Abstract
Keywords
Introduction
Chronic kidney disease (CKD) is a growing public health problem due to its high morbidity and mortality rates and greatly increased risk of chronic renal failure (CRF) and cardiovascular disease. 1 Recent data show that the global prevalence of CKD was estimated to be between 11% and 13% in 2016. 2 In addition, as CKD progresses, it can easily lead to cardiovascular disease. 3 Renal fibrosis, characterized by tubulointerstitial fibrosis and glomerulosclerosis, is a common pathway and inevitable consequence of the various progression of CKD to ESRD, which requires renal replacement therapy such as dialysis and transplantation.4–7 Therefore, it is of great medical and social value to study the treatment of CRF.
Nourishing Blood Diuretic Formula (NBDF) is an effective clinical experience prescription for treating CRF. It consists of 8 traditional Chinese medicines, including Abelmoschi corolla, Rhei Radix et Rhizoma, Astragali Radix, Angelicae Sinensis Radix, Poriae cutis, Dioscoreae Rhizoma, Hirudo, and Bombyx batryticatus. Clinical studies have shown that this formula can significantly improve the levels of blood creatinine and urea nitrogen in patients with renal failure, reduce urinary protein, protect kidney function, improve prognosis, and slow down the process of chronic kidney disease. However, due to the unclear substance basis of its efficacy and few literature reports on this prescription, it is necessary to further study the chemical components and blood components contained in this prescription. In this study, the chemical components and blood entry components of NBDF were analyzed by LC-MS technology, and the effective components of NBDF in the treatment of CRF were initially explored by combining network pharmacology and molecular docking technology, so as to clarify its pharmacodynamic material basis and provide theoretical basis and reference for clinical application research.
Materials and Methods
Instruments and Materials
In this study, we used U3000 Ultra High-Performance Liquid Chromatograph (Thermo Fisher Scientific Corporation); Q Exactive plus Orbitrap High-Resolution Mass Spectrometer (Thermo Fisher Scientific Corporation); ACQUITY UPLC I Class system Liquid Chromatographs (Waters Corporation); Synapt G2-Si Qtof Mass Spectrometer (Waters Corporation); Vortex-2 Genie Vortex Mixer (Scientific Industries Corporation); Model 5810R Cryogenic Centrifuge (Eppendorf Corporation, GER); Model WD-9415C Ultrasonic Cleaner (Liuyi Biotechnology Corporation); and Milli Q Advantage A10 Ultrapure Water Machine (Merck Corporation).
A. corolla (batch number: 210401), Rhei Radix et Rhizoma (batch number: 210501), B. batryticatus (batch number: 200701), and Hirudo (batch number: 210101) were purchased from Bozhou Jinshaotang Traditional Chinese Medicine Decoction Pieces Co., LD Astragali Radix (batch number: G03221006-07) and Angelicae Sinensis Radix (batch number: G01201117-21) were purchased from Gansu Jiuzhou Tianrun Traditional Chinese Medicine Industry Co., LD P. cutis (batch number: 211101) was purchased from Hubei Jurui Biotechnology Co., LD; Dioscoreae Rhizoma (batch number: D22090401) was purchased from Huanggang Jingui Traditional Chinese Medicine Industry Development Co., LTD; Heparin sodium (batch number: 003211009) was purchased from Chengdu Haitong Pharmaceutical Co., LTD; acetonitrile, these herbs were all characterization by Professor Duan Xueyun from Hubei University of Traditional Chinese Medicine (Wuhan, China), which conforms to the provisions of the 2020 edition of Chinese Pharmacopoeia. All storing samples were stored in the herbarium of the Hubei Institute of Traditional Chinese Medicine (Wuhan, China). Methanol and formic acid were chromatographically pure and purchased from Thermo Fisher Scientific; water was deionized water.
Animals
Fourteen SPF-grade healthy male SD rats, weighing 180∼220 g, were purchased from Wuhan Wanqian Jiaxing Biotechnology Co., LD, license No.: SCXK (Hubei) 2017-0013. Reared in the experimental animal center of Hubei Hospital of Traditional Chinese Medicine, experimental facility license: SYXK (Hubei) 2017-0095, room temperature: 22°C ± 3°C, relative humidity: 55%∼60%, no convection, artificial day and night (12 h day, 12 h night). The animal experiments involved in this study were approved by the experimental animal center of Hubei Provincial Hospital of Traditional Chinese Medicine, and the approval number is No. 2020003 of Hubei Provincial Hospital.
Chromatographic Conditions
The determination was performed on Waters ACQUITY UPLC HSS T3 column (2.1 × 100 mm, 1.8 µm) with a mobile phase consisting of 0.1% formic acid aqueous solution (A) −0.1% acetonitrile formate solution (B). Condition 1 (chemical composition identification) is gradient elution (0∼10 min, 0%∼30% B;10∼25 min, 30%∼40% B; 25∼30 min, 40%∼50% B; 30∼40 min, 50%∼70% B; 40∼45 min, 70%∼100% B; 45 min∼60 min, 100% B; 60 min∼60.5 min, 100%∼0% B; 60.5 min∼70 min, 0% B). Condition 2 (identification of blood components) is gradient elution (0∼15 min, 0%∼20% B; 15∼50 min, 20%∼100% B; 50∼60 min, 100% B; 60 min to 70 min, 100% to 0% B). The flow rate was 0.3 mL/min, the sample size was 10 μL, and the column temperature was 35°C.
Mass Spectrometry Conditions
Condition 1 (chemical composition identification) is electrospray ion source, the detection mode was Full MS-dd MS2, the positive and negative ion modes were scanned respectively, the scanning range was m/z 100-1200, the resolution of MS1 was set to 70,000, the resolution of MS2 was set to 17,500, the voltage of ion source was set to 3.2 kv, and the temperature of ion transmission tube was set to 320°C. The heating temperature of the auxiliary device was 350°C, the sheath gas flow rate was 40 L/min, and the auxiliary gas flow rate was 15 L/min. The collision energy triggering the MS2 scan was set to 30, 40, and 50 eV using the stepped fracture voltage. Condition 2 (identification of blood components): an electrospray ion source, positive and negative ion mode scanning, the mass range is 50-1500 Da, capillary voltage is 3000 V (ESI+), 2500 V (ESI−), cone hole voltage is 40 V, collision energy is 10-50 eV, and ion source temperature is 125°C.
Preparation of NBDF
According to the prescription ratio of NBDF, weigh 8 herbs such as A. corolla, Rhei Radix et Rhizoma, Astragali Radix, Angelicae Sinensis Radix, P. cutis, Dioscoreae Rhizoma, Hirudo, and B. batryticatus, add 10 times the amount of water, soak for 30 min, boil for 1 h, filter, repeat 3 times, combine the filtrate, leave overnight, concentrate the superserum, and store in the refrigerator at 4°C for later use.
Carefully remove 2 mL of NBDF solution and place it in a 15 mL centrifuge tube, add 10 mL of 50% methanol aqueous solution, ultrasound for 30 min, take 1 mL of supernatant and place it in a centrifuge tube, centrifuge at 14,000 r/min for 5 min, take the supernatant through a 0.22 μm microporous filter membrane, and put it into the sample bottle. To be analyzed by UPLC-Q-Exactive Plus-Orbitrap-MS.
Collection and Preparation of Drug-Containing Plasma
After 1 week of adaptive feeding, 14 male SD rats were randomly divided into a blank group and a drug administration group, with 7 rats in each group. The blank group was given normal saline by intragastric administration every day, and the drug administration group was given NBDF solution by intragastric administration at the dose of 12.3 g/kg per day (referring to the equivalent dose conversion method, the dose given to rats was about 6.25 times of that given to adults). After the last administration, the rats were fasted without water for 12 h. On the 8th day, blood was taken from the orbital venous plexus of the rats 15 min, 30 min, 1 h, 2 h, and 4 h after gavage, about 0.5 mL of blood was collected each time and placed into a centrifugal tube coated with heparin sodium solution. Centrifuge at 3000 r/min at 4°C for 15 min, take the supernatant and then obtain the plasma sample and store it in the refrigerator at −80°C. The plasma samples were removed from the refrigerator at −80°C and thawed on ice. The plasma samples taken from each group at various time points were mixed well, accurately absorbed 150 μL, placed in a centrifugal tube, methanol was added according to the volume ratio of 1:4, swirled for 1 min, ultrasounded in an ice bath for 15 min, and then left for 1 h in the refrigerator at 4°C. Centrifuge at 3000 r/min for 10 min, take the supernatant, dry it with nitrogen at 37°C, redissolve the residue with methanol 400 μL, swirl for 1 min, ultrasound in an ice bath for 15 min, stand in the refrigerator at 4°C for 1 h, centrifuge at 12 000 r/min for 5 min, take 200 μL of the supernatant and put it into the sample bottle. To be analyzed by UPLC-Q-TOF-MS. The blank plasma samples were treated under the same conditions.8–10
Data Processing
Compound Discover 3.2 software was used to extract characteristic peaks from raw mass spectrometry data, and the mass deviations of characteristic peak element matching, molecular formula prediction and isotope distribution matching were all set to less than 5 ppm. Characteristic peaks were identified using the mzcloud online database and mzVault database. The screening criteria for positive results were mass deviation <5 ppm, consistent isotope distribution, and matching score >70 points in mzVault best match database. The compounds were identified by referring to relevant literature and databases (Pub Chem, Mass Bank, Sci Finder, etc) and comparing them with the mass spectrum information of the screened target compounds.
Prediction of Prototype Component Targets and Disease Targets
The prototype component compounds were imported into SwissTargetPrediction (http://www.swisstargetprediction.ch/) in SMILES format to obtain the relevant targets, and the targets related to chronic renal failure disease were obtained through GeneCards (https://www.genecards.org/), and the blood component targets of NBDF as well as the disease targets of CRF were imported into Venny software to take the intersection of the intersection of the intersecting targets, which is the intersecting target of the NBDF for the treatment of CRF.
Construction of the PPI Network
The intersection targets were imported into the String (https://cn.string-db.org/) database to predict PPI, the species was set to human. The results were output in TSV format and imported into Cytoscape 3.8.2 software, and using the “cytoHubba” plugin in the Cytoscape 3.8.2 software to calculate the degree of interaction between nodes degree value, all nodes of this PPI network are presented according to the size of the degree value.
GO/KEGG Enrichment Analysis
GO includes the analysis of 3 modules: biological processes, cellular components, and molecular functions of targets, which is currently the most representative tool for describing systems biology. KEGG is a systematic database that analyzes gene functions and links genomic and functional information, which can be used not only for the discovery of differentially expressed genes but also for the discovery of network relationships and mechanisms between different biological processes. We imported the key targets into the DAVID (https://david.ncifcrf.gov/) database for analysis, with the species set to human and the P-value set to less than .05.
Molecular Docking
The 3D structures of the compounds of the prototypical components of NBDF were downloaded from PubChem (https://pubchem.ncbi.nlm.nih.gov/) and the compounds were imported into ChemDraw 20.0 for energy minimization, hydrogenation, and electric charge. The 3D structure of the screened target protein was downloaded from the PDB (http://www.rcsb.org) and imported into PyMOL 2.3.0 for the removal of nonprotein molecules from the ligand. Finally, the processed receptor and ligand were imported into AutoDockTools 1.5.7 for docking to obtain the binding energy between the ligand and the receptor, and the 6 sets of results with the best binding were plotted using PyMOL 2.3.0.
Results
Chemical Composition of NBDF
UPLC-Q-Exactive Plus-Orbitrap-MS technology was used to analyze the chemical composition of the NBDF solution. The total ion flow diagram of NBDF in positive and negative ion modes is shown in Figure 1(A) and (B). A total of 125 compounds were identified, including 39 flavonoids, 9 anthraquinones, 9 terpenoids, 16 organic acids, 22 phenylpropanoids, 4 polyphenols, 4 aldehydes, 4 alkaloids, 4 carbohydrates, 3 amino acids, and 3 phthalides. There were 2 saponins and 6 other compounds, of which hyperoside is the main component of A. corolla, emodin is the main component of Rhei Radix et Rhizoma, formononetin is a representative component of Astragali Radix, poricoic acid A is the important component of P. cutis, ferulic acid and caffeic acid are the basic ingredients of Angelicae Sinensis Radix, and the amino acids and carbohydrates are the main components of Dioscoreae Rhizoma, B. batryticatus, and Hirudo. The detailed results are shown in Table 1.

Total ion flow diagram of NBDF solution in positive ion mode (A). Total ion flow diagram of NBDF solution in negative ion mode (B).
NBDF chemical composition information table.
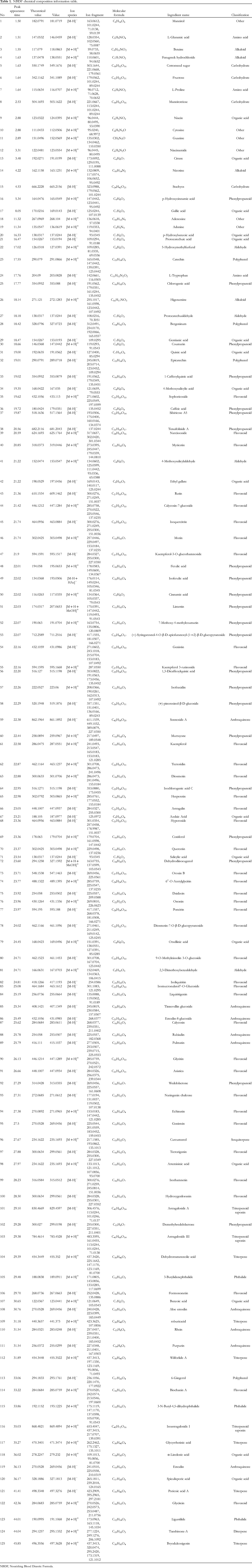
NBDF, Nourishing Blood Diuretic Formula.
Flavonoids
A total of 39 flavonoid compounds were identified in this study, which generally refers to a series of compounds made of 2 benzene rings interconnected by 3 carbon atoms, ie, a general term for a class of compounds with the C6-C3-C6 structure, which often lose neutral CH3, H2O, CO, CO2, etc, plasma fragments to form fragment ions. Taking compound No. 68 as an example, its quasi-molecular ion peak was m/z 463.0884[M-H]− with a retention time of 23.36 min, and its molecular formula was presumed to be C21H20O12. Fragmentation ions such as m/z 301.0354[M-H-C6H10O5] appeared in the secondary mass spectrum, and based on the cleavage pattern and the literature and database matching, it was hypothesized that the compound was hyperoside, an active ingredient in the A. corolla, which is hepatoprotective, cardiovascular and cerebrovascular protective, neuroprotective, antiinflammatory, analgesic, hypolipidemic, and enhances immune functions.11,12 The cleavage process is shown in Figure 2A, and the fragment ion map is shown in Figure 3A.

Cleavage pathway of hypericin (A), protocatechuic acid (B), caffeic acid (C), poricoic acid A (D), 6-gingerol (E), and emodin (F).

Fragment ions of hypericin (A), protocatechuic acid (B), caffeic acid (C), poricoic acid A (D), 6-gingerol (E), and emodin (F).
Organic Acid Compounds
A total of 16 organic acid compounds were identified in this study, which contain carboxyl groups in their molecular structures, mostly phenol carboxyl-substituted aromatic rings, fatty acids, etc. According to their structural characteristics, they can be classified into aromatic, aliphatic and terpenoid organic acids. Taking compound No. 21 as an example, its quasi-molecular ion peak was 153.0194 [M-H]− with a retention time of 16.47 min, and its molecular formula was presumed to be C7H6O4, and the secondary mass spectra yielded the characteristic fragment ions, such as m/z 109.0294 [M-H-CO2]−, m/z 91.0188 [M-H-CO2-H2O]−, etc, and based on the cleavage pattern and the literature and database matching, it was hypothesized that the compound was protocatechuic acid. The cleavage process is shown in Figure 2B, and the fragment ion map is shown in Figure 3B.
Phenylpropanoids
A total of 22 phenylpropanoid compounds were identified in this study, including caffeic acid, ferulic acid, chlorogenic acid, cinnamic acid, coumarin, and wedelolactone, most of which were coumarins. Taking compound No. 36 as an example, its quasi-molecular ion peak was 179.0351[M-H]− with a retention time of 19.72 min, and its molecular formula was presumed to be C9H8O4. The secondary mass spectra showed m/z 135.0452[M-H-CO2]− fragment ions, based on the cleavage pattern and the literature and database matching, the compound was hypothesized to be caffeic acid, the active ingredient in Angelicae Sinensis Radix, which is analgesic, antiinflammatory, antioxidant, microcirculation improvement, and antiplatelet aggregation effects. 13 The cleavage process is shown in Figure 2C, and the fragment ion map is shown in Figure 3C.
Terpenoids
A total of 9 terpenoids were identified in this study, most of which were triterpenoids, including oak cherry acid, poriaxinic acid A, and glycyrrhizinic acid. Taking compound No. 121 as an example, its quasi-molecular ion peak was 497.3277[M-H]−, with a retention time of 41.41 min, and its molecular formula was presumed to be C31H46O5. m/z 423.0909 [M-H-C3H6O2]−, m/z 395.2965 [M-H-C2H4]−, m/z 297.2181 [M-H-C7H14]− and other fragment ions, based on the cleavage pattern matched with the literature and database, it is presumed that the compound is poricoic acid A, an active ingredient in P. cutis, which has antiinflammatory, antioxidant, antitumor, hypolipidemic, and diuretic effects.14–16 The process of cleavage is shown in Figure 2D, and the fragment ion map is shown in Figure 3D.
Polyphenols
A total of 4 polyphenolic compounds were identified in this study. Taking compound No. 113 as an example, its quasi-molecular ion peak was 293.1761[M-H]− with a retention time of 33.06 min, and its molecular formula was presumed to be C17H26O4. m/z 236.1056[M-H-C4H9]−, m/z 221.1544[M-H-C4H9-CH3]−, m/z 177.0922 [M-H-C7H15O]− and other fragment ions, and the compound was presumed to be 6-gingerolby matching the cleavage pattern with the literature and the database. The cleavage process is shown in Figure 2E, and the fragment ion map is shown in Figure 3E.
Anthraquinones
A total of 9 anthraquinones were identified in this study. Taking compound No.119 as an example, its quasi-molecular ion peak was 269.0456[M-H]− with a retention time of 36.13 min, and its molecular formula was presumed to be C15H10O5. In the secondary mass spectra, there were fragment ions, such as m/z 241.0510[M-H-CO]−, m/z 225.0558[M-H-CO2]−, and m/z 210.0319 [M-H-CO2-CH3]− and other fragment ions. By matching the cleavage pattern with the literature and the database, it was hypothesized that the compound was emodin, the active ingredient in Rhei Radix et Rhizoma, which has antiinflammatory, antiviral, antitumor, and other pharmacological effects. 17 The cleavage process is shown in Figure 2F, and the fragment ion map is shown in Figure 3F.
In this study, 4 aldehydes, 4 alkaloids, 4 carbohydrates, 3 amino acids, 3 phthalides, 2 saponins, and 6 other compounds were also identified, which were analyzed and identified in the same way as the abovementioned compounds, and no more examples will be given.
Blood Entry Components of NBDF
Blank plasma from rats and plasma after gavage of NBDF solution were analyzed and identified by UPLC-Q-TOF-MS, and their total ion flow diagrams in positive and negative ion modes are shown in Figure 4(A)-(D). A total of 48 compounds were identified, among which 6 prototypical components were identified, including formononetin, emodin, epicatechin, chlorogenic acid, sennoside A, and astragaloside III. The detailed information is shown in Table 2.

Total ion flow diagram of a blank plasma sample in positive ion mode (A). Total ion flow diagram of a blank plasma sample in negative ion mode (B). Total ion flow diagram of plasma sample after administration in positive ion mode (C). Total ion flow diagram of plasma sample after administration in negative ion mode (D).
Information table of blood component of NBDF.
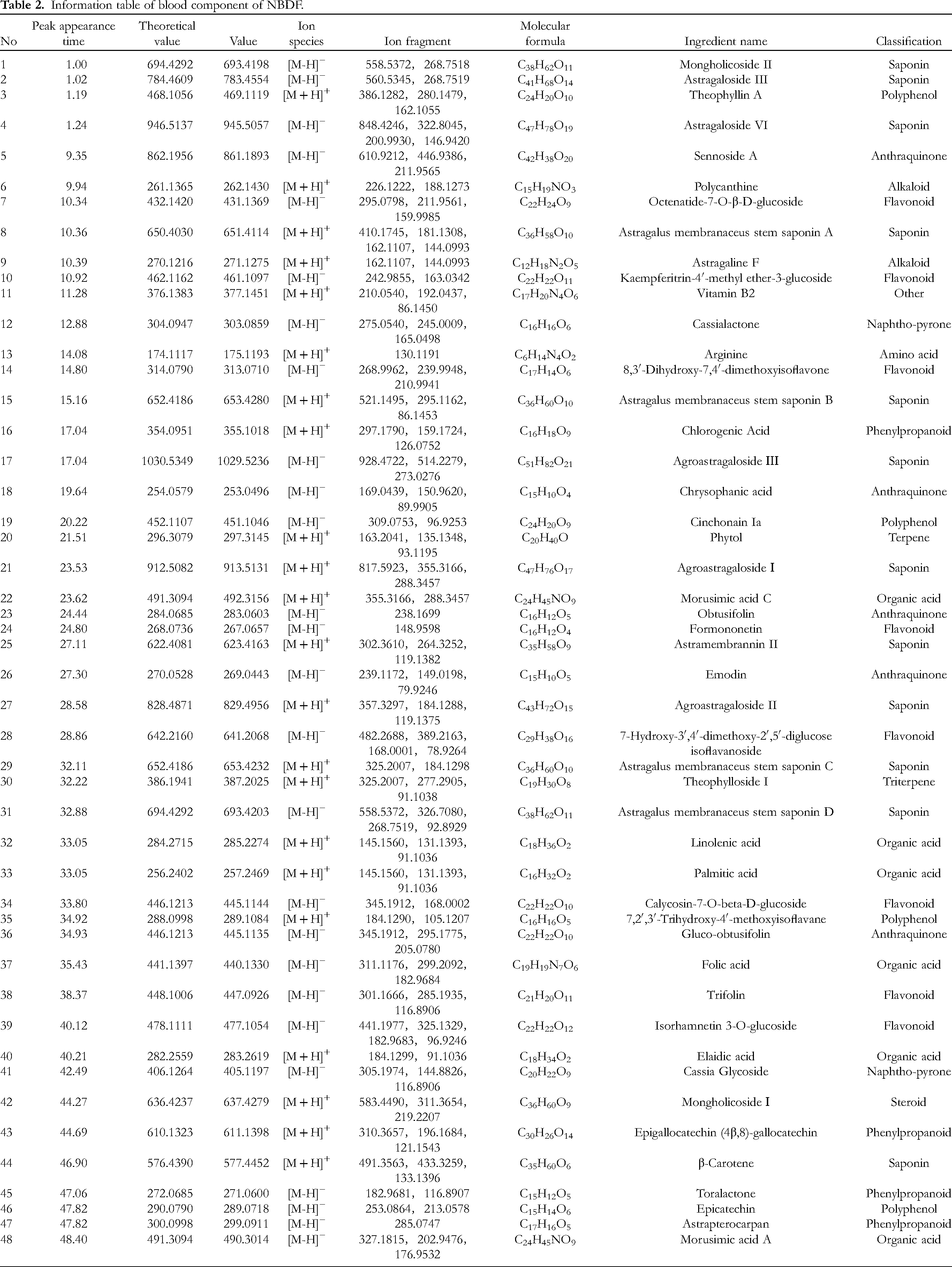
A total of 6 prototypical components were identified in this study, and compounds No. 16 and No. 24 were analyzed as examples. The quasi-molecular ion peak of compound No. 16 was 355.1018[M + H]+ with a retention time of 17.04 min, and the molecular formula was presumed to be C16H18O9, and the secondary fragment ions were m/z 297.1790[M-H-C2H2O2]+, m/z 159.1724[M-H-C7H16O6]+, m/z 126.0752[M-H-C9H9O7]+, according to the cleavage pattern and related literature, the compound was presumed to be chlorogenic acid.18–20 The cleavage process is shown in Figure 5A, and the fragment ion map is shown in Figure 6A.

Cleavage pathway of chlorogenic acid (A) and formononetin (B).

Fragment ions of chlorogenic acid (A) and formononetin (B).
The quasi-molecular ion peak of compound No. 24 was 267.0654 [M-H]− with a retention time of 24.80 min, and the molecular formula was presumed to be C16H12O4, and the secondary fragment ions was m/z 148.9598[M-H-C8H5O]−, based on the cleavage pattern and related literature, it was hypothesized that the compound is formononetin, the main component of Astragali Radix, which has antibacterial, cholesterol-lowering and hypolipidemic effects.21,22 The cleavage process is shown in Figure 5B, and the fragment ion map is shown in Figure 6B.
Screening of Prototype Component Targets and Disease Targets
Three hundred and eleven prototype component targets were obtained from the SwissTargetPrediction database, 1054 disease targets related to CRF were obtained from the GenCards database, and 102 potential targets of NBDF related to the treatment of CRF were identified by importing the prototype component targets and disease targets into Venny software to take the intersections for making a Venn diagram, as shown in Figure 7A.
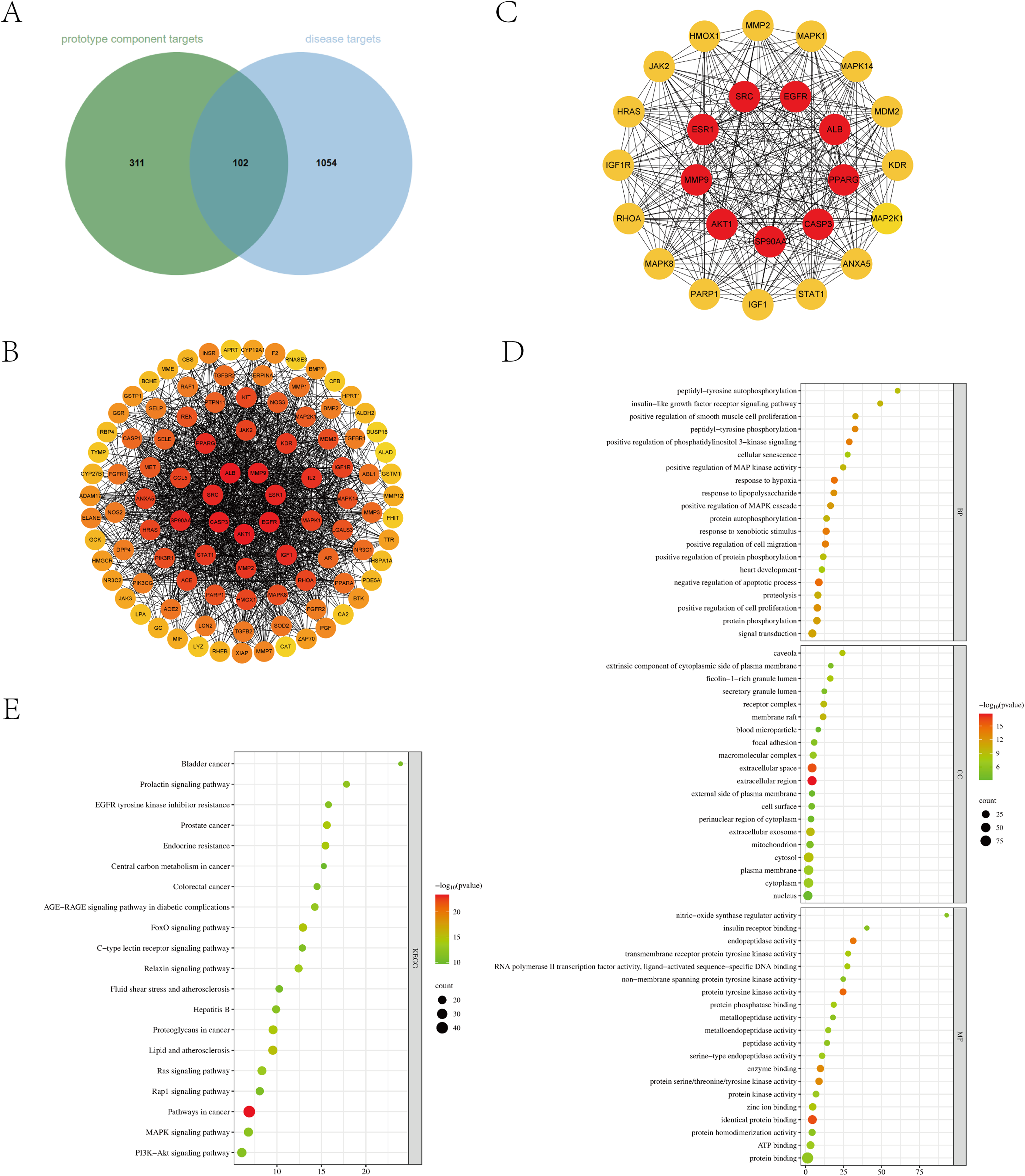
Venn diagram of prototype component targets and disease targets (A). The PPI network diagram (B-C). Bubble diagram of GO enrichment analysis (D). Bubble diagram of KEGG pathway enrichment analysis (E).
PPI Network Construction
The 102 potential targets were imported into Cytoscape 3.8.2 to construct a PPI network, and the 25 core targets with the highest degree values in the network were obtained using the plug-in Mcode in Cytoscape 3.8.2, and by comparing and analyzing these 25 core targets, we found that targets such as ALB, AKT1, and EGFR occupied a central position in the PPI network, as shown in Figure 7(B)-(C), and thus we predicted that NBDF might affect protein expression by regulating targets such as ALB, AKT1, and EGFR.
GO/KEGG Enrichment Analysis
GO enrichment analysis yielded 515 GO entries, sorted by P-value in ascending order, including 379 biological process (BP) entries, 48 cellular component (CC) entries, 88 molecular function (MF) entries, and the top 20 entries were selected for annotation analysis. BP processes such as negative regulation of apoptotic process, positive regulation of cell migration, protein phosphorylation, and signal transduction. CC processes such as extracellular space and extracellular region. MF processes such as enzyme binding and protein kinase/trans-kinase activity.
KEGG enrichment analysis showed that 133 pathways were significant, and annotation analysis was performed for the top 20 pathways, which were associated with cancer, endocrinology, glucose-lipid metabolism, cellular proliferation and differentiation, and inflammatory response, as shown in Figure 7(D) and (E), such as lipid and atherosclerosis, FoxO signaling pathway, Ras signaling pathway, and Prolactin signaling pathway.
Molecular Docking
Based on the analysis results of the PPI network, nine targets with high degree composite values: ALB, ESR1, EGFR, CASP3, AKT1, HSP90AA1, MMP9, PPARG, and SRC, as well as six prototypical constituents: formononetin, emodin, epicatechin, chlorogenic acid, sennoside A, and astragaloside III were selected and molecularly docked by the AutoDockTools 1.5.7 was used for molecular docking to obtain the binding energy between the ligand and the receptor, as shown in Figure 8, the lower the binding energy between the ligand and the receptor indicates the more solid the intermolecular binding, the higher the possibility of interaction, and the binding is better when the binding energy is ≤−5.0 kcal/mol. The docking results of the six groups with the lowest binding energy are shown in Figure 9(A)-(F).

Heat map of binding energy of composition and target docking.

Docking results of ESR1 with emodin (A), ESR1 with formononetin (B), HSP90AA1 with formononetin (C), MMP9 with emodin (D), MMP9 with formononetin (E), and PPARG with formononetin (F).
Discussion
The composition of Chinese medicine compounds is complex, and the material basis of its efficacy is difficult to be clarified. Oral Chinese medicine prescriptions exert their medicinal effects by entering into the bloodstream, so we can elucidate the material basis of the medicinal effects of Chinese medicine prescriptions by studying the components that enter into the bloodstream.
In this study, the solution of NBDF and its components before and after blood entry were analyzed using LMS technology. The samples were prepared by mixing equal volumes of rat plasma samples collected at different time points, which was able to maximize the reflection of the components of the blood entry after the administration of the compound formula. Combining the effective information of compound mass number, retention time, molecular ion peaks, fragment ions, etc, a total of 125 chemical components in the compound formula and 48 blood-entry components were identified through the analysis of cleavage pattern and database comparison. Among them, there were 6 blood-entry prototypical components, including formononetin, emodin, epicatechin, chlorogenic acid, sennoside A, and astragaloside III. Formononetin is an isoflavone phytoestrogen of Astragali Radix, which has a variety of biological activities, such as antioxidant, antiinflammatory, antitumor, lowering blood pressure, and improving cardiovascular health23–28; emodin is a natural compound taken from rhubarb, thuja and other herbs, with antiinflammatory, antiviral, antitumor, analgesic, and other pharmacological effects29,30; astragaloside has a preventive and curative effect on nephritis, so that the renal pathology is reduced, improve renal function, but also regulate the immune function of the muscles and antiaging 31 ; epicatechin has an antioxidant, lipid-lowering, antiinflammatory and antibacterial effects, etc, hypoglycemic, antiinflammatory, and antibacterial effects.32,33 These components were basically consistent with the pharmacological effects of the NBDF, so it was hypothesized that they might be the pharmacological substances of the NBDF. Through network pharmacology, we obtained the common targets of components and diseases, and constructed protein interaction network maps for these targets to obtain the potential targets of NBDF for the treatment of CRF, and after the network topology analysis, the targets with high degree values included ALB, ESR1, EGFR, CASP3, AKT1, HSP90AA1, MMP9, PPARG, and SRC, indicating that these targets are closely related to the treatment of CRF with NBDF.
AKT1, an isoform of the AKT signaling pathway, is a serine/threonine protein kinase that participates in a variety of biological processes, including cell growth, metabolism, proliferation, and insulin signaling, mediated by phosphorylation, and whose activation is dependent on P13 K signaling. Studies have confirmed that AKT1 is activated during the progression of acute kidney injury (AKI) to CKD, and the absence of AKT1 attenuates renal interstitial fibrosis as well as apoptosis during the progression of AKI to CKD, and AKT1 can be used as a potential therapeutic target to inhibit the transition from AKI to CKD. 34 CASP3 disrupts cell structure and acts on regulatory proteins FAK, PAKα, etc, causing them to lose their functions and play an important role in the process of apoptosis. EGFR can be expressed in a variety of renal cells, including endothelial cells, tethered cells, etc. It can be involved in renal fibrosis, renal interstitial as well as glomerular injury, and plays an important role in many acute and chronic kidney injuries. Its mediated overexpression of HIPK2 is thought to be a key regulatory mechanism in the transition from vancomycin-induced AKI to CKD. Increasing evidence suggests that EGFR activation is associated with a variety of physiological and pathophysiological functions and that aberrant EGFR activation is a mediator of progressive kidney injury in diabetic nephropathy.35–37
MMP9 is a member of the matrix metalloproteinase family that provides homeostasis between synthesis and degradation of extracellular matrix to maintain the structural and functional integrity of the glomerulus. It is widely expressed in renal tissues and is involved in the development of renal inflammatory responses and glomerular fibrosis. Renal tubular epithelial cells and macrophages can lead to the production of MMP9. Studies have confirmed that MMP9 is significantly elevated in urine, serum, and renal tissues of DKD patients.38–40 It has been shown that MMP9 is a key pathological mediator of renal fibrosis in diabetic rats and that MMP9 deficiency is beneficial in preventing renal fibrosis. 41 The molecular docking results showed that formononetin and MMP9 had the strongest binding ability and the lowest binding energy of −9.93 kcal/mol. Therefore, we speculate that the mechanism of NBDF for the treatment of CRF may be related to the binding of formononetin and MMP9, and through the inhibition of MMP9, thus affecting the synthesis and degradation of renal ECM, and ultimately inhibiting the formation of renal fibrosis.
In summary, using two techniques, this study predicted that the NBDF might act on the targets of ALB, ESR1, EGFR, CASP3, AKT1, HSP90AA1, MMP9, PPARG, and SRC through the key components, such as formononetin, emodin, epicatechin, chlorogenic acid, sennoside A, and astragaloside III, by reducing the inflammatory response and exerting therapeutic effects on CRF by reducing the inflammatory response. In this study, fewer prototypical components of NBDF were detected in the plasma of rats, which may be attributed to the faster metabolism of the components in the blood and the production of other substances. Therefore, the efficacy of NBDF needs to be further explored and explored. In addition, this study only explored the blood-entry components of NBDF and predicted the target and therapeutic mechanism of this formula through network pharmacology, but it was verified through experiments, and we will investigate the efficacy of different drug concentrations on chronic renal failure disease through cellular experiments and animal pharmacodynamic experiments, and investigate the mechanism of the NBDF in the treatment of CRF disease through the application of Western blotting.
Conclusion
In the present study, we analyzed the chemical composition and blood components of NBDF by LC-MS technology, and preliminarily predicted the efficacy of NBDF in the treatment of CRF by combining with the network pharmacology and molecular docking technology, so as to elucidate its efficacy material basis and therapeutic mechanism.
Footnotes
Authors’ Notes
Si-Cheng Yang, Xia Lei, and Si-Fang Xie have contributed equally to this work.
Author Contributions
Si-Cheng Yang, Xia Lei, and Si-Fang Xie: methodology, investigation, and formal analysis; Ju Huang, Feng-Qin Yue, Si-Di Chen, and Wei Peng: project administration and data curation; Si-Cheng Yang and Xia Lei: Writing-original draft; Heng Fan, Sen Li, Xue-Yun Duan, and Wan-Jin Sun: writing-review and editing.
Data Availability
Data will be made available on request.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
This study was approved by the Experimental Animal Committee of Hubei Provincial Hospital of Traditional Chinese Medicine.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the State Administration of Traditional Chinese Medicine of the People's Republic of China (Grant Nos: [2022] No. 76, [2023] No. 96).
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Statement of Human and Animal Rights
The animal experimental procedures involved in this study were conducted in accordance with the guidelines for animal experiments of the Hubei Provincial Hospital of Traditional Chinese Medicine Laboratory Animal Center and approved by the Hubei Provincial Hospital of Traditional Chinese Medicine Laboratory Animal Committee.
