Abstract
This study aimed to compare monoculture and mixed rubber plantations in terms of their soil bacterial and fungal composition. An Illumina MiSeq sequencing analysis was performed to investigate the composition and diversity of the soil bacterial and fungal communities among three different rubber (
Keywords
Introduction
Rubber (
Microbial activity plays a key role in nutrient cycling and amelioration of plant stresses and is responsible for a broad range of the ecological functions of soil (Singh et al., 2016). The soil microbial diversity is a critical marker of soil quality (He, Zheng, Chen, He, & Zhang, 2008; Kirk et al., 2004). It was reported that soil microbial diversity is affected by a variety of environmental factors (Balser, Kinzig, & Firestone, 2002; Schimel, 1995). Previous findings revealed that abiotic soil nutrients drive the soil bacterial communities on RPs in Hainan (Lan, Li, Wu, & Xie, 2017a). Soil pH is another important factor that can affect the soil bacterial community structure (Krashevska, Klarner, Widyastuti, Maraun, & Scheu, 2015). Seasonal changes are also main drivers of the bacterial composition of RPs and other tropical forests (Lan, Li, Lesueur, Wu, & Xie, 2018). In addition, other factors controlling the distribution of soil microorganisms are being studied (Rousk et al., 2010). For instance, it is important to know whether different planting patterns of RP (monoculture and mixed plantations) affect the composition and diversity of soil microbes.
Monoculture has often been criticized for reducing biodiversity, causing widespread outbreaks of pests and diseases (Erskine, Lamb, & Bristow, 2006; Lamb, Erskine, & Parrotta, 2005) and not supporting numerous conventional ecosystem services (Singh, Rai, Banyal, Chauhan, & Singh, 2018). Thus, there is an emphasis on growing mixed plantations rather than monoculture. This includes an expanded yield, better natural assurance, biodiversity protection, and remodeling (Montagnini, Gonzales, & Porras, 1995). We aimed to test if there were any differences in the composition and diversity of soil microorganisms between monoculture and mixed RPs. In this study, we examined the bacterial and fungal composition and diversity of monoculture and mixed RPs by utilizing an extensive throughput Illumina MiSeq sequencing analysis. The objectives of this study were (a) to identify the soil microbial composition and changes in community structure among monoculture, Mixed 1 and Mixed 2 plantations and (b) to understand the main factors that affect the microbial community. We aimed to test whether soil microbial diversity in mixed RPs was higher than in a pure RP.
Methods
Study Site
Hainan Province (18°10′ −20°10′N and 108°37′ −111°03′E), with a territory of 33,920 km2, is located in the south of China. Hainan Island is one of the most important hot spots of biodiversity conservation in the Indo-Burma region. Hainan Island’s tropical rainforests are situated at the northern edge of tropical Asia (Zhu & Zhou, 2002) and are recognized for their high biodiversity (Zheng, Cour, & Zou, 2002). This island has a tropical rainstorm environment with a wet period from May to October and a dry period from November to April. The annual precipitation was recorded more than 1,600 mm and the precipitation was irregularly distributed between wet and dry seasons (Francisco-Ortega et al., 2010). The lowest mean monthly temperature was recorded in January (16°C) and the highest in July 27.5°C. Precipitation frequently occurs on the eastern part of Hainan Island because of storms in the Pacific Ocean, while the western part of the island is dry (Xu, Tan, Wang, & Gai, 2016).
Sample Collection
The study site was located in Wenchang area, northeast of Hainan Island. We selected three plots (20 m × 20 m) on three RPs, that is, a monoculture RP, a Mixed 1 RP (
Soil samples were randomly collected from each RP. Six soil samples were randomly gathered within each plot by using a 5 cm diameter stainless steel cylinder. The soil samples were then separated into three sections; the first was sieved throughout a 2 mm net instantly and stored at 4°C until analysis of MBC; the second one was air-dried and passed through a 0.25 mm sieve for SOM, soil pH, and TN; and the third one remained stored at −80°C for DNA extraction. The soil pH was estimated in a 1:1 soil:water mixture. The soil moisture was determined gravimetrically. Total phosphorus (TP) was digested by molybdenum-antimony, and measurement was performed by spectrophotometric method (Soil Science Society of China, 2000). Total potassium (TK) content was measured by flame photometry. MBC was examined by chloroform fumigation and an extraction technique utilizing the correction factor 0.35 (Shen et al., 2014). The forests and soil properties are shown in Table 1.
Soils Characteristics of the Three Rubber Plantations in Wenchang, Hainan China.

DNA Extraction, PCR Amplification, and Illumina MiSeq Sequencing
Microbial DNA was extracted from 5.0 g of soil using the E.Z.N.A.® Soil DNA Kit (Omega Biotek, Norcross, GA, USA) according to the manufacturer’s protocols. For bacteria, the V3–V4 hypervariable regions of the bacterial 16S rRNA gene were amplified with primers 338 F (5′-ACTCCTACGGGAGGCAGCAG-3′) and 806 R (5′-GGACTACHVGGGTWTCTAAT-3′) by a thermocycler polymerase chain reaction system (Xu et al., 2016; GeneAmp 9700, ABI, USA). Fungal Internal Transcribed Spacer (ITS) Region 1 was amplified using the ITS1F (5′-CTTGGTCATTTAGAGG AAGTAA-3′) and ITS2 (5′-GCTGCGTTCTTCATCG ATGC-3′) primer pair (Kerfahi, Tripathi, Dong, Go, & Adams, 2016). Polymerase chain reactions (PCRs) were conducted in triplicate and the tests were conducted using a 20 μL blend consisting of 4 μL of 5 × FastPfu Buffer, 2 μL of 2.5 m dNTPs, 0.8 μL of the sample each primer (5 μM), 0.4 μL of FastPfu Polymerase, and 10 ng of template DNA. Amplicons were separated on 2% agarose gels and purified utilizing the AxyPrep DNA gel extraction kit (Axygen Biosciences, Union City, CA, USA) as indicated by the manufacturer’s instructions and measured utilizing QuantiFluor™—ST (Promega, USA). Purged amplicons were mutually equimolar and combined end-sequenced (2 × 250) on an Illumina MiSeq stage was conducted as indicated by standard conventions.
Bioinformatics and Statistical Analyses
Basic FASTQ records were demultiplexed and separated utilizing QIIME (version 1.17). The adjusted groupings were collected into operational taxonomic units (OTUs) characterized through 97% comparability (Stackebrandt & Goebel, 1994) utilizing the CD-HIT-OTU driver (Wu, Zhu, Fu, Niu, & Li, 2011). The phylogenetic attachments of all 16S and 18S rRNA gene arrangements were examined by the Ribosomal Database Project classifier of the SILVA (SSU117/119) 16S and 18S rRNA catalog utilizing a confidence threshold of 70%. We calculated the exposure proportion using Good’s technique. The Shannon diversity index and nonparametric measures of genus richness (Chao1) were designed for each illustration using Mothur (Schloss et al., 2009). Principal coordinate analysis (PCoA) based on the Bray-Curtis distance matrix of family composition was achieved. To check the microbial community’s similarity index, the similarities were statistically analyzed to test whether there was any significant variance among the RPs.
The seven ecological variables included in the analysis were the type of RP (1: monoculture RP; 2: Mixed 1 RP; 3: Mixed 2 RP), soil organic matter (SOM), TP, total nitrogen (TN), TK, water content (WC), and soil pH. To identify relationship between microbial communities and ecological factors, a redundancy analysis was conducted. Using the
Results
Taxonomic Compositions
There were a total of 1,896 OTUs belonging to 170 families and 7 phyla in the soil fungal communities from 18 soil samples (6 monoculture RPs, 6 Mixed 1 RPs, and 6 Mixed 2 RPs), among which there were a total of 1,758 OTUs belonging to 196 families and 23 phyla in the soil bacterial communities. At the phylum level, there was no difference in soil bacterial compositions between the monoculture and mixed RPs (Figure 1(a)). For fungal structure, the most abundant phylum in soils of monoculture, Mixed 1, and Mixed 2 RPs was Ascomycota. The relative abundance of Ascomycota was higher in monoculture soil than in the mixed RPs; however, the relative abundance of Basidiomycota was lower in the monoculture (Figure 1(b)). Mixed 1 and Mixed 2 RPs were fundamentally similar.

Phylum compositions of soil bacterial (a) and fungal (b) communities in the three rubber plantations (monoculture, Mixed 1, Mixed 2 RPs) *
Bacterial Diversity
Diversity estimations at the OTU level of the 16S and 18S rRNA gene banks of three communities from Illumina MiSeq sequencing analysis are shown in Figure 2. In terms of soil bacterial community, the number of OTUs in the monoculture RPs was significantly higher than in the Mixed 1 and Mixed 2 RPs, whereas Mixed 2 was higher than Mixed 1. The Chao1 diversity and Shannon diversity indexes showed that monoculture was significantly higher than the Mixed 1 RP. The number of species, Chao diversity and Shannon diversity indexes of fungal communities had no significant differences among the monoculture, Mixed 1, and Mixed 2 RPs.
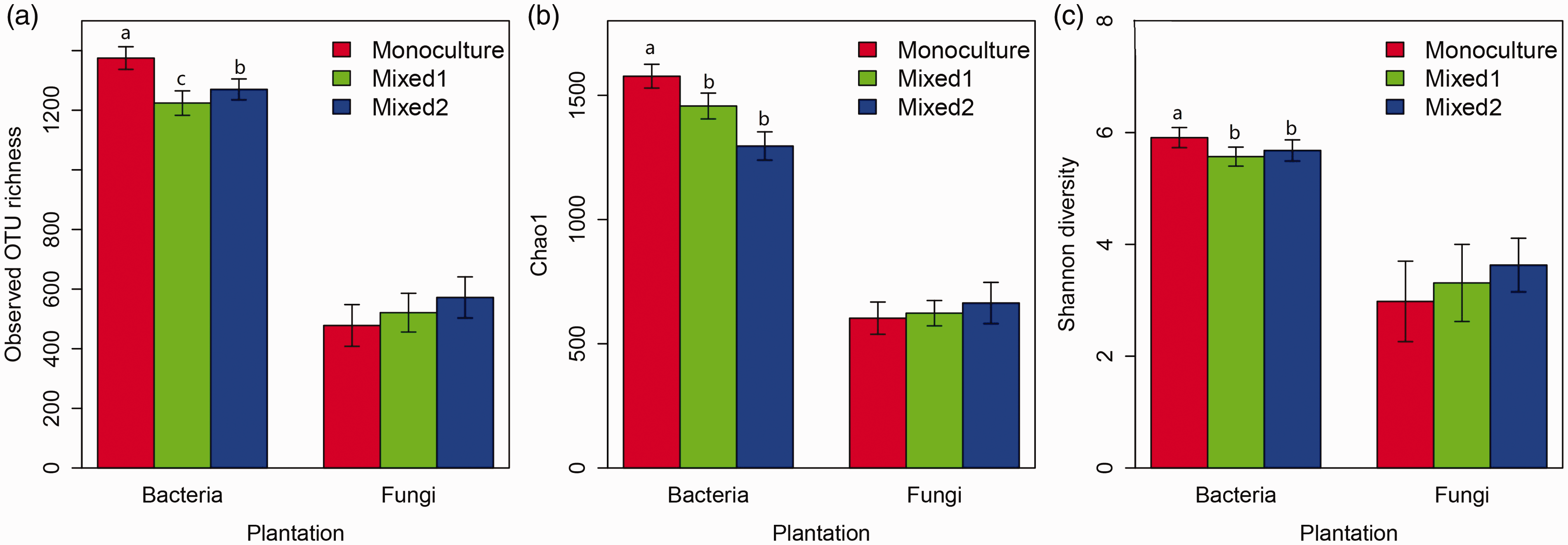
Observed OTU richness and Chao1 and Shannon diversity of soil bacteria (a–c) and fungi species level communities in the three rubber plantations (monoculture, Mixed 1, Mixed 2 RPs). a,b,c, p<0.05
PCoA of the bacterial and fungal communities at the OTU level of the three RPs showed some spatial heterogeneity of bacterial and fungal taxonomic compositions between the soils of the monoculture and mixed RPs (Figure 3). For bacteria, the analysis of similarity (ANOSIM) results showed that there was no significant difference in taxonomic composition (

PCoA of bacterial (a) and fungal (b) communities of the three rubber plantations. Points those are closer together on the ordination show communities that are more similar (monoculture, Mixed 1, Mixed 2 RPs). PCoA = principal coordinates analysis.
Affecting Factors
Compared to Mixed 1 and Mixed 2, the WC was higher in the monoculture. Furthermore, the soil pH in the monoculture was significantly higher than in the Mixed 1 and Mixed 2 (Table 1). In addition, SOM, TN, and TP did not differ between the monoculture and mixed plantations. The highest TK was measured in Mixed 1 RPs, followed by monoculture and Mixed 2 RPs. Seven variables and the abundance data of the bacterial and fungal dominant family were used for redundancy analysis. For bacteria, this variable combination explains 44.95% of the total variance of family abundance (Figure 4(a)). TN, TK, pH, and WC described 22.47%, 18.94%, 15.23%, and 15.18% of the total variance, respectively. Additional factors including Rubber Forest, TP, and SOM on the bacterial community composition had relative little effect, accounting for 6.53%, 9.74%, and 11.19% of the variance, respectively. For fungi, this variables combination explained 27.77% of the total variance of the family abundance (Figure 4(b)). The pH and WC explained 15.14% and 8.96% of the total variance, respectively. Additional factors, that is, TN, Rubber Forest, TP, TK, and SOM for fungal community composition had little effect, describing only 5.50%, 5.49%, 3.61%, 3.15%, and 2.96% of the variance, respectively.
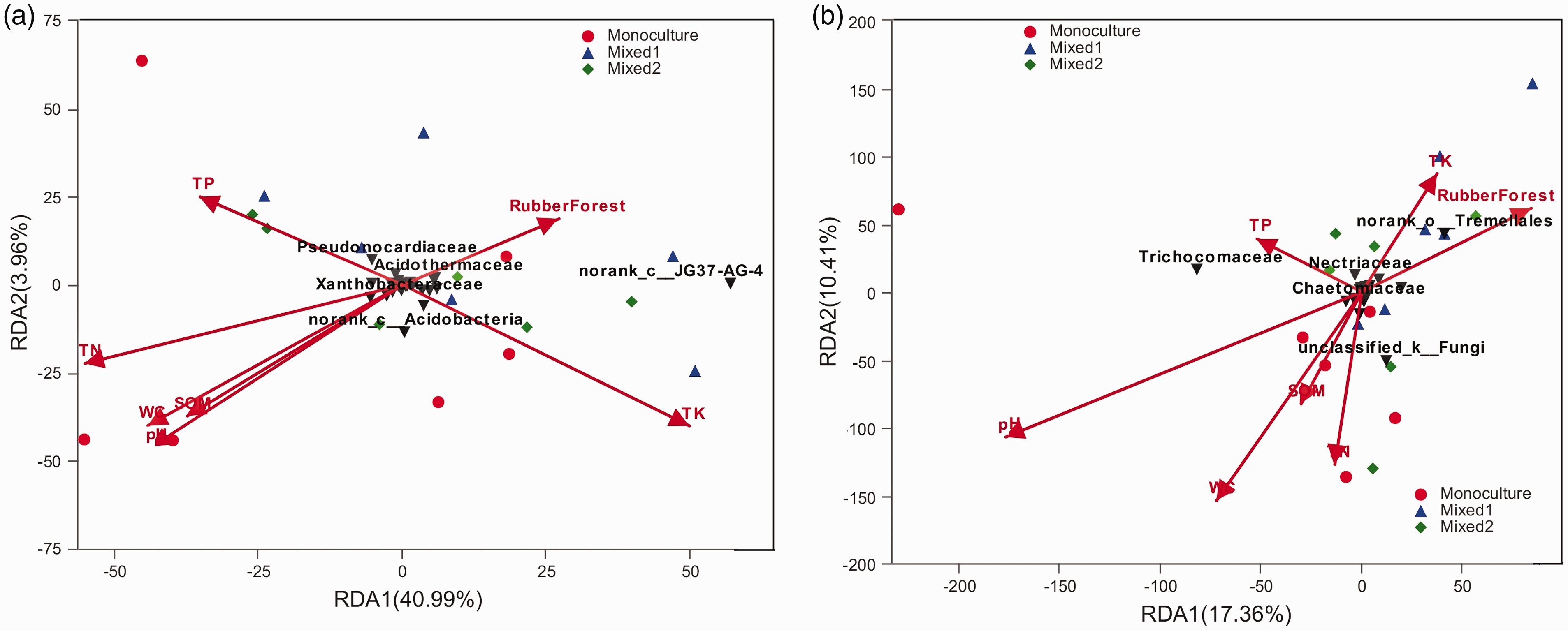
Redundancy analysis ordination of the soil samples and family compositions of bacterial (a) and fungal (b) communities across the three rubber plantations (monoculture, Mixed 1, Mixed 2 RPs). SOM = soil organic matter; TN = total nitrogen; TP = total phosphorus; TK = total potassium; WC = water content; pH = soil pH. Different values were assigned to the three rubber plantations (1: monoculture; 2: Mixed 1; 3: Mixed 2 RPs), thus the direction of the arrow of the rubber forest points to the mixed forest.
Discussion
Our results showed that the bacterial composition of the soil from the three RPs was basically similar. However, there was a significant difference in fungal communities among the three RPs at both the phylum and OTU level. The mixed RPs, that is, Mixed 1 and Mixed 2 RPs, were very similar in terms of phylum composition (ANOSIM results,
In our investigation, the species richness, Chao, and Shannon diversity of the bacterial communities of the monoculture were higher than the Mixed 1 and Mixed 2 RPs, whereas all diversity indexes of the fungal communities were comparable between the monoculture and mixed RPs. These results showed that mixed RPs did not result in a higher diversity of bacteria. A study conducted in the Amazon showed that the local taxonomic and phylogenetic diversity of the soil bacteria was higher in agroecosystems compared to natural ecosystems (Rodrigues et al., 2013). A previous study observed that soil bacteria in a bamboo forest and a fir forest showed a prominent diversity, but with a lower total sequencing number than from native forest and mixed forest. A high variety of bacterial groups in monoculture rubber forests of agricultural practices would result in a considerable variety of soil microscopic organisms (Kerfahi et al., 2016; Lan et al., 2017, 2017a, 2017b). Previous studies have shown that the percentage of Gram-negative bacterial phospholipid fatty acids in jungle RPs was lower than in monoculture RPs (Krashevska et al., 2015). Our results further confirmed that monoculture RPs have a higher bacterial diversity than mixed forests. In addition, our results also revealed that there were no significant differences in fungal diversity of soils between monoculture and mixed RPs. Our findings are similar to previous study conducted in the Jambi Province of southwest Sumatra, Indonesia (Krashevska et al., 2015), which showed that the total fungal phospholipid fatty acids of jungle RPs and monoculture RPs were comparable.
Our results showed that soil nutrition (such as TN and TK) and soil pH are the main drivers of the bacterial composition. However, soil pH and WC are the main drivers of the fungal composition. Our results are in line with a previous study in Danzhou, Hainan Island, which revealed that soil nutrients explained 43.05% of the total variance of soil bacterial composition in RPs (Lan et al., 2017a). In fact, numerous studies have shown that soil chemical properties, including soil pH, nitrogen, and carbon proportions together correspond with the bacterial community’s composition and diversity (Lee, Barbier, Bottos, Mcdonald, & Cary, 2012; Smith, Barrett, Tusnady, Rejto, & Cary, 2010; Van Horn et al., 2013; Zeglin et al., 2011). Among the soil properties, pH is an imperative variable that impacts both soil bacterial communities and their alpha diversity (Fierer & Jackson, 2006). In our study, soil pH in the monoculture was relatively higher than in the mixed RP, which resulted in higher diversity among bacterial community in the monoculture than mixed RP. Very similar results were found in Hainan (Lan et al., 2018; Lan et al., 2017a) and Xishuangbanna (Lan et al., 2017b). Increased pH presumably is a key factor favoring nutrient capture in rubber production systems (Krashevska, Klarner, Widyastuti, Maraun, & Scheu, 2015). A previous study also confirmed that soil pH was the main driver of soil microbial composition in RPs (Krashevska et al., 2015).
Implications for Conservation
Our study revealed that there was a significant difference in fungal composition between monoculture and mixed plantations. The diversity of soil bacterial communities was higher in monoculture than in mixed RPs due to the higher soil pH in the monoculture. Our results also confirmed that soil pH is the main driver for both bacterial and fungal soil structure. From the point of view of soil microbial diversity, it seems that monoculture RPs are better than mixed RPs. However, from the perspective of plant diversity above ground, the mixed RP is better than the monoculture RP. Here, we suggest that more alkaline fertilizers should be applied in mixed RPs to improve the soil pH and soil microbial diversity. In this way, the total diversity would be enhanced to some extent. At the same time, this treatment can also increase the rubber latex yield.
Appendix
Soil sampling design in three rubber plantations in Wenchang, Hainan Island.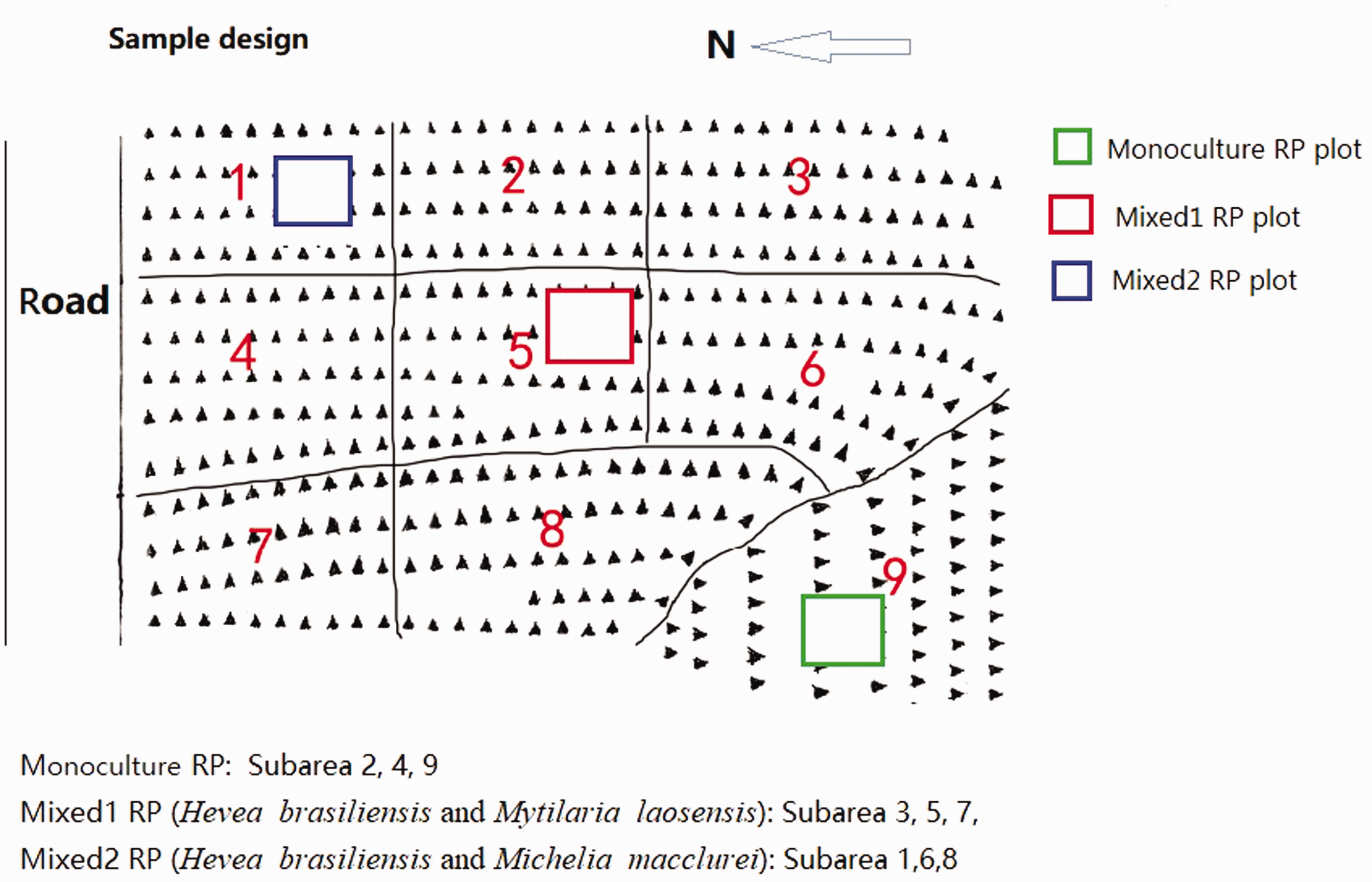
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research work was financially sponsored by the National Natural Science Foundation of China (Grant No. 31770661), the Finance Science and Technology Project of Hainan Province (ZDYF2019145), the Lancang-Mekong River International Cooperation Project (081720203994192003), and the Earmarked Fund for China Agriculture Research System (CARS-34- ZP3).
