Abstract
Objective
To systematically explore the targets and signaling pathways of sinomenine (SIN) in the treatment of osteoarthritis (OA) using integrated network pharmacology, molecular docking, and experimental validation.
Methods
The TCMSP, SwissADME, and Pharmmapper databases were used to predict SIN targets, while the databases of GeneCards, DisGeNET, OMIM, and DrugBank were selected to acquire OA targets. Subsequently, the intersection targets of SIN and OA disease were collected using the Veeny platform. Then, the protein-protein interaction (PPI) network map of “SIN-targets-OA” was established using String database and Cytoscape software. Additionally, the Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were performed through the Database for Annotation, Visualization and Integrated Discovery (DAVID). Additionally, the potential proteins of SIN against OA were validated via molecular docking technique. Finally, the experimental validation was performed in SW1353 cells induced by interleukin (IL)-1β.
Results
A total of 315 potential targets of SIN and 4300 OA-associated targets were collected from public databases, and 42 intersecting potential targets of SIN and OA disease acquired. Then, the PPI network diagram of “SIN-targets-OA” was acquired that comprised a total of 43 nodes and 82 edges. Moreover, 173 GO and 21 KEGG pathway entries were screened with a
Conclusion
This work revealed the underlying targets and signaling pathways of SIN against OA using integrated network pharmacology molecular docking, and experimental validation. These findings provide scientific evidence for the clinical application of SIN for OA treatment.
Introduction
Osteoarthritis (OA), the most common type of degenerative arthritis disease, seriously affects the daily life of people worldwide.1,2 OA is the most common form of arthritis, affecting an estimated 7% of the global population. 3 Although OA has high mortality and disability, there are still no effective therapeutic drugs for OA in clinical applications. Current clinical treatments mainly include analgesics, specific cyclooxygenase inhibitors, and nonsteroidal anti-inflammatory drugs. Embarrassingly, these clinical agents can only temporarily improve clinical symptoms, but their efficacies have been unsatisfactory for patients with enormous side effects. 3 Therefore, there is an urgent need to screen therapeutic targets for OA from natural products, especially derived from Traditional Chinese Medicines (TCMs).
Sinomenine (SIN, 7,8-didehydro-4-hydroxy-3,7-dimethoxy-17-methylmorphinane-6-one, Figure 1) is a major alkaloid purified from

The chemical structure of sinomenine and its related information.
Network pharmacology, widely used to predict the potential targets and pathways for chemical constituents against diseases based on systems biology, bioinformatics, and other disciplines. It is a novel and effective method to construct a correlation network of “drug-targets-pathways-disease”.10,11 Therefore, network pharmacology is a promising approach to systematically explore the underlying mechanisms of chemical constituents derived from chemosynthesis, TCMs, or other natural products.6,12,13 Additionally, molecular docking is a meaningful tool to validate the relationship between the drug and the target protein.2,14,15
In this study, for the first time, a novel and meaningful network pharmacology-integrated molecular docking strategy for revealing the potential targets and underlying pathways of SIN for treating OA. In addition, the relational network map of “SIN-targets-pathways-OA” between SIN and OA was constructed and analyzed. Besides, the experimental validation in (IL)-1β-induced SW1353 cells was performed. The flowchart for dissecting the potential mechanisms of SIN in the treatment of OA and their correlation network is shown in Figure 2.

The flowchart for dissecting the potential mechanisms of SIN in the treatment of OA. Abbreviations: OA, osteoarthritis; SIN, sinomenine.
Materials and Methods
Prediction of SIN Targets
The putative targets of SIN were obtained from 3 public databases: Traditional Chinese Medicine System Pharmacology (TCMSP, https://old.tcmsp-e.com/tcmsp.php), SwissADME (http://www.swissadme.ch/), and Pharmmapper (http://www.lilab-ecust.cn/pharmmapper/), with the keyword of “Sinomenine.” In brief, SIN was entered into the TCMSP database to obtain the corresponding targets directly. On the other hand, SIN was entered into the Pubchem database (https://pubchem.ncbi.nlm.nih.gov/) to obtain its smile number and 2-dimensional (2D) structure. Subsequently, the smile number and 2D structure of SIN were imported into SwissADME and Pharmmapper databases to predict the potential targets, respectively. Finally, all collected targets of OA were standardized using the Universal Protein database (UniProt, https://www.uniprot.org), with the specie limited to
Acquisition of OA Disease Targets
The 4 databases of GeneCards (https://www.genecards.org), DisGeNET (http://www.disgenet.org), Online Mendelian Inheritance in Man (OMIM, https://omim/org), and DrugBank (https://www.chemeurope.com/en/encyclopedia/DrugBank.html) were used to acquire OA-related targets using “osteoarthritis” as the keyword with limited to “Homo sapiens”. Moreover, the definitive OA-related targets were further filtered out by score values greater than the average degree value as well as ≥0.25 from GeneCards and DisGeNET, respectively. Then, all collected targets of OA were converted into standardized protein gene names using the UniProt database. Finally, all OA-related targets were merged from the 4 databases, and the same targets were removed.
Collection of Potential Targets of SIN Against OA
The intersection targets of SIN and OA disease were collected based on the platform of Venny 2.1 (https://bioinfogp.cnb.csic.es/tools/venny/), which is a meaningful approach for visualizing the overlapping targets between drug and disease.
Construction of the Protein-Protein Interaction Network Map
Firstly, the intersection targets of SIN and OA disease were imported into the String database (https://cn.string-db.org/cgi/input?sessionId=bPyuSPm1160L&input_page_show_search=on) with limited to “Homo sapiens”, and the confidence score >0.7. 12 Secondly, the protein-protein interaction (PPI) network map of “SIN-targets-OA” was established using Cytoscape software. Then, the interaction network graph was obtained and analyzed with a “Network Analyzer.” Finally, the core targets from the interaction network map of “SIN-targets-OA” were screened by the degree values more than the mean value. 12
Gene Ontology Enrichment Analysis
To explore the gene functions of SIN against OA, the core targets from the interaction network map of “SIN-targets-OA” were imported into the Database for Annotation, Visualization and Integrated Discovery (DAVID, https://david.ncifcrf.gov/) for Gene Ontology (GO) enrichment analysis, for which a threshold of
Kyoto Encyclopedia of Genes and Genomes Pathway Enrichment Analysis
The Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of the core targets from the interaction network map of “SIN-targets-OA” was also performed using the platform of DAVID and the
Verifying the Potential Target Protein by Molecular Docking
Molecular docking was performed using AutoDock Tools software.6,12 In brief, the 2D structure of SIN and 3-dimensional structure of peroxisome proliferator-activated receptor (PPAR)-γ and IL-17A were downloaded from PubChem (https://pubchem.ncbi.nlm.nih.gov/) and Protein Data Bank (PDB, https://www.rcsb.org/) databases, respectively. 12 Then, the molecular docking was carried out using AutoDock Vina, and further analysis and visual mapping were performed by PyMOL software. 6
Experimental Validation
The SW1353 cells were purchased from the American Tissue Culture Collection and were cultured in Leibovitz's L-15 medium with 10% fetal bovine serum under carbon dioxide atmosphere (5:95, v/v) at 37 °C.
The cell viability of the SIN at differential concentrations (50 and 25 μmol/L) was determined using the CCK-8 kit (Beyotime Biotechnology, Col., Shanghai, China) based on the manufacturer's instructions.
The protein expression level was carried out using a western blotting assay according to references.16–18 The proteins, including IL-17A (0.5 μg/mL, Abcam, Cambridge, UK) and PPAR-γ (1:1000, Abcam, Cambridge, UK) were conducted in this study, and recombinant glyceraldehyde-3-phosphate dehydrogenase (1:10000, Abcam, Cambridge, UK) was selected as the internal control protein.
Statistical Analysis
The data were shown as the mean ± standard deviation (n = 3), which was performed by GraphPad Prism 8.0.1 software. The statistical significance with
Results
Potential Targets of SIN
In this study, a total of 315 potential targets of SIN were predicted from the 3 different public databases, including TCMSP, SwissADME, and Pharmmapper (Figure 3A).

Collection and interaction of SIN-related and OA-related targets. (A) Potential targets of SIN, (B) number of OA-related targets, and (C) Veen plots of gene crossovers. Abbreviations: OA, osteoarthritis; SIN, sinomenine.
Potential Targets of OA Disease
A total of 4300 underlying targets associated with OA were screened from GeneCards (3602), DisGeNET (1827), Drugbank (142), and OMIM (88) databases (Figures 3B and C). Subsequently, 127 OA-related targets with a degree value exceeding the mean value (5) from the GeneCards database. Simultaneously, 93 underlying targets against OA were obtained from the DisGeNET database, and their qualification was a relevance score of more than 0.25. Finally, 404 OA-related target genes were acquired after de-duplication and a combination of these 4 public databases (Figure 3A).
Potential Targets of SIN for the Treatment of OA
Mapping the potential overlapping targets of SIN and OA disease acquired in the “Potential targets of SIN” and “Potential targets of OA disease” sections using the Veeny 2.1 platform (https://bioinfogp.cnb.csic.es/tools/venny/), and 42 intersecting potential target genes were obtained, as shown in Figure 3A and Table 1.
Forty-two Intersecting Target Genes for SIN and OA.
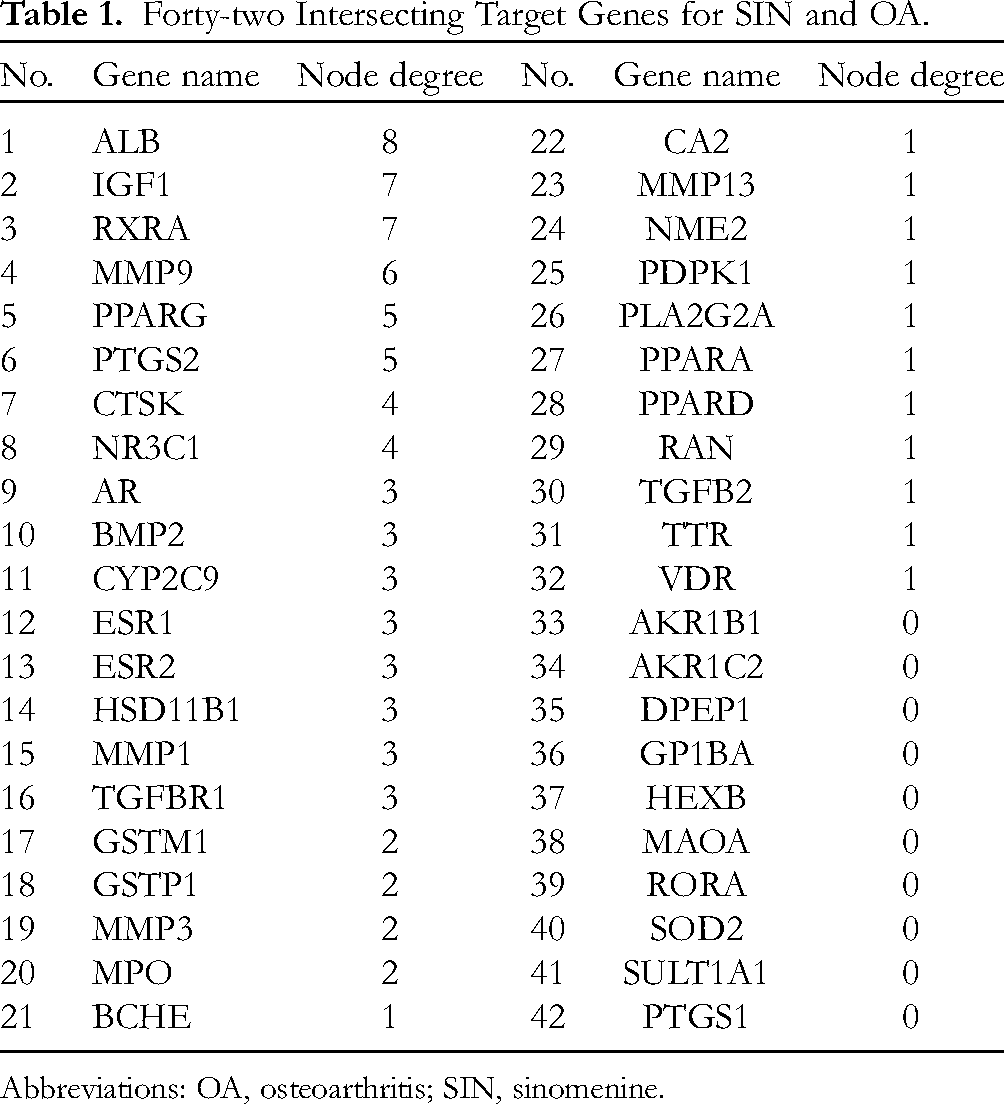
Abbreviations: OA, osteoarthritis; SIN, sinomenine.
The PPI Network Construction of SIN-OA
The 42 intersecting targets were imported into the String database, and the Cytoscape Version 3.8.2 (https://cytoscape.org/) was performed for visualization processing to construct the PPI network map of the SIN in treating OA (Figure 4A and B). Further analysis showed that the PPI network diagram comprised a total of 43 nodes and 82 edges. Among them, 16 intersecting potential target genes, including ALB, IGF1, RXRA, MMP9, PPARG, PTGS2, CTSK, NR3C1, AR, BMP2, CYP2C9, ESR1, ESR2, HSD11B1, MMP1, and TGFBR1, were considered as underlying core targets of SIN for treating OA (Table 1 and Figure 4C), with node degree values (3-8) greater than the average value (2.1). Moreover, ALB, IGF1, and RXRA are the 3 most vital target genes, with node degree values 8, 7, and 7, respectively; followed by MMP9, PPARG, PTGS2, CTSK, and NR3C1 with node degree values 6, 5, 5, 4, and 4, respectively. Meanwhile, the node degree values of AR, BMP2, CYP2C9, ESR1, ESR2, HSD11B1, MMP1, and TGFBR1 are both 3.
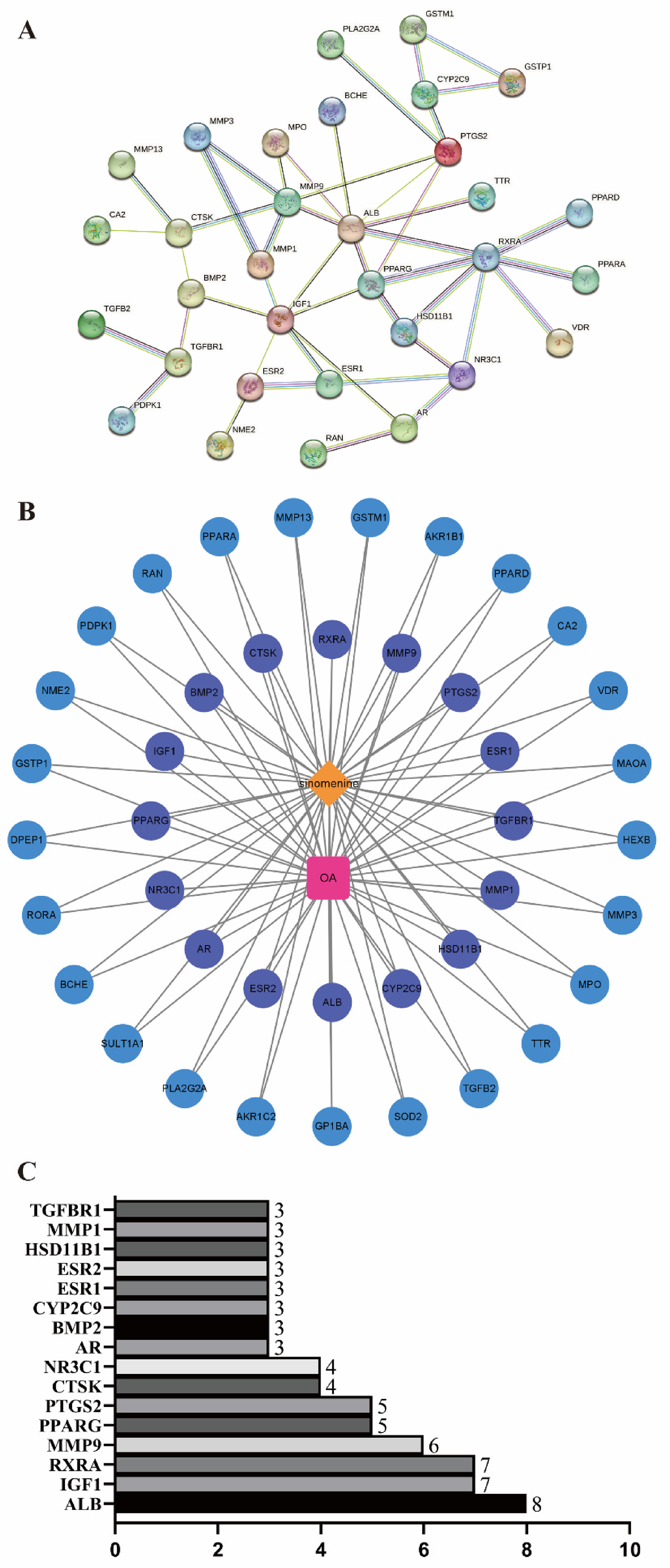
The PPI network construction of SIN-OA. (A) PPI map of SIN-OA, (B) SIN-OA-target network, and (C) the 16 core targets of SIN-OA. Abbreviations: OA, osteoarthritis; PPI, protein-protein interaction; SIN, sinomenine.
GO Functional Enrichment Analysis
To explore the biological activity of SIN in treating OA, the 16 potential core target genes were performed by the GO functional enrichment analysis using the DAVID database. As a result, a total of 173 GO gene annotation entries with the

GO functional enrichment analysis. (A) BP, (B) CC, and (C) MF. Abbreviations: BP, biological process; CC, cellular composition; GO, Gene Ontology; MF, Molecular function.
KEGG Pathway Enrichment Analysis
To reveal potential mechanisms of SIN for the treatment of OA, 21 pathways with a

Kyoto Encyclopedia of Genes and Genomes pathway enrichment analysis.
Therefore, the efficacy of SIN on OA is potentially associated with PPAR and IL-17 signaling pathways, along with 10 related target genes comprising RXRA, PPARA, PPARD, PPARG, PDPK1, MMP1, MMP3, MMP9, MMP13, and PTGS2.
Results of Molecular Docking
A molecular docking approach was performed to validate the underlying mechanism of action and asses the role of SIN in treating OA through the regulation of PPAR and IL-17 signaling pathways. The crystal structures of PPAR-γ (PDB code: 2PRG) 19 and IL-17A (PDB code: 4HR9) 12 were obtained, which were the main active form of the ligand-activated nuclear hormone of PPAR and IL-17 proteins, respectively.12,20 As a result, the SIN has a good binding site with PPAR-γ (Figure 7A) and IL-17A (Figure 7B) due to their corresponding docking scores of −6.1 and −6.3 kcal/mol, respectively, which were no more than −5 kcal/mol. 21 It indicated that SIN can be useful for treating OA through the regulation of PPAR and IL-17 signaling pathways.

The molecular docking diagrams of SIN and PPAR-γ (A) and IL-17A (B). Abbreviation: SIN, sinomenine.
Validating the Underlying Mechanism of SIN Against OA In Vitro
To measure the efficacy of SIN on the viability of SW1353 cells, a CCK-8 assay was carried out. The results showed that the concentrations at 50 and 25 μmol/L of SIN did not have toxicity to SW1353 cells, with their cell viability being 98.6 ± 3.2% and 99.5 ± 2.9% when compared with the control group.
Afterward, the concentration at 50 μmol/L of SIN was selected to explore the potential mechanism of action using western blot assay. As illustrated in Figure 8, treatment with IL-1β (10 ng/mL) significantly induced the protein expression of IL-17A and PPAR-γ (

SIN inhibited the activation of IL-17A and PPAR-γ in IL-1β-induced SW1353 cells. ###
Discussion
Although OA is one of the most common chronic arthropathy, there are currently no effective treatments and drugs for ameliorating OA in clinical applications.2,22 SIN, a phytochemical derived from
In this work, for the first time, the effect of SIN on OA and its potential mechanisms were explored using integrated network pharmacology and molecular docking approaches. Several results are acquired in the present work: (1) We first indicated that SIN has an effect against OA using a network pharmacology approach. (2) Sixteen core target genes, including ALB, IGF1, RXRA, MMP9, PPARG, PTGS2, CTSK, NR3C1, AR, BMP2, CYP2C9, ESR1, ESR2, HSD11B1, MMP1, and TGFBR1, were considered as pivotal targets of SIN for treating OA. (3) SIN possesses an effect on OA by regulating PPAR and IL-17 signaling pathways. (4) SIN has a good binding site with PPAR-γ and IL-17A based on the molecular docking analysis, further indicating that PPAR and IL-17 signaling pathways play vital parts in SIN for the treatment of OA.
Network pharmacology is a novel drug design strategy developed based on integrating systemic biology, which can comprehensively show the multichannel interactions of drugs and disease targets and build the network of “drug-targets-genes-disease.”6,12,24 Therefore, the integrated network pharmacology approach is a meaningful tool to establish a bridge of internal correlation between compounds and potential proteins/genes at the system level, thereby has been widely used to reveal pharmacodynamics and underlying mechanisms of TCMs and/or compounds in preclinical studies. In the present study, the PPI network map of the SIN for the treatment of OA was constructed using String database and Cytoscape software, and 16 underlying core targets, including ALB, IGF1, RXRA, MMP9, PPARG, PTGS2, CTSK, NR3C1, AR, BMP2, CYP2C9, ESR1, ESR2, HSD11B1, MMP1, and TGFBR1, were acquired. Among them, ALB, IGF1, RXRA, MMPs, PPARG, PTGS2, ESRs, CTSK, TGFBR1, NR3C1, and BMP2 genes are positively involved in the OA disease,25–32 while CYP2C9 and HSD11B1 genes take parts in the inflammatory response.33,34 The results indicated that these 16 major genes are potential core targets responsible for SIN to treat OA, and are closely associated with inflammatory response. Based on these, the 16 potential core genes are partly attributed to SIN for the treatment of OA, which was consistent with various previous works.
GO functional enrichment analysis displayed that SIN regulates 173 GO gene annotation entries, including 122 BP, 14 CC, and 37 MF. Meanwhile, positive regulation of transcription from RNA polymerase II promoter, positive regulation of transcription, DNA-templated, and negative regulation of apoptotic process were the major terms in BP for SIN against OA, which were consistent with previous report. 35 On the other hand, the extracellular space, extracellular region, extracellular exosome, and nucleoplasm were major potential terms in CC that take part in the process of SIN against OA, and they were intimately involved in the OA disease.24,36,37 Additionally, the top 3 terms in MF included protein binding, zinc ion binding, and DNA binding of SIN to treat OA were also reported in other research. 38 Simultaneously, PPAR and IL-17 were the core signaling pathways of SIN for treating OA based on the KEGG pathway enrichment analysis in this study. As the previous reports, the PPAR signaling pathway is positively related to OA. 39 IL-17, one of the most pivotal inflammatory factors produced by Th17, plays a vital role in cartilage and osteoblasts. 6 Therefore, PPAR and IL-17 are the core signaling pathways of SIN for the treatment of OA.
Molecular docking technology is a pivotal strategy to predict the combination between drug molecules (ligands) and receptors (proteins).12,40 Simultaneously, the binding capacity of ligand/receptor is a vital parameter expressed as a docking score, where a value of no more than −5 kcal/mol indicating good binding ability.
21
In the present work, the docking score of SIN with PPAR-γ and IL-17A were −6.1 and −6.3 kcal/mol, respectively, indicating that SIN possesses the treatment of OA due to it has good binding ability with PPAR-γ and IL-17A proteins based on molecular docking technology. In addition, the experimental verification
There are still some shortcomings in the present study: (1) The pharmacodynamics and potential mechanisms experiments of SIN against OA
Conclusion
In summary, 16 potential core genes (ALB, IGF1, RXRA, MMP9, PPARG, PTGS2, CTSK, NR3C1, AR, BMP2, CYP2C9, ESR1, ESR2, HSD11B1, MMP1, and TGFBR1) and 2 key signaling pathways (PPAR and IL-17) of SIN in treating OA were obtained using integrated network pharmacology. Moreover, SIN has a good combination with PPAR and IL-17, which were verified through molecular docking techniques and experimental verification. The deficiency of the present work is that the potential targets and pathways of SIN against OA were prediction only based on public biological platform approaches, and experimental verification
Supplemental Material
sj-docx-1-npx-10.1177_1934578X241262909 - Supplemental material for Integrated Network Pharmacology, Molecular Docking, and Experimental Validation to Explore Potential Mechanisms of Sinomenine in the Treatment of Osteoarthritis
Supplemental material, sj-docx-1-npx-10.1177_1934578X241262909 for Integrated Network Pharmacology, Molecular Docking, and Experimental Validation to Explore Potential Mechanisms of Sinomenine in the Treatment of Osteoarthritis by Shaojun Wang, Fanglin Lai, Ting Xiang and Yan Xu in Natural Product Communications
Footnotes
Author Contributions
SJW and YX are the principal investigators and conducted statistical analysis and drafted the article. SJW, FLL, and TX performed data management and analysis. SJW and YX edited and revised the article. All authors read and approved the final article.
Data Availability Statement
The original contributions presented in the study are included in the article, further inquiries can be directed to the corresponding author.
Declaration of Conflicting Interests
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Ethical Approval
This study does not contain any studies with human or animal subjects. Ethical approval does not apply to this article.
Statement of Human and Animal Rights
This study does not contain any studies with human or animal subjects. The statement of human and animal rights does not apply to this article.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
