Abstract
Background
Purpose: Baitouweng decoction (BTWD) is a well-known, classic traditional Chinese medicine prescription primarily used to treat ulcerative colitis, radiation enteritis, and other related diseases. However, the chemical constituents of BTWD remain poorly characterized, limiting understanding of its therapeutic basis, mechanisms of action, and the development of reliable quality control standards. To address this knowledge gap, the comprehensive chemical profiling of BTWD was achieved using a novel method.
Materials and methods
Ultrahigh-performance liquid chromatography-quadrupole time-of-flight mass spectrometry coupled with the Progenesis QI platform was used to rapidly identify the chemical constituents of BTWD.
Results
In total, 119 chemical constituents were successfully identified or tentatively characterized, the majority of which have not been reported in BTWD to date. These compounds were classified into major chemical groups and included 28 alkaloids, 26 terpenoids, 18 flavonoids, 16 phenylpropanoids, 13 organic acids, and 18 other compounds. Seventy-eight of these compounds were the first to be identified in BTWD in this study. This pioneering and systematic characterization could help identify the complex components of BTWD.
Conclusion
The chemical profile of BTWD was elucidated, contributing to the better understanding of its therapeutic material basis and mechanisms of action while supporting the development of modernized quality control and formulation strategies of this classic TCM.
Keywords
Introduction
Baitouweng decoction (BTWD), a commonly used traditional Chinese Medicine (TCM) formula prescription, was first documented in the ancient medical textbook Shang-Han Lun completed by physician Zhongjing Zhang during the Eastern Han Dynasty. BTWD is formulated according to the principles of TCM, and it is composed of Pulsatilla chinensis (the sovereign drug), Phellodendron chinense and Coptis chinensis (the minister drugs), and Fraxinus rhynchophylla (the assistant drug). Traditionally, BTWD is prescribed for its therapeutic effects of clearing heat, detoxifying, cooling the blood, and alleviating dysentery.1,2 These historical applications are well supported by modern pharmacological studies, which have confirmed the potent anti-inflammatory, antimicrobial, anti-endotoxin, immunoregulatory, and antitumor effects of BTWD.2,3 BTWD is widely used in clinical settings to treat several inflammatory and infectious diseases, including ulcerative colitis, bacillary dysentery, and vulvovaginal candidiasis. It also shows potential in treating colorectal and hepatocellular carcinoma.4-8 Recent studies have begun to uncover the molecular mechanisms underlying the efficacy of BTWD, demonstrating that it can alleviate colitis by enhancing autophagy and modulating key signaling pathways such as the PI3K-Akt-mTORC1 and aryl hydrocarbon receptor/IL-22 pathways.9,10 The n-butanol extract of BTWD is effective in treating vulvovaginal candidiasis via activation of the PKCδ/NLRC4/IL-1Ra axis and negative regulation of the NOD-like receptor thermal protein domain-associated protein 3 inflammasome. 5
The therapeutic material basis of BTWD remains poorly understood despite its recognized clinical applications and extensive pharmacological investigations. A comprehensive elucidation of its chemical composition is therefore a critical prerequisite for clarifying its mechanisms of action, developing robust quality control methods, and advancing BTWD-based formulations. However, previous studies have focused either on the separation and detection of a few known components or on a single class of compounds, such as flavonoids.2,11-13 While some studies have profiled the individual herbs in the formula, a systematic and holistic characterization of the final decoction, which is the clinically administered form, is conspicuously absent. Therefore, a comprehensive investigation of the chemical profile of BTWD is urgently needed to bridge this knowledge gap.
Previously, routine identification of the chemical components in TCM relied primarily on the isolation and purification of the compounds in the formulation. Then, the resultant compounds were subjected to instrumental methods of analyses such as high-performance liquid chromatography (HPLC) and thin layer chromatography. Nevertheless, these routine analytical methods are time consuming and mainly support the analysis of standard compounds. Resolving and characterizing the diverse components of TCM formulations remains highly challenging. 14 Considering the complexity of the chemical constituents, most of which are present at very low concentrations in TCM, liquid chromatography coupled with high-resolution mass spectrometry (HRMS) was chosen for this study for the analysis of BTWD. This technique has the advantages of high speed, high resolution, and high sensitivity and is considered suitable for separating and identifying the complex constituents in several TCMs. 15
Ultrahigh-performance liquid chromatography-quadrupole time-of-flight mass spectrometry (UHPLC-Q-TOF-MS) has been routinely used for the identification and characterization of the chemical constituents of TCM.16-19 HRMS-based acquisition technologies for obtaining information on MS/MS fragment ions include two acquisition modes, namely, the data-dependent acquisition (DDA) and data-independent acquisition (DIA) modes. 20 In contrast to the DDA technique, which is typically incapable of collecting fragment information from all precursor ions due to the need for a pre-screening process before fragmentation, the DIA mode harvests fragmentation ions from all detected precursor ions irrespective of their response values. 21 The MSE acquisition mode, a typical DIA technique, can be used to obtain either a full scan of the parent ions via one function at low collision energy or can obtain fragmentation information from all parent ions via another function at higher collision energy. 22
While advanced acquisition techniques, such as MSE, provide comprehensive raw data, the subsequent data-processing steps represent a significant bottleneck. Traditionally, the identification of compounds required the laborious manual extraction of molecular and fragment ions from complex chromatograms, a process that is not only inefficient but also subjective and prone to error. 23 Intelligent data-processing platforms such as Progenesis QI have been developed to overcome these challenges. Progenesis QI is an efficient and powerful intelligence platform that promotes the convenient processing of HRMS data using highly visualized and intelligent workflows. 24 This platform streamlines the analysis of complex HRMS data through a highly automated workflow that includes peak alignment, deconvolution, and compound identification via database matching. Although initially designed for metabolomics, this platform has nowadays been successfully adapted for the comprehensive characterization of TCMs, demonstrating its versatility and reliability.25,26
A rapid and comprehensive analytical strategy was established in this study by integrating the UHPLC-Q-TOF-MS system operating in the MSE mode with the Progenesis QI data-processing platform. The primary objective of this study was the systematic and holistic chemical characterization of BTWD. This foundational work will help provide a definitive chemical profile, creating a scientific basis for future investigations into the therapeutic material basis and mechanisms of action of BTWD and aid in the development of modern quality control standards. The schematic of the study is presented in Figure 1. Schematic of the study
Materials and Methods
Chemicals and Reagents
Methanol and acetonitrile (both MS grade) were acquired from Honeywell (Muskegon, MI, USA). Distilled water was purchased from Watson’s Food & Beverage Co. Ltd (Guangzhou, China). Leucine enkephalin used as a reference material for mass tuning and correction was bought from Waters Technology Co. Ltd (Milford, MA, USA). Sodium formate used to correct the mass axis was procured from Macklin Biochemical Technology Co. Ltd (Shanghai, China). Formic acid (MS grade) added to the mobile phase was obtained from Thermo Fisher (Waltham, MA, USA). The four traditional Chinese medicines, namely, Pulsatilla chinensis, Phellodendron chinense, Coptis chinensis, and Fraxinus rhynchophylla used to prepare BTWD were acquired from Changzhi Tongrentang (Changzhi, China) and authenticated by Prof. Ke Li, Modern Research Center for Traditional Chinese Medicine of Shanxi University, and are shown in Table S1. Information on the 24 reference substances (>98% purity) is presented in Table S2.
Equipment
A Waters Acquity UHPLC I-Class coupled with a Xevo G2-XS Q-TOF mass spectrometer, Waters ACQUITY UHPLC HSS T3 column (100 mm × 2.1 mm, 1.8 μm), MassLynx V4.2 workstation, and Progenesis QI platform were used in this study.
Preparation of BTWD
The following process was used: 37.5 g of P. chinensis, 15 g of C. chinensis, 30 g of P. chinense, and 30 g of F. rhynchophylla were weighed and immersed in 4 L of ultrapure water for 30 min. After heating to boiling over a flame, the water extract was allowed to simmer for 40 min. BTWD was filtered through a 100-mesh gauze to obtain the first extract. The residue was re-extracted with 1200 mL of water using the same procedure to obtain the second extract. Both filtrates were combined and concentrated under vacuum to obtain a yield of 0.5 g/mL.
UHPLC-MS Conditions
Chromatographic separation was conducted using an ACQUITY UHPLC HSS T3 column (100 × 2.1 mm, 1.8 μm, Waters Corporation) using a binary solvent manager and a sampler. The column temperature was maintained at 40°C, and the flow rate was set to 0.35 mL/min. The injection volume was 2 μL for each sample. The optimal mobile phase consisted of a linear gradient elution program wherein 0.1% formic acid in water (solvent A) and 0.1% formic acid in acetonitrile (solvent B) were used as follows: 0 min, 1% B; 0-15 min, 1%-40% B; 15-20 min, 40%-56% B; 20-24 min, 56%-99% B; 24-27 min, 99% B; 27-28 min, 99%-1% B; 28-30 min, 1% B.
High-definition MS was performed using a Xevo G2-XS Q-TOF-MS system (Waters Corporation) equipped with an electrospray ion source in both the positive and negative ion modes. The optimized conditions were as follows: capillary voltage: 2.5 kV (ESI+/ESI−), ion source temperature: 110°C, capillary voltage: 2500 V, cone voltage: 40 V, source offset: 80 V, desolvation temperature: 400°C, cone gas flow: 50 L/h, and desolvation gas flow: 800 L/h in both the positive and negative ion scan modes. The mass scanning range was m/z 50-2000 with a scan time of 0.2 s in the continuum mode during data acquisition. Resolution in the analytical process was 20000 under sensitivity mode. In the MSE mode, low and high collision energies were set as 6 eV and 30-60 eV, respectively.
Progenesis QI Data-Processing Method
Natural compounds in the four herbs constituting BTWD were identified by searching the literature, including the CNKI, WANFANG, and Traditional Chinese Medicine Systems Pharmacology (TCMSP) databases. Based on these compounds, the library for this formula was constructed using a combination of the .SDF file of each compound in BTWD. Raw data gathered using MassLynx software in the MSE acquisition mode were imported into the Progenesis QI platform for data preprocessing, comprising peak alignment, experiment grouping, peak picking and normalization, and deconvolution followed by compound identification. In identifying compounds, deconvoluted data were used to match with a self-built database when setting a series of parameters, such as precursor ion tolerance within 10 ppm, retention time < 0.2 min, and fragment tolerance < 5 ppm. The chemical composition of BTWD was characterized by combining the well-matched precursor ion and fragment ion, which was consistent with the related information in the reported literature and other databases such as the Public Chemical Substances Database and Human Metabolome Database.
Results
UHPLC-MS and Characterization of BTWD
Due to the differential response for each chemical component in BTWD in the positive and negative ion modes, data acquisition for BTWD in both modes was achieved using the MSE pattern in UHPLC-Q-TOF-MS to determine retention time and mass-related information. The chemical constituents were characterized based on the detected actual precursor ions within ±10 ppm from the theoretical value, together with the fragment ions by comparison with those reported in the literature and with the corresponding reference standards. The parent ions determined as [M+H]+ and [M+Na]+, and the other types of parent ions detected as [M-H]– and [M+FA-H]– were respectively acquired in the positive and negative ion modes. Among the 119 compounds that were identified, 28 were alkaloids, 13 were organic acids, 18 were flavonoids, 16 were phenylpropanoids, 26 were terpenoids, and 18 belonged to other classes. As shown in Figure 2, the compounds identified in the positive and negative ion modes are labeled in the total ion chromatograms of the BTWD sample. Details of the identified compounds are presented in Table S3. Comprehensive qualitative analysis of BTWD constituents by UHPLC-Q-TOF-MS. (A) Total ion chromatogram shows the identified constituents in the positive mode with MSE technique. (B) Total ion chromatogram shows the identified constituents in the negative mode with MSE technique
Characterization and Identification of Flavonoids
A total of 18 flavonoids were identified in BTWD. The precursor ion of compound 18 in the negative ion mode was m/z 463.0888 [M-H]–, from which a glucose residue was removed to obtain the fragment ion [M-H-C6H12O5]– having m/z 301.0329. Then, the loss of the neutral water molecule (molecular weight: 18.0106) from m/z 301.0329 [M-H-C6H12O5]– led to the formation of a fragment ion at m/z 283.0962 [M-H-C6H12O5-H2O]–. By comparing the predicted compounds with the available literature and related MS/MS information,
27
compound 18 was confirmed as isoquercetin. The identification process and its possible cleavage pathway are presented in Figure 3. Identification of isoquercetin. (A) MS/MS spectra show related ions in precursor ion scan (lower panel) and fragment ion scan (upper panel) modes. (B) Deductive cleavage pathway of isoquercetin
Characterization and Identification of Alkaloids
A total of 28 alkaloids, including several types of components, were identified from BTWD. As an example, the molecular ion of compound 84 in the positive ion mode was [M]+ at m/z 336.1257, which subsequently produced other observed fragment ions by cleavage at the o-dimethoxy structure. A fragment ion of [M-CH3]+ at m/z 321.0962 was generated after removal of the methyl group, followed by the production of a fragment ion of m/z 306.0738 [M-2CH3]+ on the basis of [M-CH3]+. In another fragmentation pathway, elimination of the CH4 molecule from [M]+ generated m/z 320.0923 [M-CH4]+, which continued to produce a fragment ion of m/z 292.0964 [M-CH4-CO]+, following which, m/z 277.0747 [M-CH4-CO-CH3]+ and m/z 249.0791 [M-CH4-2CO-CH3]+ were produced sequentially via the loss of a methyl group and a CO molecule based on [M-CH4-CO]+. By comparing the retention time of this compound in the total ion chromatogram of the analyzed BTWD sample in the positive ion mode with the reference standard berberine in Figure 4A and then searching the literature for the related compound,
28
compound 84 was identified as berberine. The presumed fragmentation pathway of its MS/MS information is depicted in Figure 4. As seen in Figure 4B, the abundance of fragment ions to molecular ions for berberine was obviously augmented under high collision energy compared with that under low energy. Identification of berberine. (A) Extracted ion chromatogram of berberine reference standard. (B) MS/MS spectra show related ions in precursor ion scan (lower panel) and fragment ion scan (upper panel) modes. (C) Deductive cleavage pathway of berberine
Characterization and Identification of Terpenoids
A total of 26 terpenoids were identified in BTWD. The precursor ion at m/z 1265.6230 [M+FA-H]− was determined in the negative ion mode. Meanwhile, the quasi-molecular ion of compound 92 was m/z 1219.6146 [M-H]−, which lost a molecule of rhamnose and two molecules of glucose to yield m/z 749.4529 [M-H-Rha-2Glc]−. This fragment ion continued to lose rhamnose residues to produce a characteristic fragment ion of m/z 603.3889 [M-H-2Rha-2Glc]− as noted in the mass spectrum. Moreover, the ion of the oligosaccharide chain at m/z 471.3481 can be seen in the mass spectra. It could form another ion at m/z 469.1544 via an elimination reaction. The m/z 469.1544 fragment lost a rhamnose residue to yield an ion at m/z 323.0992, which then further generated an ion at m/z 161.0452 after the loss of a glucose residue. In addition, some cross-cleavage ions, such as m/z 409.1342 and m/z 367.1248, could be seen in the mass spectrum. The ion at m/z 247.0827 was also observed with the cleavage of the glycosidic bond of m/z 409.1342. Compound 92 was speculated to be anemoside B4 by referring to the literature,29,30 analysis of the molecular weight of anemoside B4 (1220.6190) based on the monoisotopic mass for each atom using HRMS, and comparison of the retention time of compound 92 with that of the reference standard anemoside B4. Its proposed fragmentation pathway is shown in Figure 5. Identification of anemoside B4. (A) Extracted ion chromatogram of anemoside B4 reference standard. (B) MS/MS spectra show related ions in precursor ion scan (lower panel) and fragment ion scan (upper panel) modes. (C) Deductive cleavage pathway of anemoside B4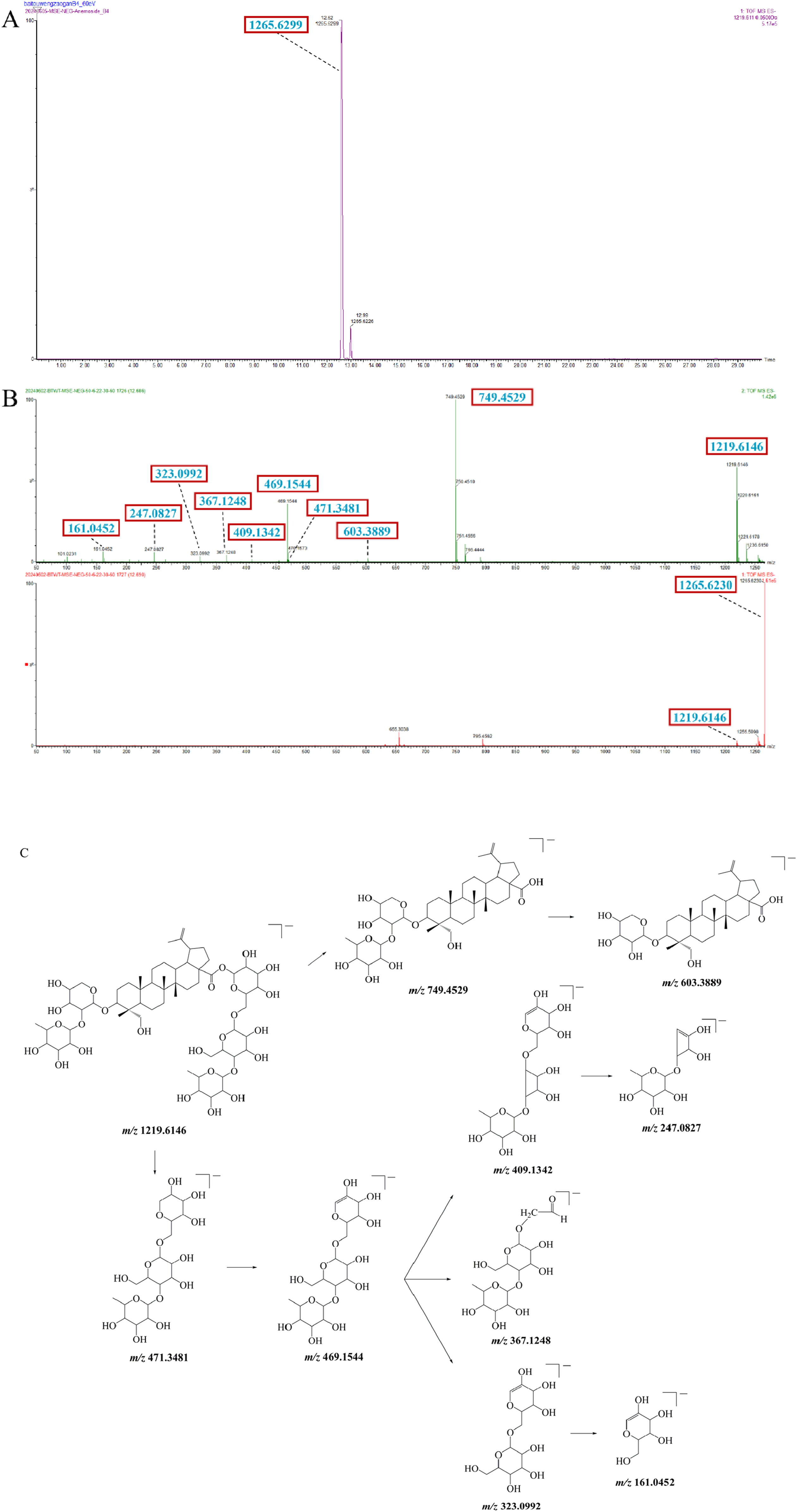
Characterization and Identification of Phenylpropanoids
A total of 16 phenylpropanoids were identified in BTWD. The excimer ion of compound 14 was m/z 177.0200 [M-H]−, which could lose a CO molecule to generate a characteristic ion of m/z 149.0231 [M-H-CO]− in the negative ion mode. Moreover, the excimer ion underwent the neutral loss of CO2 to yield the corresponding ion at m/z 133.0284 [M-H-CO2]−, which subsequently lost a CO or CO2 molecule to produce two typical fragment ions at m/z 105.0324 [M-H-CO2-CO]− and m/z 89.0379 [M-H-2CO2]−. Based on this information on the excimer ion and the fragment ions in the MSE spectra and comparison with the reference standard and the mass spectrometry information in the literature,
31
compound 14 was identified to be aesculetin. Its possible cleavage route is depicted in Figure 6. Identification of aesculetin. (A) Extracted ion chromatogram of aesculetin reference standard. (B) MS/MS spectra show related ions in precursor ion scan (lower panel) and fragment ion scan (upper panel) modes. (C) Deductive cleavage pathway of aesculetin
Characterization and Identification of Organic Acids
A total of 13 organic acids were identified in BTWD. The excimer ion of compound 4 was m/z 181.0486 [M+H]+, which produced a characteristic fragment ion of m/z 163.0607 [M+H-H2O]+ after losing a water molecule and subsequent hydrogen rearrangement owing to the easy loss of a hydroxyl moiety from the carboxyl group. Subsequently, m/z 145.0485 [M+H-2H2O]+ was generated due to the loss of another water molecule from the [M+H-H2O]+ fragment ion. In addition, the carboxyl group is mainly removed to form a fragment ion at m/z 137.0610 [M+H-CO2]+ at low collision energy based on m/z 181.0486 [M+H]+. This fragment ion continued to yield a radical ion at m/z 136.0567 with the homolytic cleavage of the covalent O-H bond, and the radical ion then further generated a fragment ion at m/z 135.0418 after the loss of a hydrogen radical. Based on the comparison of this obtained MS/MS information with the literature,32-34 compound 4 was inferred to be caffeic acid. Its MS/MS spectra and proposed fragmentation pathway are depicted in Figure 7. Identification of caffeic acid. (A) MS/MS spectra show related ions in precursor ion scan (lower panel) and fragment ion scan (upper panel) modes. (B) Deductive cleavage pathway of caffeic acid
Characterization and Identification of Other Compounds
A total of 18 other types of compounds were identified in BTWD. For example, the excimer ion of compound 5 was m/z 299.1103 [M-H]−, which generated two characteristic fragment ions at m/z 179.0473 [M-H-C8H8O]− and m/z 119.0024 [M-H-C6H12O6]− via the cleavage of glycosidic bonds under high collision energy. Compound 5 was identified as salidroside based on the MS/MS information of its fragment ions and comparison with the literature.
32
The MS/MS spectra and proposed fragmentation pathway are presented in Figure 8. Identification of salidroside. (A) MS/MS spectra show related ions in precursor ion scan (lower panel) and fragment ion scan (upper panel) modes. (B) Deductive cleavage pathway of salidroside
Discussion
BTWD is a commonly used TCM prescription that is composed of four herbs. In this study, 119 components, including flavonoids, alkaloids, terpenoids, organic acids, phenylpropanoids, and other compounds, were identified in BTWD using UHPLC-Q-TOF-MSE in combination with the intelligent Progenesis QI data-processing platform.
The MSE method used in this study was a DIA method. This technique can be used to determine the intensity of parent ions as well as the fragment ions produced from parent ions subjected to collision energy, as opposed to the DDA method, which is used only for scanning and detecting the fragment ions generated from the prescreened parent ions. 35 Thus, the MSE method can be used to elucidate the chemical profile of TCMs. Furthermore, UHPLC-HRMS coupled with UNIFI analysis software has been widely used to identify the compounds in TCMs that are composed of various complex chemical components.36-39 Nevertheless, background interference is always observed in the mass spectra of TCM samples, posing a challenge faced when using the UNIFI tool. The use of Progenesis QI can overcome this problem by importing the blank sample and processed TCM sample to eliminate background interference. Although this integrated strategy can improve the identification process, further refinement is required for the elucidation of unknown components.
BTWD is known for its properties of heat clearing, detoxification, cooling blood, and stopping dysentery.1,2 Characterization of the components of BTWD led to the identification of bioactive compounds, such as anemoside B4 (AB4), hederacoside C, and anemoside A3, which can alleviate the ailments stated above. AB4 exerts an anti-inflammatory effect to alleviate acute ulcerative colitis by downregulating the TLR4/NF-κB/MAPK signaling pathway. 40 In addition, it demonstrates potential as an antiviral agent owing to its adjustments of the immune responses. 3 Anemoside A3, a metabolite of AB4, exerts an immunomodulatory effect by modulating prostaglandin E receptor 4 signaling and inhibiting the activation of signal transducer and activator of transcription 4 and 3. 41 Studies suggest that hederacoside C alleviates colitis by recovering the function of the injured intestinal barrier by regulating S100A9/MAPK. 42 Moreover, aesculin and aesculetin, the phenylpropanoids from Fraxinus rhynchophylla, demonstrate antibacterial, immunomodulatory, and anti-inflammatory effects. 43 Phellodendrine and berberine have been identified in BTWD, with the former exerting a therapeutic effect in alleviating ulcerative colitis, inflammatory diseases, and delayed-type hypersensitivity, 44 and the latter demonstrating potential in treating ulcerative colitis. 45 Therefore, elucidation of the chemical profile of BTWD could contribute to our understanding of its mechanism of action in heat-clearing and detoxification, cooling blood, and stopping dysentery from the compound perspective.
Although our study offers valuable insights into the chemical profile of BTWD, it has a few limitations. The challenge is the analysis of complex MS data, as the isomers in BTWD can complicate the characterization of compounds, particularly without the concurrent use of ion mobility mass spectrometry. Further research is needed to explore the pharmacological effects and molecular mechanisms of these metabolites. Such studies will clarify the role of BTWD in treating related diseases and support the discovery of novel bioactive molecules for use in a clinical setting.
Conclusions
UHPLC-Q-TOF-MSE coupled with the Progenesis QI platform was used in this study for the rapid and comprehensive chemical profiling of BTWD. A total of 119 compounds, including alkaloids, flavonoids, terpenoids, phenylpropanoids, organic acids, and others, were identified or tentatively characterized. Among the identified compounds, 78 were probably the first to be identified in BTWD. Furthermore, the characteristic fragmentation pathways of the representative components were elucidated. The comprehensive chemical characterization presented herein lays the foundation for future investigations into the mechanisms of action for BTWD, discovery and validation of quality markers, and development of standardized, modern formulations of this important, traditionally used formula.
Supplemental Material
Supplemental Material - Rapid and Comprehensive Identification of the Chemical Constituents in Baitouweng Decoction Using Ultrahigh- Performance Liquid Chromatography-Quadrupole Time-of-Flight Mass Spectrometry Alongside the Progenesis QI Platform
Supplemental Material for Rapid and Comprehensive Identification of the Chemical Constituents in Baitouweng Decoction Using Ultrahigh- Performance Liquid Chromatography-Quadrupole Time-of-Flight Mass Spectrometry Alongside the Progenesis QI Platform by Kaiqiang Wang, Haoze Ping, Yaqi Wang, Xingpiao Jin, Baolian Ma, Yanbin He and Kefeng Li in Natural Product Communications.
Footnotes
Ethical Considerations
Ethical approval is not required for this study.
Consent to Participate
No human subjects or animals were used in this study; therefore, a statement on informed consent is not applicable.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: We acknowledge funding by the “Three Districts” Science and Technology Talent Support Plan of Shanxi Province (Year 2022 for Changzhi Medical College), the Applied Basic Research Program of Shanxi Province, China (Grant No. 202303021211109), and Shanxi Province College Students’ Innovation and Entrepreneurship Project (Grant No. 20240970).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The data generated in this study are available from the corresponding author on reasonable terms.
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
