Abstract
Background
SARS-CoV-2 continues to evolve, with new variants showing higher transmissibility and immune evasion. The limited effectiveness of current antiviral drugs highlights the potential of natural product compounds, such as eurycomanone as promising alternatives.
Objective
This study evaluated the antiviral efficacy of eurycomanone against SARS-CoV-2 variants through experimental and computational analyses.
Methods
The antiviral activity of eurycomanone was tested in vitro using a cytopathic effect-based assay with Vero E6 cells infected with the original Wuhan-wild-type (WT), Alpha, Delta, and Omicron variants. Cytotoxicity was measured using an ATP-based assay. Computational analysis was performed through molecular docking and dynamics simulations to analyse its binding with SARS-CoV-2 main protease (Mpro) in both WT and Omicron variants.
Results
Eurycomanone exhibited the strongest antiviral activity against the Omicron variant (EC50: 4.58 µM) among all tested variants, with low cytotoxicity (CC50 > 100 µM) as determined by a cell viability assay in Vero E6 cells. Docking studies revealed strong binding affinities to Mpro (WT: −7.7 kcal/mol; Omicron: −8.2 kcal/mol), comparable to nirmatrelvir. Molecular dynamics confirmed stable binding and MMGBSA calculations showed enhanced hydrophobic and electrostatic interactions with the Omicron variant.
Conclusion
Eurycomanone exhibited notable antiviral activity especially against Omicron, suggesting its potential as a lead compound for developing new antiviral therapies. Further in vivo studies are needed to explore its therapeutic potential.
Introduction
The COVID-19 pandemic caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) continues to present significant global health challenges. Despite achieving herd immunity through the rapid development of vaccines, the emergence of new variants of concern (VOC) has highlighted the need for ongoing antiviral treatments. The VOC of SARS-CoV-2, such as Alpha (B.1.1.7), Delta (B.1.617.2), and Omicron (B.1.1.529) have emerged since the pandemic began in early 2020. 1 Among these, the Omicron variant has shown the highest transmissibility, immune evasion capabilities, and the potential to compromise the effectiveness of existing therapeutic interventions.2,3 Several antiviral drugs such as remdesivir, molnupiravir, and nirmatrelvir have shown effective activity against SARS-CoV-2 and its VOC. 4 These drugs target conserved proteins rather than the highly variable spike protein, ensuring their sustained activity against emerging variants. However, their effectiveness may decline over time as viral mutations lead to resistance, especially in highly transmissible variants.3,5 Developing alternative drugs ensures broader treatment options and better control of the virus. During the early stages of the pandemic, numerous antiviral research strategies focused on targeting viral enzymes, such as the main protease (Mpro) to identify potential inhibitors for viral replication. 6 Mpro inhibitors such as nirmatrelvir, have been identified and proven effective in clinical settings, significantly reducing the severity and progression of COVID-19. 7
Natural products are a vital source for drug discovery due to their structural diversity and pharmacological potential.
8

Molecular structure of eurycomanone with functional groups (Pink: Hydroxyl, Blue: Carbonyl).).
Despite advancements in antiviral development, the continued evolution of SARS-CoV-2 especially mutations affecting key proteins, poses a challenge to long-term treatment success. For example, nirmatrelvir, which was once highly effective has shown signs of resistance in some variants, particularly due to mutations in the Mpro. 14 These mutations can reduce drug binding efficiency and allow the virus to escape inhibition, emphasizing the need for complementary treatment strategies. 15 Further ligand-protein interaction modelling is required to explore how eurycomanone interacts with key viral proteins in different SARS-CoV-2 variants. This drug interaction study may support in vitro testing of eurycomanone to explore its potential against certain SARS-CoV-2 variants, especially when existing antiviral drugs are less effective.
The present study investigates the antiviral potential of eurycomanone against SARS-CoV-2 and its VOC through experimental and computational methods. We tested its efficacy against the original Wuhan-wild-type (WT), Alpha, Delta, and Omicron variants using in vitro assays in Vero E6 cells. To explore its potential mechanisms of action, we performed molecular docking and dynamics studies to evaluate its binding affinity to Mpro targets (PDB IDs: 6LU7 for WT variant and 7TLL for Omicron variant). These approaches provide a clear characterisation of the antiviral properties of eurycomanone. This research enhances the understanding of natural product-derived antivirals and positions eurycomanone as a promising lead for developing therapeutics against SARS-CoV-2 and its variants.
Material and Methods
Chemicals
The stocks of the tested compound eurycomanone (99.91% purity, TargetMol, USA) and the control drug nirmatrelvir (Cayman Chemical, USA) were prepared by dissolving in 100% dimethyl sulfoxide (DMSO) at a concentration of 20 mM and stored at −20 °C prior to use. The authenticity of eurycomanone was verified using liquid chromatography-mass spectrometry (LCMS) to confirm its purity prior to further use in experiments. The LC-MS profile of eurycomanone is shown in Supplemental Figure S1.
Cells and Virus
Clinical SARS-CoV-2 variants, including Wuhan wild-type (WT), Alpha, Delta, and Omicron (B.1.1.529), were previously isolated and cultured by the Virology Unit, Institute for Medical Research, National Institutes of Health, Malaysia. Viral stocks were cultured in Vero E6 cells (ATCC® CRL-1586™) maintained in Dulbecco's Modified Eagle Medium (DMEM) supplemented with 2% fetal bovine serum (FBS) at 37 °C under 5% CO2 in a biosafety level 3 (BSL-3) containment facility. Virus isolation from human samples was not considered human subject research, and the requirement for written informed consent was exempted by the Medical Research and Ethics Committee (MREC), Ministry of Health, Malaysia.
Cytotoxicity Assays
The cytotoxicity of eurycomanone was assessed in Vero E6 cells using an ATP-based luminescence assay, following the CellTiter-Glo Luminescent Cell Viability Assay (Promega, USA) protocol described in the product insert. 16 Briefly, Vero E6 cells were seeded into 96-well plates and incubated overnight. After cell attachment, cells were treated with a series of eurycomanone concentrations ranging from 0.098 µM to 25 µM. Untreated cells served as the positive control for cell viability, while cells treated with 0.05% Triton X-100 served as the negative control. After 72 h of incubation at 37 °C in a 5% CO2 incubator, the CellTiter-Glo reagent was added to quantify intracellular ATP as an indicator of metabolically active cells. Luminescence was measured using a FLUOstar Omega microplate reader (Germany). The results were used to generate dose-response curves and to determine the half-maximal cytotoxic concentration (CC50).
Anti-SARS-CoV-2 Activity
The antiviral activity of eurycomanone was evaluated against SARS-CoV-2 variants of concern (VOCs), including the WT, Alpha, Delta, and Omicron strains using infected Vero E6 cells. Infected cells were treated with a range of eurycomanone concentrations (0.098 µM to 25 µM) and incubated for 72 h at 37 °C with 5% CO2. Infected cells with 0.5% DMSO served as the negative control, while healthy cells with 0.5% DMSO were used as the positive control. Nirmatrelvir was included as a reference antiviral drug and was tested in parallel using the same concentration range and assay conditions. The Viral ToxGlo Assay (Promega, USA) protocol was used to assess the cytopathic effect. 17 After incubation, the Viral ToxGlo reagent was added as described in the assay protocol, and luminescence was measured using a GloMax® Discover microplate reader (Promega, USA). The luminescence data were used to generate dose-response curves and to determine the half-maximal effective concentration (EC50) through nonlinear regression analysis in GraphPad Prism (version 10.0). The selectivity index (SI) was then calculated.
Microscopic Observation of Infected Cells
To assess cytopathic effects, microscopic analysis was performed on wells infected with the WT variant after eurycomanone treatment. Images were captured using an inverted phase-contrast microscope (Olympus CKX53, Olympus Corporation, Tokyo, Japan) at 100× magnification, focusing on the centre of each well. Cell morphology was evaluated to assess virus-induced damage and the protective effects of the compound, following the protocol outlined by the American Society for Microbiology for cytopathic effect observation. 18 Eurycomanone was tested at concentrations ranging from 3.67 µM to 100 µM, with the lowest dose selected based on initial antiviral screening. The highest concentration was included to evaluate potential cellular recovery or saturation effects at elevated doses. The WT variant was used in this analysis as it was expected to display cytopathic effects representative of all tested variants, allowing a reliable assessment of morphological outcomes.
Data Analysis
The dose-response curves were generated using nonlinear regression analysis in GraphPad Prism (version 10.0), applying the inhibitor versus normalized response model with a variable slope (XY analysis method). From these curves, the effective concentration (EC50) and cytotoxic concentration (CC50) values were calculated. Both EC50 and CC50 represent the concentration of compound that produces 50% of the maximal response, based on the four-parameter logistic equation:
ADMET and Molecular Docking
Prior to docking, ADMET (Absorption, Distribution, Metabolism, Excretion, and Toxicity) screening of both compounds was performed using the SwissADME web tool. 23 This predictive analysis was used to evaluate physicochemical properties, pharmacokinetics, and drug-likeness to support their potential as drug candidates.
The 3D structure of eurycomanone was obtained from PubChem in SDF format, and nirmatrelvir was included as a reference Mpro inhibitor. The crystal structures of the SARS-CoV-2 main protease were retrieved from the Protein Data Bank (PDB). Two structures were used: the wild-type (WT) variant complexed with the N3 inhibitor (PDB ID: 6LU7) and the Omicron variant complexed with PF-07321332 (PDB ID: 7TLL). Nirmatrelvir was included as a reference Mpro inhibitor. Protein preparation followed the AutoDock Tools (ADT) protocol, which involved removal of water molecules and heteroatoms, addition of hydrogen atoms, and assignment of Gasteiger charges. 24 Protonation states were adjusted using PROPKA3 to reflect physiological conditions. 25 The docking grid was cantered on the active site (x = −13.00 Å, y = 15.90 Å, z = 68.20 Å) with a box dimension of 20 Å on each side. Docking simulations were performed using AutoDock Vina through the AMDock graphical interface. 26 The docked complexes were visualized and analysed using PyMOL and Discovery Studio Visualizer to evaluate protein-ligand binding interactions.
Molecular Dynamic Simulation
Molecular dynamics (MD) simulations were performed using GROMACS version 2022. 27 The protein-ligand complexes were taken from the docking results. The CHARMM27 all-atom force field was applied for protein topology, while ligand topologies and parameters were generated using the SwissParam web toolvto ensure compatibility with the CHARMM force field.28,29 Each complex was placed in a dodecahedron box filled with TIP3P water molecules, maintaining a 1.0 nm distance from the box edge. Sodium ions (Na+) were added to neutralise the system. Energy minimisation was carried out using the steepest descent algorithm for up to 50,000 steps until the maximum force fell below 10.0 kJ/mol. The systems were then equilibrated under NVT (constant volume) and NPT (constant pressure) ensembles for 1 ns each, maintaining 300 K temperature and 1 bar pressure using the Berendsen thermostat. 30 Electrostatic interactions were treated using the Particle Mesh Ewald (PME) method with a 10 Å cut-off. 31 The production MD run was conducted for 100 ns, and snapshots were saved every 10 ps. post-simulation analyses were conducted using GROMACS tools: gmx rms for root mean square deviation (RMSD), gmx rmsf for root mean square fluctuation (RMSF), and gmx hbond for hydrogen bond analysis, using a standard cut-off of 3.5 Å (0.35 nm). Visual Molecular Dynamics (VMD) and PyMOL were employed for trajectory visualisation. Additionally, analysis plots were generated using the gnuplot program. The average RMSD and RMSF values were derived from the final 100 ns of molecular dynamic simulation trajectories.
Results
Antiviral and Cytotoxicity of Eurycomanone Against SARS-CoV-2 Variants
The antiviral activities of eurycomanone were assessed against the original WT strain and three SARS-CoV-2 variants of concern: Alpha, Delta, and Omicron. As shown in Figures 2a to 2d, eurycomanone inhibited viral replication in a dose-dependent manner across all tested variants, with EC50 values of 6.88 µM (WT), 8.34 µM (Alpha), 11.94 µM (Delta), and 4.58 µM (Omicron). Cytotoxicity testing in Vero E6 cells showed no significant toxicity up to 100 µM, confirming high cell viability across all treatment concentrations. Based on the CC50 (>100 µM), the corresponding selectivity indices (SI) were >14.53 (WT), >11.99 (Alpha), >8.38 (Delta), and >21.83 (Omicron), indicating a favourable safety and efficacy profile. Among all tested variants, Omicron was the most responsive to eurycomanone treatment. In comparison, the reference compound nirmatrelvir demonstrated greater potency across all strains, with an EC50 of 1.11 µM and a selectivity index exceeding 90.10, as shown in Supplemental Figure S2.
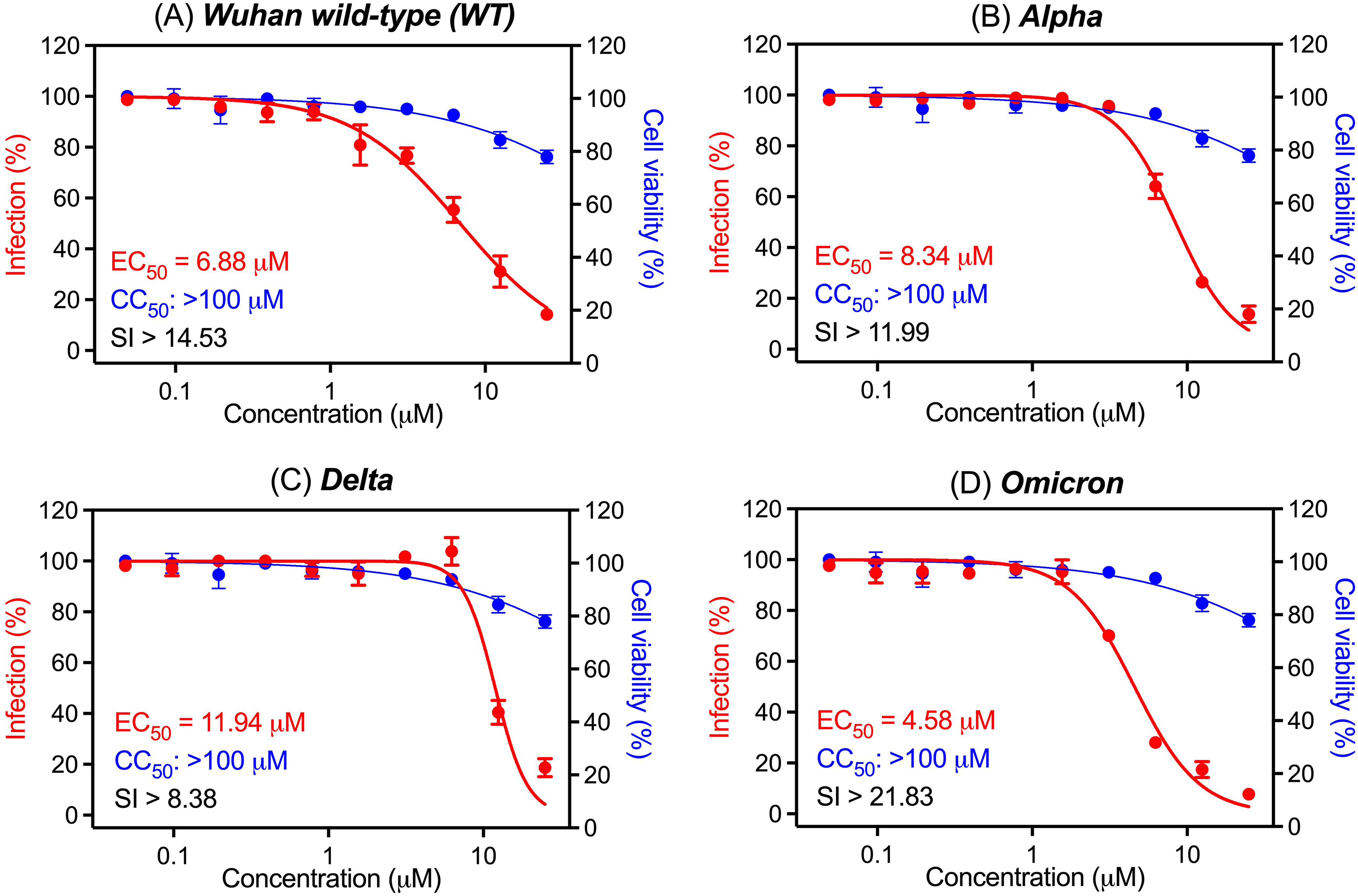
Antiviral and cytotoxic activities of eurycomanone against SARS-CoV-2 variants: (a) Wuhan wildtype (WT), (b) Alpha, (c) Delta, and (d) Omicron variants. Dose-response curves show the percentage of infection (red) and cell viability (blue) in Vero E6 cells treated with increasing concentrations of eurycomanone and incubated for 72 h at 37 °C in a 5% CO2 incubator. The infection was measured by CPE inhibition assay, while cell viability was assessed using an ATP-based assay. All data points represent mean values, with error bars showing the standard error of the mean (SEM) from three independent experiments.
Morphological Evaluation of Eurycomanone Treatment in SARS-CoV-2 WT-Infected Cells
To assess the morphological effects of eurycomanone treatment, microscopic analysis was conducted on Vero E6 cells infected with the WT SARS-CoV-2 variant. As shown in Figure 3, cell morphology was examined following infection and treatment with increasing concentrations of eurycomanone. Uninfected control cells displayed healthy, dense, and well-organized monolayers with no signs of cytopathic effects (CPE). In contrast, infected but untreated cells exhibited pronounced CPE, including cell rounding, detachment, and a large central area of cell loss. Eurycomanone treatment led to a concentration-dependent improvement in cell morphology. At 3.67 µM, limited recovery was observed, with partial cell attachment and continued central disruption. Morphological restoration became more noticeable at 6.13 and 12.50 µM, where CPE was visibly reduced and cell density improved. At higher concentrations (25, 50, and 100 µM), the monolayers appeared largely intact and closely resembled the uninfected control, indicating strong protection against virus-induced damage without visible cytotoxicity. These findings support the concentration-dependent protective effect of eurycomanone on infected Vero E6 cells.

Microscopic analysis of cytopathic effects in SARS-CoV-2 WT-infected Vero E6 cells treated with eurycomanone. Vero E6 cells were infected with the SARS-CoV-2 wild-type (WT) variant and treated with increasing concentrations of eurycomanone (3.67-100 μM) for 72 h. Cell morphology was observed to assess virus-induced damage and the protective effects of treatment. Non-infected and untreated infected cells served as controls. Images were captured using an inverted phase-contrast microscope (Olympus CKX53, Japan) at 100× magnification.
Microscopic evaluation was limited to the WT strain, as cytotoxicity remained consistent across all tested variants, and the WT represents the original reference strain used in early antiviral screening. Additionally, the dose-response curves for Alpha, Delta, and Omicron showed similar patterns of inhibition, supporting the use of WT as a representative model for visual comparison of virus-induced morphological changes.
ADMET, Molecular Docking and Binding Affinity Analysis
Eurycomanone shows good drug-like characteristics as shown in Table 1, including no violations of Lipinski's rule, a suitable molecular weight, and excellent water solubility. However, its high polarity (TPSA = 153.75 Å2) may limit oral absorption, suggesting the need for specialized formulation strategies. Nirmatrelvir displays a better pharmacokinetic profile, fulfilling most key drug-likeness criteria, with favourable membrane permeability and higher gastrointestinal absorption. These comparisons make nirmatrelvir as a useful reference for optimizing the oral bioavailability and drug development potential of eurycomanone. The detailed ADMET and drug-likeness properties of eurycomanone and nirmatrelvir are provided in Supplemental Table S1.
Drug-Likeness Evaluation of Eurycomanone and Nirmatrelvir.

Eurycomanone had a binding affinity of −7.7 kcal/mol for the WT variant (6LU7) and −8.2 kcal/mol for the Omicron variant (7TLL), while nirmatrelvir showed −7.7 kcal/mol for 6LU7 and −8.8 kcal/mol for 7TLL, as shown in Table 2. Although nirmatrelvir showed a slightly higher binding affinity for the Omicron variant, the comparable binding affinities of eurycomanone suggest its significance in investigating interactions with viral proteases.
Binding Affinities of Eurycomanone and Nirmatrelvir to the SARS-CoV-2 Mpro Targets.

The plausible binding of eurycomanone and nirmatrelvir within the active sites of WT variant and Omicron highlights their similarity in binding, as shown in Figure 4. Eurycomanone occupied the catalytic cleft of both WT and Omicron Mpro but engaged distinct residues. In the WT complex, its carbonyl formed a hydrogen bond with H163, while hydroxyl groups interacted with N142 and H172, supported by hydrophobic contact with L141. In Omicron, the carbonyl shifted toward E166, with hydroxyl groups binding N142 and H164, further stabilized by proximity to C145 and hydrophobic packing with M49 and L141. Nirmatrelvir, as the reference inhibitor, consistently interacted with key residues H41, C145, N142, E166, and Q189 across both variants. The catalytic dyad (H41–C145) was more prominently engaged in Omicron, while less exposed in the WT, and electrostatic mapping confirmed both ligands were well accommodated within the catalytic cleft. These observations support the similarity in binding mode between eurycomanone and nirmatrelvir and suggest a conserved inhibitory mechanism despite structural differences between the two compounds. Since the WT represents the original viral strain and eurycomanone already shows promising experimental activity against multiple variants, docking against WT and Omicron is sufficient to evaluate its antiviral potential without requiring simulation of all variant structures.
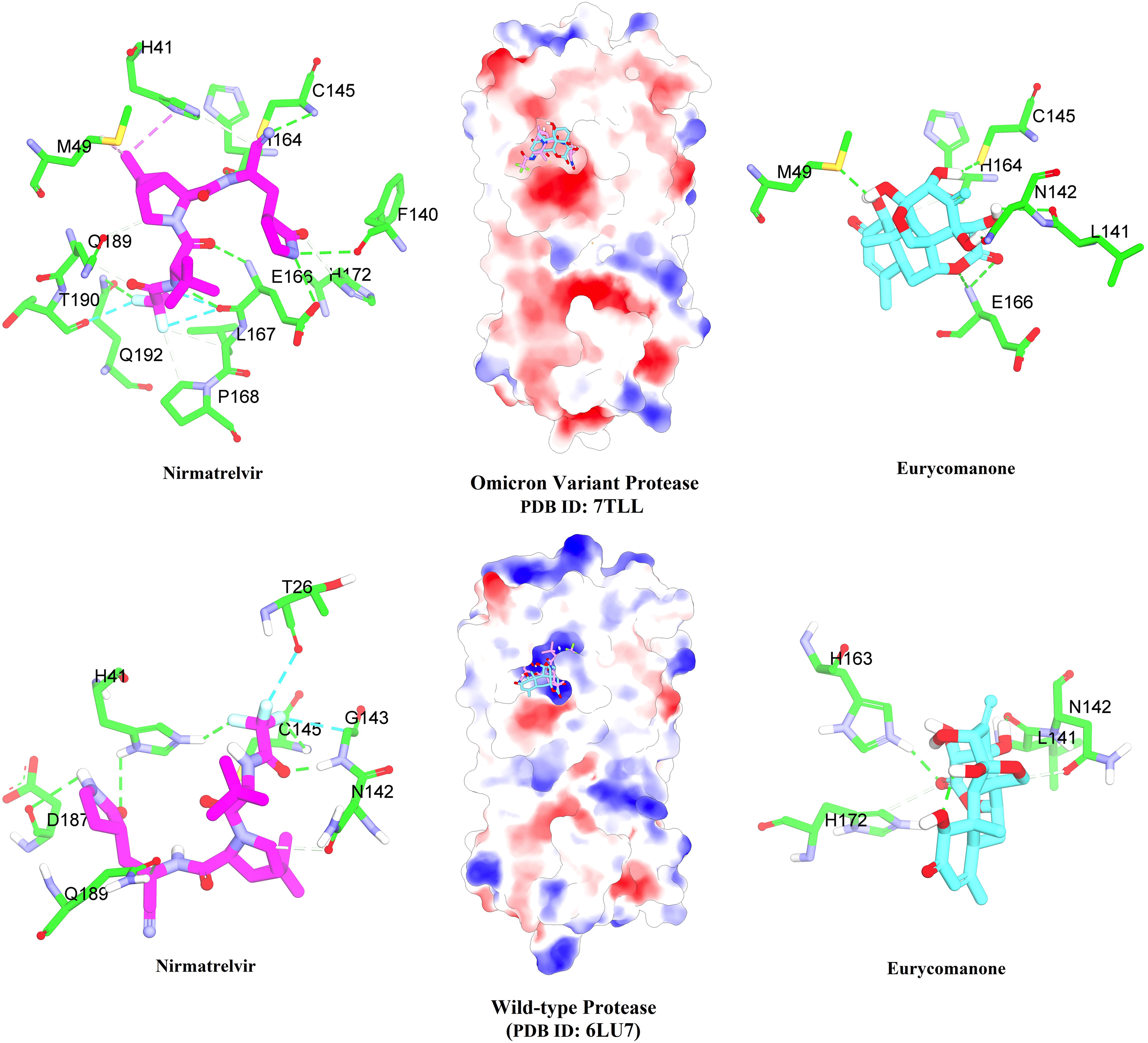
Molecular docking interactions of eurycomanone and nirmatrelvir with the SARS-CoV-2 main protease (Mpro). Overview of the positioning of eurycomanone and nirmatrelvir in the binding sites of the Omicron variant (PDB ID: 7TLL) and wild-type (WT) protease (PDB ID: 6LU7). Eurycomanone (cyan) and nirmatrelvir (purple) are shown interacting with key active-site residues (green). Hydrogen bonds are depicted as green dashed lines. Surface models at the centre illustrate ligand binding within the protease cavities.
Molecular Dynamics Simulation and Stability Analysis
The stability of the eurycomanone-protease complexes was analysed through molecular dynamics (MD) simulations, focusing on root mean square deviation (RMSD), root mean square fluctuation (RMSF), and solvent accessible surface area (SASA) (Supplemental Figure S3-S8). In the analysis of the 6LU7, the RMSD for the eurycomanone-bound complex was measured at 0.21 nm during the 100 ns simulation, which is slightly higher than that of the nirmatrelvir-bound complex, recorded at 0.15 nm. This finding indicates that the eurycomanone complex had slightly greater flexibility. However, the RMSF values were similar for both compounds, suggesting they interact with the protease residues comparably. For the 7TLL, the RMSD for the eurycomanone complex was 0.17 nm, while that for the nirmatrelvir complex was 0.14 nm. The minimal differences in RMSD and SASA values between eurycomanone and the reference compound indicate that eurycomanone also potentially has stable binding interactions with the Omicron Mpro. Although nirmatrelvir showed slightly better stability in terms of RMSD, eurycomanone still demonstrated favourable dynamics, supporting its continued evaluation as an antiviral compound.
Molecular Mechanics with Generalised Born and Surface Area Solvation (MMGBSA) Free Energy Calculations
The MMGBSA analysis highlights promising antiviral potential of eurycomanone particularly against the Omicron variant compared to the WT variant protease. As shown in Table 3, eurycomanone demonstrates a stronger overall binding interaction with the 7TLL, indicating a greater affinity that aligns well with its potential as an antiviral candidate. This difference in affinity between the two variants is partly attributed to the favorable van der Waals (ΔG_vdW) contributions observed in the 7TLL complex, which suggests enhanced hydrophobic interactions in the Omicron protease. Additionally, the electrostatic energy (ΔG_elec) and solvation free energy (ΔG_solv) contributions further support stability of eurycomanone in the Omicron active site, where ΔG_elec shows significant ionic interactions, and ΔG_solv stabilizes the complex in the solvent environment. Eurycomanone shows competitive binding characteristics compared to nirmatrelvir, particularly in the ΔG_vdW and ΔG_solv components, highlighting its potential efficacy. The total binding free energy (ΔTotal) consistently favours eurycomanone in the 7TLL variant, suggesting its suitability as a candidate for targeting SARS-CoV-2 Omicron variant infections.
The Binding Free Energies (ΔG) for Eurycomanone and Nirmatrelvir for 6LU7 and 7TLL Proteases.

Per-Residue Interaction Analysis
A per-residue energy decomposition analysis was conducted to identify key interactions of eurycomanone with Mpro (6LU7 and 7TLL). Only residues within proximity of ≤ 4 Å of each compound were considered, as these close-contact residues typically play significant roles in ligand binding and stability. Eurycomanone exerted stronger interactions with key residues of 7TLL than 6LU7, with significant energy contributions from residues GLN189, GLU166, and MET165, which are critical for binding stability (Figure 5a). Figure 5b shows that eurycomanone forms stronger interactions than nirmatrelvir, especially with the Omicron variant protease. Eurycomanone maintains a stable binding profile, with GLU166 and MET165 exhibiting higher energy decomposition values. This suggests stronger binding in Omicron compared to the WT protease. Unlike nirmatrelvir, which also interacts substantially, eurycomanone shows comparable or even better affinity at specific 7TLL residues. This finding highlights the potential of eurycomanone as a promising alternative and justifies further discovery of its antiviral properties, especially against Omicron variant.

Per-residue interaction energy decomposition of eurycomanone and nirmatrelvir with SARS-CoV-2 main protease (Mpro). (A) Interaction energies with the wild-type protease (PDB ID: 6LU7). (B) Interaction energies with the Omicron variant protease (PDB ID: 7TLL). Total energy contributions (kcal/mol) from key active site residues are shown for eurycomanone (maroon) and nirmatrelvir (grey). Negative values indicate stabilizing interactions.
Discussions
Global health continues to face challenges from the persistent threat of SARS-CoV-2 and its rapidly evolving variants, particularly those that are highly transmissible and immune-evasive. Existing antiviral drugs, such as remdesivir, molnupiravir, and nirmatrelvir, have played crucial roles in managing COVID-19 but they have significant limitations. 4 These drugs often lose effectiveness against new variants due to the SARS-CoV-2 mutations. Nirmatrelvir, a well-known antiviral approved by the FDA is recognized for its high potency against SARS-CoV-2. However, immune-evasive variants adapt to resist protease inhibitors and make nirmatrelvir less effective. 32 These limitations highlight the urgent need for alternative drugs mainly from natural compounds with broader antiviral potential. Given these challenges, researchers are using natural product compounds for new antiviral solutions.
Natural product compounds offer unique chemical diversity and have historically provided the foundation for many effective medicines.
8
Our study focused on eurycomanone, a quassinoid from
The cytotoxicity profile of eurycomanone (CC50 > 100 µM) supports its potential as a safe antiviral compound. In comparison, a previous study using the same ATP-based cell viability method reported that andrographolide displayed cytotoxicity with a CC50 value of 40.19 µM, chloroquine with a CC50 value of 54.08 µM, while remdesivir exhibited no cytotoxic activity with a CC50 value greater than 200 µM. 22 In our experiment, the control drug nirmatrelvir also showed minimal cytotoxicity with a CC50 value greater than 200 µM. These results indicate that eurycomanone demonstrates a more favourable cytotoxicity profile compared to andrographolide and chloroquine and shows a tolerable safety margin that is comparable to established antiviral agents such as remdesivir and nirmatrelvir. Previous toxicological evaluation in animal models also showed that low-dose exposure to eurycomanone did not cause significant genotoxic effects or liver damage, further supporting its safety at biologically active doses. 35
According to our current evidence, this is the first study to comprehensively demonstrate the antiviral potential of eurycomanone across multiple SARS-CoV-2 variants. Previous research conducted by Choonong et al highlighted the antiviral activity of eurycomanone and other quassinoids against unspecified SARS-CoV-2 variants, utilizing in-cell ELISA assays with Vero cells. 10 This study reported an IC50 value of 14.96 μM for eurycomanone, while other quassinoids showed IC50 values ranging from 0.43 to 9.39 μM. Furthermore, eurycomanone demonstrated notable efficacy against HCoV-C43 (common cold coronavirus) with an IC50 value of 4.47 μM, emphasising its potential beyond SARS-CoV-2. 10 Building on these findings, our study extended the analysis to specific SARS-CoV-2 variants, including the WT, Alpha, Delta, and Omicron variants.
The dose-response curves in Vero E6 cells show that eurycomanone reduced SARS-CoV-2 infection in a concentration-dependent manner, with EC50 values ranging from 4.58 to 11.94 µM across tested variants. Since Vero E6 cells lack type I interferon production, the antiviral effect is unlikely to involve immune modulation. 36 The data suggest that eurycomanone acts by disrupting key stages of viral replication after cell entry. These include polyprotein processing, viral protein expression, and virion assembly, all of which are essential for producing new infectious particles. 37 Consistent with this mechanism, cells treated with eurycomanone maintained better structural integrity than untreated infected controls, as observed under microscopic evaluation. At higher concentrations, the treated cells exhibited clearer outlines, reduced cytopathic effects such as rounding or detachment, and preserved monolayer structure. These morphological features closely resembled those of uninfected cells, suggesting that eurycomanone mitigated the extent of virus-induced cellular damage in a dose-responsive manner. This observed improvement supports its potential protective role during SARS-CoV-2 infection. Although direct evidence of eurycomanone inhibiting viral RNA synthesis is not available, the activity of similar antiviral agents provides supporting rationale. For instance, enisamium is an antiviral drug derived from isonicotinic acid and has been reported to reduce SARS-CoV-2 RNA levels by inhibiting viral RNA polymerase. 38 While their chemical structures differ, both compounds share the potential to disrupt critical steps in the viral replication cycle. Taken together, these findings suggest that eurycomanone acts on internal steps of the viral life cycle rather than the spike protein.
The broad antiviral potential of eurycomanone is further supported by earlier studies that highlight its ability to inhibit the dengue virus, showing its potential as a versatile compound for antiviral drug development.
11
This versatility in targeting both coronaviruses and flaviviruses highlights its structural adaptability for multi-variant efficacy. Several studies have demonstrated the antiviral activity of natural product compounds against multiple SARS-CoV-2 variants. For instance, Lancemaside A from
To support the therapeutic potential of eurycomanone, we performed molecular docking and molecular dynamics (MD) simulations against the SARS-CoV-2 Mpro. We selected both the wild-type (WT) structure (PDB: 6LU7) and the Omicron-specific structure (PDB: 7TLL) to evaluate its binding performance against original and mutated forms of the virus. The SARS-CoV-2 Mpro is a vital enzyme for viral replication, making it an excellent target for antiviral drug discovery due to its conserved substrate-binding pocket across coronaviruses.6,39 Omicron-specific mutations such as P132H keep Mpro catalytically active but slightly reduce its thermal stability. 40 These changes highlight the need to design inhibitors that adapt to evolving structural features. Computational modelling showed that inhibitors targeting of this conserved pocket could achieve effective antiviral activity against both WT and mutated strains, including the highly transmissible Omicron variants. The crystal protein with PDB: 6LU7 represents the crystallized structure of the SARS-CoV-2 Mpro in its WT variant form and widely used benchmark in inhibitor studies due to its well-characterized active site and relevance in drug discovery research. 39 The Omicron-specific structure (PDB: 7TLL) shows unique adaptations in the viral protease associated with this highly mutated lineage, making it suitable for evaluating interactions with antiviral candidates targeting emerging variants. 41 These structural selections ensure that the study addresses both baseline interactions and variant-specific binding characteristics.
Molecular docking showed that eurycomanone effectively binds to the Mpro of both WT and Omicron SARS-CoV-2 variants. The binding affinity was stronger for the Omicron variant (−8.2 kcal/mol) compared to the WT strain (−7.7 kcal/mol), suggesting enhanced interactions with the mutated protease. Since Mpro is essential for viral polyprotein processing, strong binding to this site may disrupt viral replication. 42 A comparison between eurycomanone and nirmatrelvir in the Mpro of the Omicron variant revealed similar key binding interactions with adjacent residues. Both compounds formed hydrogen bonds with critical C145 residue-eurycomanone through its 12-OH group and nirmatrelvir by way of a covalent bond involving its nitrile group. Additionally, while nirmatrelvir's cyclic amide ring interacted with F140 and E166 residues, the lactone ring of eurycomanone engaged in multiple hydrogen bonding interactions with E166, further stabilized by hydrogen bonds with L141 via its 15-OH group. Notably, the trifluoroacetyl group of nirmatrelvir participated in multiple halogen and van der Waals interactions with P168, Q192, Q189, T190, L167, E166 residues, whereas eurycomanone did not exhibit these unique interactions. Furthermore, nirmatrelvir's aza-bridge engaged in extensive alkyl interactions with residues H41, along with M165 and M49 residues, while eurycomanone demonstrated similar interactions with H41 and M49 through hydrogen bonds and van der Waals forces via its 11-OH and 12-OH groups. Moreover, the rigid hydrophobic tetracyclic scaffold of eurycomanone stabilizes the ligand through van der Waals contacts with surrounding residues. These molecular features are essential for stable occupancy of the catalytic cleft. The slightly higher binding affinity observed for the Omicron variant corresponds to its lower EC50 values, indicating improved compatibility of the quassinoid scaffold with the mutated protease pocket. Eurycomanone exhibited a binding profile nearly identical to that of the the Mpro of the Omicron variant in the WT. The structural similarity in binding poses between eurycomanone and nirmatrelvir supports the potential of eurycomanone as a promising lead for further antiviral development.
Per-residue interaction analysis highlighted significant contributions from GLN189, GLU166, and MET165. These residues are key to stabilizing inhibitor binding in the Mpro active site. 39 In the Omicron variant, these residues have been implicated in important roles, likely due to structural changes in the binding site that increased electrostatic and van der Waals contributions, as confirmed by MMGBSA free energy calculations. MD simulations confirmed the stability of eurycomanone binding to Mpro, with consistent RMSD values across simulations. Although nirmatrelvir showed slightly better stability, eurycomanone demonstrated comparable performance. MMGBSA analysis highlighted its strong van der Waals and electrostatic interactions against the Omicron variant and its adaptability to structural changes in viral proteins. 43 A previous study using the same methods and crystal structure evaluated natural product inhibitors, and identified acteoside as a potential inhibitor of SARS-CoV-2 Mpro in both the wild-type and Omicron variants. 44 In that study, acteoside formed stable interactions and showed comparable binding energies with both variants, suggesting broad-spectrum potential through hydrogen bonding and hydrophobic interactions. These findings highlight the importance of discovering natural compounds capable of adapting to evolving viral targets.
Despite its promising results, this study has several limitations. It was conducted in vitro using Vero E6 cells, which are commonly used in viral research, including studies on SARS-CoV-2. 45 However, these cells lack the physiological complexity found in human tissues, including key features like TMPRSS2 expression, mucus secretion, and coordinated ciliary activity. 46 This simplified nature may reduce the clinical relevance of the findings. Computational models provide insights but cannot fully replicate biological interactions, highlighting the need for more advanced approaches to validate therapeutic potential. Moreover, the precise mechanism of action of eurycomanone remains unclear. While the compound shows promising antiviral activity, its exact pathway and mode of action have not been fully elucidated and limiting our understanding of its therapeutic potential. These limitations emphasize the necessity of in vivo studies to confirm the safety and efficacy of eurycomanone in more representative biological environments. Although eurycomanone demonstrates broad-spectrum antiviral activity against multiple SARS-CoV-2 variants, its poor bioavailability remains a challenge. Our recent ADMET analysis confirms that eurycomanone has limited oral absorption, as indicated by its low GI absorption. This is supported by previous studies showing low permeability across biological membranes, which reduces its effectiveness in vivo. 47 Therefore, using eurycomanone in its pure form may not be ideal. Further development of novel formulations and targeted drug delivery systems is necessary to enhance its pharmacokinetic properties and improve therapeutic outcomes.
This research extends beyond COVID-19 treatment strategies. By exploring the antiviral potential of a natural compound, we emphasize the importance of combining experimental virology, computational modelling, and natural product chemistry. This approach can accelerate the discovery of resilient antiviral strategies. Recommendations include conducting toxicology studies, testing efficacy against other viral infections, and expanding research into related natural products. Eurycomanone offers a significant step toward addressing the need for adaptable therapeutic interventions. By demonstrating its ability to inhibit viral replication across SARS-CoV-2 variants, this research contributes to the development of COVID-19 treatments and advances the discovery of antiviral drugs, as well as the management of emerging viral threats.
Conclusion
This study identifies eurycomanone as a promising antiviral candidate against SARS-CoV-2 variants, showing broad-spectrum activity and low cytotoxicity. In vitro assays confirmed antiviral effects across all tested variants, with the strongest response observed against the Omicron variant. Eurycomanone reduced viral replication and interacted well with key residues in the Omicron Mpro, suggesting a targeted mechanism of action. Molecular docking and dynamics simulations supported the stability of its binding to the protease. Although its predicted binding affinity was slightly lower than nirmatrelvir, the results support its potential as a complementary treatment, especially for highly mutated variants. While this study is limited to in vitro and computational models, these findings support the need for in vivo validation and further investigation into how mutations may affect treatment. The low absorption of eurycomanone also underscores the need for a more effective formulation to enhance its clinical utility. This study highlights the potential of natural products in antiviral development, particularly for addressing emerging SARS-CoV-2 variants.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X261416683 - Supplemental material for Evaluating Eurycomanone as a Potential Antiviral Agent Against SARS-CoV-2 Variants
Supplemental material, sj-docx-1-npx-10.1177_1934578X261416683 for Evaluating Eurycomanone as a Potential Antiviral Agent Against SARS-CoV-2 Variants by Norazlan Mohmad Misnan, Nur Hannani Ahmad Rozani, Fatahiya Mohamed Tap, Maizatul Hasyima Omar, Kok Wai Lam and Mohd Ridzuan Mohd Abd Razak in Natural Product Communications
Footnotes
Acknowledgment
The authors would like to thank the Director-General of Health, Ministry of Health, Malaysia, and the Director of the Institute for Medical Research for permitting the publication of this paper. NMM also acknowledges the Ministry of Health Malaysia for funding postgraduate studies in the Faculty of Pharmacy, Universiti Kebangsaan Malaysia, through the Educational Scholarship for Doctor of Philosophy program.
Ethical Considerations
This research involved only in vitro experiments and did not include any human subjects or direct human interventions. As such, the study was exempted from ethical review by the Medical Research and Ethics Committee (MREC), Ministry of Health Malaysia (Ref: NMRR ID-23-00540-UHV). The virus isolation part in this study was performed by the Virology Unit, Institute for Medical Research (IMR), and was covered under ethical approval granted by MREC (Ref: KKM/NIHSEC/P20-1584(4). All viral isolates used in this study were fully anonymized and contained no identifiable patient information. All experimental procedures were conducted in accordance with the principles outlined in the Declaration of Helsinki.
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Funding
This study was funded by the Ministry of Health Malaysia (NMRR-23-00540-UHV).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
