Abstract
The methyl-CpG binding proteins (MBPs) interpret the methylation of DNA and its components. The number of MBPs in the human body currently stands at 15, which are split into 3 branches, a reflection of the intricate mechanisms of gene regulation. Each branch utilizes a different mechanism for interacting with methylated DNA or its components. These interactions function to direct gene expression and maintain or alter DNA architecture. It is these functions that are commonly exploited in human disease. For this review, we will focus on each protein and any roles it may have in initiating, promoting, progressing, or inhibiting cancer. This will highlight common threads in the roles of these proteins, which will allow us to speculate on potentially productive directions for future research.
Keywords
Introduction
The field of epigenetics and cancer is broad and extensive. The extent of these studies is a reflection of how much eukaryotic biology is reliant on appropriate DNA methylation. A key aspect of the field involves the study of the proteins that directly interpret the methylation of DNA and its components. The methyl binding protein (MBP) family plays a pivotal role in this interpretation. Since the identification of the first MBP in 1989,
A Brief History of MBPs
In mammalian DNA, the MBPs were originally characterized by their interactions with methylated DNA, namely CpG dinucleotides. The CpG is a cytosine-guanosine (CG) dinucleotide DNA sequence, in which the cytosine undergoes chemical modification to contain a methyl group. This methylation of cytosine was first detected in 1925 in the tubercle bacillus. 4 It was another 23 years before it was observed in eukaryotes when it was detected in calf thymus DNA in 1948. 5 The importance of methylation became apparent in the 1970s following key findings such as the fact that the doublet CpG sequence occurs at approximately one fifth of the expected frequency within the genome6,7 but represents the main target of methylation.8,9 These findings suggested that the regulation of genes had to be due to the interaction of genomic DNA with the cytoplasmic enzymes that it encoded.10,11 By the end of the 1970s, it had been demonstrated that an important aspect of eukaryote gene regulation came from protein-methylated DNA interactions12,13 and that DNA methylation patterns are inheritable.14,15
Today, the nature of DNA methylation and many of its roles within eukaryotic cells have been elucidated. The enzymes that catalyze the methylation of a CpG dinucleotide have been identified and characterized into the DNA methyltransferase family. 16 These enzymes take a methyl group, donated from S-adenosyl-L-methionine, and cova-lently attach it to the fifth carbon position of cytosine to form a symmetrical CpG dinucleotide. In the mammalian genome, approximately 60% to 90% of CpGs are methylated. 17 The remaining unmethylated CpGs are clustered into what are termed CpG islands, which comprise around 1% of the genome.18,19 These CpG islands are located within the promoter, first exon, or 3′UTR regions of approximately 60% of the RNA polymerase II–transcribed genes in the vertebrate genome, consistent with their role in gene expression.16,20 Indeed, methylated CpGs (mCpGs) are now synonymous with a repressed chromatin environment and silencing of gene expression. They play an important role in expression of patterns of imprinted genes, X chromosome inactivation, suppression of transposable elements, developmental biology, and many disease mechanisms. 17
One of the key aspects of the relationship between methylation of DNA and the silencing of gene expression was established in the late 1980s and early 1990s, with the identification of proteins that could bind to mCpGs and recruit protein complexes. These complexes contained histone-modifying enzymes, which leads to heterochromatin formation and gene silencing.
21
The initial identification of a methyl-CpG binding protein1,22 (
The MBP Superfamily and Cancer
As previously mentioned, the MBP family is divided into 3 branches: 1) MBD containing proteins, 2) methyl-CpG binding zinc fingers, and 3) the SRA domain containing proteins. 30 This section will cover the functions of the individual members of each family that are present in humans and highlight any roles in cancer (Table 1). Much of the data available on the roles of these proteins in cancer are related to their functions as transcriptional repressors or chromatin remodelers (Fig. 1). There is little evidence in the literature that deficiency of any of these proteins results in elevated cancer predisposition, a notion supported by a lack of any entries in the Catalogue of Somatic Mutations in Cancer (COSMIC) database 31 implicating their inactivation.
Types of Cancer in Which Methyl-CpG Binding Proteins (MBPs) Have Been Implicated to Play a Role

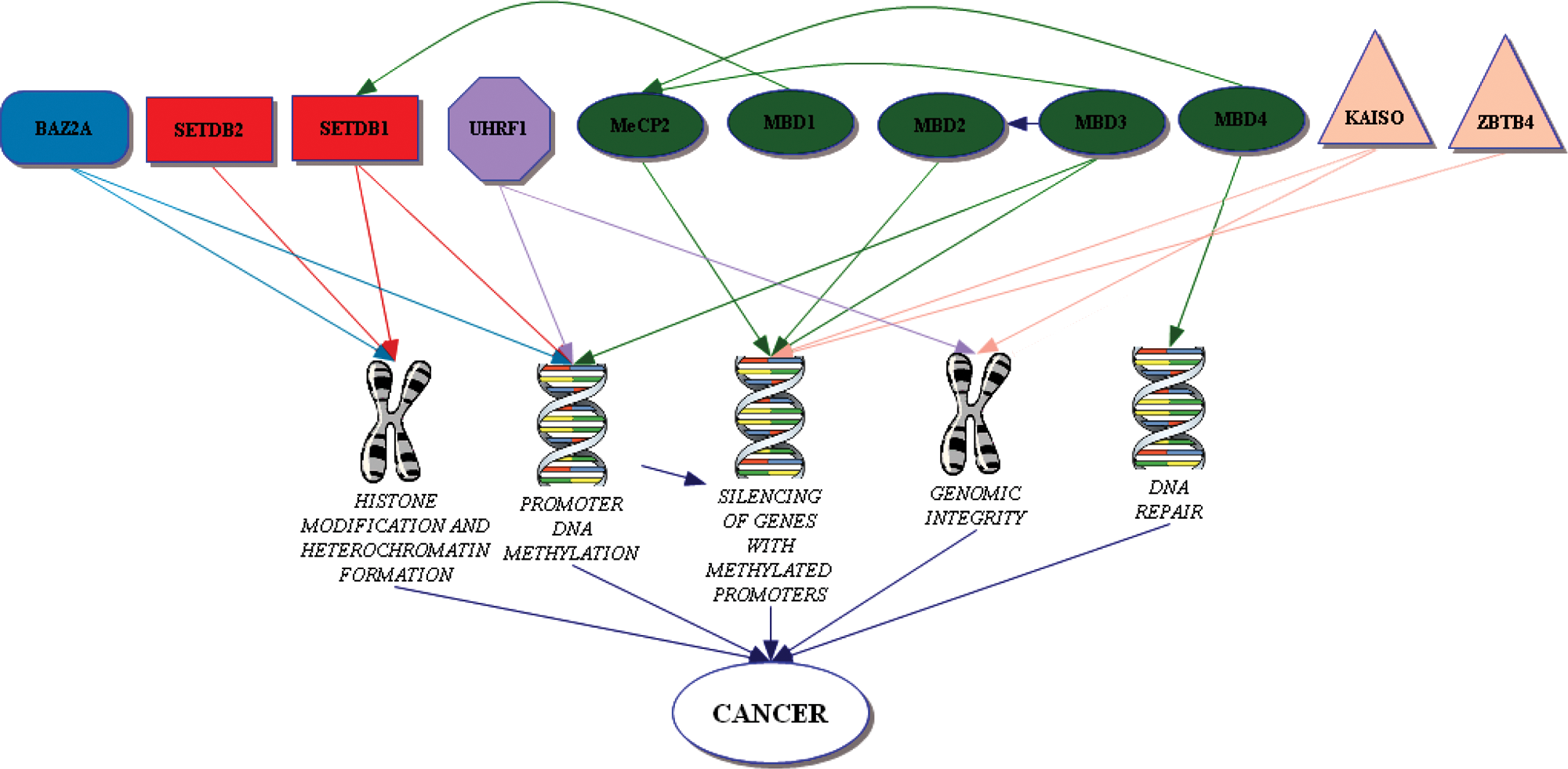
Mechanisms by which methyl-CpG binding proteins (MBPs) are implicated in cancer.
MBD-Containing Proteins
These proteins comprise the largest branch of the MBP family. Currently, the NCBI Conserved Domain Database (CDD)32-34 lists 11 human proteins containing the MBD derived from
HMT_MBD
There are 2 members of this family: SETDB1 (also known as ESET) on chromosome 1 and SETDB2 (also known as CLLD8) on chromosome 13. As well as containing a MBD, they carry a PreSET and bifurcated SET domain, which mediates protein-protein interactions.
As may be expected from their function, their roles in cancer are defined by the genes that they repress and interact with. Currently, the only evidence implicating
The SETDB1 interactions with MCAF1 and DNMT3A link promoter CpG hypermethylation to histone methylation, a crucial precursory event to heterochromatin formation. Its interaction with MCAF1 also links it to another transcriptional repressor, MBD1 (another MBP).
43
Through MBD1, it is directed to mCpGs, where the MBD1:SETDB1: MCAF1 complex converts the dimethyl H3-K9 to trimethyl H3-K9, resulting in heterochromatin formation and transcriptional repression.40,44-46 In cancer, this MCAF1:MBD1 complex is associated with the maintenance of telomerase activity,
47
but this function seems independent of SETDB1. The interaction with the DNMT3A protein is relevant as it is a
The link with heterochromatin formation is further supported by its interaction with KAP1, which also provides a link to global gene regulation as KAP1 is a universal cofactor involved in regulating the Kruppel-associated box domain zinc finger proteins (KRAB-ZFP), the largest group of transcriptional repressors in higher organisms. 57 Surprisingly, for such a large group, there is a paucity of information on the targets of KRAB-ZFP proteins. Along with the ability of KAP1 to mediate long-range repression through heterochromatin spreading, it is difficult to assess any specific roles for its interaction with SETDB1 in cancer. 38
It is perhaps SETDB1 interaction with Akt that is currently of greatest significance as the aberrant activation of the Akt/PI3K pathway is being targeted for treatment in a variety of cancers.
58
The role of SETDB1 on Akt, the main downstream effector molecule of the PI3K pathway,
59
is not well understood, although it has been demonstrated
MeCP2-MBD
This represents the largest group of the MBD branch of the MBP superfamily. It comprises the original MeCP2 protein and 6 others termed MBD1 to MBD6.19,62,63 The roles of many of these proteins in cancer were reviewed in 2007 2 and 2008. 3 We will provide an update of the latest developments for the regulation of these proteins that have, or may have, roles in cancer.
MeCP2
The founder protein of the MBP superfamily MeCP2 is a global transcriptional repressor. It is a 50-kDa protein encoded by a gene on the X chromosome. It can bind to a single methylated CpG and recruit the Sin3A repressor complex to silence transcription via histone deacetylation.
26
As well as its role in Rett syndrome, a neurodevelopmental disorder,
64
it has also been shown to have a role in tumorigenesis2,3 as many cancers rely on genes becoming silenced via hypermethylation of their promoter. Recently, it has been shown to target genes
MBD1
The MBD1 protein is the largest member of the family. It has a complex expression profile as there are 13 isoforms of the gene expressed from chromosome 18. The main difference between the isoforms is the presence of 2 or 3 CXXC-type zinc fingers present in the protein.
76
The isoforms containing the first 2 CXXC domains preferentially repress methylated promoters, whereas those with the third CXXC domain are capable of DNA binding regardless of methylation status.
77
These isoforms containing the third zinc finger have been shown to repress the transcription of genes with unmethylated promoters
In relation to cancer, polymorphisms in MBD1 have been shown to be significantly associated with lung cancer risk in a Chinese population,
80
both positively and negatively. The effect of these SNPs on the protein or its expression has yet to be investigated. In relation to its transcriptional roles, most of the genes silenced by MBD1 in cancer have been identified
The capability of these MBD1 isoforms to play a role in cancer is likely to be effected by factors such as redundancy and target specificity. As MBD1 binds with greater efficiency to genes in which the promoter has the sequence TCMGCA or TGCMGCA, 78 and a comparison of cancer cell lines of different origins demonstrated that the promoters of many silenced genes were occupied by more than one MBD protein. 79
MBD2
The gene for
Despite its attractiveness as a therapeutic target in colorectal cancer, no drugs that specifically target the MBD2 gene or protein have been identified to date. However, the drug B1 (N-[2-[dimethylamino]ethy]l-2-aminothiazonaphthalimide) has been shown to alleviate MBD2 repression of the 14-3-3σ gene in promyelocytic leukemia cells and induce their apoptosis. Further characterization of this ability would be desirable. An alternative way of targeting MBD2 is offered by targeting the PRMT1 or PRMT5 protein arginine methyltransferases that have been shown to methylate the MBD2 arginine residues. This methylation inhibits the ability of MBD2 to associate with its repressive complexes and silence genes. 104
As attractive as MBD2 is as a therapeutic target, caution must be exercised in an application that will deregulate global gene transcription. By way of illustration, MBD2 is known to play a role in the silencing of the human telomerase reverse transcriptase promoter (hTERT) in somatic cells. Given that upregulation of this gene is a key event for the ability of many cancer cells to overcome replicative senescence associated with telomere shortening, 105 there are clear potential negative ramifications for inactivation of MBD2.
MBD3
The chromosome 19 gene,
Roles in human cancer that are specific to MBD3 are not commonly observed.
107
There are MBD2-independent reports of a role for the MBD3/NuRD complex shown in both prostate cancer cells
108
and leukemias. In acute promyelocytic leukemia, it interacts with the fusion PML-RARa oncoprotein that is generated from the 15;17 translocation,
109
and in acute myeloid leukemia, it interacts with the
A different role has been shown in mouse lymphosarcoma cells in which Mbd3 interacts with Brg1 and Dnmt3a 115 to silence the methylated MT-I gene. It remains to be identified which complex mediates the silencing as the association of Mbd3/NuRD repressor with Dnmt3a can link it to methylated DNA. Alternatively, the SWI/SNF complex, of which Brg1 is a central component, could also be used as it associates with the Mecp2/Sin3a/HDAC complex.116,117 Its roles in the SWI/SNF complex are not well understood but may make MBD3 relevant to many more cancers than previously suspected. 118
The successful development of drugs to target this protein will again be dependent on tumor-specific targeting as MBD3 has recently been shown to play a significant role in maintaining intestinal homeostasis through the interaction with nonphosphorylated c-Jun.119,120 The N-terminal phosphorylation of c-Jun inhibits this interaction and results in upregulation of its target genes in intestinal homeostasis and tumorigenesis. In the absence of MBD3, the intestines are characterized by hyperproliferation and an increased susceptibility to colitis-induced tumorigenesis. Nevertheless, therapeutically targeting MBD3 may prove to have benefits in cancer treatment. Potentially, we could develop drugs that are specific for the role in silencing methylated genes. The proteins of 2 other genes, MBD3L1 and MBD3L2, which show substantial homology to MBD2 and MBD3 but lack a MBD, 121 may give us a clue to how to approach this as MBD3L2 can interact with the MBD2/MBD3/Mi2/NuRD complex in a way that displaces the complex from methylated DNA and reactivate transcription. 122
MBD4
MBD4 is primarily encoded from a single transcript on chromosome 3 62 and stands out from the other MBD proteins due to its interactions with the DNA repair machinery. 123 The protein uses its N-terminal MBD to bind methylated CpG and its C-terminal glycosylase (which is homologous to base excision repair proteins) to mediate repair of the hypermutable CpG, which is susceptible to mismatch formation following either the hydrolytic deamination of 5-methylcytosine to thymine or deamination of cytosine to uracil. Thus, MBD4 plays a key role in maintaining methylated DNA gene regulation and suppressing mutation at CpG sites.
These abilities, along with its interaction with the mismatch repair protein MLH1 (mutL homolog 1), potentially impact chemotherapy as MBD4 has been shown to be required for the maximal apoptotic effects of many of these chemotherapeutic drugs,124,125 a role that may in part be mediated through an interaction with Fas-associated death domain protein (FADD).
126
This indicates that MBD4 function is crucial for normal mutation suppression and optimal response to chemotherapy; however, there is also evidence that its absence can be beneficial, as it has been shown
The significance of MBD4 in cancer is largely indirect as deficiency results in a mutator phenotype and subsequent alterations to
Although it is best known for its repair activities, MBD4 has also been shown to repress
MBD5 and MBD6
MBD5 on chromosome 2 and MBD6 on chromosome 6 were initially 2 uncharacterized proteins that were renamed in 2003 due to them containing a MBD domain.19,63 There is a limited amount of data available on the functions of these proteins. Neither of them is likely to bind methylated DNA, but they do associate with heterochromatin, where they may contribute to the formation or function of heterochromatin. 137 As with other MBP proteins, MBD5 deficiency is linked to a developmental disorder, the 2q23.1 microdeletion syndrome,138,139 but there have been no reports linking either of them to cancer.
HAT_MBD
BAZ2A and BAZ2B
There are 2 proteins in this final group: BAZ2A (also known as TIP5), encoded by the
The main identified role of these proteins is in the epigenetic silencing of ribosomal DNA (rRNA) within the nucleolus.141-144 The BAZ2A protein is a major component of the nucleolar remodeling complex (NoRC) that represses ribosomal gene transcription via histone deacetylation. This NoRC complex associates with the histone deacetylase HDAC1
There are very few reports of these proteins having a role in cancer. It has been shown that overexpression of miRNAs, which can regulate expression of
Methyl-CpG Binding Zinc Fingers
This branch of MBPs is based on a 3–zinc finger motif found in the C-terminus of the Kaiso protein, which is capable of binding a pair of methylated CpG dinucleotides. 27 The protein also binds unmethylated DNA and has a greater affinity for CTGCNA than mCGmCG. 148 The Kaiso protein is the archetypal member of this branch, and the zinc finger motif has been used to identify 2 other members, ZBTB4 and ZBTB38, 149 that are also capable of binding methylated DNA.
Kaiso
Kaiso is encoded by the
Given its multiple functions, it is perhaps not surprising that altered Kaiso activity is a feature of some cancers, particularly given that Kaiso functions as a regulator of the target genes of the canonical and noncanonical Wnt pathways.150,156-158 In Wnt-driven colon cancer cell lines, Kaiso mediates the silencing of the tumor suppressor genes
Much of the current work focuses on the identification of the relevant Kaiso-silenced genes in cancer. Given Kaiso’s ability to repress transcription at both methylated and unmethylated promoters, it is currently difficult to assess how important its mCpG binding role is in cancer. For example, in lung cancers, Kaiso mediates silencing of both the methylated
Beyond its role in transcriptional repression, Kaiso has also been localized to mitotic spindles and centrosomes at different stages of the cell cycle. 163 Altering the ability of Kaiso to locate to these regions resulted in defective centrosomes, whereas overexpressing Kaiso resulted in mitotic arrest, and knocking it down led to elevated proliferation. Given that Kaiso is often mislocalized in cancer, clearly these observations are of high potential significance to cancer; combined with the observation that the Kaiso-deficient mouse is healthy and viable, it makes a potentially attractive therapeutic target. However, given its diversity of roles, great care will need to be taken in the use of such a therapeutic.
ZBTB4 and ZBTB38
The genes for ZBTB4 and ZBTB38 are located on chromosomes 17 and 3, respectively. Unlike Kaiso, these proteins can bind a single methylated CpG
With respect to cancer, there is a single report of ZBTB4 associating with the Sin3/HDAC complex and silencing the
A p53-independent apoptotic role has also been confirmed for ZBTB38 in the mouse through its interaction with CtBP. 166 A role in apoptosis for ZBTB38 is also suggested as it is a substrate for caspase-3, a feature of mitochondrial-driven apotosis. 167 However, the significance of either of these genes in cancer therefore remains largely undetermined.
SRA Domain–Containing Proteins
There are 2 members of this family, UHRF1 and UHRF2, which contain a SET and RING finger–associated (SRA) domain as well as a ring finger motif, PHD finger, and ubiquitin-like domain. The ability of the SRA domain to bind methylated CpG was first recognized in human breast cancer cells
28
as decreased expression and increased promoter methylation of
UHRF1
Prior to the discovery that UHRF1 (also known as Np95 and ICBP90) on chromosome 19 could bind methylated DNA, it had been identified as a protein of many functions. These include regulation of
In human cancer, all of these functions have been shown to be relevant to different types of cancer. The maintenance of methylation plays a significant role in cancers by maintaining promoters in a hypermethylated state. In lung cancer, UHRF1 controls the cell cycle through maintenance of promoter methylation at
The many roles for UHRF1 in cancer have marked it for therapeutic targeting.193,194 However, as for all the MBPs in this review, their wide-ranging functions in multiple tissues again urge caution when targeting them. In this respect, the role played by UHRF1 in maintaining DNA methylation patterns required for progenitor maintenance and self-renewal of somatic tissue is of particular importance. 195
UHRF2
UHRF2 (also known as NIRF or Np97) was identified due to its role in the cell cycle.
168
Its expression is highest in proliferating cells, where it ubiquitinates PCNP,168,196 and its overexpression results in G1 arrest, presumably through an interaction with Cdk2.
197
Any other roles have yet to be described for this protein, but it may play a role in lung cancer cells as UHRF2 expression has been shown to be repressed by let7-a miRNA, which subsequently led to elevation of p21waf1 levels
Summary
This review has highlighted the many and intricate roles that MBPs play in human biology. Primarily, this involvement in gene regulation has led to them being utilized by normal cells to exert transcriptional control but has also led to them being exploited by cancer cells to escape such control. The dramatic effects of suppressing or enhancing their roles in cancer make them eminently suitable targets for therapeutic intervention. However, their many basic regulatory roles in a variety of tissues in the human body also present us with limitations and hazards of any such targeting. For this reason, we now need a much better understanding of the specific changes that these proteins induce in cancer cells to permit their propagation. The hope is that we can identify a subset of targets that are highly specific to cancer cells and so begin to develop a range of novel targeted therapies.
Footnotes
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) received no financial support for the research, authorship, and/or publication of this article.
