Abstract
Objective
This study aims to explore the potential molecular mechanisms of puerarin in alleviating non-alcoholic fatty liver disease (NAFLD) by combining network pharmacology and experimental analysis methods.
Methods
The study identified the active chemical components of puerarin and its potential targets by integrating databases, followed by constructing a target gene database for NAFLD. Through the analysis of the protein-protein interaction (PPI) network and identification of core genes, the focus was on the inhibitory effects of puerarin on inflammatory mediators such as TNF, IL6, IL1B, and STAT3. Laboratory experiments using the HepG2 cell line further validated the hepatoprotective effects of puerarin, demonstrated by reduced levels of ALT and AST, as well as the regulation of stress responses. Quantitative polymerase chain reaction (qPCR) experiments supported these findings, confirming the impact of puerarin on the expression of key inflammatory genes.
Results
The active components of puerarin and corresponding targets were identified through ETCM2.0 and HERB databases, resulting in the identification of 189 target genes. A total of 2317 target genes related to NAFLD were integrated from GeneCards, DisGeNET, and OMIM databases. Using Venn diagram tools, 188 intersecting target genes between puerarin and NAFLD were determined, and a PPI network was constructed. Gene Ontology (GO) functional analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis showed that NAFLD is closely related to biological processes such as insulin resistance, inflammatory response, lipid metabolism disorders, and apoptosis. Experimental validation results showed that puerarin treatment significantly reduced ALT and AST levels in the cell culture supernatant (by 47% and 29%, respectively), decreased MDA levels by 71%, and increased SOD levels by 23%. qPCR experiments indicated that the relative expression levels of IL6, TNF, IL1B, and STAT3 genes in the PA + puerarin treatment group were significantly lower than those in the PA-only treatment group, reduced by 2.0-fold, 2.1-fold, 2.9-fold, and 2.5-fold, respectively.
Conclusion
This study suggests that puerarin may alleviate the progression of NAFLD by targeting inflammatory pathways and regulating oxidative stress. The combined results of network pharmacology analysis and experimental validation indicate that puerarin modulates the inflammatory response by inhibiting the expression of genes such as IL6, TNF, IL1B, and STAT3, thereby reducing the production and release of inflammatory factors, alleviating liver inflammation, reducing liver fat accumulation, and slowing the progression of NAFLD. Our research provides fundamental data for puerarin as a potential drug for the treatment of NAFLD. With further research and development, puerarin can be designed as an oral medication or other forms of administration or combined with other NAFLD treatments to improve liver health in patients.
Keywords
Introduction
Non-Alcoholic Fatty Liver Disease (NAFLD) is increasingly recognized as a predominant chronic liver disorder worldwide. Epidemiological studies reveal a global adult prevalence of approximately 23–25%, with marked regional disparities, being particularly prevalent in the Middle East and South America, and less so in Africa. Notably, a rising trend is observed in pediatric and adolescent obesity-related NAFLD cases. 1 Beyond its direct health implications, NAFLD significantly contributes to the burden on public health infrastructure and economic systems due to its association with various metabolic disorders. The pathogenesis of NAFLD is multifactorial, predominantly characterized by abnormal hepatic fat accumulation, influenced by factors like obesity, insulin resistance, dyslipidemia, and genetic predisposition. Progression of NAFLD can lead to Non-Alcoholic Steatohepatitis (NASH), with potential complications including liver fibrosis, cirrhosis, and hepatocellular carcinoma, thus posing serious liver and systemic health threats. 2 Recent decades have seen a paradigm shift in NAFLD research, aiming at more effective treatments. Traditional pharmacological treatments offer limited efficacy and are often associated with adverse effects. In this context, natural compounds, particularly those with unique pharmacological attributes, minimal side effects, and wide biological activities, have come to the forefront. Puerarin, a key constituent of traditional Chinese medicine, known for its anti-inflammatory and antioxidative properties, is a case in point. 3 Nonetheless, a comprehensive understanding of how puerarin modulates NAFLD at the molecular level remains elusive. This study leverages network pharmacology and experimental validation to demystify the therapeutic potentials and molecular mechanisms of puerarin in NAFLD management.
Although previous studies have shown that puerarin exerts certain therapeutic effects on NAFLD by regulating oxidative stress and inflammatory responses, the specific molecular mechanisms remain incompletely.4,5 Current research primarily focuses on the overall efficacy of puerarin in NAFLD, while its specific action pathways and targets at the cellular level, such as reducing lipid accumulation and oxidative stress by regulating the c-Jun N-terminal kinase (JNK) signaling pathway, lack in-depth exploration. 5 Additionally, while some studies have investigated the impact of puerarin on the expression of certain key genes such as Srebf1, Chrebp, and Acaca, the broader scope of gene regulation and signal pathway interactions requires further clarification.5,6 Research also indicates that puerarin can improve metabolic dysfunction-associated fatty liver disease by inhibiting ferroptosis and inflammatory responses. 4 Moreover, studies on puerarin in neurological diseases have shown that it exerts protective effects through the PI3K/Akt signaling pathway, a mechanism that may also be applicable in NAFLD. 6 However, these findings still require more experimental and clinical research to verify the specific mechanisms of puerarin in the treatment of NAFLD.
Methods
Identification of Active Components and Targets of Puerarin
A systematic search of traditional medicine databases, including ETCM2.0 (http://www.tcmip.cn/ETCM2/front/#/) 7 and HERB (http://herb.ac.cn/), 8 was conducted to gather information on the reported active components of puerarin. This information was further organized, and potential targets of these components were identified through searches in public databases such as PubChem (https://pubchem.ncbi.nlm.nih.gov/), 9 ETCM2.0, and HERB. Subsequently, the identified targets were matched in the UniProt (http://www.uniprot.org/) 10 database to determine their unique gene names.
Identification of NAFLD Targets
The NAFLD-related target genes were derived from a combination of three databases: GeneCards (https://www.genecards.org/), 11 DisGeNET (http://www.disgenet.org/), 12 and Online Mendelian Inheritance in Man (OMIM, https://omim.org/). 13 Following the integration of target genes from these databases, redundant genes were removed to establish a comprehensive target gene database for NAFLD.
Intersection Gene Network
The intersection genes between NAFLD and puerarin were determined using the online tool Interactive Venn (http://www.interactivenn.net/). 38 A Venn diagram was created to visualize the intersecting genes.
PPI Network Construction and Identification of Core Target Genes
The list of intersecting genes was exported and uploaded to the STRING database (http://string-db.org/) 14 using the Multiple proteins tool, with the species limited to Homo sapiens. This process generated a Protein-Protein Interaction (PPI) network for the intersecting genes, saved as a TSV format file. The TSV file was then imported into Cytoscape 3.7.2 software 15 for network topology analysis, constructing the “Puerarin-Target-Pathway-NAFLD” network. The top 10 genes most relevant to the modulation of NAFLD by puerarin were identified as core genes using the cytohubb plugin in Cytoscape.
GO Function Analysis and KEGG Pathway Enrichment Analysis
Following the identification of intersecting targets of puerarin and NAFLD, Gene Ontology (GO) function and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were conducted using the Metascape database (https://metascape.org/gp/index.html). 16 The top 20 entries in biological processes (BP), molecular functions (MF), KEGG pathways, and cellular components (CC) were analyzed, with a significance threshold set at P < 0.05.
Molecular Docking Methods and Results Presentation
Molecular Docking Methods
Molecular docking techniques were employed to verify the affinity of puerarin active components with the crystal structures of core target proteins. The 3D structures of the top 10 proteins from the puerarin and NAFLD intersecting genes were downloaded from the RCSB PDB database (https://www.rcsb.org/). 17 The 3D structure of puerarin (as the ligand) was retrieved from the PubChem database. PyMOL 2.4.2 software was used to dehydrate the receptor proteins. Proteins and ligands were uploaded to AutoDock Tools 18 for water molecule removal, hydrogenation, and calculation of protein charges. Docking sites on receptor proteins were set to include the active pocket binding sites of original small molecule ligands. Finally, docking between the receptor proteins and puerarin was conducted using AutoDock Vina 4.2, 19 with a binding energy threshold of ≤ −5 kJ/mol as the criterion for successful docking.
Protein-Ligand Structural Interaction Presentation
To detail the interactions between the proteins and ligands, LigPlot2.2.8 software 20 was employed to generate two-dimensional interaction diagrams of protein-ligand complexes. The PDB format files of docking complexes generated via PyMOL 2.4.2 software were inputted into LigPlot2.2.8. The software automatically identified ligands, surrounding amino acid residues, and analyzed hydrogen and hydrophobic interactions. Hydrogen bonds were represented by dashed lines, with the length inversely proportional to the strength of the interaction, and hydrophobic contacts were illustrated by arc lines, where the width of the arc indicated the size of the contact area. All interaction parameters were set to default in LigPlot2.2.8 to ensure consistency of results. The final graphics were adjusted for clarity to showcase all key interactions for presentation in the paper.
Experimental Validation
Cell Experiments
In the initial phase of our study, we systematically evaluated the effects of puerarin, a compound of interest, on the viability of HepG2 cells, a hepatocyte-derived cell line. We administered six different concentrations of puerarin (10, 20, 30, 40, 50, 80 μM) to determine the concentration that optimally promotes cell viability without inducing cytotoxicity. Detailed results of these screenings are accessible in Supplementary Document 1. Notably, concentrations of 10 μM and 20 μM of puerarin substantially enhanced HepG2 cell activity, while higher concentrations (50 μM and above) appeared to diminish this effect or were potentially cytotoxic.
To further validate the therapeutic effects of puerarin on non-alcoholic fatty liver disease (NAFLD), we conducted experiments using the HepG2 cell line, which was sourced from the Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences. The cells were cultured in DMEM enriched with 10% fetal bovine serum and incubated with 1 mM palmitic acid (PA). Treatments were conducted over 48 h with and without 20 μM puerarin, under a 5% CO2 atmosphere at 37 °C. Post-treatment, we analyzed the cell supernatants for alanine aminotransferase (ALT) and aspartate aminotransferase (AST) using specific ELISA kits. 21 Additionally, levels of malondialdehyde (MDA) and superoxide dismutase (SOD) were quantified using respective assay kits.
Quantitative PCR (qPCR) Experiments
In this study, we focused on several genes closely associated with lipid metabolism and inflammatory responses, including IL10, IL1B, IL6, STAT3, and TNF. These genes were selected due to their critical roles in regulating inflammation and metabolic abnormalities during the progression of non-alcoholic fatty liver disease (NAFLD). We conducted our qPCR experiments using the RT² Profiler PCR Array system from QIAGEN (product number 330231), a commercial kit designed for standardized experiments to ensure the reproducibility and comparability of results.
For normalization of our experimental data, we selected GAPDH as the reference gene because its expression remains relatively stable under various treatment conditions, making it suitable as an internal control to ensure the accuracy and reliability of our data.
Statistical Analysis
In this study, appropriate statistical methods were chosen based on the distribution characteristics of the data to compare continuous variables between groups. Grouped t-tests were used when the data demonstrated normal distribution and homogeneity of variance. For data that did not meet these criteria, the Wilcoxon rank-sum test was utilized. Additionally, to comprehensively assess the effects of different treatments on key biomarkers (such as AST, ALT, MDA, SOD) and inflammatory factors (such as TNF, IL6), one-way analysis of variance (ANOVA) was employed. All statistical analyses were conducted using SPSS 25.0 and GraphPad Prism 9.5.2 software, with a significance level set at P < 0.05.
Results
Network Pharmacology Study Results
Identification of Active Components and Targets of Puerarin
The active components of puerarin and their corresponding targets were retrieved from the ETCM2.0 and HERB databases. After eliminating duplicate genes, a total of 189 target genes were identified, with PubChem CID 5281807.
Identification of NAFLD Targets
Target genes related to NAFLD were compiled from three databases: GeneCards, Disgent, and OMIM. Upon removal of duplicates, 2317 NAFLD targets were identified.
Establishment of Intersection Genes
Using an online Venn diagram tool, the intersecting genes between puerarin and NAFLD were established, resulting in 188 intersection targets. See Figure 1A for details.

A) This Venn diagram illustrates the overlap of two datasets: one representing puerarin with 189 identified targets, and the other depicting NAFLD with 2317 targets. It highlights 126 common targets shared between puerarin and NAFLD. B) A complex protein-protein interaction (PPI) network is shown, constructed using the STRING database. In this network, nodes symbolize different targets, and connecting lines represent their interactions or associations. C) Displayed here are the top 10 core targets, identified using the Cytohub plugin in Cytoscape, indicating their central role in the network.
Construction of the PPI Network
The list of intersecting genes was uploaded to the STRING database to construct a Protein-Protein Interaction (PPI) network for the intersecting genes. Refer to Figures 1B and 2A for the network diagrams.

A) This image depicts the PPI network of intersecting genes, assembled in cytoscape 3.7.2. Each red circle denotes a target, with the intensity of the red hue correlating to the degree value; darker shades indicate higher degrees. B) This network diagram presents an intricate mapping of active components, targets, pathways, and diseases, showing their interconnectedness and influence.
Compound-Target-Pathway-Disease Network and Core Target Genes
A network linking compounds (puerarin), targets, pathways, and NAFLD was constructed using Cytospace 3.7.2, as shown in Figure 2B. This network displays the various targets and pathways involved in puerarin and NAFLD, as well as their complex interactions.
GO Function Analysis and KEGG Pathway Enrichment Results
From the 189 intersecting genes, 194 KEGG pathways (P ≤ 0.01) were identified, and GO function analysis included 1973 biological processes (BP), 176 molecular functions (MF), and 89 cellular components (CC). The top 20 entries in BP, MF, and CC by enrichment are presented in Figure 3.
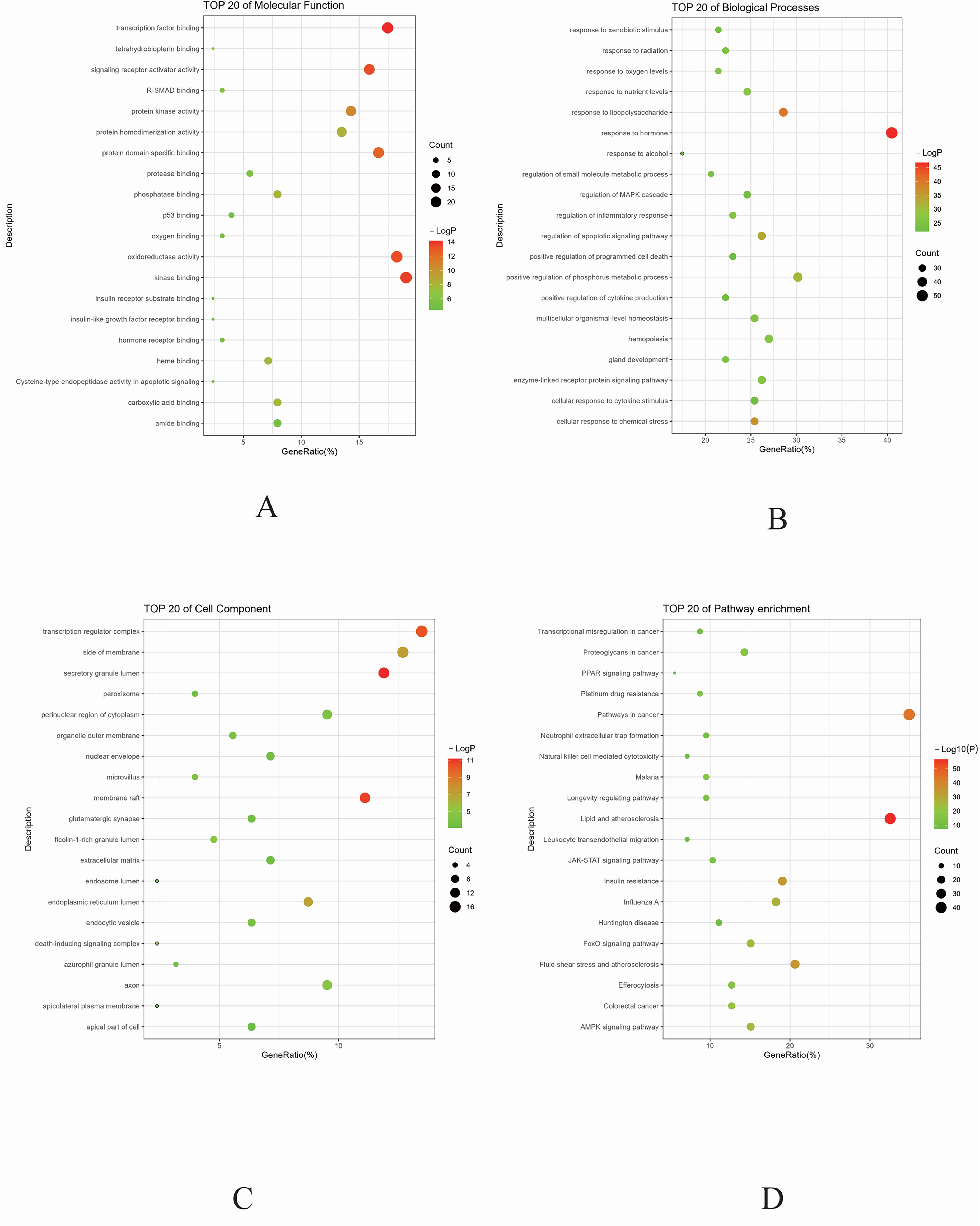
This figure presents the results of gene ontology (GO) function analysis and Kyoto encyclopedia of genes and genomes (KEGG) pathway enrichment analysis for the intersecting genes. The bubble size corresponds to the number of genes enriched in each component, while the color intensity signifies the P-value significance. A) Molecular Function (MF). B) Biological Processes (BP). C) Cellular Component (CC). D) Pathway enrichment (KEGG).
Non-Alcoholic Fatty Liver Disease (NAFLD) is a complex metabolic disorder involving multifaceted biological changes, including biological processes, cellular components, molecular functions, and various metabolic pathways. The enrichment analysis highlights the association of NAFLD with critical biological phenomena such as insulin resistance, 22 inflammatory response, 23 dysregulation of lipid metabolism, and cellular apoptosis. For instance, reduced hepatic sensitivity to insulin, a hallmark of insulin resistance, precipitates alterations in glucose and lipid metabolism, culminating in hepatic fat accumulation. Moreover, the aberrant activation of inflammatory pathways plays a vital role in NAFLD's progression to Non-Alcoholic Steatohepatitis (NASH), marked by the dysregulation of cytokines and other mediators. 24
Concerning cellular components, the roles of cell membranes, membrane-associated regions, receptor complexes, and secretory granules are pivotal in NAFLD's pathology. Variations in cell membrane lipid composition mirror disruptions in lipid metabolism, while alterations in membrane regions could influence cellular signaling and intracellular transport, essential aspects in the development of NAFLD. 25 Additionally, changes in receptor complexes may affect the transmission of hormonal signals, such as insulin and glucocorticoids, thereby modulating cellular hormonal responses.
At the molecular function level, the activities of transcription factors, nuclear receptors, oxidoreductases, kinases, and antioxidants are integral to NAFLD progression. Transcription factors and nuclear receptors exert a direct influence on genes governing lipid metabolism and inflammatory responses. Oxidative stress, a key aspect of NAFLD, is modulated by oxidoreductases, crucial for the cellular response to oxidative stress; alterations in kinase and antioxidant functions can impact signal transduction and cellular defense mechanisms against oxidative stress.
In the realm of metabolic pathways, the KEGG pathway analysis delineates several critical pathways in NAFLD pathogenesis, including lipid and atherosclerosis pathways, insulin resistance pathways, AMPK signaling, HIF-1 signaling, PPAR signaling, and JAK-STAT signaling. These pathways illustrate shifts in lipid metabolism and energy homeostasis, encompassing processes such as inflammation, cell proliferation, and apoptosis. Disruptions in the insulin resistance pathway could lead to increased hepatic lipid accumulation and heightened inflammatory responses, while anomalies in the JAK-STAT signaling pathway might contribute to metabolic imbalances, furthering NAFLD progression.
Identification of Core Targets and Molecular Docking Results
Using the Cytohub plugin in Cytospace 3.7.2, the top 10 core target genes of the intersecting genes were identified as HIF1A, IL6, TNF, IL1B, ALB, PTGS2, STAT3, NFKB1, INS, MMP9. Details are provided in Table 1 and Figures 4, 5, 6.

These figures comprise molecular docking diagrams, illustrating the two-dimensional and three-dimensional structures of interactions between puerarin and its top 9 core targets, including IL6, TNF, IL1B, STAT3. The diagrams detail the specific bonds between each compound and surrounding amino acid residues, depicted through hydrogen bonds, hydrophobic interactions, and more.

Molecular docking results revealed that puerarin successfully docked with the top 9 targets, with binding energies all below −5.0 kcal/mol.
Experimental Validation Results
Cell Experiment Results
The cell experiments demonstrated that puerarin treatment significantly reduced the levels of Aspartate Aminotransferase (AST) and Alanine Aminotransferase (ALT) in the cell culture supernatant. Specifically, in the PA + puerarin treated group, ALT levels decreased significantly by 47%, from 103.68 ± 3.22 U/L to 54.77 ± 1.40 U/L, and AST levels decreased by 29%, from 90.85 ± 1.886 U/L to 64.41 ± 3.041 U/L (P < 0.05), indicating a notable hepatoprotective effect of puerarin, as shown in Figure 7A and 7B.
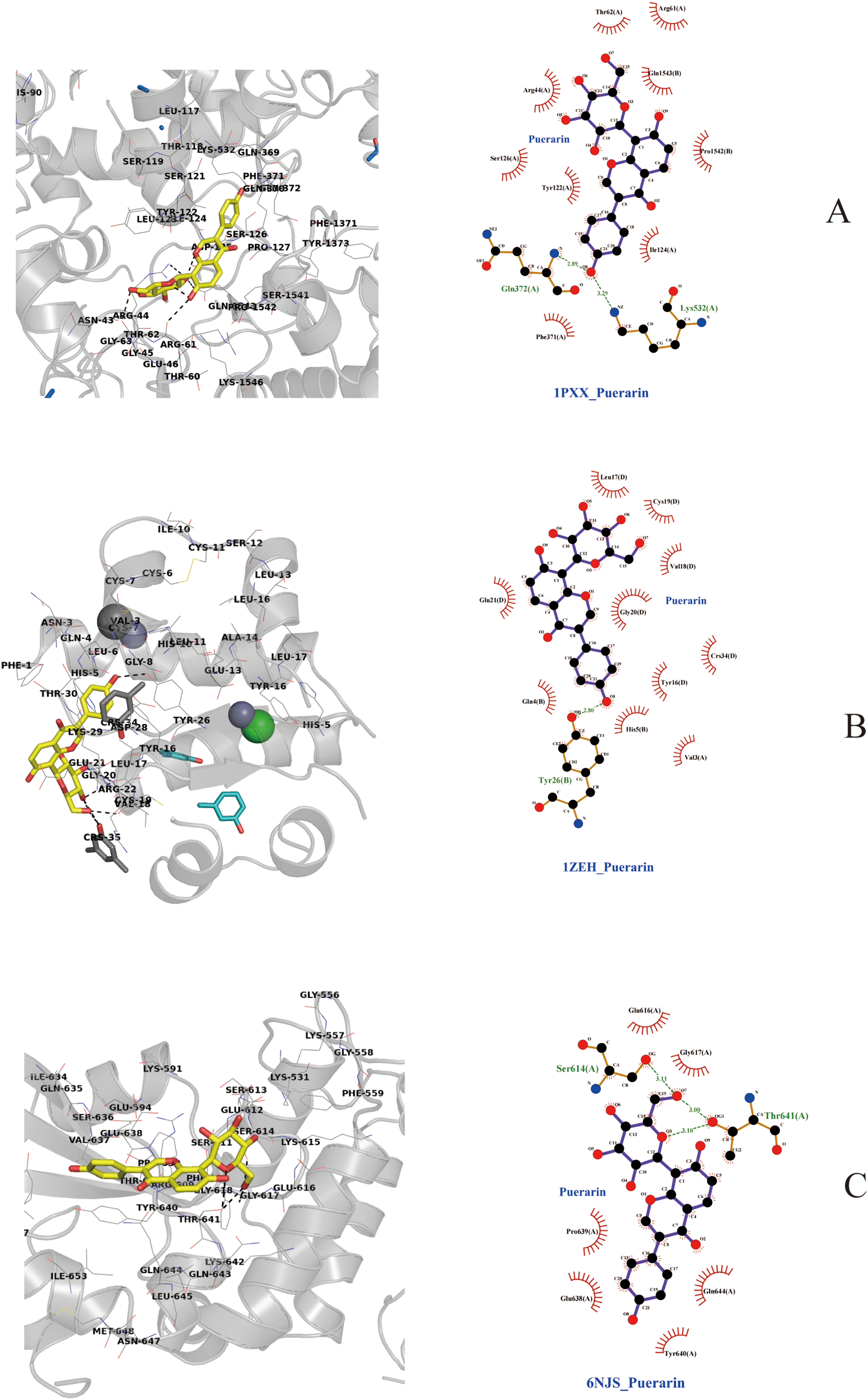
These figures comprise molecular docking diagrams, illustrating the two-dimensional and three-dimensional structures of interactions between puerarin and its top 9 core targets, including IL6, TNF, IL1B, STAT3. The diagrams detail the specific bonds between each compound and surrounding amino acid residues, depicted through hydrogen bonds, hydrophobic interactions, and more.
In the PA-induced HepG2 cell model, compared to the model group, the PA + puerarin treated group exhibited a significant reduction in Malondialdehyde (MDA) levels by 71%, from 1.7 ± 0.15 nM/mL to 0.5 ± 0.06 nM/mL (P < 0.05), and a notable increase in Superoxide Dismutase (SOD) activity by 23%, from 145.59 ± 12.03 U/L to 178.68 ± 11.47 U/L (P < 0.05). These findings suggest that puerarin effectively mitigates oxidative stress induced by PA,as shown in Figure 7C and 7D.
Overall, the research indicates that puerarin exerts a protective effect on liver cells in vitro, potentially through the reduction of ALT and AST levels, alleviation of oxidative stress, and downregulation of key inflammatory genes
qPCR Experiment Results
The results from quantitative Polymerase Chain Reaction (qPCR) experiments further elucidate the impact of puerarin on gene expression. In the group treated with both PA and puerarin, there was a significant reduction in the relative expression levels of key genes such as IL6, TNF, IL1B, and STAT3 compared to the group treated with PA alone, with a P value of less than 0.01 indicating statistical significance. This observation suggests that the therapeutic effects of puerarin may be attributed to its modulation of these genes, which are among the top 10 targets identified in the study, as shown in Figure 8. Furthermore, the treatment with puerarin significantly reduced key inflammatory gene expressions: TNF decreased by 2.1-fold, IL6 by 2.0-fold, STAT3 by 2.5-fold, and IL1B by 2.9-fold.

These figures comprise molecular docking diagrams, illustrating the two-dimensional and three-dimensional structures of interactions between puerarin and its top 9 core targets, including IL6, TNF, IL1B, STAT3. The diagrams detail the specific bonds between each compound and surrounding amino acid residues, depicted through hydrogen bonds, hydrophobic interactions, and more.
In our comprehensive study, we expanded our quantitative PCR (QPCR) investigations to include an additional 55 genes involved in lipid metabolism, aiming to deepen our understanding of dendrobine's molecular effects, as shown in Figure 9.

A) Graph depicting ALT concentrations across three different treatment conditions: control group (blue), PA-treated group (red, concentration at 1 mm), and PA plus Puerarin-treated group (green, Puerarin concentration at 20 μm). Significance levels are indicated with asterisks (* for P < 0.05, ** for P < 0.01, *** for P <0.001). B) Graph showing AST concentrations under the same treatment conditions as in A, with corresponding color coding and significance indicators. C) Graph illustrating MDA concentrations for the same treatment groups, following the same color and significance coding as in A. D) Graph depicting SOD concentrations under the same treatment conditions as in A, with the respective color coding and significance indicators.

This figure displays the relative expression levels of various proteins under three treatment conditions, normalized against GAPDH: control group (blue), PA-treated group (red, PA concentration at 1 mm), and PA plus puerarin-treated group (green, puerarin concentration at 20 μm). The significance of differences is indicated as in Figure 7.
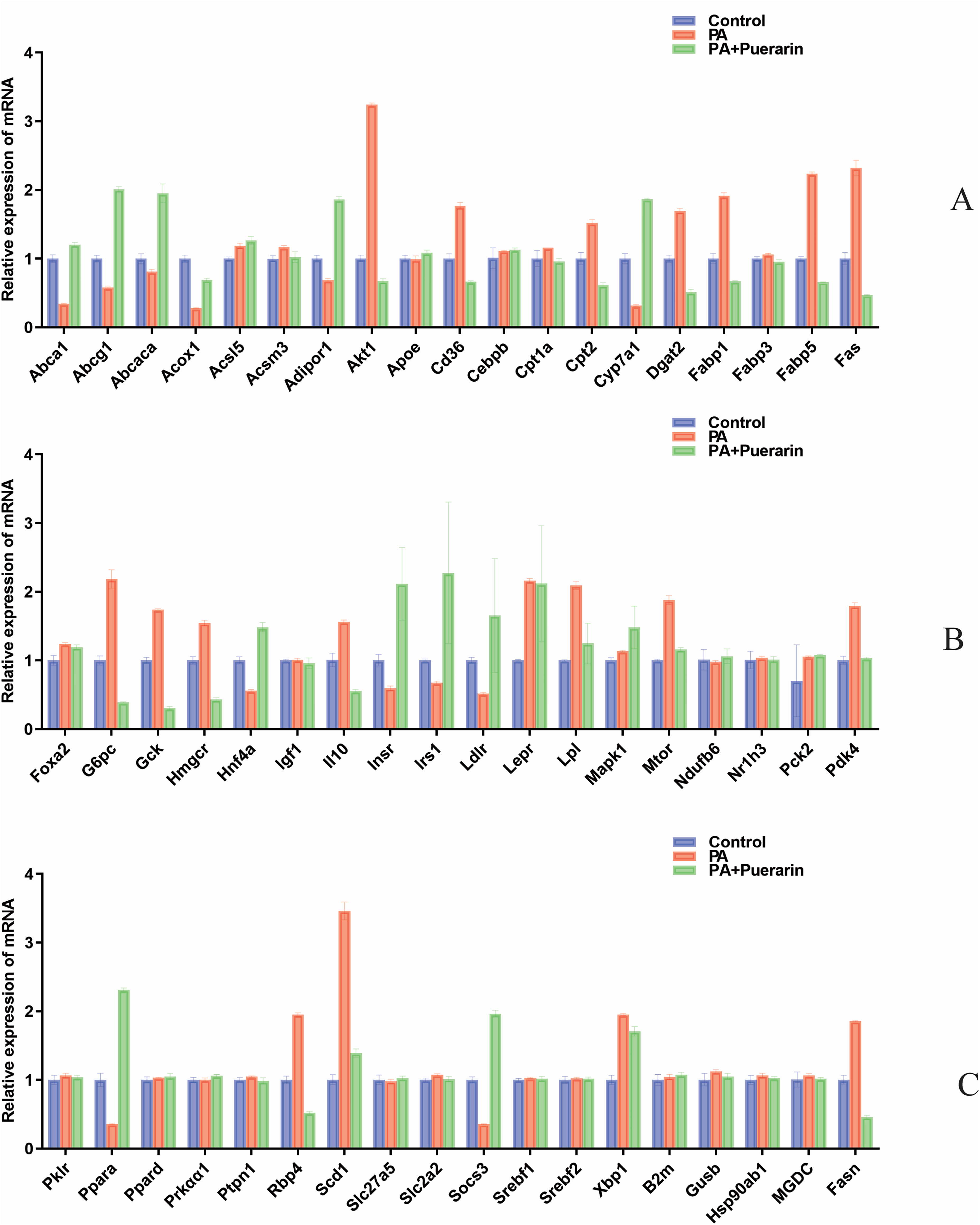
This figure illustrates the relative expression levels of additional proteins of interest, excluding IL6, TNF, IL1B, STAT3, under the same treatment conditions as in figure 8. The data are normalized to GAPDH, with color coding and significance indicators consistent with previous figures, and includes treatment conditions: Control group (blue), PA-treated group (red, PA concentration at 1 mM), and PA plus Puerarin-treated group (green, Puerarin concentration at 20 μM).
Discussion
Existing literature reports that puerarin has protective effects on various metabolic diseases. For instance, the study by Kořínková 2 et al demonstrated that puerarin significantly protects against high-fat diet-induced Non-Alcoholic Fatty Liver Disease (NAFLD) by regulating lipid metabolism and improving insulin sensitivity.
In this study, we combined network pharmacology and experimental validation to further elucidate the molecular mechanisms of puerarin. We found that puerarin significantly reduces the expression of inflammatory mediators such as IL6, TNF, and IL1B and alleviates liver inflammation by modulating the STAT3 signaling pathway. This is consistent with existing reports on the anti-inflammatory effects of puerarin. 5
Additionally, our literature review indicates that puerarin can mitigate the pathological progression of NAFLD by inhibiting ferroptosis and oxidative stress. 4 Compared to other flavonoids like resveratrol and quercetin, puerarin exhibits a unique multi-target mechanism. Resveratrol primarily improves NAFLD through the SIRT1/AMPK pathway, while quercetin exerts protective effects via Nrf2-mediated oxidative stress reduction.6,26 In contrast, puerarin demonstrates a unique advantage in comprehensively improving NAFLD through multiple pathways, such as regulating inflammatory mediators and oxidative stress.
Oxidative stress and inflammatory responses play crucial roles in the development and progression of NAFLD.5,27 Our study further confirms this, showing that puerarin significantly reduces the expression of inflammatory mediators such as IL6, TNF, and IL1B, and alleviates liver inflammation by modulating the STAT3 signaling pathway. 5
Dysregulated lipid metabolism is a major characteristic of NAFLD. Studies have shown that puerarin can regulate the expression of several lipid metabolism-related genes, such as Srebf1, Chrebp, Acaca, and Fasn, which are key in hepatic lipid synthesis and accumulation. By modulating these genes, puerarin can reduce hepatic lipid accumulation and alleviate the pathological changes associated with NAFLD.5,28
Insulin resistance is a critical mechanism in the development and progression of NAFLD. Literature indicates that oxidative stress and inflammatory responses exacerbate insulin resistance, promoting the progression of NAFLD. Our study found that puerarin improves insulin sensitivity by reducing the production of inflammatory mediators and oxidative stress levels, thereby alleviating NAFLD symptoms. 5
The progression of NAFLD can lead to liver fibrosis, cirrhosis, and hepatocellular carcinoma. Oxidative stress plays an important role in liver fibrosis formation. Our study shows that puerarin can reduce the risk of liver fibrosis by alleviating oxidative stress and inhibiting lipid peroxidation. 27 Furthermore, puerarin can regulate the expression of fibrosis-related genes, thereby further reducing the degree of liver fibrosis5.
Through network pharmacology analysis, we identified relevant targets of puerarin and further delineated 10 key targets associated with the intervention of Non-Alcoholic Fatty Liver Disease (NAFLD), including HIF1A, IL6, TNF, IL1B, ALB, PTGS2, STAT3, NFKB1, INS, and MMP9.Molecular docking experiments showed that, except for NFKB1, the binding energies of puerarin with these targets were all below −5.0 kcal/mol, suggesting their potential molecular-level efficacy against NAFLD. Further investigations were conducted using the HepG2 cell line induced by palmitic acid (PA) and treated with puerarin. The results demonstrated that puerarin effectively reduced ALT and AST levels in cells and improved oxidative stress conditions. qPCR experiments revealed significantly reduced expression levels of IL6, TNF, IL1B, and STAT3 in the PA + puerarin treated group compared to the PA-only group. As a chronic liver disease closely associated with metabolic disorder, the development of NAFLD involves various inflammatory factors,29,30 particularly cytokines such as TNF, IL1B, and IL6, which play crucial roles in the progression and severity of the disease.
KEGG enrichment analysis indicated that the JAK-STAT signaling pathway is one of the critical pathways, particularly involving IL6 and STAT3, in NAFLD. IL6, a multifunctional cytokine, can activate the STAT3 signaling pathway by stimulating JAK2. STAT3, a transcription factor, plays roles in numerous cellular functions, including cell growth and survival. In the context of NAFLD, upregulation of IL6 may promote inflammatory responses and affect hepatic metabolism and function through the JAK-STAT pathway. These findings highlight the importance of IL6 and STAT3 in the pathogenesis of NAFLD, suggesting the JAK-STAT signaling pathway as a potential therapeutic target for NAFLD.31,32
Research indicates that puerarin, a flavonoid compound, reduced the expression of inflammatory proteins IL-6 and IL-8 in atherosclerosis models, suggesting that its anti-inflammatory pathway might alleviate the progression of NAFLD. 33 Our qPCR results corroborate this finding.
While this study reveals potential molecular mechanisms of puerarin in alleviating NAFLD through network pharmacology and experimental validation, there are several limitations that need further exploration.
Firstly, network pharmacology analysis relies on the quality and accuracy of databases. Although multiple databases were used to identify active components and targets of puerarin, the information may be biased or incomplete. Additionally, network pharmacology methods cannot fully simulate complex biological systems and may overlook critical biological processes or interactions. Thus, the results based on network pharmacology require further experimental validation.
Secondly, the HepG2 cell line was used as the experimental model to verify the effects of puerarin. While HepG2 cells are widely used in liver disease research, they cannot fully represent the complex environment of the human liver. A single cell type model cannot reflect the interactions among various cell types in vivo, and the genetic stability and phenotype of cell lines may change with passage, affecting the reliability of experimental results. Furthermore, experiments used high concentrations of palmitic acid (PA) to induce oxidative stress and inflammatory responses, conditions that might differ from physiological conditions. In reality, the pathological process of NAFLD is influenced by various factors such as diet, genetics, and metabolic status, necessitating further studies in more physiologically relevant experimental environments.
Although cell experiments showed significant effects of puerarin, its actual therapeutic efficacy in humans needs to be validated through clinical trials. Results from cell experiments cannot be directly extrapolated to humans. The absorption, distribution, metabolism, and excretion of puerarin in the human body, as well as its potential side effects and drug interactions, need systematic study in clinical trials. Additionally, as a natural compound, puerarin has the characteristics of multiple components and multiple targets. This complexity is both an advantage and a challenge. Multiple components may bring synergistic effects, but they may also cause mutual interference. The multi-target mechanism is complex and difficult to fully elucidate through a single experiment. Therefore, a comprehensive application of various research methods, combined with techniques such as systems biology and metabolomics, is needed to fully reveal the mechanism of action of puerarin.
In summary, a combined analysis of network pharmacology and experimental validation suggests that puerarin may ameliorate Non-Alcoholic Fatty Liver Disease (NAFLD) through the following mechanisms: puerarin potentially directly or indirectly inhibits the expression of genes such as IL6, TNF, IL1B, STAT3, regulating inflammatory responses, reducing the production and release of inflammatory factors, thereby mitigating hepatic inflammation, reducing hepatic fat accumulation, and consequently slowing the progression of NAFLD.
Conclusion
In this study, we explored the therapeutic potential of puerarin in alleviating the inflammatory responses and oxidative stress associated with Non-Alcoholic Fatty Liver Disease (NAFLD) interventions. Our hypothesis centered on the premise that puerarin interacts with specific molecular targets, such as IL6, TNF, IL1B, and STAT3, significantly altering the activity or expression of these proteins at the molecular level. This interaction has profound implications on cellular functions and states. In the HepG2 cell model, puerarin treatment was observed to effectively reduce ALT and AST levels and significantly enhance oxidative stress conditions, indicating its notable anti-inflammatory and hepatoprotective properties. These effects are attributed not only to direct molecular interactions but also to the modulation of signaling pathways.
The alterations in cellular state and function brought about by puerarin subsequently impact intracellular signaling pathways, notably the modulation of the JAK-STAT signaling pathway. This mechanism acts as a response to the molecular-level changes, further influencing cellular metabolism and inflammatory responses. However, it is important to note that our current experimental data and literature review are not exhaustive. Comprehensive clinical research and detailed in vivo and in vitro experiments are essential to accurately ascertain the therapeutic efficacy and safety of these components.
Additionally, our literature review indicates that puerarin has significant anti-inflammatory and antioxidant effects, making it a promising candidate for the treatment of various metabolic diseases.3,6 These findings provide a theoretical basis for developing puerarin-based treatments for Non-Alcoholic Fatty Liver Disease (NAFLD). As a natural compound, puerarin's relatively low toxicity and broad pharmacological effects enhance its potential as an effective NAFLD therapeutic agent.
In the treatment of NAFLD, puerarin can exert its effects through multiple mechanisms, such as regulating lipid metabolism, improving insulin resistance, and reducing liver fibrosis. Studies have shown that puerarin significantly reduces the expression of inflammatory mediators such as TNF-α, IL-6, and IL-1β, while increasing the activity of antioxidant enzymes like superoxide dismutase and glutathione peroxidase, thereby reducing oxidative stress and inflammatory responses. 34
Despite the promising results from our in vitro cell experiments demonstrating the potential of puerarin in treating NAFLD, several challenges remain in translating these findings into clinical applications. Firstly, puerarin has low bioavailability and poor oral absorption. 3 Secondly, in vitro experimental results often face uncertainties when applied to humans due to biological complexity and individual differences. The pharmacokinetic and pharmacodynamic properties of puerarin in humans need to be validated through systematic clinical trials. These trials should include assessments of the drug's safety, efficacy, dose-response relationship, and potential side effects. 35
Toxicological studies have shown that puerarin exhibits a certain level of safety in acute and subchronic toxicity tests, but its long-term safety requires further investigation. Future research should focus on the long-term toxicity and potential chronic side effects of puerarin, including its effects on the liver, kidneys, and cardiovascular system.36,37 Additionally, the interactions and combined toxicity of puerarin when used with other drugs need to be evaluated. 26
In summary, our study reveals the potential molecular mechanisms of puerarin in the treatment of NAFLD, highlighting its broad application prospects as a therapeutic agent. However, overcoming several challenges is necessary to translate these in vitro findings into clinical applications, including improving drug bioavailability, validating safety and efficacy in humans, and developing personalized treatment regimens. Future research should further explore these aspects to provide a more comprehensive therapeutic strategy.
Supplemental Material
sj-xlsx-1-npx-10.1177_1934578X241293660 - Supplemental material for Investigating Puerarin's Role in Counteracting Non-Alcoholic Fatty Liver Disease: A Synergistic Study of Network Pharmacology and Experimental Analysis
Supplemental material, sj-xlsx-1-npx-10.1177_1934578X241293660 for Investigating Puerarin's Role in Counteracting Non-Alcoholic Fatty Liver Disease: A Synergistic Study of Network Pharmacology and Experimental Analysis by Feng Li, Junlan Chen, Ye Xu, Miao Wang, Shuyan Sun and Shigao Zhou in Natural Product Communications
Supplemental Material
sj-xlsx-2-npx-10.1177_1934578X241293660 - Supplemental material for Investigating Puerarin's Role in Counteracting Non-Alcoholic Fatty Liver Disease: A Synergistic Study of Network Pharmacology and Experimental Analysis
Supplemental material, sj-xlsx-2-npx-10.1177_1934578X241293660 for Investigating Puerarin's Role in Counteracting Non-Alcoholic Fatty Liver Disease: A Synergistic Study of Network Pharmacology and Experimental Analysis by Feng Li, Junlan Chen, Ye Xu, Miao Wang, Shuyan Sun and Shigao Zhou in Natural Product Communications
Footnotes
Acknowledgments
We extend our sincere gratitude to Professors Miao Wang and Shigao Zhou for their invaluable academic support and guidance throughout the preparation of this manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Statement
This research did not involve human participants, animals, or clinical trials; therefore, ethical approval and informed consent were not applicable. This study was conducted in strict accordance with the scientific and ethical guidelines pertaining to the research scope.
Funding
This research was supported by two grants. The first grant, titled “Application of AI Technology in the Construction of Traditional Chinese Medicine Compatibility Platform and Clinical Validation Research Based on Reverse Network Pharmacology,” was assigned Project Number 2021–007. The second grant, part of the “Long Scholar” Clinical Science and Technology Innovation Project, was allocated Project Number KC2022008.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
