Abstract
Objective
This study set out to examine the phenylpropanoid pathway gene expression and phenolic compound accumulation in different organs of Chelidonium majus (C. majus).
Methods
Total RNA was extracted and expression of phenylpropanoid biosynthesis genes was assessed using quantitative real-time polymerase chain reaction in different organs of C. majus. In addition, the phenylpropanoid compounds were analyzed from different organs of C. majus by using high-performance liquid chromatography.
Results
The results demonstrate the variance in the expression levels of phenylpropanoid pathway genes in the flowers, leaves, stems, and roots of C. majus. CmPAL and CmC4H were expressed at their greatest levels in the stem, while CmCHS and CmCHI were strongly expressed in the leaf, and CmF3H, CmFLS, Cm4CL, and CmDFR were more abundant in the flower. The total phenolic contents in the different organs also varied significantly. The highest value was recorded in the flowers (2152.89 ± 78.71 μg/g dry wt.), which was 54.26 times higher than the lowest values obtained from roots (39.68 ± 0.05 μg/g dry wt.). Except for roots, rutin showed the highest level in all other organs. According to a heat map, the majority of the phenolic compounds were highest in the flower. By using variable importance in projection analysis, the most significant metabolite was found to be chlorogenic acid, which had a positive association with all phenolic chemicals, but it showed a negative correlation with trans-cinnamic acid. Except for trans-cinnamic acid, all other phenolic compounds likewise showed positive correlation with other phenolic compounds.
Conclusions
This work offers pertinent data on the expression analysis of genes involved in the phenylpropanoid biosynthetic pathway and the presence of phenolic compounds in C. majus. These findings will be helpful for the researcher to understand the relationship between the expression level of phenylpropanoid pathway genes and phenylpropanoid compound accumulation in C. majus.
Introduction
Chelidonium majus (C. majus) of the family Papaveraceae can be found extensively worldwide in Asia, Europe, North America, and Northwest Africa. 1 This perennial herbaceous plant is also called by many names such as greater celandine, swallow-wort, or rock poppy. 2 In Chinese traditional medicine, C. majus is used to treat obstruction of blood circulation and to reduce pain edema and icterus. 3 Furthermore, the latex of C. majus has long been used to cure several disorders such as stomach ulcers, liver diseases, oral infections, tuberculosis, and skin rashes in many European, Asian, and African countries. 3 Isoquinoline alkaloids, flavonoids, carotenoids, saponins, organic acids, and vitamins A and C are the distinct biologically active compounds of C. majus.4,5 According to the metabolites identified, it has been found to possess a range of pharmacological effects such as antimicrobial activity, 6 anti-inflammatory, analgesic, 7 natriuretic, antidiuretic, 8 and anticancer 3 effects.
Phenylpropanoids are a large class of plant secondary metabolites and conduct plenty of important functions in plants such as signaling caused by biotic or abiotic stresses, attraction for reproduction, and protection against harmful factors such as UV irradiation, excessive water loss, herbivore, and pathogens. 9 Also, they have numerous health benefits related to the aforementioned properties.
The phenylpropanoid biosynthetic pathway begins with the deamination of phenylalanine to cinnamic acid, catalyzed by the enzyme phenylalanine ammonia-lyase (PAL) (Figure 1). The enzymes cinnamate 4-hydroxylase (C4H) and 4-coumarate-CoA ligase (4CL) catalyze, respectively, 2 consecutive conversions of cinnamic acid to 4-coumaroyl CoA. Several phenylpropanoid products, including, lignins, coumarins, stilbenes, and flavonoids, are synthesized from the precursor p-coumaroyl CoA. 10 The condensation of one p-coumaroyl CoA and 3 malonyl-CoA molecules derived from the Krebs cycle turns into naringenin chalcone through the catalyzation of chalcone synthase (CHS). In the following step, chalcone isomerase (CHI) transforms naringenin chalcone into naringenin. Then, by the enzyme flavanone 3-hydroxylase (F3H), naringenin is switched to dihydrokaempferol, which can be further converted into kaempferol or dihydroquercetin by the enzymes flavonol synthase (FLS) and flavonoid 3′-hydroxylase (F3′H), respectively. Lastly, the enzyme FLS and dihydroflavonol 4-reductase (DFR) transforms subsequently dihydroquercetin into quercetin and leucocyanidin, the precursor of epicatechin and catechin hydrate.
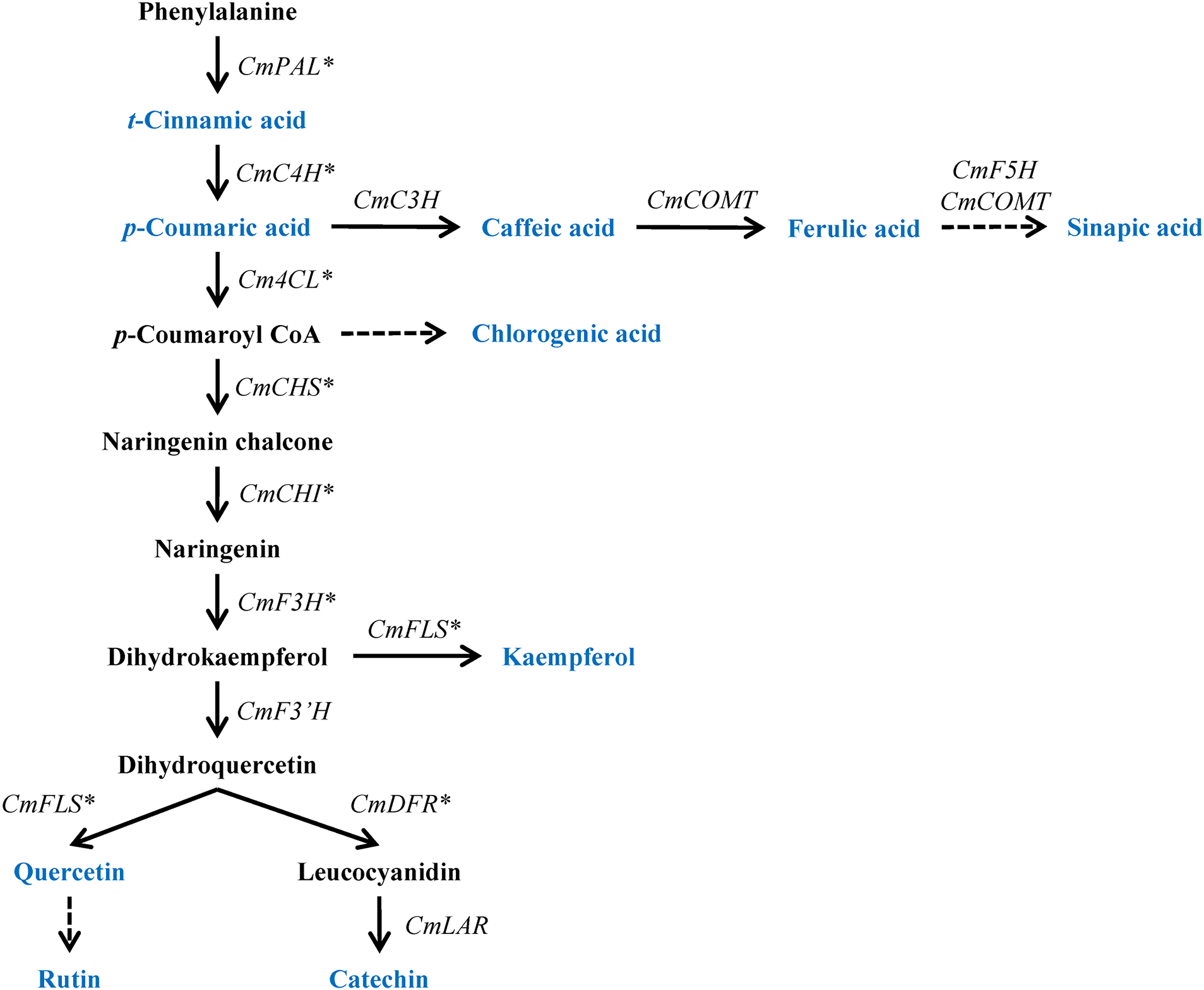
Schematic representation of phenylpropanoid biosynthetic pathway in Chelidonium majus. The blue color indicates the phenylpropanoids utilized in this study via HPLC analysis. Asterisk denotes the target genes selected for gene expression via qRT-PCR. C3H, 4-hydroxycinnamate 3-hydroxylase; COMT, caffeic acid 3-O-methyltransferase; F5H, ferulate 5-hydroxylase; 4CL, 4-coumaroyl CoA ligase; LAR, leucocyanidin reductase; qRT-PCR, quantitative real-time polymerase chain reaction; HPLC, high-performance liquid chromatography.
Many prior studies have addressed the isoquinoline alkaloids of C. majus, which are the principal secondary metabolites in this plant. 11 On the other hand, there has been few research dealing with the phenolic compounds in C. majus. In addition, metabolic profiling could be used to determine diverse cell- or organism-specific responses to biological or environmental conditions through the identification and quantitation of low-molecular-weight intermediates produced from many catabolic reactions and the biosynthetic pathways in plants. Therefore, this study is most important for the researchers. This current study will fulfill the lack of comprehensive research on the changes in flowers, leaves, stems, and roots of C majus. Therefore, the purpose of this study was to identify and quantify phenolic compounds as well as analyze the expression profiles of the biosynthetic genes related to the production of phenolic compounds in different parts (flowers, leaves, stems, and roots) of C. majus plants. The correlation between the phenolic compounds of various parts of the C majus plants and individual phenolic compounds and other phenolic compounds was also investigated using multivariate analysis.
Results
Expression of Phenylpropanoid Biosynthetic Genes in Different Organs of C. majus
Quantitative real-time polymerase chain reaction (qRT-PCR) was performed to determine the expression levels of phenylpropanoid biosynthetic genes in the flowers, leaves, stems, and roots of C. majus. Figure 2 displays the levels of expression of CmPAL, CmC4H, Cm4CL, CmCHS, CmCHI, CmF3H, CmFLS, and CmDFR in the flowers, leaves, stems, and roots of C. majus. In various parts of the plant, the expression levels of the genes varied greatly. The expression of CmPAL was significantly higher in the stem followed by the leaf, flower, and root. The greatest expression of CmC4H was likewise seen in the stem. In comparison, the C. majus root and flower showed a moderate expression, while the leaf of the plant showed the least expression of CmC4H. Furthermore, Cm4CL showed a significant variance. The flower had the greatest amount of Cm4CL expression, which was followed by stem, leaf, and root. Although CmCHS was not expressed in the root, it was most highly expressed in the leaf as compared to the flower and stem. The maximal expression of CmCHI, which is statistically comparable to the amount of expression in flowers, was also seen in leaves as opposed to other plant parts. However, the root showed the least level of expression among all plant sections. The highest level of CmF3H expression was in the flower, followed by moderate levels in the leaf, and lowest levels in the root. Similar patterns of expression were followed by CmFLS as well. In relation to various plant components, a considerable fluctuation in CmDFR expression level was also seen. According to Figure 2, the flower had the greatest degree of expressiveness, followed by the root, stem, and leaf.
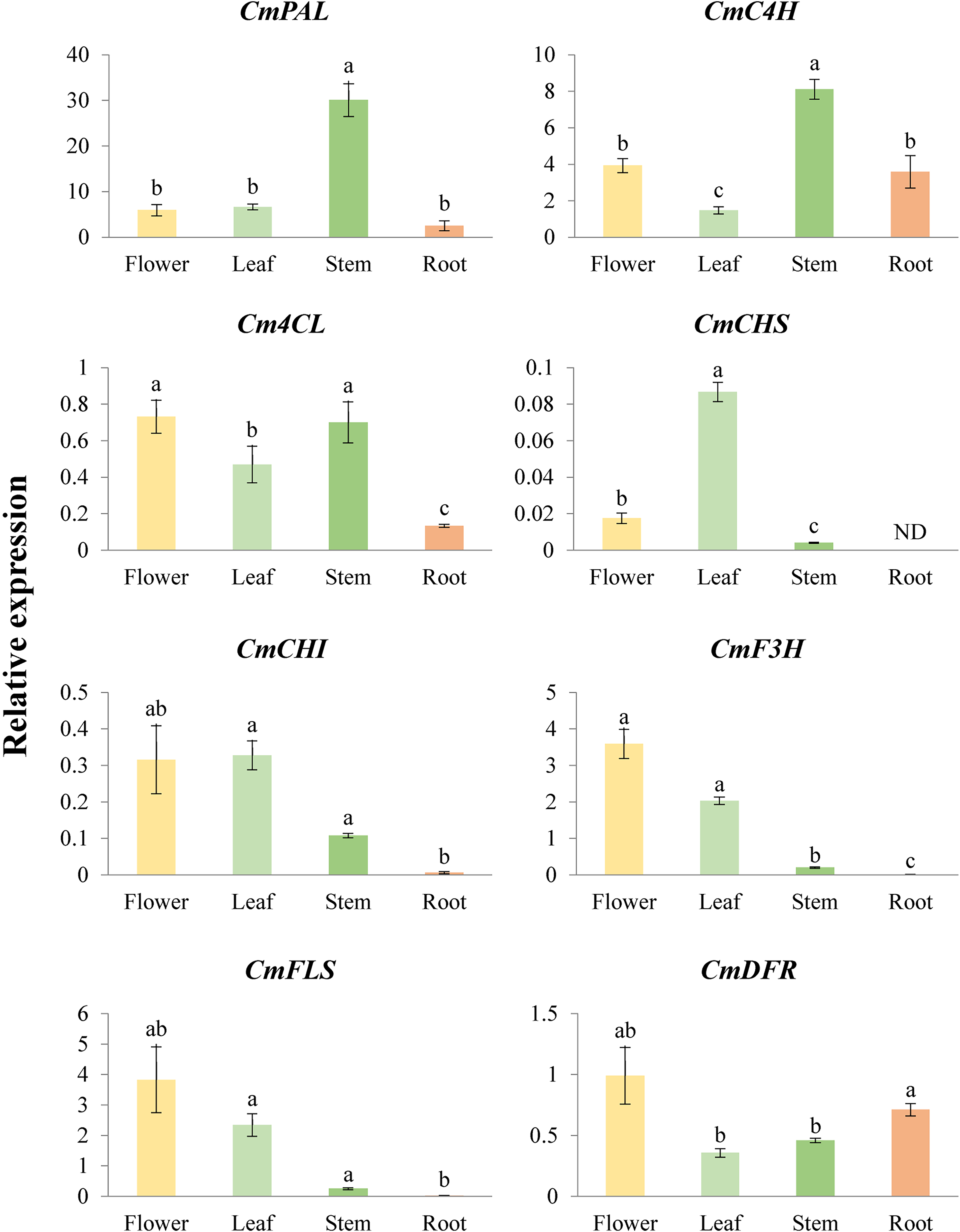
Relative expression levels of phenylpropanoid biosynthetic genes in different organs of Chelidonium majus. Each value and error bar indicates the mean and standard deviation (SD) obtained from 3 replicates. Letters a-c denote significant differences (P < .05) by Tukey's honestly significant difference (HSD) test (if data fulfilled homoscedasticity) or Welch's one-way ANOVA followed by Tamhane's T2 test (if data did not fulfill homoscedasticity) for multiple comparisons with an α = 0.05. ND, not detected; ANOVA, analysis of variance.
Phenylpropanoid Content in Different Organs of C. majus
High-performance liquid chromatography (HPLC) and ultra-HPLC coupled to quadrupole Orbitrap mass spectrometer (UHPLC-Q-Orbitrap-MS) analysis showed the presence of 10 phenylpropanoid-derived compounds in C. majus parts, including trans-cinnamic acid, p-coumaric acid, caffeic acid, ferulic acid, sinapic acid, chlorogenic acid, kaempferol, quercetin, rutin, and catechin (Table 1, Figures S1-S7). The total phenolic compound contents in the different organs of C. majus were distinct largely. The highest value was recorded in the flowers (2152.89 ± 78.71 μg/g of dry wt.), which was 54.26 times higher than the lowest values obtained from roots (39.68 ± 0.05 μg/g of dry wt.). Especially, rutin was identified as the largest content in different organs except for roots. All the phenolic compounds except trans-cinnamic acid were identified in the flower. Ferulic acid, sinapic acid, kaempferol, and catechin were not detected in the leaf. The presence of only 4 phenylpropanoid-derived compounds, trans-cinnamic acid, p-coumaric acid, rutin, and catechin, was identified in the stem of C. majus. In the root, only 2 phenolic compounds (trans-cinnamic acid and kaempferol) were detected in a small amount.
The Accumulation of Phenolic Compounds in the Flowers, Leaves, Stem, and Root of Chelidonium majus.
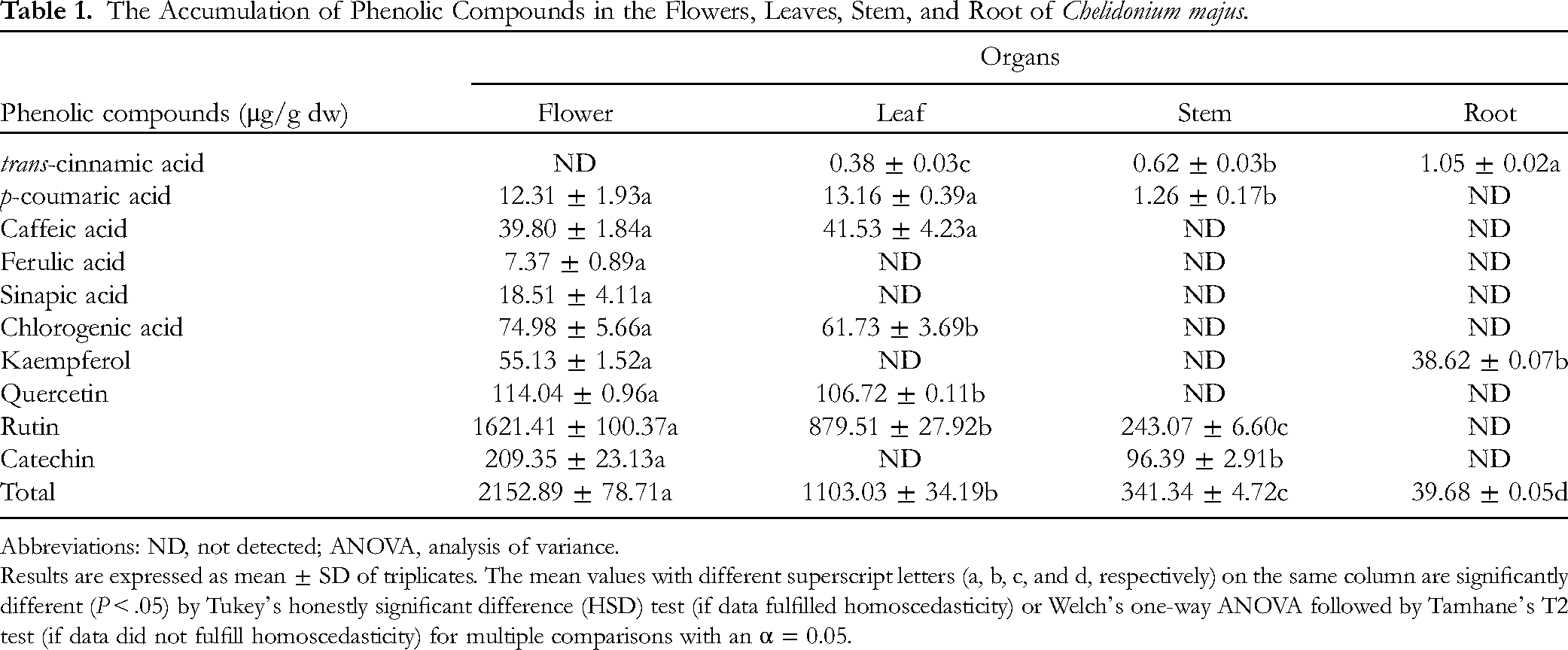
Abbreviations: ND, not detected; ANOVA, analysis of variance.
Results are expressed as mean ± SD of triplicates. The mean values with different superscript letters (a, b, c, and d, respectively) on the same column are significantly different (P < .05) by Tukey's honestly significant difference (HSD) test (if data fulfilled homoscedasticity) or Welch's one-way ANOVA followed by Tamhane's T2 test (if data did not fulfill homoscedasticity) for multiple comparisons with an α = 0.05.
Metabolic Profiling of Identified Metabolites
Ten phenolic compounds were identified and quantified from the different organs of C. majus. A heat map result showed that most of the individual phenolic compounds were highest in the flower (Figure 3). In detail, the heatmap was mainly clustered into 2 groups, namely group 1 and group 2. Group 1 consisted of the phenolic compounds highly present in the flower and leaf (rutin, p-coumaric acid, caffeic acid, chlorogenic acid, quercetin, and total phenolics), whereas group 2 was separated based on most of the individual phenolic compounds highly present only in the flower, except trans-cinnamic acid. In the root, the trans-cinnamic acid and kaempferol contents were only present, whereas other phenolic compounds were not detected. However, in the heatmap, these 2 compounds (trans-cinnamic acid and kaempferol) formed a separate node in cluster 2. Similarly, the catechin content was significantly high in the flower followed by the stem, whereas it was not detected in the leaf and the root also formed a separated node in cluster 2. Interestingly, ferulic and sinapic acid content were only detected in the flower which formed a separate node in cluster 2. Significantly higher (P ≤ .05) levels of phenolic compounds in the different organs of C. majus (flower, leaf, stem, and root) are shown in Figures S6 and S7, which demonstrates that distinctive metabolite signature features were found for the different organs. Considering these results, the different organs showed differential accumulation of phenolic compounds in C. majus.

A heatmap showing the changes in the relative metabolite content in different organs of Chelidonium majus. The relative metabolite concentrations are shown on the color scale (2 to −2) at the right, with high and low concentrations represented by red and blue, respectively.
According to the principal component analysis (PCA), PC1 and PC2 showed 76.1% and 18.0% variance, respectively (Figure 4). Among the different organs, all the organs showed clear separation. This clear separation between the different organs of C. majus might be due to the trans-cinnamic acid, and the only compound that showed a positive eigenvector value was 0.32053, whereas all other compounds such as total phenolics, rutin, chlorogenic acid, quercetin, ferulic acid, caffeic acid, p-coumaric acid, sinapic acid, catechin, and kaempferol, exhibit negative eigenvector values −0.34432, −0.34394, −0.32674, −0.31653, −0.30192, −0.30095, −0.30006, −0.29736, −0.25176, and −0.017289, respectively (Figure 4A). In addition, partial least-squares discriminant analysis (PLS-DA) was carried out to exploit the separation between the different organs, the result showed a far group separation between each organ, which showed 76.1% (PC1) and 7.0% (PC2). Similar to the PCA model, the PLS-DA analysis showed the positive eigenvector value for trans-cinnamic acid (0.31986). All other phenolic compounds showed negative eigenvector values such as rutin, total phenolics, chlorogenic acid, quercetin, caffeic acid, p-coumaric acid, ferulic acid, sinapic acid, catechin, and kaempferol, and the associated eigenvector values were −0.34584, −0.3458, −0.33131, −0.32161, −0.30652, −0.30497, −0.30115, −0.29658, −0.24647, and −0.17488, respectively (Figure 4B). In variable importance in projection (VIP) analysis, the most important predicted metabolites for the far separation were chlorogenic acid, quercetin, rutin, caffeic acid, and p-coumaric acid (Figure 5). This supports the heatmap results, which showed that the different organs showed differential accumulations of individual compounds in C. majus.

Principal component analysis (PCA) (A) and PLS-DA (B) scores and loading plots of the metabolites identified in different organs of Chelidonium majus, respectively. PLS-DA, partial least-squares discriminant analysis.

The main components, separating different organs of Chelidonium majus, are based on the VIP scores obtained via the PLS-DA model. PLS-DA, partial least-squares discriminant analysis; VIP, variable importance in projection.
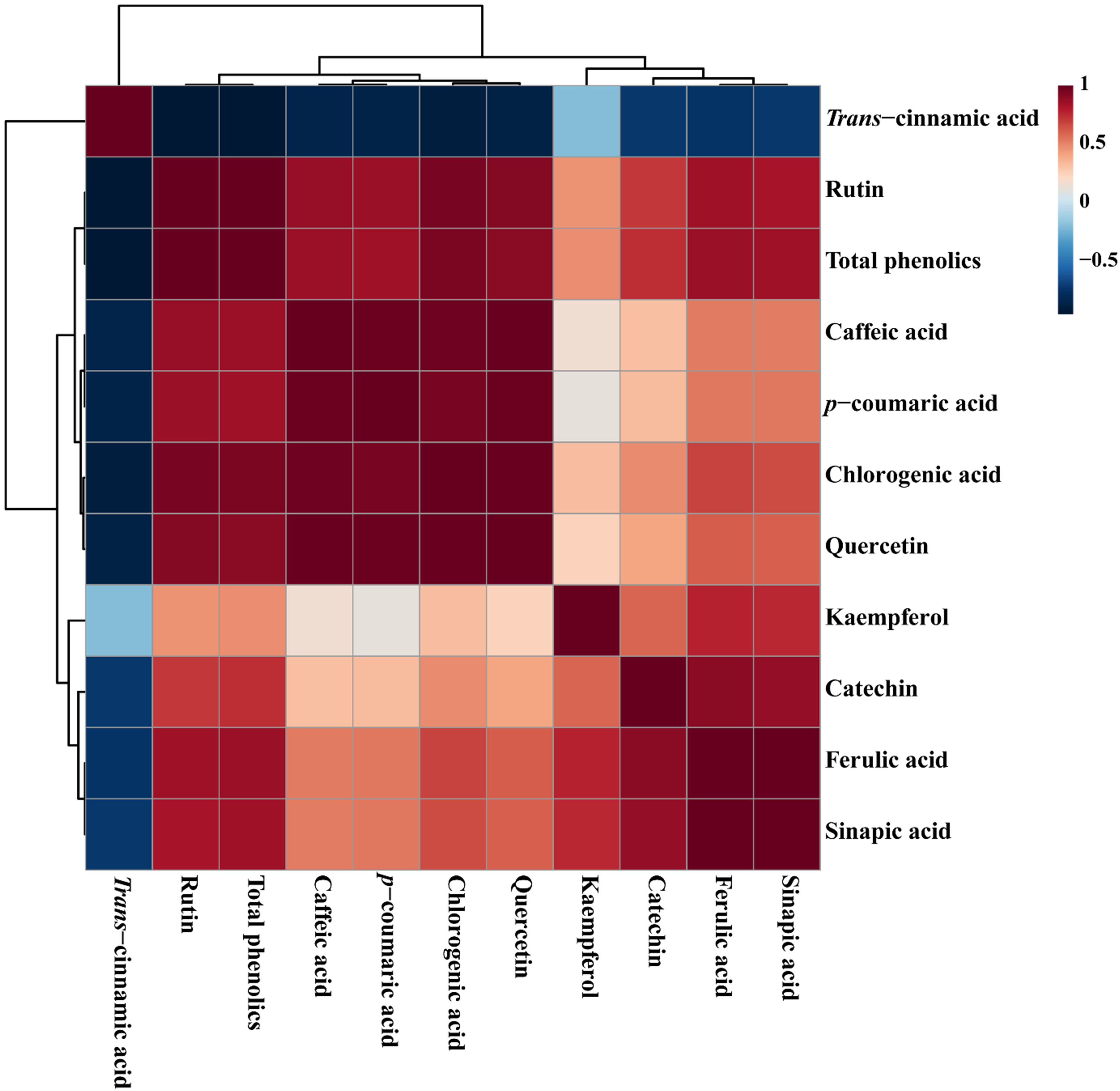
Correlation matrix of metabolites identified in different organs of Chelidonium majus. Each colored box signifies Pearson's correlation coefficient for a given pair of compounds. The correlation coefficient values are indicated by the intensity, with a high and low concentration appearing in red and blue, respectively, as shown on the right-side color scale (1 to −0.5).
Correlation matrix analysis of 10 identified individual phenolic compounds from different organs of C. majus (Figure 6) was divided into 3 main groups based on a positive correlation. Interestingly, the trans-cinnamic acid alone formed a separate cluster, this separation might be due to the only phenolic compound which showed a positive eigenvector value in both PCA and PLS-DA models is trans-cinnamic acid. The trans-cinnamic acid showed a negative correlation with all individual phenolic compounds and the total phenolics. In contrast, the most important metabolite identified by VIP analysis was chlorogenic acid, which exhibited a negative correlation with trans-cinnamic acid (r = −0.88495, P = .00013), whereas it showed a positive correlation with all phenolic compounds. Similarly, all other phenolic compounds also showed positive correlation with other phenolic compounds, except trans-cinnamic acid.
Discussion
Eight genes (CmPAL, CmC4H, Cm4CL, CmCHS, CmCHI, CmF3H, CmFLS, and CmDFR) related to phenylpropanoid biosynthesis were identified in different parts of C. majus plant. The expression patterns of CmPAL, CmC4H, Cm4CL, CmCHS, CmCHI, CmF3H, CmFLS, and CmDFR were analyzed in the root, stem, leaf, and flower of C majus.
In the root of the C. majus plant, the CmC4H gene was expressed more than the other genes followed by CmPAL, CmDFR, Cm4CL, CmFLS, CmF3H, and CmCHI. The root of C. majus did not have CmCHS gene expression. In another investigation, it was shown that buckwheat roots had the greatest expression of Fe4CL1, whereas stems had the highest levels of Fe4CL2 expression. 12 Additionally, Lee et al noted that IdCHS was hardly expressed in the root. 13 However, in our study, it was found that the CmPAL gene was expressed most strongly in the stem of C. majus. When compared to CmPAL and CmC4H's expression level in the stem was fairly low, but it was greater than that of other discovered genes, which showed very low levels of expression. Three isoforms of PAL were discovered in Scutellaria baicalensis, and Xu et al noticed that they had various expression patterns in various organs. 14 The stem, leaf, and root, respectively, had the greatest amounts of SbPAL1, SbPAL2, and SbPAL3 transcripts. 14 According to research by Lee et al, Ixeris dentata var. albiflora leaf had the greatest levels of expression for IdPAL1, IdC4H, Id4CL1, IdCHS, and IdDFR. 13 In the leaf, all the genes were expressed, and there was a noticeable fluctuation in the amount of expression, as shown in Figure 2. The CmPAL gene expressed at the greatest level in this organ similar to the stem and was followed by the CmFLS, CmF3H, CmC4H, Cm4CL, CmDFR, CmCHI, and CmCHS genes. Lee et al reported that leaf expression of IdCHS was 2.5-, 6.9-, and 32.0-fold higher than that in the bud, stem, and flower, respectively, whereas it was rarely expressed in the root of Ixeris dentata var. albiflora. 13 The C. majus flower likewise showed expression of every gene, with CmPAL showing the greatest level of expression. Moderate expression was seen in CmC4H, CmFLS, and CmF3H. The expression of other genes, such as CmDFR, Cm4CL, CmCHI, and CmCHS, was minimal. So far, multiple researchers have looked at the ways that the genes responsible for phenylpropanoid biosynthesis are expressed in diverse plants.12–17 According to Richet et al, exposure to ozone may affect the expression of phenylpropanoid genes in several popular organs. 18 Additionally, treatments with methyl jasmonate, wounding, yeast extract, and silver nitrate enhanced the expression of genes involved in the synthesis of phenylpropanoid in S. baicalensis and Agastache rugosa cell culture.14,19
The presence of 10 phenylpropanoid-derived compounds in C. majus parts, including trans-cinnamic acid, p-coumaric acid, caffeic acid, ferulic acid, sinapic acid, chlorogenic acid, kaempferol, quercetin, rutin, and catechin, was detected in this study. The flower accumulated 54.26 times higher phenylpropanoid-derived compounds than the roots. Except for roots, rutin was found to have the highest concentration in all other organs. However, Lee et al found that the leaf had the greatest concentrations of catechin, chlorogenic acid, ferulic acid, and quercetin, whereas benzoic acid and rutin were not found in the leaf of Ixeris dentata var. albiflora. 13 The leaf of F. esculentum was found to have the greatest concentration of rutin, followed by the stem, seed, and root. 12 Rutin is an effective antioxidant and a good source of dietary fiber. The kind of plant and environmental variables affect its composition. 20 There was no trans-cinnamic acid found in the flower, although there was a little amount of ferulic acid buildup. In contrast, the accumulation of trans-cinnamic acid was extremely little in the leaf, stem, and root, but neither ferulic acid nor sinapic acid was found there. Catechin and kaempferol did not accumulate in the leaf either. In the stem only 4 (catechin, p-coumaric acid, rutin, and trans-cinnamic acid), and in the root only 2 (trans-cinnamic acid and kaempferol) phenylpropanoid-derived compounds were detected. These phenylpropanoid-derived chemicals may have accumulated as a result of the genes expressed in various plant sections. Numerous investigations have revealed a connection between PAL, C4H, and 4CL gene expression and flavonoid accumulation in numerous species.21–24 Additionally, it has been shown that the expression of the genes AgPAL and AgC4H was associated with the generation of secondary metabolites in A. gigas. 21
The PAL and C4H are the initial transcript genes in the phenylpropanoid biosynthetic pathway. These 2 genes were highly expressed in the stem, however, a significant amount of total phenolics was achieved in the flower. This temporal variation was independent of CmPAL and CmC4H expression (Figure 2); therefore, these 2 genes does not play a significant role in the phenylpropanoid biosynthetic pathway in C. majus. This might be because of the 2 hypotheses (1) RT-PCR using degenerated primers failed to catch major CmPAL and CmC4H ortholog players in phenylpropanoid biosynthesis in different organs of C. majus and (2) the level of CmPAL and CmC4H transcripts might be enough to allow the pathway to operate without coordinated regulation.25,26 In addition, it has been reported that the phosphorylation of PAL and ubiquitination of PAL by KFB proteins were identified in French bean and Arabidopsis thaliana, respectively.27,28 This might be due to the conservation of the phosphorylation site in PAL from diverse species, which implies that phosphorylation of PAL may be a ubiquitous regulatory mechanism in higher plants; hence the expression of CmPAL and CmC4H might be regulated by phosphorylation and other posttranscriptional modifications. To clarify these hypotheses, transcriptome-wide analysis of different parts of C. majus might be helpful to elucidate this part.
Najid et al study reported that the CHI gene is involved in the isomerization of naringenin chalcone to naringenin. 29 The main key point for rutin production is naringenin. Therefore, we hypothesized that there might be a close relationship between CmCHI gene expression level and rutin production in different organs of C. majus. Our result proves the hypothesis, that among the different organs, only flower, leaf, and stem showed greater expression of CmCHI genes which leads to the highest production of rutin content. In rice, it has been reported that CHI could interact substantially with F3H, F3′H, DFR, and ANS1 by forming a flavonoid multienzyme complex. This result was consistent with our study that the expression levels of CmCHI, CmF3H, and CmDFR were high in the flower, leaf, and stem. This might be due to the CmCHI gene interacting with the CmF3H, and CmDFR thus leading to the highest expression of all those genes in the flower, leaf, and stem which results in the highest production of rutin content.
Previously, several studies have reported that F3H and FLS regulates the production of quercetin and kaempferol by catalyzing related metabolic reactions.30–34 This was consistent with our result that the CmF3H and CmFLS genes were significantly expressed in flowers, which leads to the highest accumulation of quercetin and kaempferol in flowers. In the present study, the phenylpropanoid biosynthesis genes in C. majus demonstrated organ-specific expression patterns, suggesting that they might have different physiological processes for biosynthesis depending on the organ.
The CHS gene is the initial enzyme specific for the flavonoid biosynthetic pathway. In Sophora flavescens, the expression level of SfCHS transcripts was substantially present in the leaf. 35 This result was consistent with our study that the gene expression of CmCHS was significantly high in the leaf. From this result, it is shown that in most of the plant genera, the highest gene expression of CmCHS was achieved in the leaf.
Previous studies reported that the PAL gene expression and PAL enzymatic activity were highest during developing fruits.26,36–38 In contrast, in our study, the CmPAL gene expression was high in the stem, this is because C. majus does not produce fruit. Hence, we did not analyze the CmPAL gene expression in the fruit of C. majus.
Conclusions
In addition to revealing the presence of phenolic compounds in C. majus, this study also offers significant information on the expression of genes crucial for the generation of phenylpropanoid chemicals. The study discovered variations in the expression level of CmPAL, CmC4H, Cm4CL, CmCHS, CmCHI, CmF3H, CmFLS, and CmDFR in the flowers, leaves, stems, and roots of C. majus. The majority of the phenolic compounds were significantly present in the flower. In addition, by VIP analysis, we found that an important metabolite for the separation of different organs was chlorogenic acid, which had a positive correlation with all phenolic compounds, but it showed a negative correlation with trans-cinnamic acid. This research offers crucial details on the expression of genes important to the synthesis of phenylpropanoid compounds as well as the presence of phenolic compounds in each organ of C. majus that will be useful for future studies.
Materials and Methods
Plant Materials
Chelidonium majus seeds were obtained from the greenhouse house farm of Chungnam National University, Daejeon, Republic of Korea. The seeds were sown in a pot and then grown for 3 months in the greenhouse of Chungnam National University. Every 2 days the sowed seeds were sprayed with water. The plantlets that emerged with different plant organs (flowers, leaves, stems, and roots) were harvested at the fully flowered stage, and then immediately frozen in liquid nitrogen, and stored the materials at −80 °C for further analysis. Each sample was collected in triplicate.
RNA Extraction and cDNA Synthesis
Total RNA was extracted from the different organs (flowers, leaves, stems, and roots) of the C. majus plants. All the samples were ground into a fine powder with the help of liquid nitrogen using a pestle and mortar. Then, 100 mg of each fine powdered sample was transferred to a new 2.0 mL Eppendorf tube. To extract the total RNA Plant Total RNA Mini Kit (Geneaid, Taiwan) was used, according to the manufacturer's protocols. The concentration and the quality of RNA were determined using NanoVue Plus spectrophotometer (GE HealthCare Life Sciences) and 1% agarose gel electrophoresis, respectively. The isolated total RNAs were converted into cDNA using a ReverTra Ace-α-kit (Toyobo Co. Ltd), according to the manufacturer's protocols. The transcribed cDNA was diluted 20-fold with RNase/DNase-free water for further experimental analysis.
Phenylpropanoid Biosynthetic Genes Expression
For qRT-PCR, the actin gene was used as an internal control. Specific primers for the PAL, C4H, 4CL, CHS, CHI, F3H, FLS, DFR, and actin genes were designed using Gene Runner version 5 software. The PCR was performed in triplicate on a CFX Real-Time System combined with a C1000 Thermal Cycler (Bio-Rad) using a 2× Real-Time PCR Master Mix (Including SFC green® I) (BioFACT). Thermal cycling conditions were as follows: pre-denaturation at 95 °C for 15 min, 40 cycles of denaturation at 95 °C for 15 s, and annealing/extension at 57 °C for 1 min 15 s. The expression of each phenylpropanoid biosynthetic gene relative to the actin gene was calculated using the 2−ΔCt method. 39
Phenylpropanoid Extraction and HPLC Analysis
The extraction and analysis of phenylpropanoids in different organs of C. majus were carried out using a previously reported HPLC method with slight modification. 40 A 100 mg of ground sample was extracted with 1 mL of 80% aqueous MeOH (v/v), vortexed for 30 s, and sonicated for 1 h. Subsequently, the supernatant was collected and filtered through a 0.45-µm PTFE syringe filter after centrifugation at 14 810 g for 20 min at 4 °C. The phenolic compounds were analyzed by using an HPLC system (NS-4000, Futecs) connected to an OptimaPak C18 column (250 × 4.6 mm, 5 µm, RStech), a UV-Vis detector, and an autosampler (NS-6000, Futecs). The HPLC analysis conditions and gradient program were taken from a previous study. 41 Each phenylpropanoid was identified based on the spike test results comparing it with the retention time of individual standards and quantified by the corresponding calibration curves. The identified peaks were further confirmed by UHPLC-Q-Orbitrap-MS (Thermo Fisher Scientific). The mobile phase and gradient program are shown in Table S1. All the standard chemicals were purchased from Sigma-Aldrich with ≥95% purity.
Statistical Analysis
All experiments were done in triplicate and data were analyzed using IBM SPSS Statistics 26 (IBM; AsiaAnalytics), and the data are expressed as the mean ± SD of triplicate determinations. The significant differences between the organs were analyzed using a one-way analysis of variance (one-way ANOVA) followed by Tukey's honestly significant difference test (if data fulfilled homoscedasticity) or Welch's one-way ANOVA followed by Tamhane's T2 test (if data did not fulfill homoscedasticity) for multiple comparisons with an α = 0.05. Heat map, PCA, PLS-DA, VIP, and Pearson correlation analysis of all the phenolic compounds identified from different organs of C. majus were performed using MetaboAnalyst 5.0 (http://www.metaboanalyst.ca/) with autoscaling.
Supplemental Material
sj-doc-1-npx-10.1177_1934578X241239835 - Supplemental material for Expression Analysis of Phenylpropanoid Pathway Genes and Phenylpropanoid Accumulation in Different Organs of Chelidonium majus L.
Supplemental material, sj-doc-1-npx-10.1177_1934578X241239835 for Expression Analysis of Phenylpropanoid Pathway Genes and Phenylpropanoid Accumulation in Different Organs of Chelidonium majus L. by Haejin Kwon, Ramaraj Sathasivam, Jiwon Yoon, Chanung Park, Nam Il Park, Yong Suk Chung and Sang Un Park in Natural Product Communications
Footnotes
Authors Contributions
YSC and SUP designed the experiments and analyzed the data. HK, RS, JY, CP, and NIP performed the experiments and analyzed the data. HK and RS wrote the original draft of the manuscript. RS performed metabolomic analysis and edited the manuscript. All authors have read and agreed to the published version of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This research was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education, (grant number 2019R1A6A1A11052070, 2022R1I1A3054240).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
