twenty-four hours after transfection, cell lysates were prepared using passive lysis buffer (Promega). Cell extracts containing equal amounts of protein were assayed for firefly and Renilla luciferase activities as described earlier. 25
Western blot analysis
Western blot analysis was carried out as described previously.47,48
Antibodies
Antibody against MDM2 (Ab-1 Calbiochem, San Diego, California) was used in 1:100 dilutions. NF-κB2 antibody (Upstate, Burlingame, CA) was used in a 1:5000 dilution, and β-actin antibody (Sigma, St. Louis, Missouri) was used in a 1:200 dilution.
Colony formation assay
Paclitaxel sensitivity was determined by colony formation assay as described by us earlier. 25 H1299 cells stably transfected with MDM2 expression plasmid (or expression vector alone) described above were infected with lentiviruses expressing either shLuc or shRNA against p100/NF-κB2 (sh NF-κB2, Add Gene, Burlingame, CA). Equal numbers of infected cells were seeded and treated with 25 nM paclitaxel (Sigma) or the vehicle, DMSO, for 48 hours. The surviving cells were allowed to form colonies with periodic changes of media. Colonies were fixed with methanol, stained with methylene blue, and counted. The percentage survival was determined considering the number of colonies generated in DMSO treated plates to be 100%. All experiments were performed in triplicates and repeated multiple times.
Cell growth assay
Cell growth assays were performed as described earlier with some modifications. 29 Cells were seeded at a density of 5 × 104 cells per 60-mm plate 16 hours after transfection of NF-κB2 or control siRNA. Triplicate sets of plates were harvested at 24-hour intervals and counted using a Coulter counter.
Statistical analysis
Relationships between the MDM2 expression (independent variable) and p100 expression levels (dependent variables) were determined using separate linear regression models. The relationships were also examined by grouping the lung cancer samples into WT and mutant p53 harboring populations.
Footnotes
Acknowledgements
We thank Michelle Anderson for technical help and V. Ramakrishnan for statistical analysis
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was initiated by pilot project funds from the VCU Massey Cancer Center (P30CA016059), and was supported by funds from Jeffress Memorial Trust to Swati Palit Deb and by and NCI (CA121144) to Sumitra Deb.
