Abstract
Decellularized extracellular matrix (dECM) plays an important role in tissue engineering by preserving native biochemical and structural cues while removing immunogenic cellular components. Addressing donor shortages, this study develops a standardized, reproducible protocol for producing cell-derived dECM for bone and cartilage applications, focusing on effective deoxyribonucleic acid (DNA) removal to prevent immune responses in 3D-bioprinted hydrogels. We also evaluate dECM’s impact on cell viability and differentiation potential. Human dermal fibroblasts were decellularized using nonidet P-40, a nonionic detergent (nonyl phenoxypolyethoxylethanol) (NP-40) lysis buffers (1% or 10%) for 1 or 3 h. Decellularization efficacy was assessed via double-stranded DNA (dsDNA) Qubit assay, gel electrophoresis, immunofluorescence, and bicinchoninic acid protein assay. Hydrogels (5 wt% alginate, 3 wt% gelatin) with/without 1% dECM were extrusion bioprinted. Structural and mechanical properties were analyzed using Raman spectroscopy and rheology. Fibroblast viability within bioprinted constructs was monitored over 21 days. Hoechst staining and Qubit assay confirmed residual DNA after 1-h incubations, but complete removal (<50 ng dsDNA) occurred after 3 h with both NP-40 concentrations. The 10% NP-40/3-h protocol yielded the highest protein content. dECM incorporation did not compromise scaffold properties. Significantly enhanced cell viability and glycosaminoglycan (GAG) content (up to day 6) were observed in dECM hydrogels versus controls. Mechanical testing showed a 33% increase in Young’s modulus in dECM-containing hydrogels. Raman spectroscopy confirmed successful dECM integration via a characteristic GAG peak (895 cm−1). We established an optimized decellularization protocol (10% NP-40, 3 h) that effectively eliminates cellular/nuclear material (DNA <50 ng, RNA undetectable) below immunogenic thresholds while preserving essential extracellular matrix components. Fibroblast-derived dECM significantly enhanced alginate-gelatin hydrogel performance, improving cell viability, GAG synthesis, and early osteogenic markers without compromising structural integrity. This protocol provides a robust and standardized source of bioactive dECM, offering a viable alternative to tissue-derived matrices for advanced bone and cartilage tissue engineering bioinks. While the method demonstrates potential for scale-up, further validation following internationally recognized International Organization for Standardization (ISO) standards would be necessary before production-level implementation.
Impact Statement
This study establishes a reproducible, optimised protocol for generating fibroblast-derived decellularised extracellular matrix (dECM) and integrating it into 3D bioprinted hydrogels. By standardising detergent exposure and incubation conditions, the approach achieves effective cell and nucleic acid removal while preserving bioactive ECM components. Incorporation of dECM enhanced cell viability, glycosaminoglycan deposition, and early osteogenic activity without compromising mechanical integrity. These findings provide a methodological framework for scalable, cell-derived dECM production and advance the development of bioactive, patient-specific bioinks for bone and cartilage tissue engineering.
Introduction
The extracellular matrix (ECM) is a complex three-dimensional network primarily composed of proteoglycans (PGs) and fibrous proteins. 1 PGs dominate the extracellular interstitial space, forming a hydrated gel that buffers, retains water, binds molecules, and resists mechanical forces, thereby maintaining ECM structural and functional integrity. 2 Key fibrous proteins include collagens, elastin, fibronectins (FN), and laminins. 3 Collagens provide mechanical strength, support cell adhesion, chemotaxis, migration, and guide tissue development, while elastin imparts resilience to cyclic stretching through its interaction with collagen fibrils.4,5 FN organizes the interstitial ECM and acts as a mechano-regulator, converting physical forces into biological signals that influence cell behavior. 6
Embedded within the complex ECM network are nanovesicles, carrying enzymes, microRNAs, growth factors, and various other cell-signaling molecules, positioning the ECM as a dynamic reservoir regulating cellular function.7,8
Replicating ECM complexity
However, standardized protocols for cell-derived DECM are still limited, especially in balancing nucleic acid removal with ECM preservation. While it lacks the full 3D architecture of tissue-derived ECM, it retains the complex biochemical cues vital for regeneration.
Current methodologies frequently lack detailed protocol standardization and robust criteria for evaluating decellularization efficacy, often overlooking RNA quantification despite evidence that residual nucleic acids may elicit immune responses. 14 Most studies rely on qualitative techniques such as immunofluorescence staining, 15 including 4′,6-diamidino-2-phenylindole (DAPI), 16 Hoechst, 17 hematoxylin and eosin 18 staining, and SEM imaging 19 to confirm DNA removal, which may be insufficient for comprehensive nucleic acid assessment.
To address these limitations, our study systematically refines the protocol established by Harris et al. 20 by optimizing detergent concentration (1% vs. 10% NP-40) and incubation duration (1 vs. 3 h). Unlike previous work, critically, we incorporate both DNA quantification via Qubit assay and RNA assessment by gel electrophoresis, ensuring comprehensive nucleic acid clearance. We employ human dermal fibroblasts as the ECM source, selected for their mesenchymal stem cell-like plasticity—including demonstrated osteogenic and chondrogenic differentiation potential—as well as practical advantages such as robust collagen and FN synthesis, resilience to bioprinting processing conditions, and immunomodulatory properties. 21 Collectively, this method provides a robust, reproducible framework for generating cell-derived DECM with preserved bioactivity, supporting its translation for regenerative medicine and 3D bioprinting applications.
Effective bone and cartilage regeneration remains a major challenge, as degenerative diseases such as osteoarthritis and traumatic injuries impair mobility, cause chronic pain, and create significant global socioeconomic burdens. 22
Current regenerative techniques, such as autologous chondrocyte implantation, microfracture, and osteochondral grafting, are limited by donor site morbidity and inconsistent long-term outcomes, often producing fibrocartilage formation or endochondral ossification that fails to restore native tissue function. Conventional 3D-bioprinted hydrogels allow precise spatial control but lack the biochemical and mechanical cues needed for proper differentiation and ECM deposition.23,24 Integrating decellularized ECM into bioprinted hydrogels offers a promising solution, combining native-like biochemical and structural features with the spatial precision of bioprinting. 23
DECM delivers essential signaling molecules, growth factors, and matrix proteins that enhance cellular responses, promoting tissue-specific regeneration. This study develops and standardizes a modified protocol to produce cell-derived DECM from human dermal fibroblasts and evaluates its use as a bioactive additive in alginate–gelatin hydrogels for bone and cartilage tissue engineering. Integrating fibroblast-derived DECM into these hydrogels links standardized decellularization with functional 3D bioprinting, harnessing fibroblasts’ ECM remodeling. This strategy addresses the shortage of chondrocyte- and MSC-MSC-compatible bioinks, offering a scalable, reproducible, and bioactive platform for bone and cartilage regeneration.
Materials and Methods
ECM preparation
Cell culture
Normal human dermal fibroblasts (adult; PromoCell, Heidelberg, Germany, Cat. No. C-12302, RRID: CVCL_9212) were cultured under aseptic conditions in Dulbecco’s Modified Eagle Medium (DMEM; Thermo Fisher Scientific, Waltham, MA, USA) supplemented with 50 U/mL penicillin, 50 µg/mL streptomycin (Sigma–Aldrich, Merck KGaA, Darmstadt, Germany), 2 mM Glutamax (Invitrogen, Thermo Fisher Scientific, Waltham, MA, USA), and 10% fetal bovine serum (Sigma-Aldrich, Cat. No. F9665, Merck KGaA, Darmstadt, Germany; non-USA origin). Cells were maintained at 37°C, 5% CO2, and 95% humidity and seeded at 1 × 105 cells/mL in six-well plates (2 mL/well). When cultures reached 90–100% confluence, decellularization was initiated using a mild nonionic detergent protocol. 16
Decellularization of fibroblast culture
Confluent fibroblast layers were washed three times with prewarmed PBS (37°C) to remove serum residues and treated with Nonidet P-40 Substitute (Sigma-Aldrich, Cat. No. 74385; Merck KGaA, Darmstadt, Germany) at either 1% or 10% (v/v) in PBS for 1 h (short incubation) or 4 h (long incubation; solution refreshed after 1 h). Plates remained stationary during detergent exposure to preserve ECM structure. Matrices were then washed three times with PBS (10 min each) and twice with deionized water at room temperature, followed by overnight incubation in DNase/RNase-free PBS at 4°C. Samples were lyophilized to obtain dry decellularized ECM (dECM). Control groups underwent identical steps without detergent treatment. A schematic of the process is provided in Figure 1.
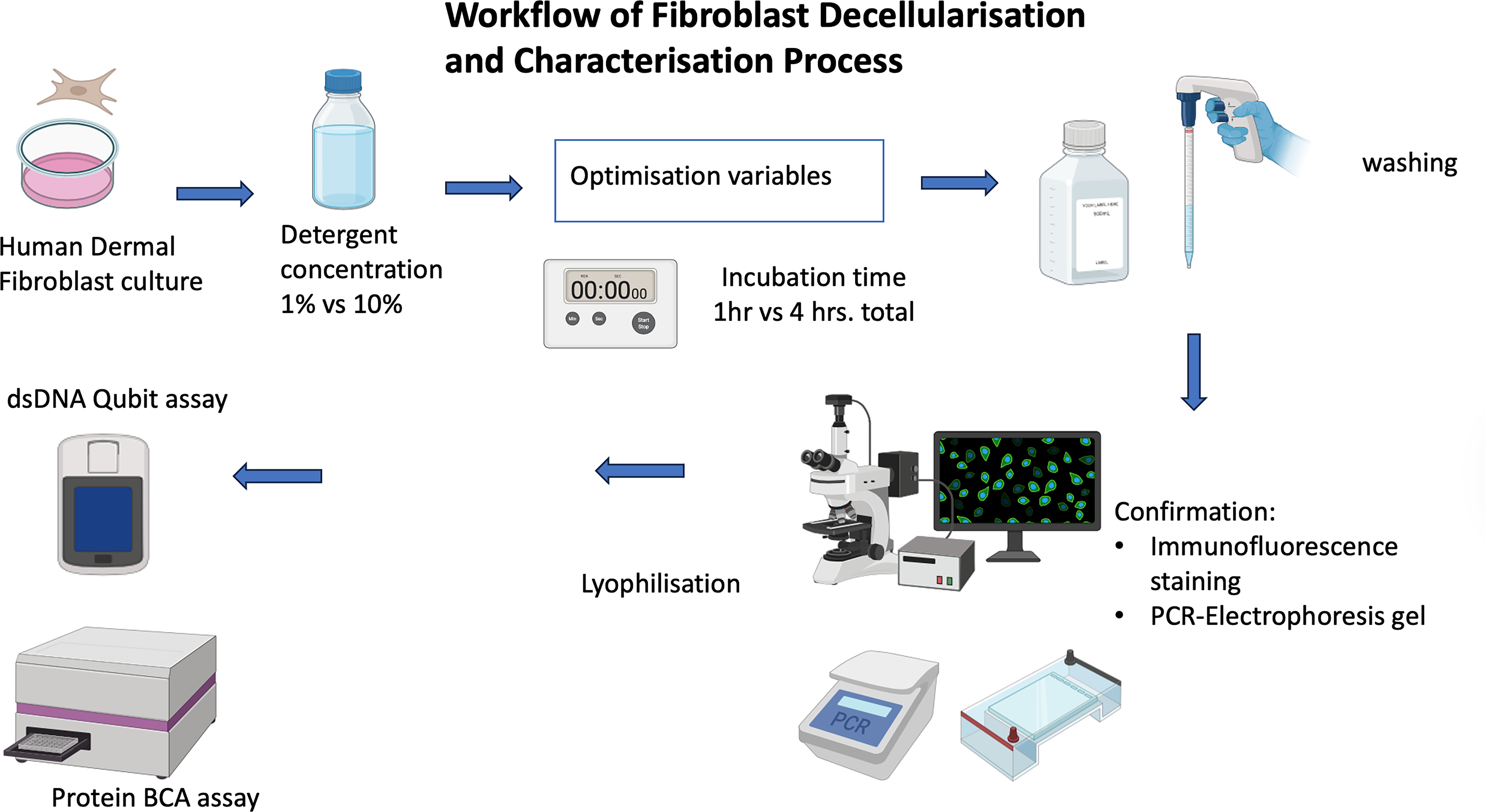
Workflow of fibroblast decellularization and characterization process. Human dermal fibroblasts were decellularized using Nonidet P-40 detergent (1% or 10%) for varying incubation durations (1 or 4 h total). Following washing and lyophilization, decellularization efficacy was confirmed by immunofluorescence staining and PCR-electrophoresis. Quantitative assays (dsDNA Qubit and protein BCA) were used to assess residual nucleic acid and protein content. BCA, bicinchoninic acid; dsDNA, double-stranded DNA.
Immunofluorescence staining
Residual DNA and ECM quality were assessed by fluorescence microscopy. Samples were fixed with 4% paraformaldehyde for 15 min, permeabilized with 0.1% Triton X-100 for 10 min, and blocked with 3% FCS (Gibco, Thermo Fisher Scientific, USA) for 1 h at room temperature. Primary anti-FN antibody (Sigma-Aldrich, Cat. No. F3648; Merck KGaA, Darmstadt, Germany) was applied overnight at 4°C (1:500), followed by FITC-conjugated secondary antibody (Invitrogen, Ref,656111 Thermo Fisher Scientific, USA) for 1 h (1:200). Nuclei were counterstained with Hoechst 33342 (1:1000). Images were acquired using an Olympus IX51 microscope (Olympus Life Science, Tokyo, Japan).
DNA and RNA extraction, quantification, and PCR
Genomic DNA from decellularized and control samples was isolated using TRIzol™ and quantified with a Qubit 3.0 Fluorometer (Thermo Fisher Scientific, USA). RNA was extracted and reverse-transcribed to complementary DNA (cDNA) for amplification of the RPL-19 housekeeping gene. PCR products were resolved on 2% agarose gels and visualized under UV light. Full procedures, reagent sources, and specifications are provided in Supplementary Data S1, S2, S3, and S4.
Protein quantification (bicinchoninic acid assay)
Protein content was measured using the bicinchoninic acid assay (Thermo Fisher Scientific, USA). Lyophilized dECM and control samples were incubated with 200 µL of reagent mixture (A:B) at 37°C for 30 min, and absorbance was read at 562 nm (Opsys Mr™, Dynex Technologies, USA).
DECM incorporation into 3D-bioprinted hydrogel
Bioink components—sodium alginate (5% w/v; Sigma-Aldrich, Merck KGaA, Darmstadt, Germany) and gelatin (3% w/v; porcine skin type A, ∼300 Bloom; Sigma-Aldrich, Merck KGaA, Darmstadt, Germany)—were prepared in sterile high-glucose DMEM at 55°C with magnetic stirring (150 rpm). Solutions were centrifuged (200 rpm, 5 min) to remove bubbles and stored at 4°C until use. Before printing, hydrogel stocks were warmed to 37°C.
Cell-laden bioinks were prepared by gently mixing 6 × 106 cells with 6 mL of hydrogel. For treated groups, lyophilized dECM powder (1% w/v; 6 mg per 6 mL hydrogel) was incorporated for minimal double-stranded DNA (dsDNA) and optimal protein concentration. 25 dECM was mixed thoroughly for homogeneous distribution. Bioinks (±dECM) were loaded into 5 cc syringes and centrifuged (300 rpm, 3 min) to remove air bubbles.
Bioprinting was performed using a Regemat3D extrusion bioprinter (Granada, Spain) with 0.41 mm tips. Constructs (12 mm diameter, 2.1 mm height, 0.2 mm pore size, 0.35 mm fiber width) were printed at 32°C and 6 mm/s. Six scaffolds were printed simultaneously for consistent cooling and shape fidelity. Postprinting, scaffolds were cross-linked in 100 mM CaCl2 for 40 min per side, rinsed with PBS, and transferred to 24-well plates for culture.
Cell-laden hydrogels (±dECM) were cocultured with human dermal fibroblasts in growth medium. Viability and proliferation were assessed up to 21 days using AlamarBlue™. Chondrogenic and osteogenic differentiation were induced with standard supplements, followed by glycosaminoglycan (GAG) and ALP assays. Absorbance was read using a microplate reader. Experimental details and analyses are described in Figure 2 and Supplementary Data S5.
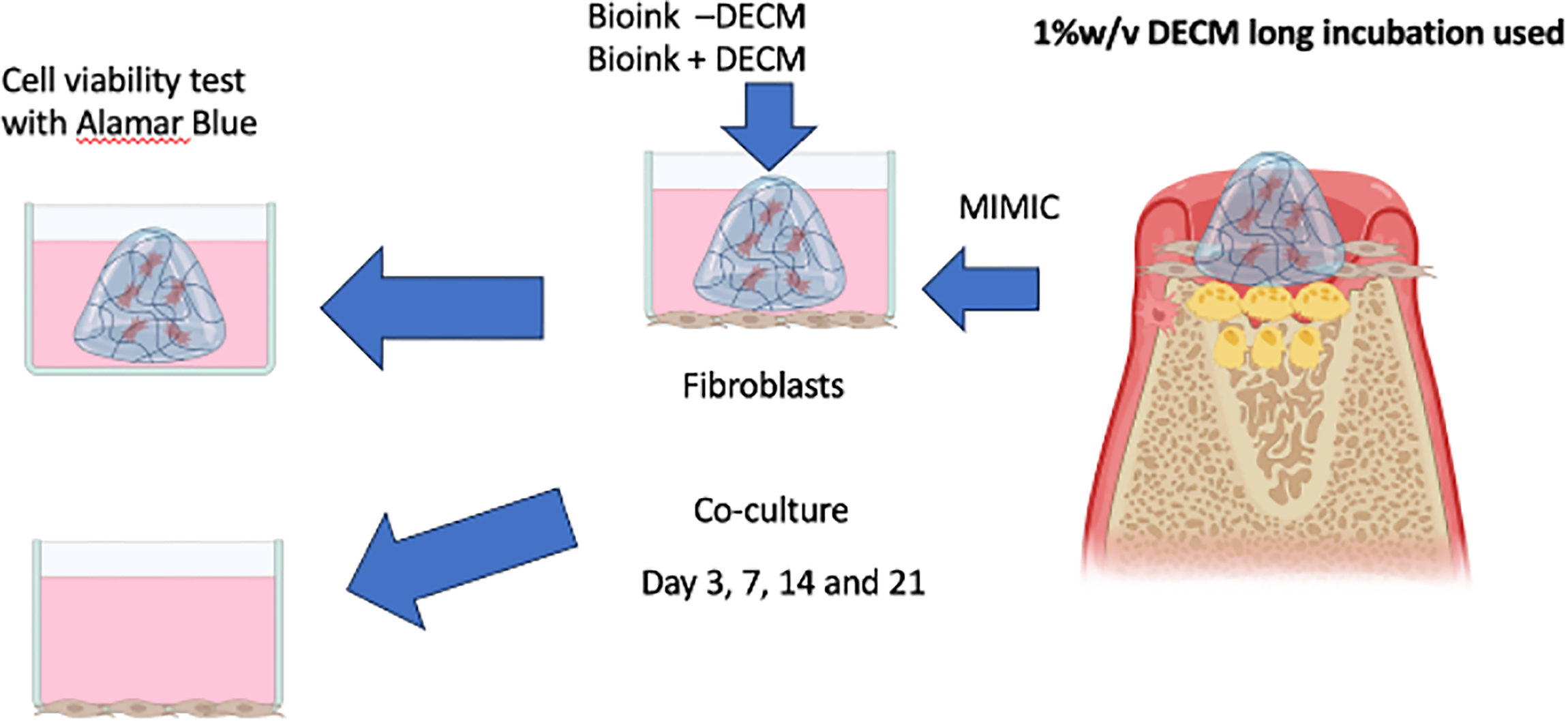
Schematic illustration of the fibroblast-laden 3D bioink coculture model and assessment of cellular responses. Bioinks containing 1% (w/v) long-incubated dECM (Bioink + dECM) or bioink without dECM (Bioink − dECM) were fabricated and cultured to mimic the osteochondral microenvironment. Cocultures were maintained for 3, 7, 14, and 21 days and assessed for cell viability using the Alamar Blue assay. Osteogenic differentiation was evaluated by alkaline phosphatase (ALP) activity, while chondrogenic matrix deposition was measured by glycosaminoglycan (GAG) content. dECM, decellularized extracellular matrix.
Statistical analysis
Each condition included
Raman spectral and mechanical analyses
Raman spectra of bioprinted hydrogels (±dECM) were obtained using a confocal Raman microscope (532 nm laser). Baseline correction, smoothing, and normalization were performed in MATLAB R2019 (MathWorks, USA). Peak assignments followed the reference. 26 Compressive stress–strain testing was conducted using an Instron 5967 Universal Testing Machine under unconfined compression (∼25% strain min−1). Young’s modulus (E = σ/ε) was derived from the linear elastic region to compare stiffness between groups.
Results
Immunofluorescence microscopy
Fluorescence imaging of untreated controls revealed a dense FN matrix and abundant nuclei, confirming intact cellular presence (Fig. 3). Samples treated with 1% detergent (short incubation) retained strong FN staining but showed residual nuclei, indicating incomplete decellularization. Prolonged incubation improved DNA removal with minimal matrix loss, maintaining FN integrity.
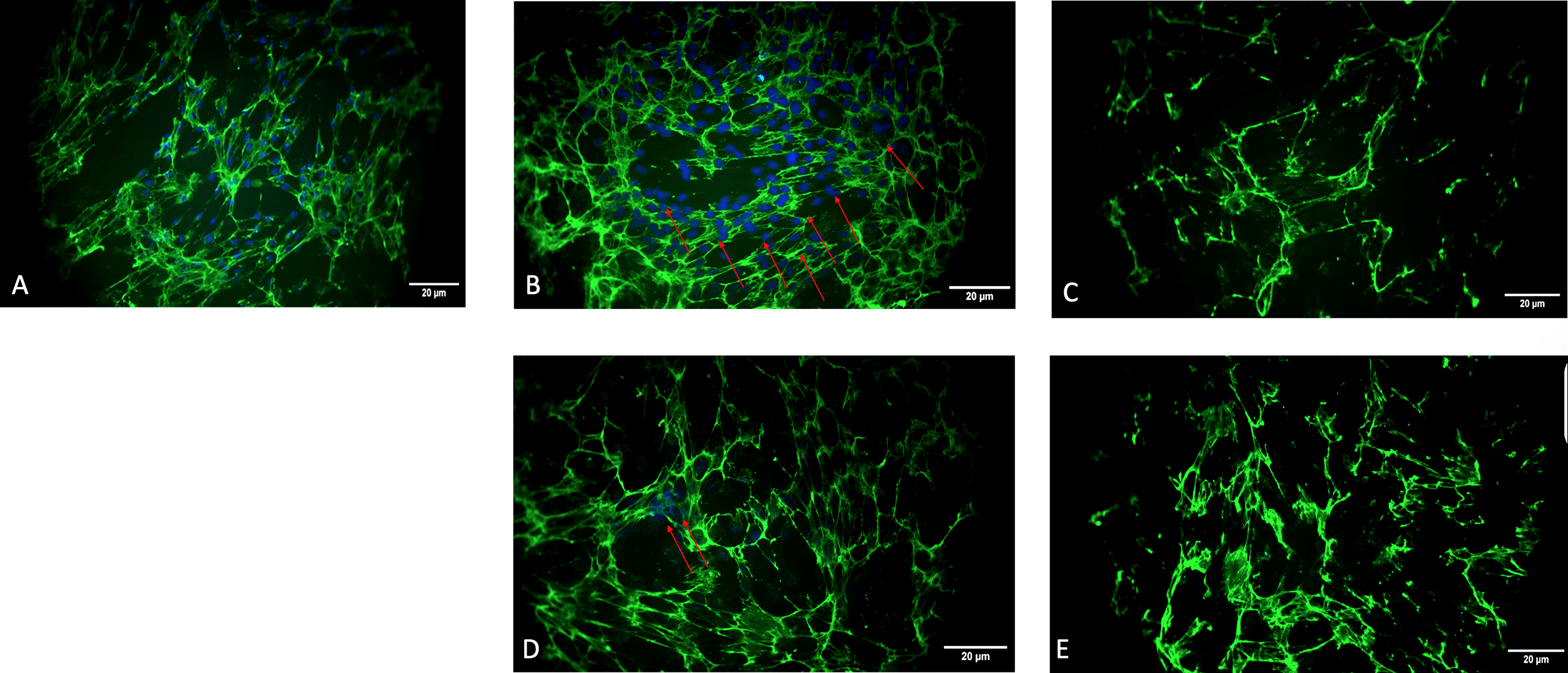
Immunofluorescence staining of fibronectin in control and decellularized treated with different NP-40 concentrations and incubation times. Representative fluorescence images showing fibronectin (
dsDNA quantification
DNA content varied markedly across six treatment groups (Fig. 4). The highest concentrations occurred in control and short-incubation groups (up to ∼180 ng/mL), while long-incubation treatments—especially 1% and 10% detergent—showed minimal residual DNA (∼30 ng/mL). Two-way ANOVA (Table 1) identified significant effects of incubation time (
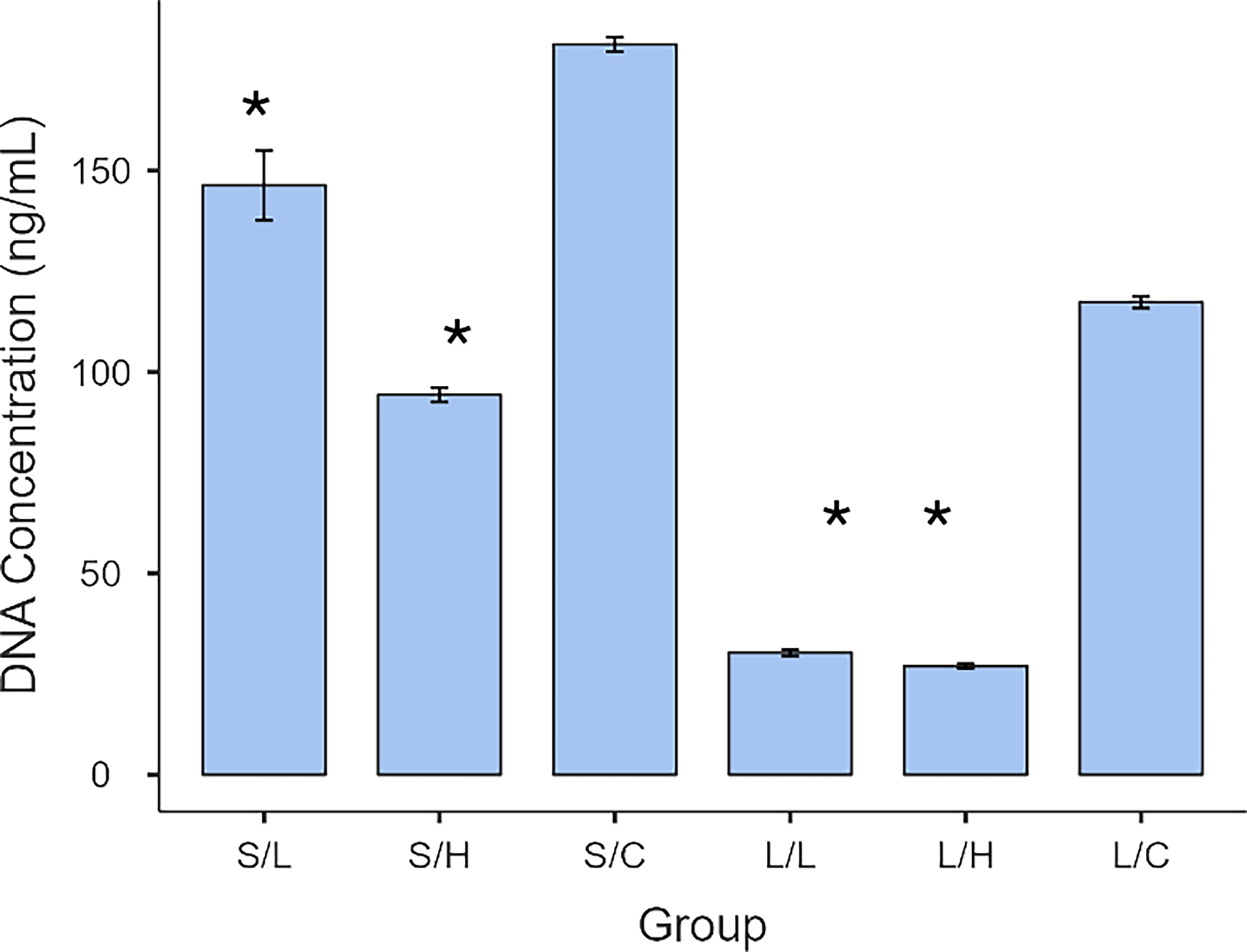
dsDNA concentration across experimental groups following decellularization. DNA concentration was quantified using the Qubit 3.0 Fluorometer and Qubit High-Sensitivity dsDNA Assay Kit after Trizol reagent extraction. Data are presented as mean ± standard deviation (
Two-Way Factorial ANOVA and Post Hoc Tukey’s and Bonferroni Tests Were Conducted to Investigate the Interaction of Incubation and Concentration Variables on dsDNA Concentration
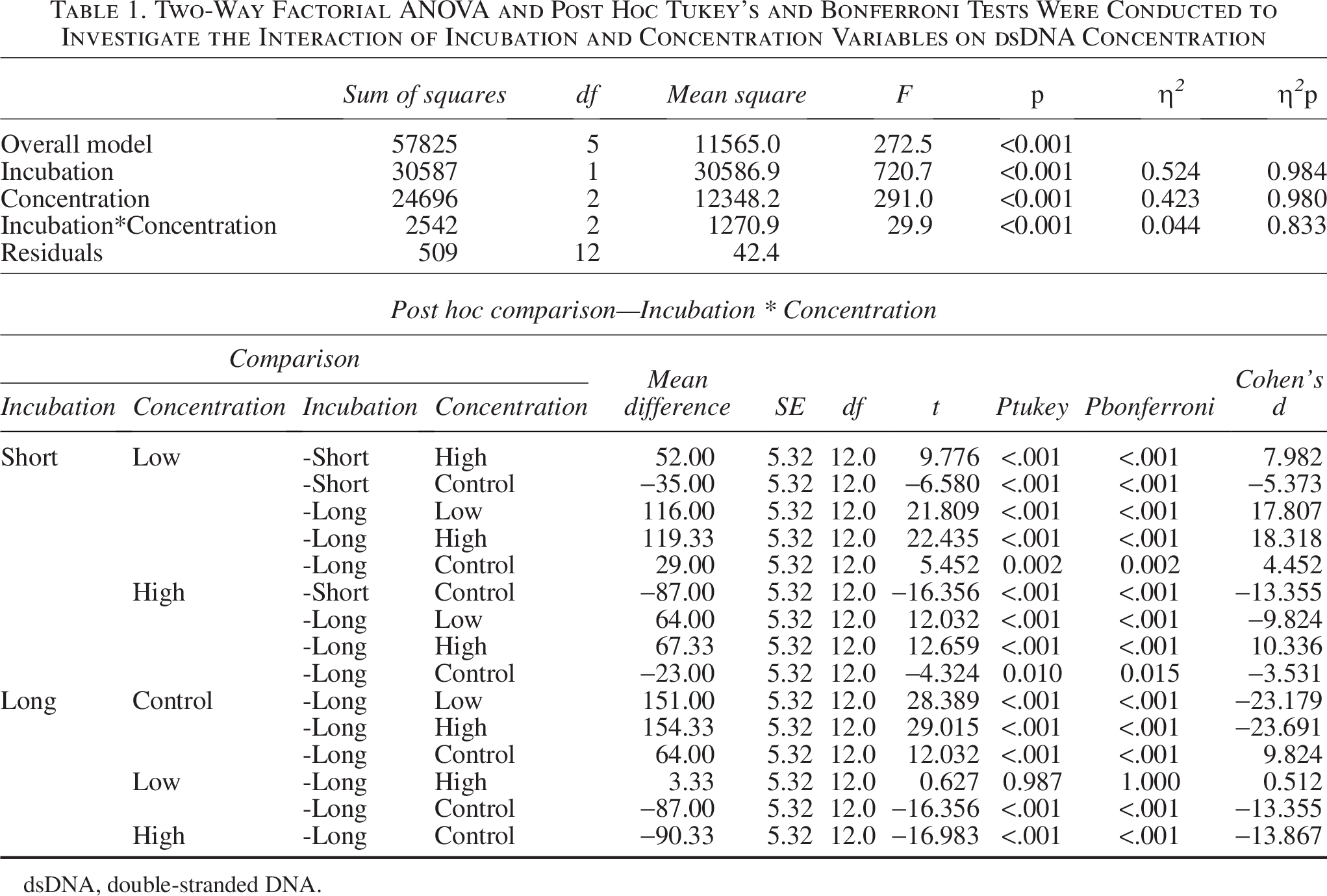
dsDNA, double-stranded DNA.
Gel electrophoresis
Agarose gels (Fig. 5) supported the quantitative data: strong DNA bands appeared in controls, while long-incubation, detergent-treated samples showed faint or absent bands, confirming efficient DNA removal while retaining matrix structure.
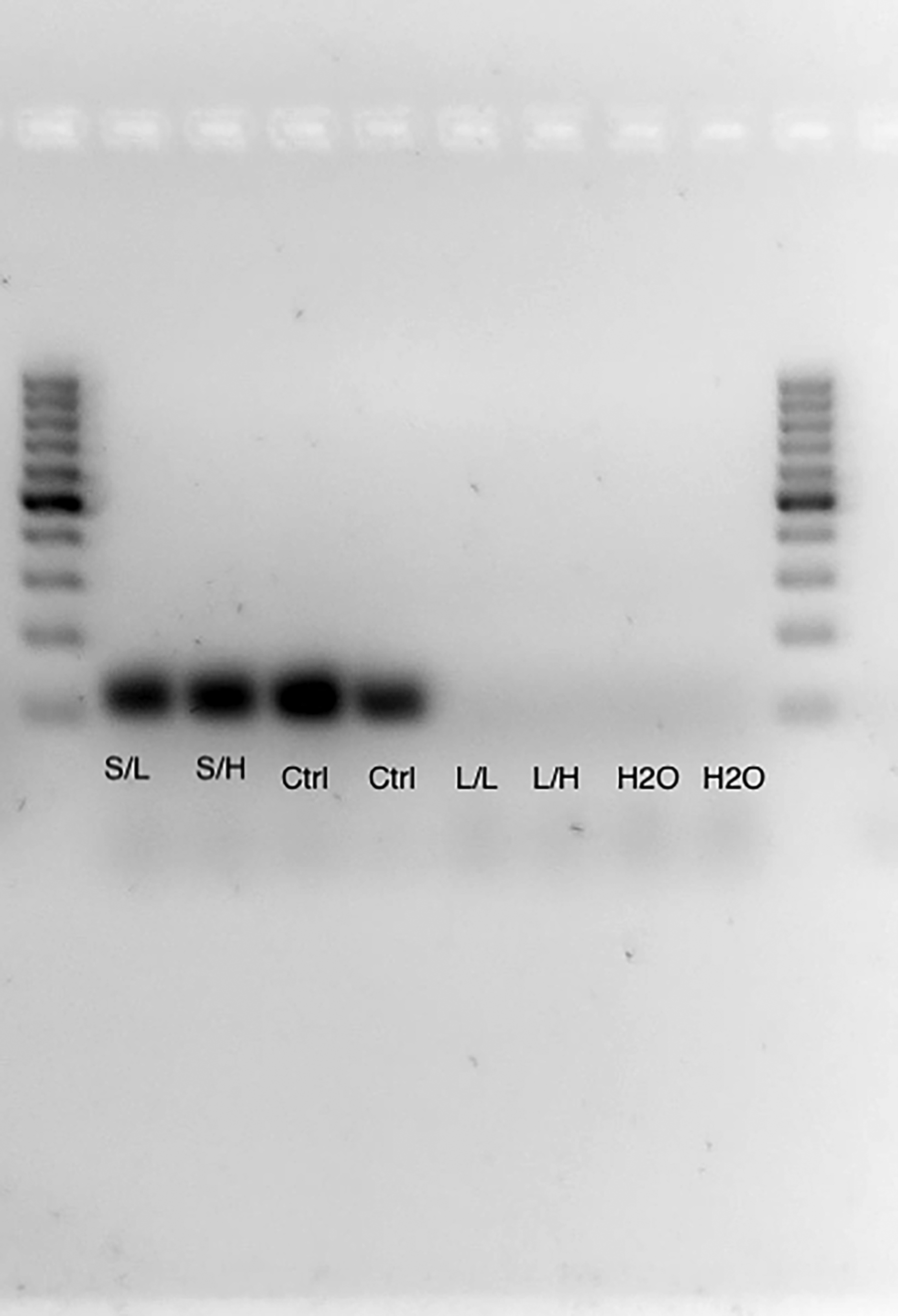
Amplification of the housekeeping gene ribosomal protein, large (60S) subunit (RPL) from decellularized and nondecellularized fibroblast matrices. Representative 2% (w/v) agarose gel electrophoresis showing PCR products of RPL (expected amplicon ≈150–200 bp). Nondecellularized controls display clear RPL bands, confirming the presence of cellular DNA, whereas decellularized samples (1% short, 1% long, 10% short, 10% long incubations) show markedly reduced or absent amplification, indicating effective removal of nuclear material. Negative controls (H2O) show no amplification. DNA ladder at right denotes molecular-weight reference.
Total protein quantification
Protein levels (Fig. 6) were highest in control groups (∼150 µg/mL) and dropped sharply in all decellularized samples (<10 µg/mL). Two-way ANOVA (Table 2) showed a significant main effect of detergent concentration (
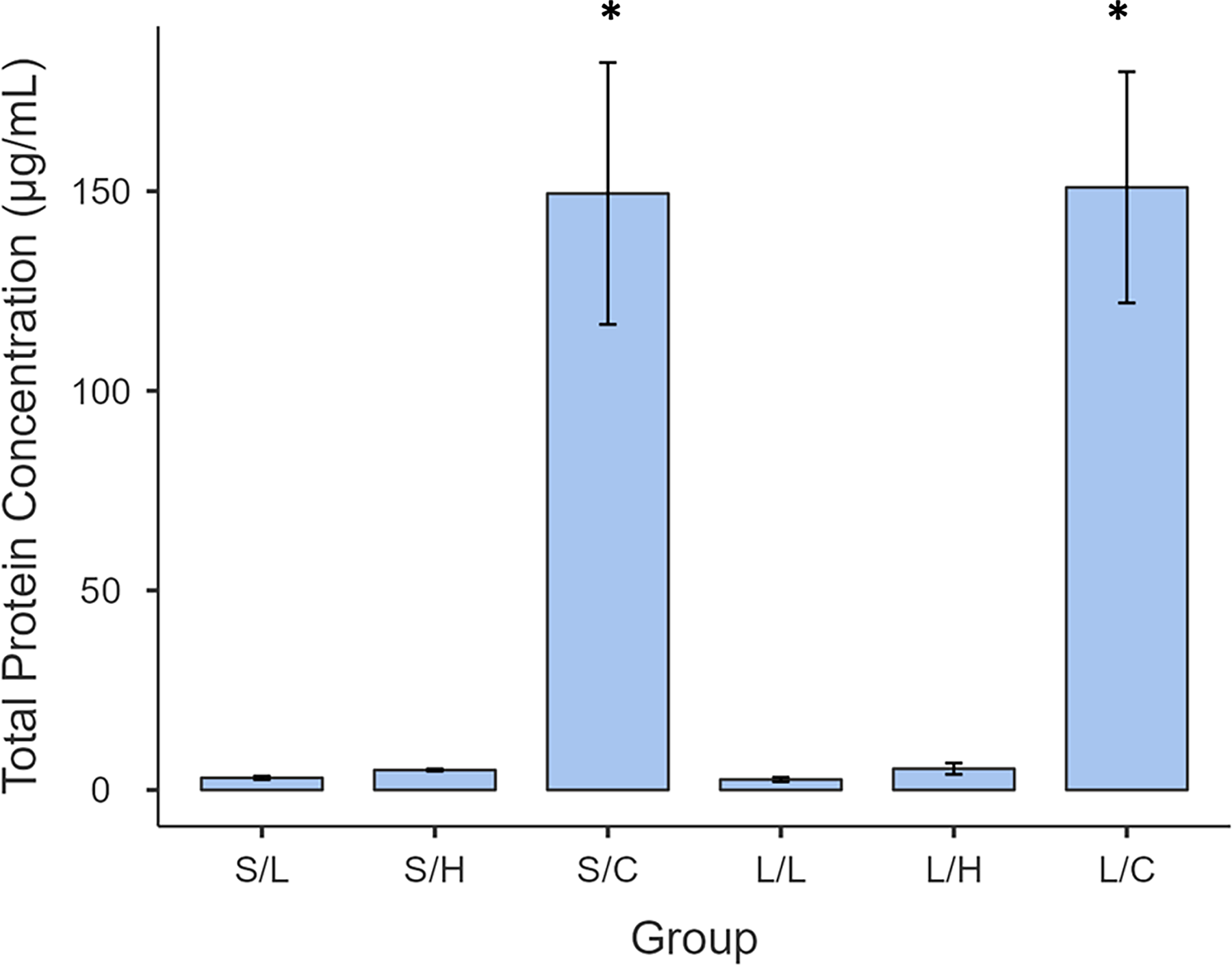
Total protein concentration in each group was determined using BCA following lyophilization. Data presented as a mean. S/L = 1% + Short incubation; S/H =10% + Short incubation; S/C = Short incubation, control non-dECMl; L/L = 1% + Long incubation; L/H = 10% + Long incubation; L/C = Long incubation, control non-dECMl. Asterisks (*) indicate statistically significant differences (
Two-Way Factorial ANOVA and Post Hoc Tukey’s and Bonferroni Tests Were Conducted to Investigate the Interaction of Incubation and Concentration Variables on Protein Concentration
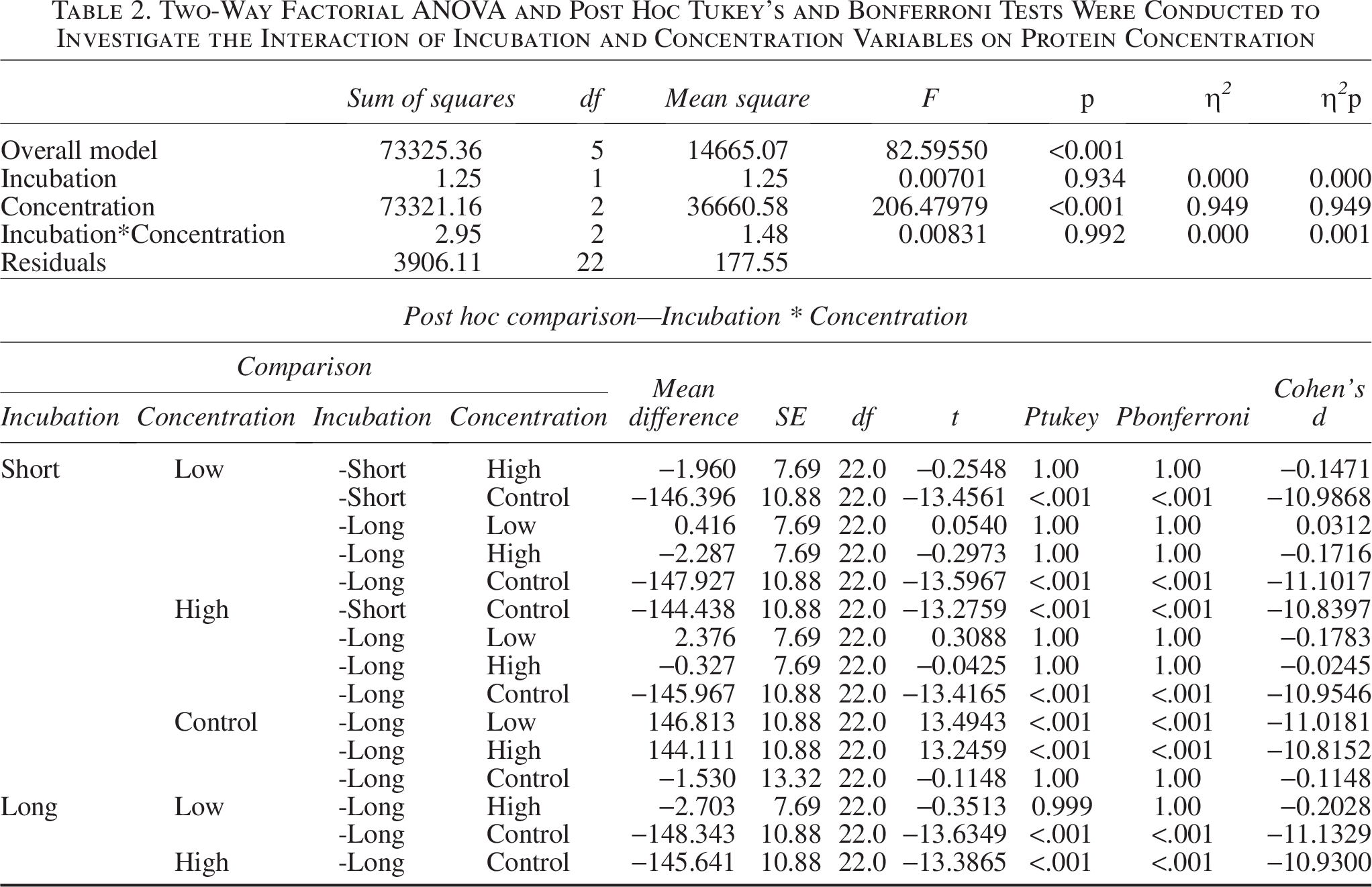
dECM weight
No significant changes in scaffold weight were found between treatments (Supplementary Data). The long-incubation, low-concentration condition was selected for 3D bioprinting due to its low DNA and preserved protein content.
Cell viability
AlamarBlue™ analysis (Fig. 7) confirmed cell survival and proliferation up to day 21, peaking on day 14. dECM-containing bioinks showed significantly higher viability on day 7 (
Results of One-Way ANOVA for the Effect of dECM Presence on Day 7 Cell Viability
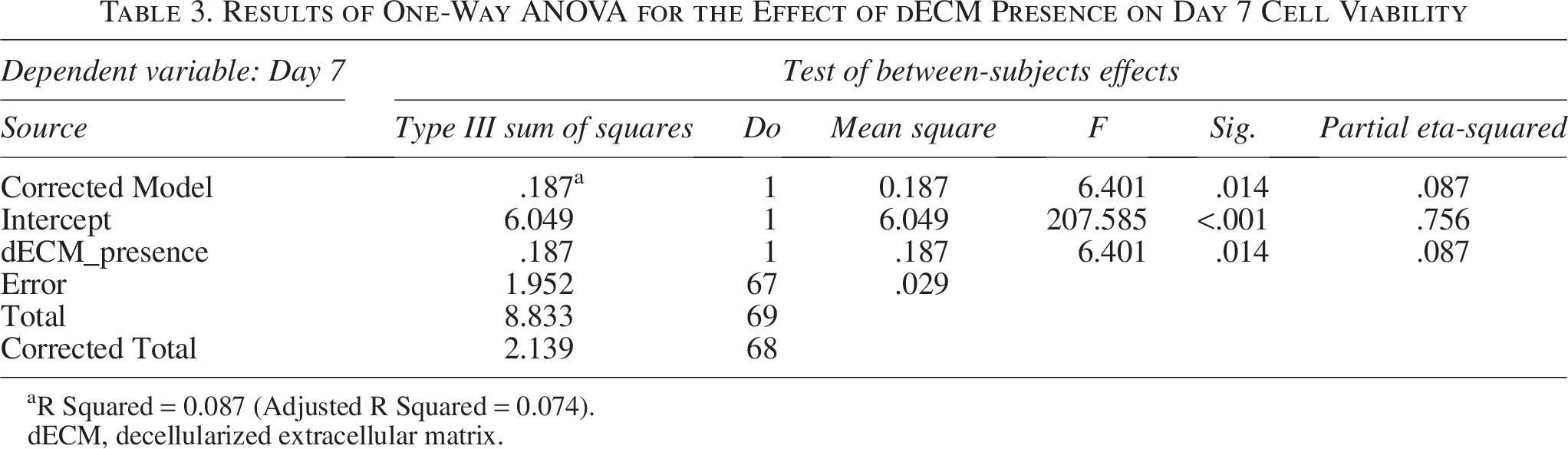
R Squared = 0.087 (Adjusted R Squared = 0.074).
dECM, decellularized extracellular matrix.
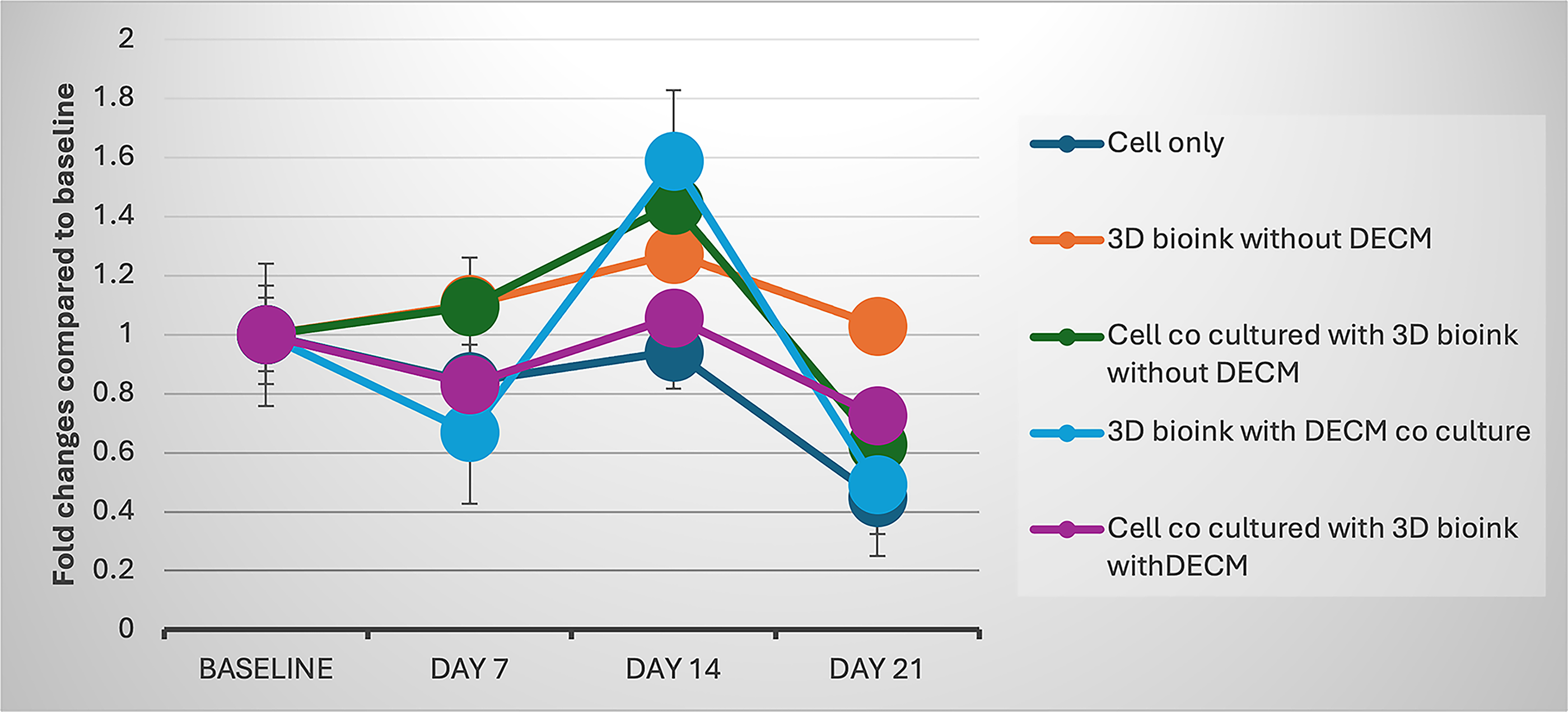
Cell viability of different culture conditions over 21 days, expressed as fold change compared with baseline (Day 3). Data represent mean ± standard deviation (SD). Each group consisted of triplicate samples (
GAG and ALP activity
Chondrogenic cultures with dECM (dECM CM) showed the greatest GAG accumulation (
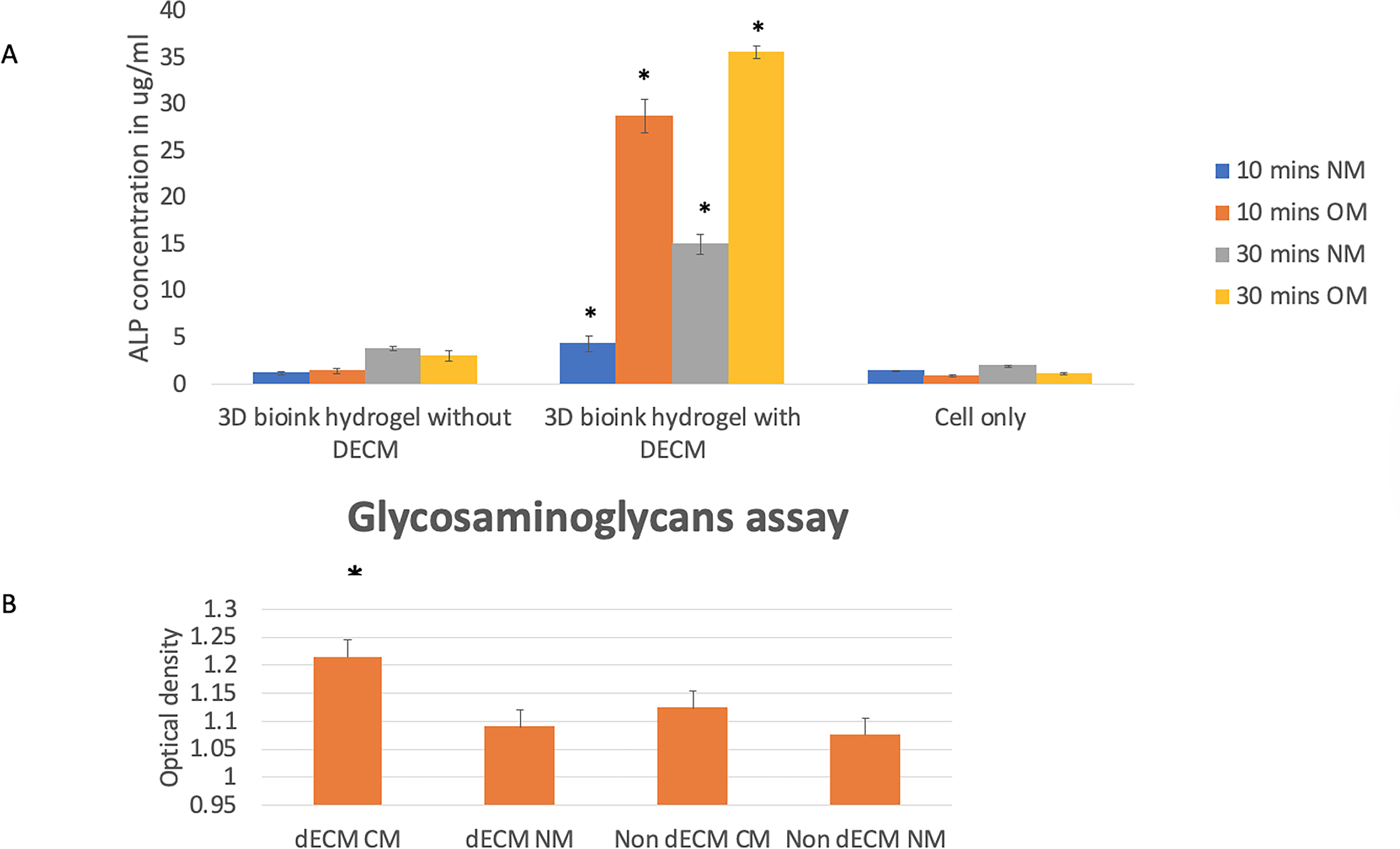
Influence of dECM on osteogenic and chondrogenic differentiation of 3D bioink hydrogels.
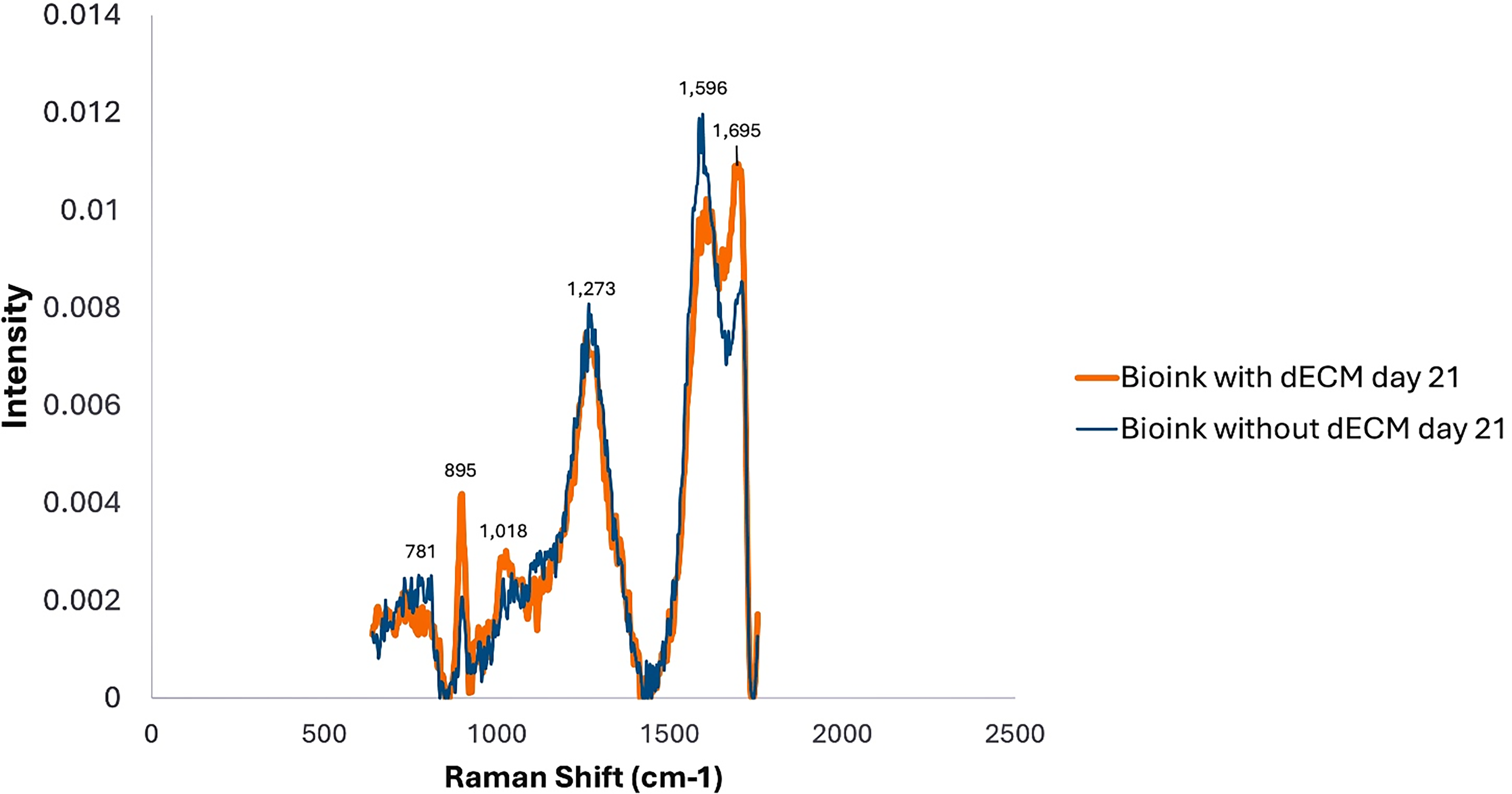
Raman spectroscopy profiles of bioink with and without dECM at day 21. Representative Raman spectra of bioink formulations containing dECM (
Raman spectral analysis
The Raman spectra of bioink formulations showed characteristic peaks within the fingerprint region (600–1800 cm−1), with major bands at 781, 895, 1018, 1273, 1596, and 1695 cm−1 (Fig. 10). Both formulations exhibited broadly similar spectral profiles, indicating comparable molecular composition.
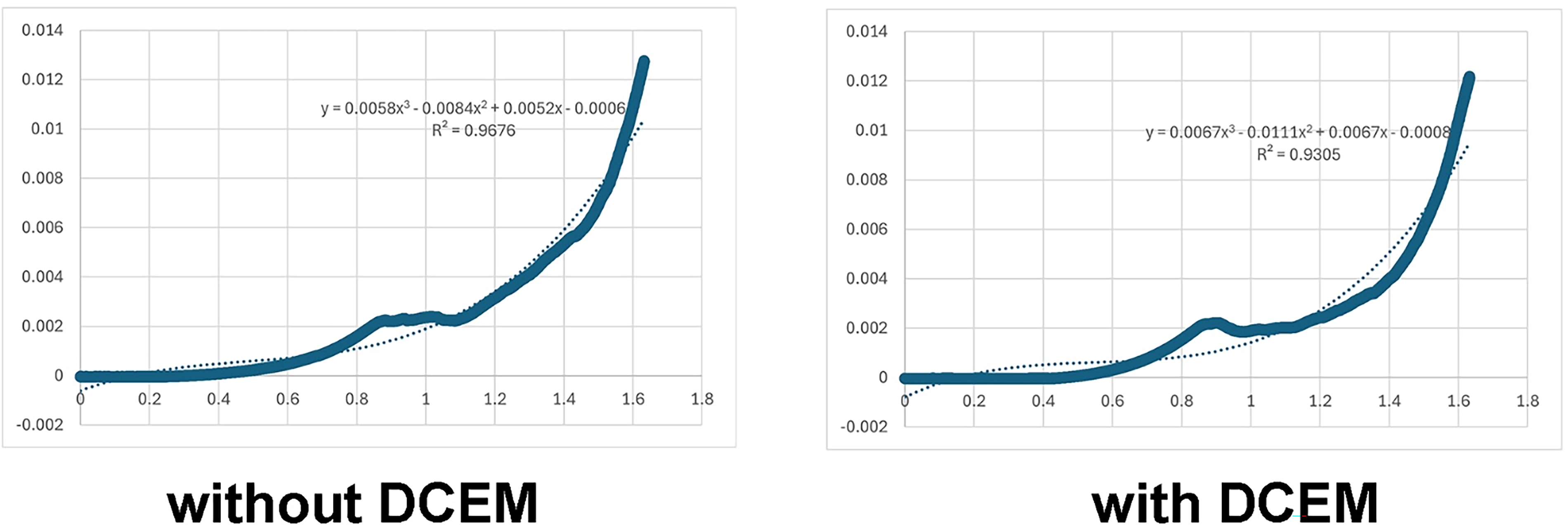
Stress–strain curve of 3D-bioprinted hydrogel with/without dECM. Representative stress–strain profile of the 3D-bioprinted hydrogel sample fabricated without dECM, obtained from uniaxial compression testing using an Instron universal testing machine.
The 781 cm−1 band, of low intensity in both samples, corresponds to DNA vibrations. The 895 cm−1 peak showed greater intensity in the dECM-containing bioink, consistent with enhanced carbohydrate and GAG content. The 1018 cm−1 and 1273 cm−1 peaks, attributed to C–O stretching and the amide III band (C–N and N–H vibrations of proteins, particularly collagen), appeared similar in both groups, suggesting comparable protein presence at day 21.
Prominent bands at 1596 cm−1 (DNA) and 1695 cm−1 (amide I, C = O stretching in proteins) distinguished the two bioinks. The dECM formulation showed stronger protein-associated amide I intensity, while the non-dECM bioink exhibited a higher DNA-related signal, suggesting a relative shift toward greater protein structural content in the dECM hydrogel.
Stress–strain analysis
Mechanical testing demonstrated distinct stress–strain behavior between hydrogels with and without dECM. In the linear elastic region, both formulations exhibited comparable initial stiffness, with Young’s moduli of approximately 0.12 MPa (Fig. 11). However, analysis of covariance analysis (SPSS) confirmed that the inclusion of dECM significantly affected stress and strain responses (
Tests of Between-Subjects Effects for Stress–Strain Model
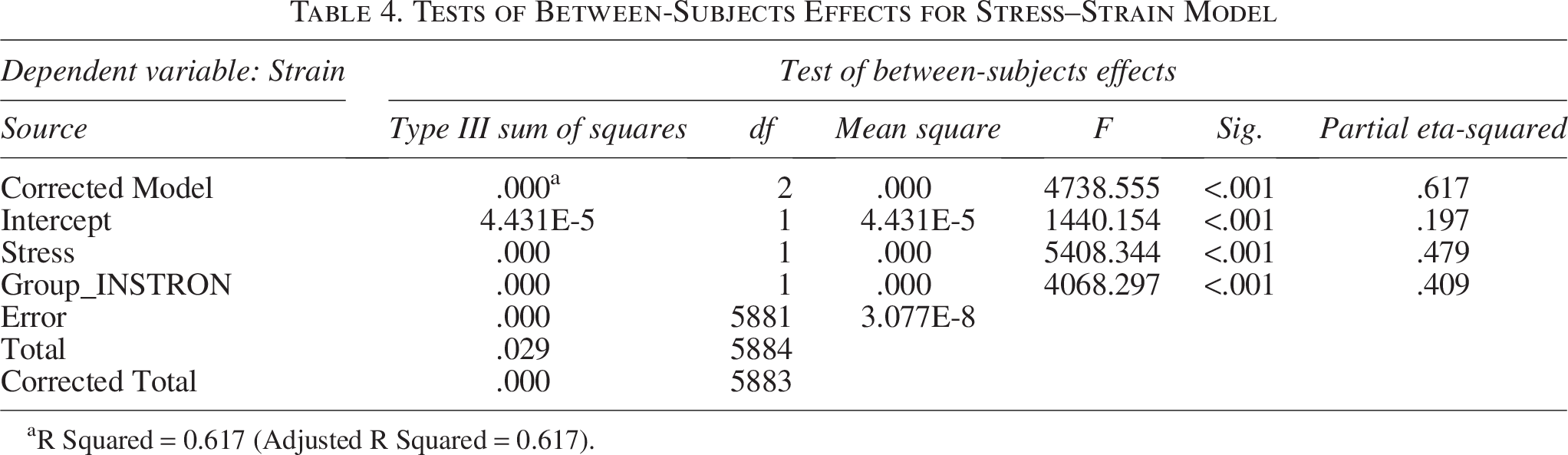
R Squared = 0.617 (Adjusted R Squared = 0.617).
Parameter Estimates for Stress–Strain Model
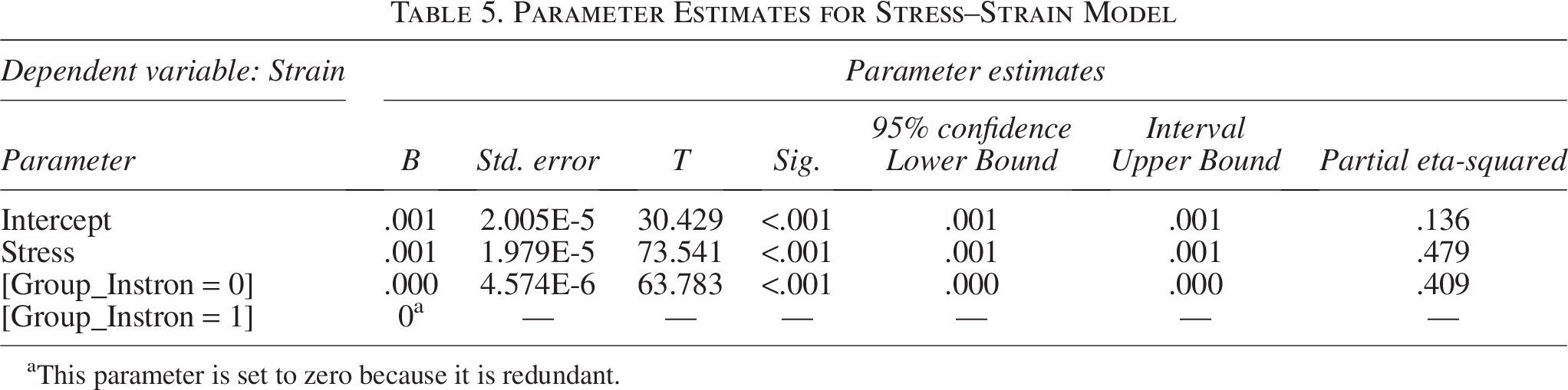
This parameter is set to zero because it is redundant.
Discussion
Kim et al. presented a comprehensive decellularization protocol detailing DNA extraction, imaging, and residual dsDNA quantification while omitting RNA analysis—an omission common across studies using whole organs, tissues, or cell-derived dECMs. 24 This widespread focus on DNA is largely influenced by the criteria proposed by Crapo et al., 25 which, despite their broad adoption, lack experimental validation and have been criticized as arbitrary in recent literature. 26 RNA exclusion is typically justified by its single-stranded instability and presumed degradation alongside DNA. However, recent evidence challenges this assumption: a forensic study showed RNA undergoes less depurination than DNA under identical degradation conditions and remains amplifiable via quantitative polymerase chain reaction. 27 Although the degradation conditions differed, these findings highlight RNA’s potential resilience and the importance of assessing both residual DNA and RNA in DECM to improve decellularization quality metrics and biomaterial safety.
To strengthen RNA quantification, we supplemented polymerase chain reaction (PCR)-based assessments with NanoDrop spectrophotometry (see Supplementary Data). Unlike DNA analysis, which included gel electrophoresis, NanoDrop provided precise RNA concentration data across all groups. Figure 7 presents PCR products from cDNA synthesized from extracted RNA, confirming the presence or absence of amplifiable transcripts. This parallels Fattorini et al., who showed that even degraded RNA can be reverse transcribed and amplified, establishing PCR absence as a reliable marker of RNA removal. 27 Integrating NanoDrop data with PCR visualization provides robust evidence of effective RNA elimination, underscoring the rigor of our protocol and minimizing immunogenicity risks.
Unlike the fixed 15-min intervals of Harris et al., 20 our dual incubation system (Table 1) enables progressive debris removal while minimizing ECM disruption (Fig. 3). Postdecellularization lyophilization further enhances ECM stability, addressing previous concerns about matrix fragility.
Fibroblasts, sharing CD90/CD105 markers with mesenchymal stem cells and exhibiting immunomodulatory roles critical for bone and cartilage repair, 28 contribute ECM enriched with pro-osteogenic factors, consistent with elevated ALP activity (Fig. 9). The FN–rich matrix (Fig. 3) may also promote osteo- and chondrogenesis through mechanotransductive pathways. 29
To evaluate chondrogenic potential, we cultured cell-laden hydrogel bioinks with and without DECM in both chondrogenic (CM) and normal media. This design circumvents chondrocyte encapsulation challenges and offers a practical alternative for chondrogenic hydrogel engineering. The combination of DECM and CM significantly enhanced GAG expression, confirming an augmented chondrogenic response and demonstrating an optimized integrative approach for functional tissue regeneration.
Extended cultures incorporating histological analyses (Alizarin Red, Safranin O) and comparative studies of fibroblast- versus chondrocyte-laden hydrogels under identical CM conditions are warranted to validate osteo/chondrogenic potential. Such comparisons may clarify whether fibroblast differentiation can serve as a practical alternative to direct chondrocyte bioprinting, overcoming limitations in chondrocyte viability and proliferation.
Spectroscopic analysis revealed distinct molecular signatures between bioink formulations after 21 days. The enhanced 895 cm−1 peak in DECM bioinks indicates successful incorporation of GAG components, while the 1695 cm−1 peak reflects preserved ECM protein structures (Fig. 11). These molecular features likely contribute to improved bioactivity, mechanical properties, and cellular interactions. The presence of native ECM components within DECM-enriched bioinks provides a more physiologically relevant microenvironment supporting adhesion, proliferation, and lineage-specific differentiation. 30
Retention of native ECM signatures supports effective integration and stability within the bioink, highlighting DECM’s potential to recapitulate native microenvironments crucial for tissue regeneration. 31 Future work should correlate spectroscopic data with mechanical characterization and cellular responses to advance structure—function understanding and biomimetic bioink design. 32
Although DECM and non-DECM hydrogels displayed similar initial Young’s moduli (∼0.10–0.15 MPa), their stress–strain profiles differed. DECM hydrogels sustained higher stress and deformation under load, consistent with reports that DECM hydrogels, while retaining similar baseline stiffness, exhibit nonlinear mechanical responses due to fibrous architecture and biochemical heterogeneity.
33
Mechanistically, DECM may enhance structural resilience by providing a network of native ECM proteins (collagens, GAGs, FN) and cross-linking or entanglements that are absent in simpler hydrogel formulations. These retained macro- and microscale features can modulate how the network reconfigures under strain: for example, fiber alignment, recruitment of hidden network elements, or fiber–matrix interactions can cause strain-dependent stiffening or delay structural softening. In composite hydrogels and tissues, this behavior has been described under the “strain-stiffening” paradigm, in which the filler–polymer interactions and network architecture dominate the mechanical response at higher strain.
34
Thus, the two gels began with nearly identical elastic slopes; the DECM network appears to better resist structural deformation or rearrangement under load. In practical terms, the results suggest that incorporating DECM into your hydrogel scaffold confers mechanical advantages under deformation beyond the initial elastic region—potentially making the material more suitable for applications where modest loads or deformations occur (e.g., bioprinted constructs, soft tissue plugs, dynamic loading
From a translational perspective, the integration of volumetric bioprinting methodologies with automated decellularization systems holds significant potential to address current challenges related to scalability and reproducibility, while preserving the bioactivity of the ECM. Furthermore, establishing standardized reporting metrics for decellularization efficacy across various cell types and detergent protocols is crucial. Such standardization would facilitate cross-study comparisons, thereby accelerating the clinical translation of decellularized ECM-based therapies.
The present work was intentionally designed as a methodological optimization and proof-of-concept study. Our primary objective was to establish a reproducible decellularization and hydrogel integration protocol rather than to provide a complete biological validation. Comparable studies in the field12,23,33 have similarly focused on biochemical and early functional endpoints before undertaking gene or protein-level analyses. Future investigations will, therefore, include molecular (RUNX2, SOX9, COL1A1, ACAN) and histological assessments to substantiate lineage-specific differentiation and tissue maturation.
Conclusion
This study presents an optimized decellularization approach using human dermal fibroblasts that effectively removes cellular and nuclear material while maintaining key ECM components suitable for incorporation into 3D-bioprinted hydrogel scaffolds. Through a strategic combination of extended incubation and calibrated detergent concentrations, DNA content was reduced below established immunogenic thresholds, with qualitative confirmation of RNA removal, ensuring biosafety in downstream applications.
Importantly, incorporation of fibroblast-derived DECM into alginate–gelatin hydrogels did not compromise structural integrity or mechanical properties. Instead, it significantly enhanced cellular viability, increased GAG deposition, and elevated early osteogenic markers relative to controls, underscoring its bioactive potential. This study presents a reproducible protocol for generating cell-derived dECM bioinks with enhanced bioactivity for regenerative medicine. Although the approach demonstrates potential for future scale-up, further validation under ISO-compliant frameworks would be required to ensure suitability for production and clinical translation.
Leveraging the accessibility and functional plasticity of human dermal fibroblasts further supports the feasibility of standardized DECM production for bioink formulation. While the current study focuses on process optimization and early functional outcomes, future work will extend biological validation at gene, protein, and histological levels to confirm the osteo- and chondrogenic potential of the optimized dECM system. While additional investigations are warranted to assess long-term regenerative capacity and
Footnotes
Funding Information
The study was funded by The Friends of Guy’s and St Thomas’s Hospital. LRM was supported by King’s Undergraduate Fellowship program.
Disclosure Statement
No competing financial interests exist.
