Abstract
Objective
Actinomycetes have been known to be the great natural sources to explore antibiotics for the treatment of tuberculosis (TB). The isolation of actinomycetes from the samples in Vietnam followed by the screening of their antimycobacterial activity was performed in this study. The metabolites isolated from the most active strain were further evaluated for their antimycobacterial, antimicrobial and cytotoxic activity.
Methods
Actinomycetes were growth in culture media, isolated and identified by colony, spore chain morphology and 16S rRNA gene sequencing. Agar diffusion assay was used for the screening of the isolated strains against Mycobacterium smegmatis, a safety surrogate for Mycobacterium tuberculosis. The metabolites produced from the most active strain were investigated by actinomycete fermentation, extraction and isolation from biomass and cultures. The structures of the isolated compound were elucidated by spectral data and comparison with the reported literatures.
Results
181 strains were isolated from nine regions along the north to central Vietnam. The five most active strains against Mycobacterium smegmatis were detected. Following the bioassay-guided result, the strain A121 (Streptomyces alboniger) was selected for further isolation of the bioactive metabolites. As a result, obscurolide B2β (
Conclusions
Five strains possessing potential antimycobacterial activity were identified from the samples collected in Vietnam. Two compounds including obscurolide B2β (
Introduction
Tuberculosis (TB), one of the most global health burdens, is an infectious disease caused by Mycobacterium tuberculosis. According to the World Health Organization (WHO), TB has become the thirteen leading cause of death worldwide, with 1.6 million deaths and 10.6 million new infectious cases in 2021. 1 The diagnosis and treatment for TB patients were interrupted during the Covid 19 epidemic, leading to the increasing numbers of TB death and transmission to the community. 1 WHO reported that in 2021 Vietnam ranked 11th out of 30 countries with the highest burden of tuberculosis and multidrug-resistant TB in the world. Each year, there are approximately 170 000 new cases and about 10 400 deaths of TB in Vietnam. 2 The prevalence of drug-resistant TB has increased considerably, with approximately 8400 new cases of rifampicin-resistant (first-line drug) and multidrug-resistant tuberculosis each year.3–5 The cost for treatment of multidrug-resistant tuberculosis is ten times more expensive than that of common TB, causing a heavy burden on TB patients. 3 Therefore, drug-resistant TB is currently considered as a serious public health problem in Vietnam. Therapeutic treatments for multidrug-resistant tuberculosis (MDR) have been extensively developed by researchers and pharmaceutical companies. However, the spread of drug-resistant TB is still the major challenge for disease control. Hence, finding new antibiotics and more effective drugs are urgently needed to overcome this problem.
Actinomycetes are known to be the largest group of microorganisms for production of antibiotics. 6 Streptomycin, 7 kanamycin, 7 azomycin, 8 rifamycin, 9 etc. are the antituberculosis drugs derived from a great number of actinomycetes species. It was proven that the secondary metabolites produced from the actinomycetes that survived in extreme or unique environment may present a wide range of antimicrobial activity. 6 Vietnam has a great diversity of natural ecosystem including tropical ever-green forest, mangrove forest, coastal island, high mountains, beach, etc. which are the potential resources to discover actinomycetes diversity and natural antibiotics. However, up to now, there are only few reports on the antituberculosis metabolites from actinomycetes in Vietnam. 10
As a part of our ongoing research on antituberculosis agents from actinomycetes, we have collected the samples from various regions along the North to Central Vietnam. The cultivation of actinomycetes from different ecological habitats led to the isolation of 181 strains, which were subsequently screened for their antimycobacterial activity against Mycobacterium smegmatis. This bacteria was used as a surrogate for Mycobacterium tuberculosis in our study, due to its safety, non-pathogenic and eco-friendly properties.11–13 Unlike other Mycobacterium species, M. smegmatis grows rapidly in cell culture laboratories, and does not live in mammals.11–13 Thus, this organism does not cause infection in humans.11,12 It is noteworthy that M. smegmatis contains substantial genome sequences sharing the similarities to M. tuberculosis.11,12 Therefore, M. smegmatis has been extensively served for antimycobacterial activity assay.14–21
As the result, the five most active strains were detected including Streptomyces avidinii (A002), Streptomyces spiroverticillatus (A067), Streptomyces wuyanensis (A079), Streptomyces hawaiiensis (A105) and Streptomyces alboniger (A121). We reported herein for the first time the antimycobacterial properties of the above strains against M. smegmatis. From them, the most active one Streptomyces alboniger was fermented in large scale for further investigation of antimicrobial metabolites. Consequently, obscurolide B2β (
Results and Discussion
Isolation of Actinomycetes
As shown in Table 1, all the samples were collected in May 2020, in the following regions: Area 1: Mangrove forest in Phu Long commune, Cat Hai district, Hai Phong city; Area 2: Cat Ba National Park, Cat Hai district, Hai Phong; Area 3: On the top of May Bac mountain, Cuc Phuong National Park, Nho Quan district, Ninh Binh province (648 m above sea level); Area 4: On the way to the top of May Bac mountain, Cuc Phuong National Park, Nho Quan district, Ninh Binh province; Area 5: The confluence of Day river and Vac river, Thuong Kiem commune, Kim Son district, Ninh Binh province; Area 6: Mangrove forest in Kim Son district, Ninh Bình province; Area 7: Quynh Luong Mangrove Forest, Quynh Luong commune, Quynh Luu district, Nghe An province; Area 8: Quan Son lake, My Duc district, Ha Noi. Area 9: Yen Stream, Huong Pagoda, My Duc district, Ha Noi. A total of 181 actinomycete strains were recognized based on their differences in colony characteristics such as color, colony size, filamentous structure, mycelium growth, and pigment secretion (Figure 1). Preliminary assessment of actinomycete morphology showed that of 181 total strains there were 135 strains belonging to the genus Streptomyces (74.6%), while the rest one (46 strains) were rare actinomycetes (25.4%). It was observed that the rare actinomycetes were mostly found in the areas 1 and 2, where the mud and decaying leaf samples gave a significant actinomycetes population than those in the soil.

Colony characteristics of some actinomycetes isolated from different locations.
The Strain Codes, Natural Source and Numbers of the Strains Isolated from Different Areas Along the Northern to Central Vietnam.

Antimycobacterial actinomycetes Against Mycobacterium Smegmatis
Agar plate assay was used for the screening of actinomycete strains against Mycobacterium smegmatis. The evaluation result showed that of 181 actinomycete strains, 14 strains (7.7%) were able to inhibit M. smegmatis. Of which, the most active actinomycete strains against M. smegmatis were A002 (from area 1), A067 (from area 5), A079 (from area 7), A105 (from area 4), and A121 (from area 3) where the ecological environment has been rarely explored by humans. The location, sampling coordination (latitude and longitude), colony color, spore and inhibition zone diameter (mm) of the strains A002, A067, A079, A105 and A121 were illustrated in Table 2. The strains A002, A067 and A079 showed an equivalent activity with an inhibition zone of 9 mm diameter, while the strains A105 was active against M. smegmatis with 8 mm of inhibited zone. The strain A121 was the most active one with an inhibition zone of 11 mm in diameter (Figure 2). Compared with strain A105 whose sample collecting location was also in Cuc Phuong National Park, but in the lower position (on the way to the top of May Bac mountain), the activity of strain A121 was much higher. This may be due to the reason that strain A121 was isolated from a less explored region (on the top of a high mountain) where the ecological environment has not yet been much affected by the activity of humans. Previous literatures reported that many actinomycetes discovered from poorly explored habitats or unusual environments are the sources for production of potential active metabolites. 22 Living in such regions may allow the actinomycetes to produce large genomes with many transcription factors that are possible to respond to the particular environments.23,24 Therefore, the secondary metabolites produced by these organism as well as those biological activity could potentially be increased. 25
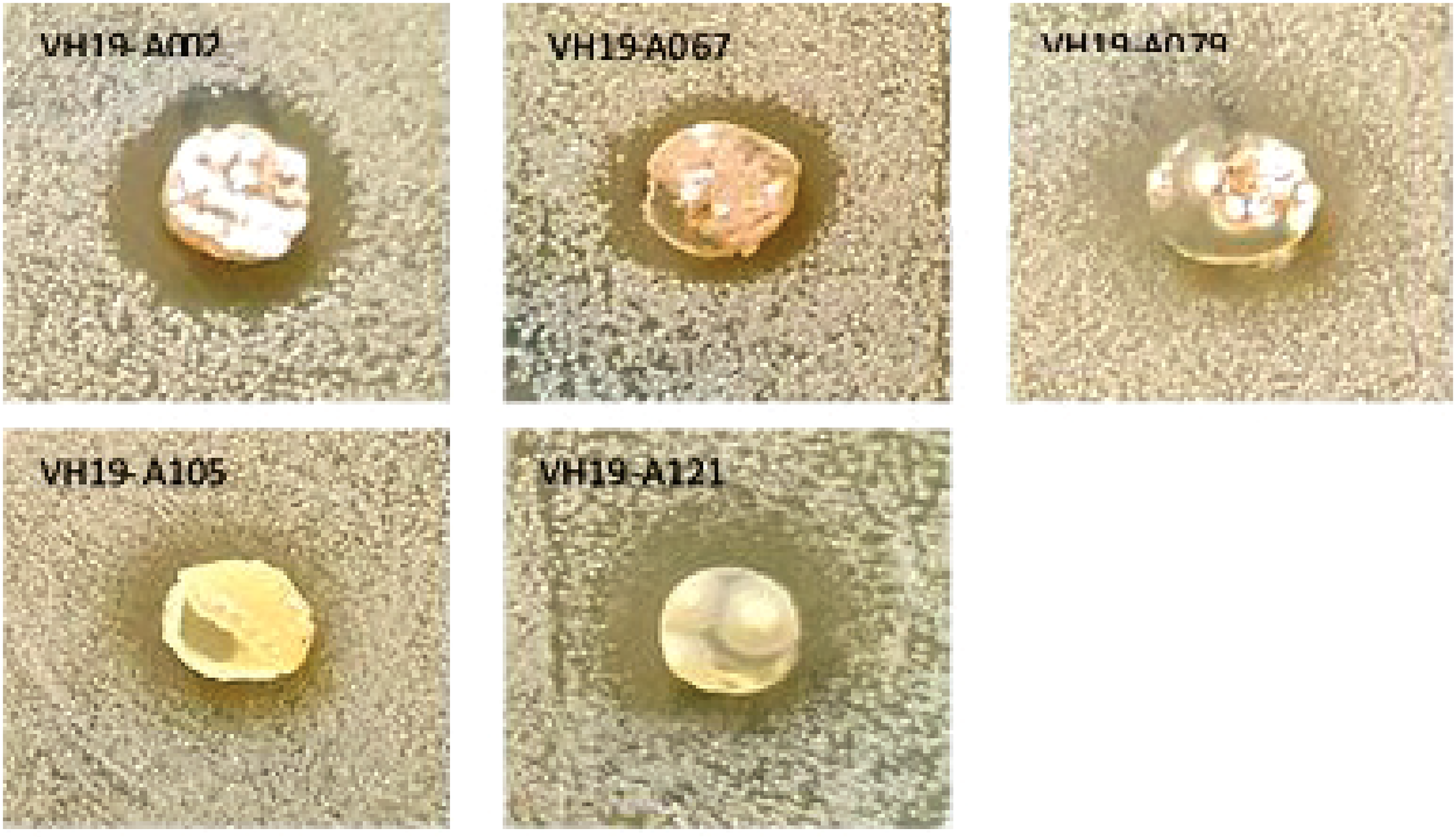

Antimicrobial activity illustration of the five most active strains (A002, A067, A079, A105, A121) against M. smegmatis.
Location for Sampling Sites, Colony, Spore Morphology and Inhibition Zone Diameter of the Five Most Active Strains Against M. smegmatis.

Classification of the Five Most Active Strains
The most active strains (A002, A067, A079, A105, A121) against M. smegmatis were classified based on their cultural characteristic, the color of the substrate mycelia, the colony morphology, the ability to produce pigment, the morphology of the spore chain and 16S rDNA sequences. By analysis of the above data, the actinomycetes A002, A067, A079, A105 and A121 were identified as Streptomyces avidinii, Streptomyces spiroverticillatus, Streptomyces wuyanensis, Streptomyces hawaiiensis and Streptomyces alboniger, respectively (Figure 3, Table 2). All of them belong to the genus Streptomyces, which have been known as promising sources for exploration of antituberculosis drugs. Of the five antimycobacterial strains above, Streptomyces wuyanensis (A079) was first discovered in Wuyan province, China in 2013 26 and has not yet been investigated on biological activity and metabolites up to now. This is the first time Streptomyces wuyanensis has been investigated for its antimycobacterial activity against M. smegmatis. Streptomyces avidinii (A002), Streptomyces spiroverticillatus (A067), Streptomyces hawaiiensis (A105) were found to possess antimicrobial, antibiotic and antitumor metabolites.27–30 However, our study reported for the first time antimycobacterial activity of Streptomyces avidinii and Streptomyces spiroverticillatus against M. smegmatis. Streptomyces hawaiiensis was noted for its production of acyldepsipeptides (ADEPs), the prospective antituberculosis agents against M. tuberculosis. 31 Streptomyces alboniger (A121) was first isolated in 1954 from the soil samples in Illinois, America 32 and was found to exhibit potential cytotoxic effect on many cancer cell lines. 33 In our study, Streptomyces alboniger (A121) was the most active strain against Mycobacterium smegmatis. It is a light brown substrate bacteriophage, a gray white color, and is capable of forming colonies. The investigation of metabolites from these strains led to the isolation of pamamycin, an antibiotic resistance toward many bacterial pathogens, including Mycobacterium smegmatis and Mycobacterium phlei. 34

Colony morphology of the five most active actinomycetes (A002, A067, A079, A105, A121) against M. smegmatis.
Structural Elucidation of the Compounds Isolated from Streptomyces Alboniger (A121)
Streptomyces alboniger (A121) was chosen for further isolation and characterization of the bioactive metabolites. Compound

The structures of compound

HMBC, COSY and NOESY correlations of compounds
Compound
1H and 13C NMR Spectral Data of Compound
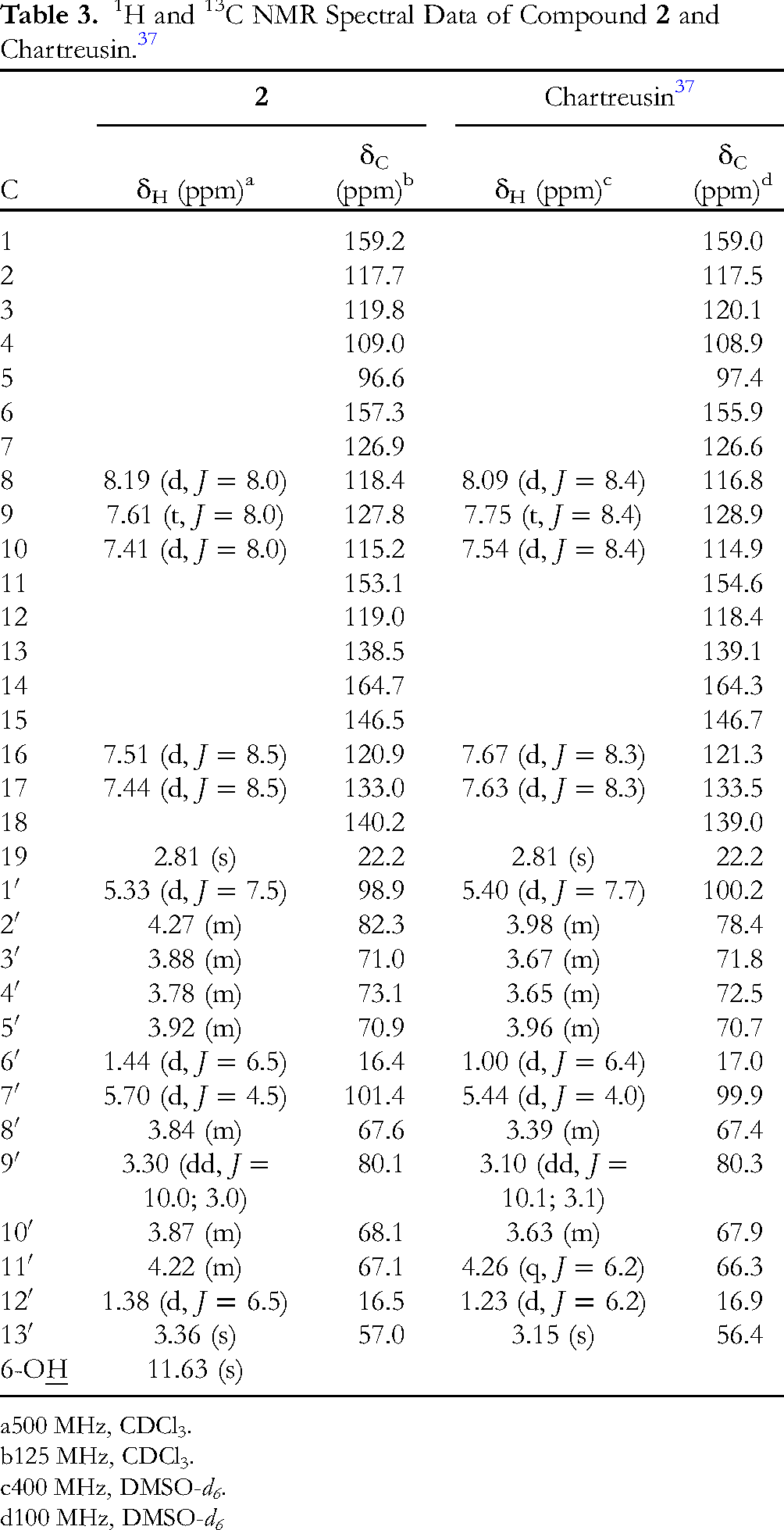
500 MHz, CDCl3.
125 MHz, CDCl3.
400 MHz, DMSO-d6.
100 MHz, DMSO-d6
Antimycobacterial Activity
Compounds
Antimycobacterial Activity of Compound

*: Inhibition zone diameter (mm)
Additionally, compound
Antibacterial Activity of Compound

Cytotoxic Activity
Compound
Cytotoxic Activity of Compound

Conclusion
In conclusion, we have discovered and reported for the first time the potential antimycobacterial activity of the actinomycetes collected in the regions along the north to the central of Vietnam, including Streptomyces avidinii (A002), Streptomyces spiroverticillatus (A067), Streptomyces wuyanensis (A079), Streptomyces hawaiiensis (A105), Streptomyces alboniger (A121). Obscurolide B2β (
Materials and Methods
Chemical and Equipment
NMR spectra were recorded on a Bruker Avance 500 MHz spectrometer, using deuterated solvents and tetramethylsilane as internal standard. ESI-MS spectra were measured on an Agilent LC-MSD-Trap SL instrument. Polar rotation was obtained on a Jasco P-2000 Polarimeter serial A060161232 equipment. Merck 60F254 pre-coated silica gel plates (0.2 mm thickness) were used for thin layer chromatography (TLC). Silica gel 60 (0.040-0.063 mm, 230-400 mesh), Sephadex LH-20 were purchased from Merck. Other chemicals were purchased from Merck and used without purification unless otherwise needed.
Sample Collection
Soils and sediments were collected along the areas from Northern to Central Vietnam (nine areas in Table 1). The coordinates of the collection areas were recorded by GPS navigation device. Moisture, salinity and temperature of the soil samples were determined by soil testing equipment. Each sample was placed in a 50 mL falcon tube and kept in the refrigerator (0-5 °C) during transport to the laboratory.
Isolation and Identification of Actinomycetes
The samples were left drying naturally for 2–3 days and powdered. Each sample (1 g) was diluted to 10−3 and 10−4 concentrations, then spread onto a Petri dish using two different media, NaST21 (solution A: 750 mL sterilized seawater, 1 g K2HPO4, 16 g Bactogar; solution B: 250 mL sterilized seawater, 1 g KNO3, 1 g MgSO4, 1 g CaCl2.2H2O, 0.2 g FeCl3, 0.1 g MgSO4.7H2O) and HV (1 g Humic acid, 0.02 g CaCO3, 0.01 g FeSO4.7H2O, 1.7 g KCl, 0.05 g MgSO4.7H2O, 0.5 g Na2HPO4, 5 mL B-group vitamins, 16 g Agar, 1 L distilled water, pH 7). Both culture media above were supplemented with nalidixic acid (20 mg/L) and cycloheximide (50 mg/L). The isolation plates were incubated at 28–30 °C for 14 to 21 days. Colonies with different morphology were cultured on YS medium (2 g yeast extract, 10 g agar, 1 L distilled water) and kept at − 80 °C. Identification and classification of actinomycetes were determined based on colony, spore chain morphology and 16S rRNA gene sequencing. Amplifications of the 16S rRNA gene from total DNA were carried out through PCR using the primers pair 27F (5′-AGAGTTTGATCCTGGCTCAG-3) and 1492R (5′-GGTTACCTTGTTACGACTT-3). The PCR product was analyzed by 1% agarose gel electrophoresis, purified by PCR purification Kit (Bioneer, Korea) and finally sequencing. The sequence of the 16S rRNA fragment was used to detect the species by the EZ Taxon tool. Actinomycete strains obtained from the purification step were kept in glycerol (20%) at − 80 °C for further studies.
Antimycobacterial Activity Assay
Antimycobacterial activity was determined by the agar diffusion assay. 42 Actinomycete strains were cultured on YS agar medium at 30 °C for 7 days. Mycobacterium smegmatis mc2155 KCTC 9108 strain was obtained from Center for Nutraceutical and Pharmaceutical Materials Myongji University, Korea. This strain was grown on 219 liquid medium (g/L: peptone – 10, yeast extract – 5, malt extract – 5, casamino acid – 5, beef extract – 2, glycerol – 2, tween 80-50 mg, MgSO4.7H2O – 1) for 24 h at 30 °C. Bacterial culture was added to the sterilized 219 agar medium with the ratio of 1% and poured into a petri dish. Cultured pieces of agar containing actinomycetes with 5 mm in diameter were placed onto plate. For testing the extract or compounds obtained from fermentation and isolation, holes in the agar plate were made by a plastic tube, following adding a solution of extract or compound (50 µL) to these holes. The agar plates were incubated at 30 °C for 24 h. Antimycobacterial activity was determined by inhibition zone diameter on the agar plate. Agar pieces without bacteria were used as negative control. All experiments were performed in triplicate and the data were obtained as mean ± SD. Rufomycin was used as a reference substance with the test concentration of 10 µg/mL.
Antimicrobial Activity Assay
Antimicrobial activity was determined by Broth microdilution method. 43 The test microorganisms included three Gram-positive bacteria Staphylococcus aureus ATCC 13709, Bacillus subtilis ATCC 6633, Lactobacillus fermentum N4 and three Gram-negative bacteria Salmonella enterica, Escherichia coli ATCC 25922, Pseudomonas aeruginosa ATCC 15442. MHB (Mueller-Hinton Broth), MHA (Mueller-Hinton Agar); TSB (Tryptic Soy Broth); TSA (Tryptic Soy Agar) were used as culture media. The microbacteria were kept at − 80 °C and activated in culture medium before use, to reach the concentration of 5 × 105 CFU/mL. The test samples were diluted in DMSO and sterilized distilled water to obtain the concentrations of 128.0, 32.0, 8.0, 2.0 and 0.5 µg/mL. 10 µL of sample solution at different concentrations were added into a 96-well plate, following the addition of 200 µL of microbacterial solution. The place was incubated at 37 °C for 24 h. All experiments were tested in triplicate and the data were obtained as mean ± SD. Wells with bacteria suspension in growth medium were used as positive control, while the one containing culture medium without bacteria was considered as negative control. Ampicillin and cefotaxime were used as the reference substance. The MIC (Minimum Bactericidal Concentration) value was determined at the well with the lowest concentration of the test sample that inhibited the growth of bacteria. The IC50 (50% inhibitory concentration) value is calculated by TECAN spectrometer and raw data software.
Cytotoxic Activity Assay
The cytotoxic activity was performed based on MTT asaay. 44 This method assessed the cell viability through the discoloration of tetrazolium salt (MTT - (3-(4,5-dimethylthiazol-2 - yl)- 2, 5 - diphenyltetrazolium)). The samples were evaluated against carcinoma cancer cells (KB), liver cancer cells (Hep-G2), lung cancer cells (Lu-1) and breast cancer cells (MCF-7). The sample (20 µL) was introduced into the wells of the 96-well plate at the following concentration: 128 µg/mL; 32 µg/mL; 8 µg/mL and 2 µg/mL. The cells were cultured in Dulbecco's modified eagle's medium with 10% fetal bovine serum, 1% penicillin and streptomycin, 1% L-glutamine at 37 °C, 5% CO2 for 72 h. Cells were separated with trypsin and seed to reach the concentration of 3.104 cell/mL. After that, 180 µL of cells was put onto each well of the 96-well plate containing the sample. The plate was incubated at 37 °C, 5% CO2 for 72 h. After incubation, 10 μL of MTT solution in the phosphate buffer was added and the plate was further incubated for 2–4 h. MTT was removed and 70 μL of DMSO was added. Positive control wells contained culture medium with cells, while the one containing only culture medium served as negative control. Ellipticine was used as the reference substance. All experiments were conducted in triplicate and the data were obtained as mean ± SD. The results were determined by the OD values (optical density), which were measured on a Microplate reader EPOCH 2 spectrophotometer, Biotech, USA at 540 nm. The IC50 value was calculated based on the percentage inhibition value by Table curve software.
Fermentation, Extraction and Isolation of the Metabolites from Strain A121 (Streptomyces alboniger)
Strain A121 was cultured on agar medium for 5–7 days, then it was inoculated into a 1000 mL conical flask containing 350 mL of YS medium, shaking at 160 rpm, 28–30 °C for 72 h. The actinomycete culture broth was inoculated into the 50-liter fermentation equipment with the ratio of 2.5–3% (v/v). The culture strain reached the highest cell density after 144 h, indicating that the growth of actinomycetes was at the equilibrium phase. When the fermentation process ended, the actinomycete culture was collected. Biomass and actinomycete culture were separated by centrifugation, which was used immediately for the extraction and isolation process.
The culture of strain A121 (50 L) was centrifuged at 10 000 rpm (20 min, room temperature). The filtrate (40 L) was extracted with EtOAc (3 × 24 h × 30 L). The EtOAc extracts were grouped and the solvent was evaporated in vacuo to give 6.2 g EtOAc residue (121E). The aqueous fraction remaining after extraction with EtOAc was condensed to yield 10.2 g aqueous residue (121CN). The EtOAc extract (6.2 g) was subjected to a Sephadex LH-20 column (MeOH 100%) to afford five subfractions (121E.1 → 121E.5). Fraction 121E.2 (3.4 g) was purified by a silica gel column (n-hexane/CH2Cl2 98:2→ 1:1) to yield four fractions (121E.2.1 → 121E.2.4). Fraction 121E.2.3 (30 mg) was separated over a silica gel column (CH2Cl2/Acetone 9:1→8:2) to give compound
The cell residue (biomass) was extracted with MeOH/H2O (9:1, 3 × 4 h × 1 L), then centrifuged at 10 000 rpm at room temperature to remove the residue. The solvent was evaporated to provide 7 g residue (121X). The fraction 121CN and 121X were examined by thin layer chromatography. As a result, these fractions contained similar compounds, which were therefore grouped together to give fraction 121CX (17 g). Fraction 121 CX was separated on a Sephadex LH-20 column (MeOH/H2O 9:1) to obtain 8 subfractions (121CX1 → 121CX8). The fraction 121CX5 (0.1 g) was subjected to a Sephadex LH-20 column (MeOH/H2O 9:1) to obtain 5 fractions (121CX.5.1 →121CX.5.5). Fraction 121CX5.4 (101 mg) was repeatedly purified on a Sephadex LH-20 column (MeOH/H2O 9:1) to afford compound
Footnotes
Acknowledgments
The authors are grateful to the Ministry of Science and Technology of Vietnam (MOST) for financial support for this study (Vietnam-Korea International bilateral cooperative project, Project Nr: NĐT.47.KR/18). PhD Huynh Thi Ngoc Ni and Master Ngo Van Hieu acknowledge the financial support by the Domestic Master/Ph.D. Scholarship Programme of Vingroup Innovation Foundation.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Our institution does not require ethical approval for reporting individual cases or case series.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Vietnamese Ministry of Science and Technology, (grant number NDT.47.KR/18)
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Supplemental Material
Supplemental material for this article is available online.
