Abstract
Introduction:
Biobanks of specimens of human origin have accumulated millions of specimens. Their storage is costly, while many of them may not be useful and should be culled.
Objectives:
Our objective was to develop, pilot test, and evaluate a quantitative culling tool.
Methods:
We developed a culling tool based on a series of parameters with a quantitative score attributed to each. The parameters of the culling tool correspond to different aspects of the value of collections, such as the richness of the associated data, the types of samples, their conservation mode, and regulatory constraints.
Results:
The culling tool was adapted and independently applied by the Foundation for Innovative New Diagnostics and the Biological Resource Center of Institut Pasteur biobanks. The cumulative final score supported evidence-based and standardized decision-making. A “diagnostic” threshold could be established for the “diagnosis” of collections of low value.
Conclusion:
The culling tool is an algorithm developed to assess the value of legacy collections of biological resources of human origin and help establish culling plans. Biobanks can use this culling tool when they periodically assess the value of stored collections and need to decide or advise to cull them, and also when deciding whether to accept requests to host new collections previously stored elsewhere.
Introduction
Biobanks are strategic research infrastructures supporting research in medical and life sciences and biotechnological applications. The immense success of biobanks worldwide in collecting samples has led to a crisis in sustainability, with numbers of samples in store, and hence costs, soaring but distribution rates stubbornly low. 1 Key Performance Indicators of biobanks usually include numbers of samples in stock, numbers distributed, quality control metrics, number of supported scientific projects, and cost recovery as a proxy for sustainability. While the number of samples in stock is not the most relevant in indicating the value of a biobank,2,3 sustainability, discussed since 2008, 4 is the most pressing indicator, as biobank closures become a real possibility. 5
High costs are incurred for energy, maintenance, and space, especially when the biobank is implemented in campuses with limited capacity or when storage is externalized in commercial repositories that typically charge 1500–2000 USD per freezer per month.
Hence, culling of unused or unusable specimens has become critical. The ISBER Best Practices includes a special section on financial sustainability and suggests reducing storage costs by establishing methodologies for consolidation and/or disposal (culling) of stored material. 6
Specimens of human origin may be unusable for contractual reasons, when material deposit agreements explicitly limit the scope of use or when the informed consent template used is not broad enough, or be unusable when national regulations are incompatible with biobank research infrastructure practices. The latter is the case in France, where regulations only recognize the concept of project-specific collections, but not biobank open-ended collections. 7
Culling is relevant to social responsibility (energy consumption) and to operational and financial sustainability 8 (cost recovery). Storing millions of samples that, for any reason, cannot be reused or have low chances of being reused becomes difficult to justify. We developed a quantitative tool to “diagnose” collections or groups of biological specimens to be culled, especially when there is limited storage space or resources. This tool can be adapted to different biobank contexts and integrated into a Quality Management System, with its application described in a standard operating procedure (SOP). It is applicable to biobanks with more than one collection.
The FIND (Foundation for Innovative New Diagnostics) Specimen Bank is the infectious disease biobank of the nongovernmental organization FIND, whose primary objective is to support development and validation of diagnostics for specific infectious diseases. CRBIP (Biological Resource Center of Institut Pasteur) is an institutional biobank whose primary objective is to support research at Institut Pasteur. The focus of research is mostly on infectious diseases, but there are also research groups working on chronic diseases. CRBIP includes both collections of microbial strains and collections of human specimens, the latter preserved at the CRBIP CHIP (Collections for Human Health of Institut Pasteur) Unit. A previously developed ranking tool focused on the prioritization of microbial strains for a given application (e.g., for sequencing and integration into the biobank’s catalogue). 9 Here, we apply an unbiased scoring system to identify the collections of human specimens for subsequent culling. This scoring system has been configured for clinical disease-oriented biobanks, not population biobanks.
Methods
Principles of the culling tool
A “collection” was defined as a set of biological samples of a specific type, collected in the context of a specific project. A collection was scored higher with smaller size; younger age; lower storage temperature; broader informed consent; legal or contractual requirements preventing its destruction; richer associated data; wider scope of fitness-for-purpose; better sample quality control data; higher usage rate; more difficult replacement; better organized data management; more user groups; and more associated publications or patent applications. The usage rate corresponded to the number of samples distributed relative to the number of samples collected. The fitness for purpose was evaluated based on available information on the processing and storage of the samples, biospecimen science literature 10 and information based on actual successful or unsuccessful usage of the samples.
The culling tool, with a score range between −15 and 59, is available as supplementary data (Supplementary Table S1). Negative scores were implemented for parameters corresponding to bad quality of specimens or data to strongly discriminate against such collections.
FIND and CRBIP independently adapted the culling tool to their individual contexts. The culling parameters, the options for each parameter, the corresponding scores, and the rationale for the options and the scores are shown in Table 1 for both FIND and CRBIP.
Parameters and Scores of the Culling Tool

The parameters are scored for each collection.
LN, liquid nitrogen; BIMS, Biobank Information Management System; RDT, rapid diagnostic test; NAAT, nucleic acid amplification test; IHC, immunohistochemistry.
Some of the parameters of the culling tool can be used as qualitative eliminatory criteria for a collection, and in this case, a user would not need to go to the effort of calculating a score for that collection. During the development phase and first implementations of the culling tool, described in this article, we calculated the scores for all collections to obtain comprehensive data and to evaluate the impact on the performance of the culling tool of including or removing collections on the basis of eliminatory criteria.
Implementation by FIND
FIND applied the culling tool to 174 collections, making decisions separately for each of its four strategic disease groups: tuberculosis, malaria, hepatitis C, and fever syndromes. Specimen types included serum, plasma, urine, sputum, saliva, whole blood, DNA, and culture isolates. For FIND, the range of the score values, based on the applied parameters, was from 0 to 41 (Table 1).
After the culling tool scores were calculated for all collections, FIND applied a set of eliminatory criteria: inability to find the patient informed consent form (“no informed consent”), unsatisfactory associated data (“only demographic information”), and missing data about the age of the collection (“unknown age”). Collections, failing any one of the above criteria, were tagged for elimination. Another eliminatory criterion was examined: the expiry of the maximum allowable storage duration, as provided by the informed consent.
For each disease group, we employed a k-means clustering approach (Version 2502, Microsoft 365, 2025), which is a method that aims to group a number of observations (scores) into clusters, in which each observation (score) belongs to the cluster with the nearest mean. For this, we divided the final scores into five categories: near average, slightly below average, well below average, slightly above average, and well above average. A collection was considered to have failed the evaluation if it fell into the “well below average” category. First, we selected five arbitrary centroids. Then, we applied the ABS() function to calculate the absolute distance between each collection score and the five centroids. Next, we used a combination of MATCH() and MIN() functions to identify the closest centroid to each collection score. Afterwards, we calculated the actual centroids using the AVERAGEIF() function. We repeated the process to determine the true absolute distance between each collection score and these new centroids and identified the nearest centroid using MATCH() and MIN() again. Finally, each collection score was assigned to one of the five score groups based on its proximity to the actual centroids.
For FIND’s biobank, distribution is the most important indicator of value, with specimens that are of interest to diagnostic manufacturers having the highest value. All collections tagged for elimination, based on either the eliminatory criteria or the quantitative approach, were evaluated based on their usage rate before making final culling determinations. The whole process is shown in Figure 1.
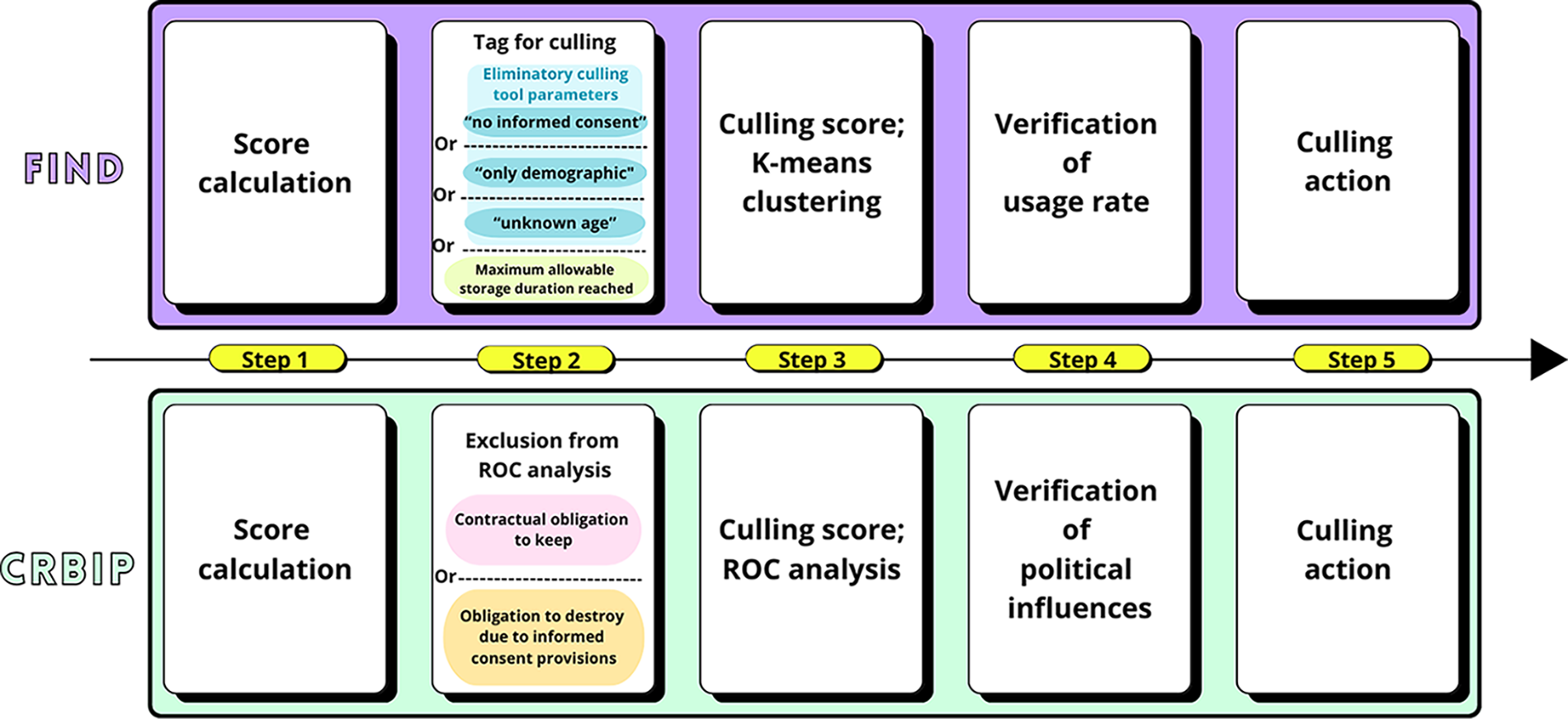
Schematic representation of the processes that FIND and CRBIP followed to apply and evaluate the culling tool. CRBIP, Biological Resource Center of Institut Pasteur; FIND, Foundation for Innovative New Diagnostics.
We performed frequency analysis for the different score values in GraphPad Prism version 10.4.2. This frequency analysis was performed for all 174 collections.
Implementation by CRBIP
A total of 198 different collections of the CRBIP CHIP Unit were evaluated. These collections corresponded to the following disease areas: tuberculosis, HIV, neurological infections, listeriosis, leukemia, COVID-19, influenza, familial adenomatous polyposis, Alzheimer’s disease, malaria, Verneuil’s disease, tick bites, papillomavirus infections, head and neck cancer, anosmia, stomatodynia, or infectious syndromes. They also included healthy donor collections, mostly used as matching controls. Specimen types included whole blood, buffy coat, PAXgene RNA blood, serum, plasma, PBMCs (peripheral blood mononuclear cells), cerebrospinal fluid, saliva, urine, different types of swabs, gastric aspirates, stool, different types of biopsies, DNA, and RNA.
The culling scores were calculated for all 198 collections, and a frequency analysis for the different score values was performed in GraphPad Prism version 10.4.2. For CRBIP, the range of the total possible score values, based on the applied parameters, was from −15 to 58 (Table 1).
We then removed 31 collections from the analysis to which a legal obligation applied: either a contractual obligation to keep the collection or an obligation to destroy the collection based on informed consent conditions (maximum allowable time of storage). The whole process is shown in Figure 1.
A committee, including the CRBIP quality manager and curators, independently examined all the information related to the remaining 167 collections, without seeing the scores of the culling tool, and came to independent conclusions and recommendations as to whether to cull or not. The committee was asked to provide recommendations excluding “political factors” (see below). These recommendations were considered as “true positive” in the sense of “cullable” and “true negative” in the sense of “non-cullable.”
A receiver operating characteristic (ROC) analysis was performed with AnalyseIT version 6.15, using the Wilcoxon-Mann-Whitney method for estimation of the area under curve (AUC) and the DeLong method for standard error and confidence interval estimations and with a type I error of 0.05. 14 We also used the online tool easyROC version 1.3.1 (http://biosoft.erciyes.edu.tr/app_direct/easyROC/) to obtain the performance characteristics for different threshold values. The ROC analysis is used to show the performance of a binary classifier model (in our case, “cullable” vs. “non-cullable”) at different threshold levels (in our case, different score thresholds).
Before final culling decisions were made, “political factors” linked to the status of the principal investigator (PI) in the institute and/or to an explicit request from the PI going against the culling tool-based recommendation were considered.
Results
Results of the application of the culling tool at FIND
FIND’s adaptation of the Culling Tool resulted in a streamlined version with only 9 variables, producing final scores from 13 to 32 (Table 1). FIND applied the tool to a total of 174 collections. After the culling tool scores were calculated, 18 collections were tagged for elimination, based on the k-means clustering analysis. When the eliminatory criteria were also applied, a total of 53 collections were tagged for elimination. Overlap between criteria brought the number of collections to 45. The average usage rate across the FIND Specimen Bank was 8%, and it was decided not to cull collections of which the usage rate exceeded 8%, even if these collections were tagged for elimination. Hence, the final number of collections to cull was narrowed to 38 collections (Table 2). This was equivalent to approximately 70,000 samples.
Results of the Application of the Eliminatory Criteria (in Italics) and the Culling Tool Scores, in Terms of Numbers of Collections at FIND
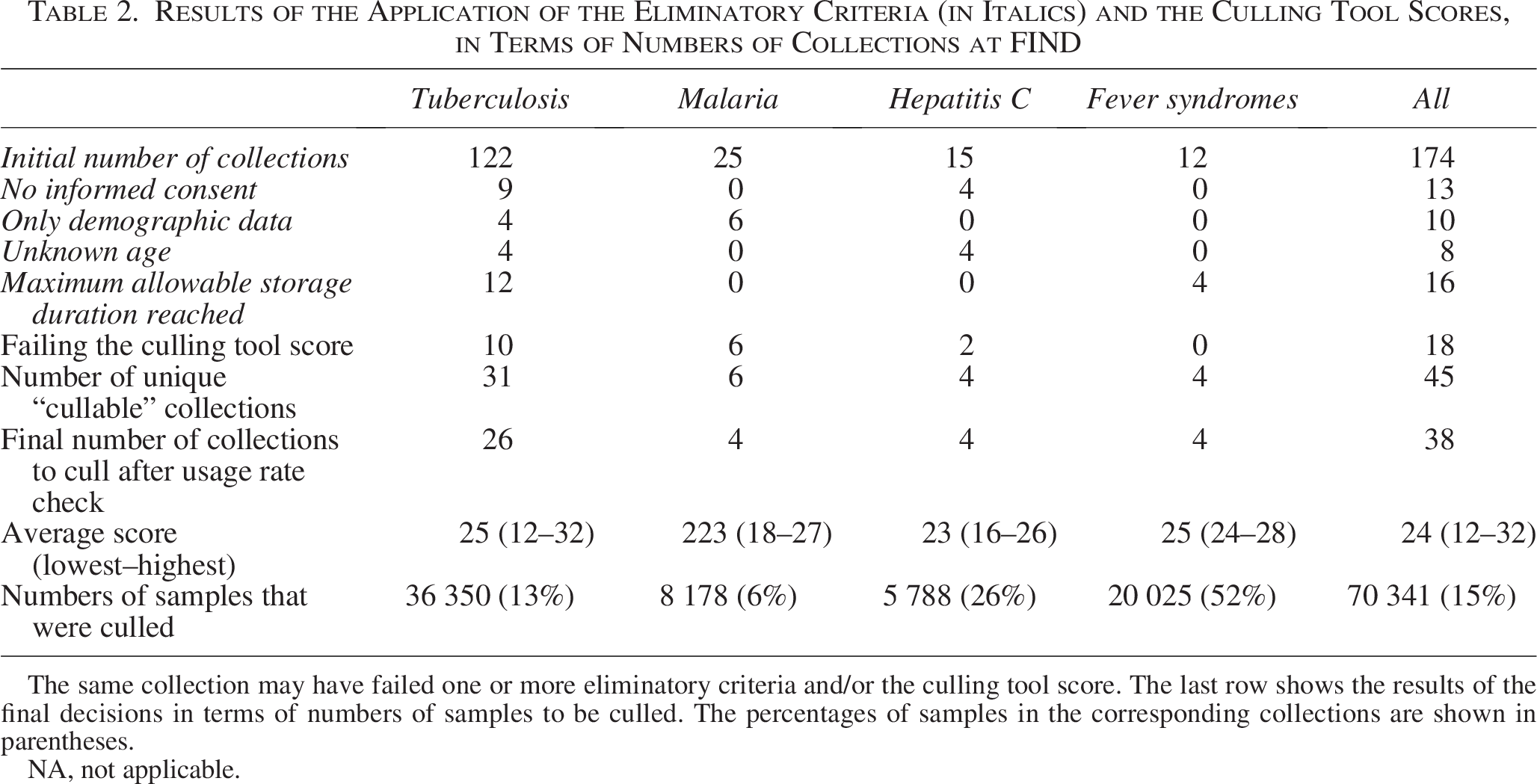
The same collection may have failed one or more eliminatory criteria and/or the culling tool score. The last row shows the results of the final decisions in terms of numbers of samples to be culled. The percentages of samples in the corresponding collections are shown in parentheses.
NA, not applicable.
A frequency analysis for the score values of all collections showed an uneven, non-Gaussian distribution (Fig. 2A). This analysis indicated that if, for example, there was a management decision to cull 20% of the FIND Specimen Bank collections, then all collections with a score lower than 22 should be culled.
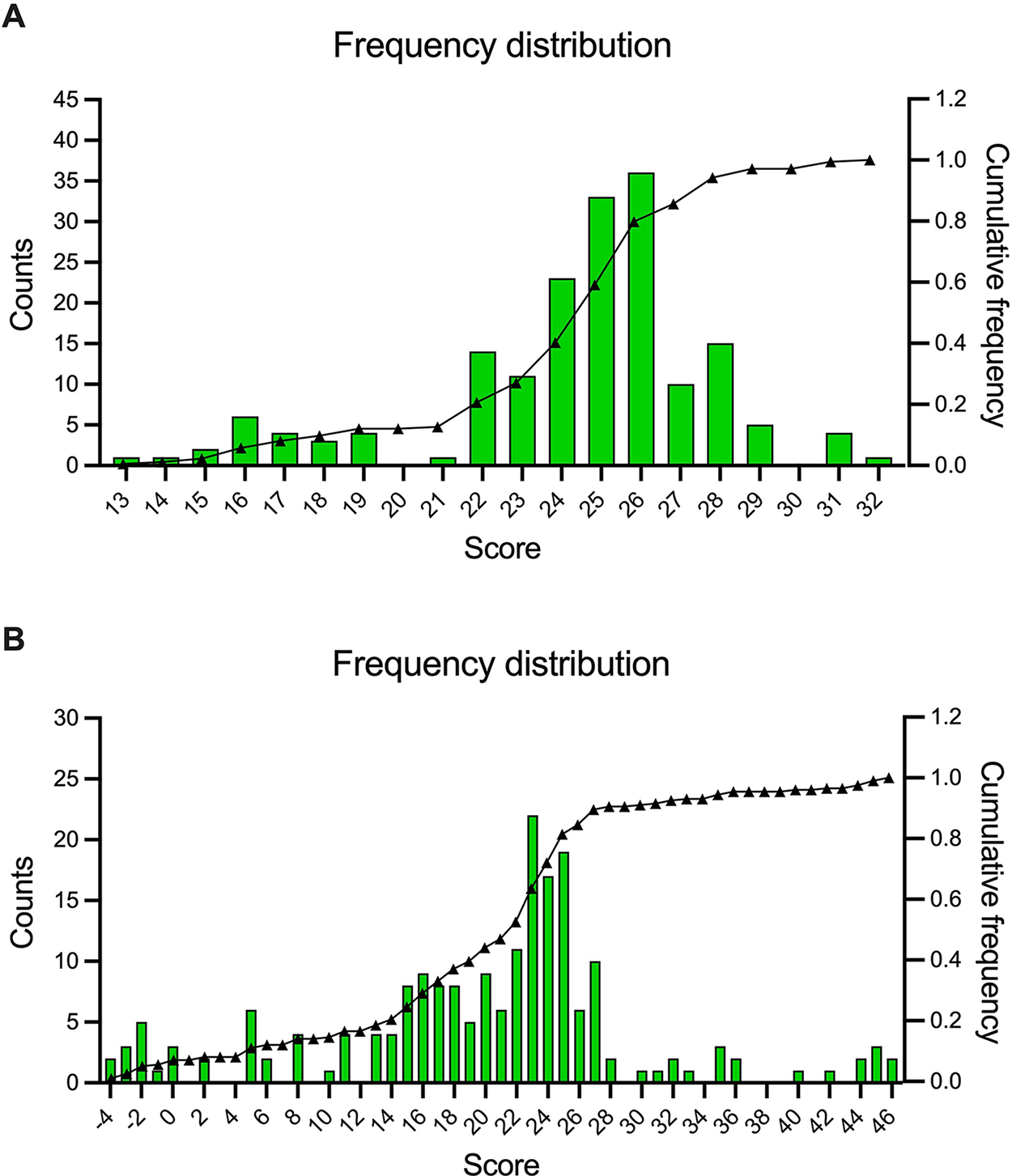
Frequency plots of the culling tool scores that were obtained for all FIND
Results of the application of the culling tool at CRBIP-CHIP
The frequency analysis of the score values showed a non-Gaussian distribution, with final scores from −4 to 46 (Fig. 2B). The majority (72%) of the collections had intermediate scores, between 10 and 30, while 5% had scores higher than or equal to 40 and 15% had scores lower than or equal to 10. This analysis indicated that if, for example, there was a management decision to cull 20% of the CRBIP CHIP collections, then all collections with a score lower than 14 should be culled. The frequency analysis also showed a cluster of collections with very low scores. These collections corresponded to HIV, tuberculosis, and malaria disease areas, and also to healthy donors. All of them were older than 10 years, had no associated clinical data, or were not included in any data management system, and interestingly, all dated to before the implementation of a Quality Management System.
ROC analysis of the culling scores at CRBIP-CHIP
Our purpose was to determine a threshold value of the culling tool that would have a good “diagnostic” value in identifying “cullable” collections when the tool would be applied periodically by biobank managers as an SOP. In other words, we wanted to identify a cut-off below which a collection is of such low value that it can be culled with confidence.
ROC analysis of the scores of the 167 collections gave an AUC of 0.79 (0.71–0.87) (Fig. 3). The threshold to be applied is a management decision. A threshold of 20 corresponds to a sensitivity (probability that the culling tool correctly identifies a collection to be culled) of 77%—hence, the tool will miss 23% of the “cullable” collections. The threshold of 20 corresponds to a specificity of 78%—hence, there will be approximately 22% of the collections identified as “cullable” that would be worth keeping. A threshold of 16 corresponds to a sensitivity of 62% and a specificity of 90%. At the cut-off of 20, which was the cut-off giving the optimal combination of sensitivity and specificity, we achieved the removal of 65 collections, equivalent to approximately 60,000 specimens, and an approximate cost saving of 25,000 USD per year. CRBIP was congratulated for this result by the Environmental, Social, and Governance Department of the Institute.
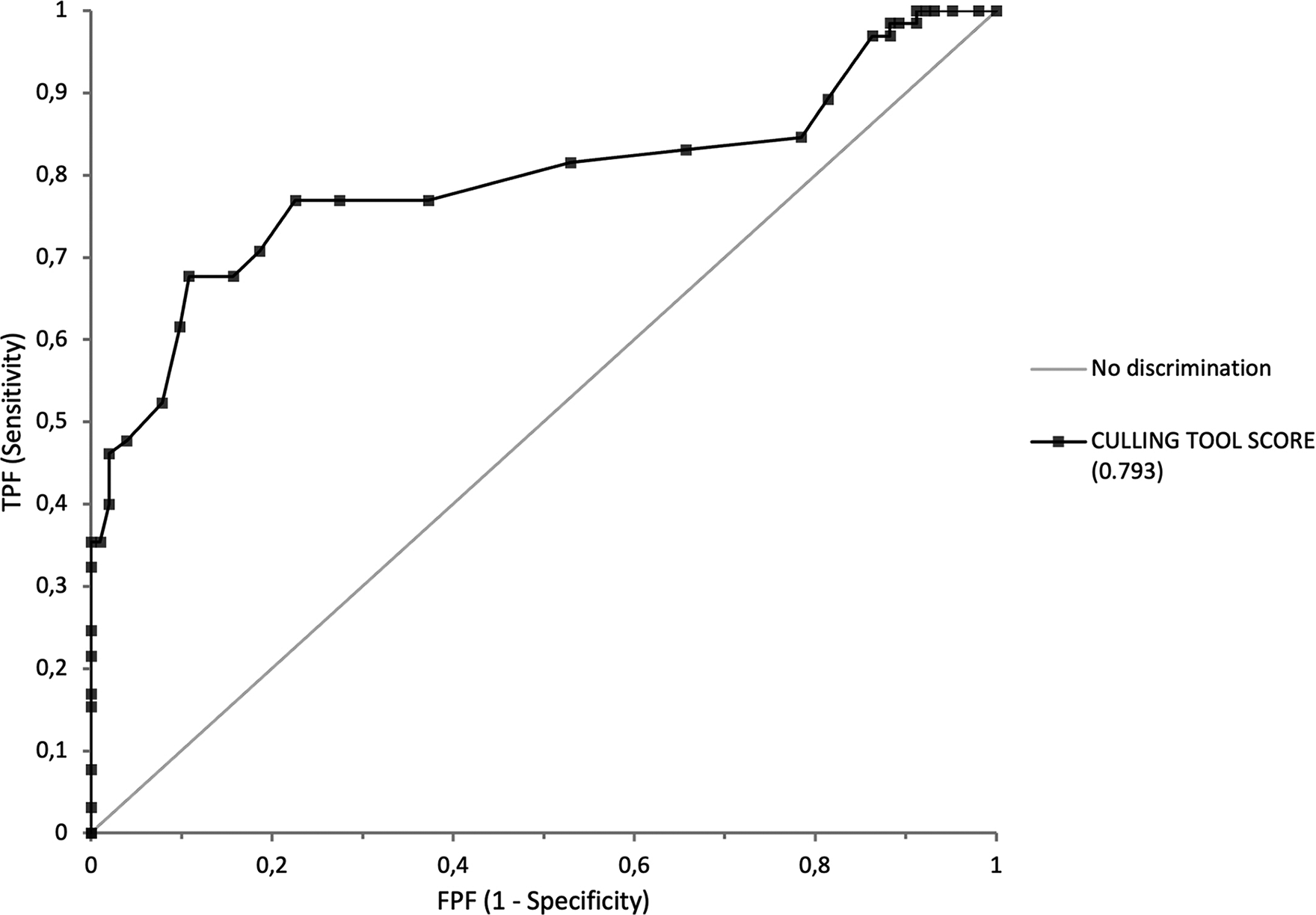
ROC curve analysis of the CRBIP culling tool scores, evaluated against independently made expert recommendations. A collection was considered to be “positive” if it was “cullable.” This ROC analysis corresponds to the 167 CRBIP collections, after removal of the 31 collections with contractual or informed consent-based obligations. The independently made expert recommendations corresponded to the “true positive” and “true negative” collections. The ROC curve is the plot of the true positive fraction (TPF) against the false positive fraction (FPF) at each threshold setting. CRBIP, Biological Resource Center of Institut Pasteur; ROC, receiver operating characteristic.
As explained in the Methods section, we excluded the 31 collections to which a contractual obligation applied. In these cases, the decision to keep or cull is not based on the objective quality criteria of the culling tool; hence, reducing its diagnostic performance. Indeed, if we did not remove these collections and repeated the ROC analysis of the scores of all 198 collections, the “diagnostic” value was slightly lower, with AUC = 0.74 (0.66–0.82). Interestingly, there were four collections with scores between 23 and 25, corresponding to healthy donors, that were removed because of the limited scope of the corresponding informed consent forms.
Discussion
Long-term storage in biobanks incurs significant costs linked to space and energy consumption. While for prospective collections, biobanks can preemptively balance the acquisition of new collections and expected demand, institutional strategies may change over time, and significant volumes of historical collections may remain unused. In this case, “stock is waste.” Regular evaluation and culling are therefore essential for sustainability; however, deciding which samples to discard remains challenging, as most biological materials of human origin are nonrenewable. The scoring that we applied was strongly weighted against poor biospecimen quality and in favor of rich and well-managed annotated data. We saw that different biobanks have different distributions of culling scores of their collections; these distributions could be Gaussian, multimodal, or flat. The score distribution is expected to have a direct impact on the “diagnostic” performance of the culling tool. A limitation of the culling tool may be that sensitivity and specificity can never be 100%, so utilization of the tool for “automatic” culling decisions would not be possible. Culling decisions should always be vetted by a decision-making body, according to a culling SOP.
Apart from the financial, logistical, and ecological considerations, there is also an ethical component to culling: it can be argued that it is unethical to maintain collections where the quality and fitness for purpose of the specimens is compromised and knowingly distribute them for research. Even if the donors of the specimens had donated them with the assumption that they would be used in research, the primary responsibility of the biobank is to provide resources fit-for-purpose.
We have also shown that political considerations may interfere and might lead to the retention of collections that should otherwise be culled. For example, at CRBIP, we were only able to obtain authorization for partial destruction of a fully “cullable” collection. Conversely, we have seen otherwise valuable collections with high scores being removed because of informed consent conditions requiring destruction. The importance of as broad as possible informed consent cannot be overstated.
Before any disposal, the PI’s opinion and approval were requested. In most cases, the CRBIP culling tool-based assessment was in line with the judgement of the PI of the project-driven collections. However, we have encountered situations where this was not the case. For example, in the case of one collection, the PI decided to resume his research after a break of several years and wanted to use the historical collection, despite its age and our reservations about its fitness for purpose. Another scientist asked us to postpone the destruction of a collection to give her time to finalize publications in the event of editor requests for further analyses.
Another aspect is the impact of the contractual provisions on any culling decisions. While biobank managers cannot change regulations, they can and should be involved in the contractual discussions at the beginning of new collection projects. Their input concerns not only technical and pre-analytical aspects but also contractual and/or protocol-related aspects of duration of storage and secondary use.
The Culling Tool represents an innovative and quantitative approach to developing strategies for culling collections of human specimens. In this work, we evaluated its applicability and performance in two different biobanks. For this first evaluation, we went to the effort of calculating the scores of all collections to better evaluate the tool. In the future, the procedure that a biobank is expected to follow when applying this or a similar culling tool would include elimination, or tagging for elimination, of collections based on predefined eliminatory criteria, followed by calculation of the culling scores for the remaining collections only, especially when resources for evaluation operations are limited. Each biobank can define eliminatory criteria that best correspond to its context; for example, easy replaceability could also be a criterion. Finally, application of the culling tool may not be appropriate for collections for which an evaluation should be made at the level of each individual sample, based on criteria such as specific bioanalytical results, even if in some cases such an approach might introduce selection bias in the collection. This would be the case, for example, if only specimens that are positive for antibodies against a specific pathogen should be kept.
This Culling Tool is not intended to become a standard. Each biobank is expected to adapt it to its specific circumstances, adding or removing parameters, adjusting the weight of the scores and/or the process of implementation. If statistical calculations appear difficult, a simple way to define a culling score threshold would be simply based on visual examination of the score distribution.
The conclusions of a recently published guideline on the evaluation of collections 15 are quite compatible with the culling tool described here. The culling tool guides culling decisions in biobanks other than population biobanks with a single population cohort and represents a scalable and adaptable solution supporting sustainable biobank management. It can also be used in other contexts, such as in quality management as a quantitative indicator related to the performance of specific sites (highlighting site-specific collections with systematically lower scores) or supporting decision-making on which collections to include in national catalogues (if catalogues should include only collections with higher scores). Finally, it can support sustainability by showing to funders that the biobank follows a structured and evidence-based approach to stock management.
In summary, we provide a flexible and proven approach to the challenge of identifying legacy collections of biological resources of human origin and establishing culling plans.
Authors’ Contributions
I.E.I.: Applied the tool at FIND, did data analysis, and contributed to the writing of the article; V.P.: Applied the tool at CRBIP, did data analysis, and contributed to the writing of the article; W.F.: Supervised the work at FIND; E.R.: Supervised the work at CRBIP; C.M.: Participated and coordinated the independent committee at CRBIP; M.L.F.: Critically reviewed the article and contributed with data analysis; E.A.: Critically reviewed the article; and F.B.: Developed the culling tool, did data analysis, and wrote the article.
Footnotes
Author Disclosure Statement
No competing financial interests exist.
Funding Information
No funding was received for this article.
Supplemental Material
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
